Abstract
Key Message
Coordinated regulation of amylose and amylopectin synthesis via manipulation of SSII-2, SSII-3 and Wx expression in endosperm can improve rice eating and cooking quality.
Abstract
With increasing rice consumption worldwide, many researchers are working to increase the yield and improve grain quality, especially eating and cooking quality (ECQ). The rice ECQ is mainly controlled by the expression of starch synthesis-related genes (SSRGs) in endosperm. Although the Wx and SSII-3/SSIIa/ALK genes, two major SSRGs, have been manipulated to improve rice ECQ via various breeding approaches, new methods to further improve ECQ are desired. In our previous study, we enhanced rice ECQ by knocking down SSII-2 expression in the japonica Nipponbare cultivar (carrying the Wxb allele) via RNA interference. Herein, the SSII-2 RNAi was introduced into two Nipponbare-derived near-isogenic lines (NILs), Nip(Wxa) and Nip(wx), carrying Wxa and wx alleles respond for high and no amylose levels, respectively. Analysis of physicochemical properties revealed that the improved grain quality of SSII-2 RNAi transgenic lines was achieved by coordinated downregulating the expression of SSII-2, SSII-3 and Wx. To further confirm this conclusion, we generated ssii-2, ssii-3 and ssii-2ssii-3 mutants via CRISPR/Cas9 technique. The amylopectin structure of the resulting ssii-2sii-3 mutants was similar to that in SSII-2 RNAi transgenic lines, and the absence of SSII-2 decreased the amylose content, gelatinisation temperature and rapid visco-analyser profile, indicating essential roles for SSII-2 in the regulation of amylopectin biosynthesis and amylose content in rice endosperm. The effect of SSII-2 was seen only when the activity of SSII-3 was very low or lacking. Our study provides novel approaches and valuable germplasm resources for improving ECQ via plant breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is a major staple food worldwide, and its yield has been greatly increased over recent decades, but grain quality still requires improving. Rice grain quality includes eating and cooking quality (ECQ) (Huang et al. 2020a), nutrition quality (Yang et al. 2019; Zhu et al. 2018), grain appearance (Zhao et al. 2018) and milling quality (Wang et al. 2016). Among these, ECQ is of foremost consideration for both consumers and rice breeders. Starch, accounts for ~ 90 % of the edible part of milled rice, and is considered the primary determinant of rice ECQ because it influences the palatability, appearance, hardness, stickiness and digestibility of cooked rice. Starch is a semi-crystalline particle consisting of two glucose polymers, linear amylose and highly branched amylopectin. Both the composition and fine structure of starch determine its physicochemical properties, and hence the eventual rice ECQ. Previous studies demonstrated that rice cultivars with low-to-intermediate amylose content (AC) tend to show better palatability, stickiness and hardness than those of both glutinous and high-AC rice types (Huang et al. 2020b; Tian et al. 2009; Zhang et al. 2019a; Zhou et al. 2020). In addition, an increase in the ratio of amylopectin A chains (Ap-A) with degree of polymerization (DP) 6–12 to B1 chains (Ap-B1) with DP 13–24 may reduce gelatinisation temperature (GT) and hardness, and improve the stickiness of cooked rice, resulting in better texture and cooking properties (Li et al. 2016, 2020; Zhu 2018). Therefore, a moderate reduction in AC and an increase in ratio of amylopectin A chains to B1 chains are keys to improving rice ECQ.
The physicochemical properties of starch, including the apparent amylose content (AAC), gel consistency (GC), gelatinisation temperature (GT) and rapid visco-analyser (RVA) score, have been utilised to evaluate rice ECQ (Tan et al. 1999; Tian et al. 2009). Starch synthesis-related genes (SSRGs), including genes encoding ADP glucose pyrophosphorylase (AGPase), soluble starch synthase (SS), granule-bound starch synthase (GBSS), starch branching enzyme (SBE) and starch debranching enzyme (DBE), function cooperatively to determine these traits (Jeon et al. 2010). The Waxy (Wx) gene encoding granule-bound starch synthase I (GBSSI) is the major determinant of AC and GC, and a minor contributor to GT (Sano 1984; Su et al. 2011; Tian et al. 2009; Wang et al. 1990; Zhang et al. 2019a). SSII-3 (also known as ALK and SSIIa) is a major gene regulating GT, and it also contributes to AC and GC (Tian et al. 2009; Umemoto et al. 2002). Wx and SSII-3 genes are closely located on chromosome 6, hence they have been co-selected during rice domestication (Tian et al. 2009).
To increase the proportion of amylopectin short chains with DP 6–12, SSII genes encoding soluble starch synthase II (SSII), which elongates amylopectin intermediate chains with DP 13–24 by extending amylopectin short chains with DP 6–12, have been knocked down with some success (Chen et al. 2020; Commuri and Keeling 2001; Fujita et al. 2006; Morell et al. 2003). There are three SSII isoforms in rice; SSII-1 (SSIIc), SSII-2 (SSIIb) and SSII-3 (SSIIa). The amino acid sequence identity among the three isoforms is greater than 50 %, and they share the consensus motif KXGGL, which is believed to form part of the ADPG binding site of SSII (Furukawa et al. 1990; Jiang et al. 2004; Ohdan et al. 2005b). These three SSII genes in rice are thought to result from two replications of large genomic fragments; the first replication produced two SSII genes, one of which differentiated into SSII-1, and the other subsequently produced SSII-2 and SSII-3 in a second replication (Jiang et al. 2004). The gene organisation near the SSII-2 and SSII-3 loci is similar; both contain the PLAGXNVMNX motif that is conserved in many starchy crops (Harn et al. 1998; Li et al. 1999). Studies showed that the ssii-3 mutant contained more short amylopectin A chains (DP 6−12) and fewer medium-length amylopectin B1 chains (DP 13−24), leading to beneficial starch properties for improving rice ECQ (Gao et al. 2003; Miura et al. 2018; Umemoto et al. 2002). Key amino acid variations in SSII-3 can affect its enzyme activity and cause GT differences among rice cultivars, and at least three SSII-3 alleles have been reported (Bao et al. 2006; Chen et al. 2020; Nakamura et al. 2005; Umemoto et al. 2002). The activity of japonica types SSIIaj is only ~ 10 % that of indica type SSIIai/ALKc, resulting in a lower GT (Chen et al. 2020; Nakamura et al. 2005). The recent study showed that the SSIIaj gene from japonica cultivars could be further divided into two alleles ALKa and ALKb, and the GT of ALKb is lower than that of ALKa (Chen et al. 2020). The ssiiaj-deficient mutant displayed a lower GT than those of both ALKa and ALKb (Chen et al. 2020; Miura et al. 2018). In addition to SSII-3, the other two SSII genes, especially SSII-2, also have potential as targets for regulating the chain-length distribution (CLD) of amylopectin in rice endosperm (Li et al. 2018).
AC is the chief determinant of rice ECQ (Tian et al. 2009). Natural allelic variation in Wx, including Wxlv, Wxa, Wxin, Wxb, Wxmw, Wxmp, Wxmq, Wxop/hp and wx, has led to a wide variety of AC values, ranging from 30 to 0 % in different rice cultivars (Cai et al. 1998; Liu et al. 2009; Mikami et al. 1999, 2008; Sato et al. 2002; Zhang et al. 2019a). Wxa (AC ~ 25 %) and Wxb (AC 15−18 %) are the two major Wx alleles in indica and japonica rice cultivars, respectively. The wx allele is a null Wx type present in glutinous rice (Cai et al. 1998; Hirano et al. 1998). Recent studies showed that the Wx alleles have undergone a high-to-low AC selection trend during rice domestication (Anacleto et al. 2019; Zhang et al. 2019a). Application of the elite Wx alleles, particularly Wxin, Wxb and Wxmp corresponding to intermediate-to-low AC, has been the major method employed to improve rice ECQ in modern rice breeding programs (Phing Lau et al. 2016). For example, introgression of Wxmp or Wxmq alleles (AC 8−12 %) into commercial rice cultivars can be used to breed elite ‘soft rice’ with high ECQ and soft grains after cooking (Zhang et al. 2019b).
RNA interference (RNAi) is a powerful tool for knocking down target gene expression by inducing the degradation of corresponding mRNAs (Fire et al. 1998). More recently, CRISPR/Cas9 has been used to knockout target genes efficiently and specifically (Miao et al. 2013; Zhang et al. 2014). Gene knockout is typically achieved by frame-shift mutation via CRISPR/Cas9, and the inserted transfer DNA (T-DNA) can be removed by separation, hence this simple approach has been widely adopted in modern molecular plant breeding programs (Chen et al. 2019; Donohoue et al. 2018).
In our previous studies, we suppressed the expression of SSII-2, a neglected SSRG, in japonica rice cultivar Nipponbare using RNAi, which markedly reduced AC and GT, and increased GC, thereby significantly improving rice ECQ (Li et al. 2018; Xu et al. 2020b). SSII-2 RNAi transgenic rice exhibited similar ECQ as traditional soft rice, but the appearance was much better than that of soft rice. Thus, this represents a promising approach for generating novel elite soft rice. To confirm this notion and further evaluate its potential for cooperatively improving rice quality using various Wx alleles, we herein introduced the SSII-2 RNAi construct into Wxa and wx near-isogenic lines (NILs) derived from Nipponbare (Wxb). In addition, we further created ssii-2 and/or ssii-3 single and double mutants using CRISPR/Cas9 to explore the underlying mechanism by which SSII-2 influences the regulation of rice grain quality, and to provide novel germplasm resources for breeding high-ECQ rice.
Materials and methods
Plant materials
The japonica rice cultivar Nipponbare (Nip) and its derived NILs, transgenic lines and mutants were used in this study (Table 1). Nipponbare itself has the Wxb allele with a relate low AC (~ 16 %), herein named Nip(Wxb) or wild-type (WT). The two NILs with Wxa and wx alleles in the Nipponbare background are called Nip(Wxa) and Nip(wx), respectively, corresponding to high and zero AC (Zhang et al. 2019a). Nip(Wxa) (BC8F7) and Nip(wx) (BC9F8) were generated after at least eight rounds of backcrossing with the recurrent Nip and their genetic backgrounds were genotyped by whole-genome sequencing (Zhang et al. 2019a). The Wxa allele is from the indica cultivar Longtefu, while the wx allele is from the japonica waxy cultivar Suyunuo. Nip(Wxb)_Ri-1 and Nip(Wxb)_Ri-2 are two representative SSII-2 RNAi transgenic lines in the Nipponbare background, corresponding to SSSII-2 RNAi Line 3 and SSSII-2 RNAi Line 4 in our previous study, respectively (Li et al. 2018). SSII-2 RNAi was introduced into the NILs Nip(Wxa) and Nip(wx) by crossing, and four homozygous lines, Nip(Wxa)_Ri-1, Nip(Wxa)_Ri-2, Nip(wx)_Ri-1 and Nip(wx)_Ri-2, were selected in their selfing progenies by genotyping (Table 1).
Nipponbare was also used to generate the related ssii-2 and ssii-3 mutants via CRISPR/Cas9 (Table 1). All the above rice lines were grown under the same climatic and management conditions during the summer in a paddy field at Yangzhou University, Yangzhou, China (32°23′N). Each line was grown in triplicate and planted randomly in each plot.
Construction of CRISPR/Cas9 vectors and screening of homozygous mutants
The CRISPR/Cas9 system used in this study included two vectors, intermediate vector SK-gRNA and the CRISPR-Cas9 binary vector pC1300-Cas9 (Wang et al. 2015). The target sites in SSII-2 and SSII-3 genes (Fig. S1A, B) were designed via the CRISPR-GE (http://skl.scau.edu.cn/) online toolkit and individually cloned into SK-gRNA (Xie et al. 2017). Genomic DNA (gRNA) fragments containing the target sites were then ligated into pC1300-Cas9 to obtain the SSII-2 or SSII-3 single-knockout vectors and the SSII-2SSII-3 double-knockout vector via the isocaudamers-based system (Fig. S1C, D). Confirmed constructs were introduced into Nipponbare by Agrobacterium-mediated transformation. The primers used are listed in Table S1.
The target DNA region of SSII-2 and SSII-3 were verified by sequencing, and the sequencing data were analysed by DSDecodeM (http://skl.scau.edu.cn/dsdecode/) or manual decoding (Liu et al. 2015). The homozygous ssii-2 and/or ssii-3 mutants without the T-DNA insertion were screened and selected for following analysis. Six homozygous mutated lines, two from each construction, were selected for further experimental analysis, and named s2-1, s2-2, s3-1, s3-2, s23-1 and s23-2 (Table 1), where s2 and s3 are single null ssii-2 or ssii-3 mutants, while s23 indicates double mutants with both ssii-2 and ssii-3 mutations.
Total RNA extraction and quantitative real-time PCR (qRT-PCR) expression analysis
Total RNA was extracted from rice caryopses (with glumes removed) 10 days after flowering (DAF) using an RNAplant Plus Reagent kit (Tiangen, Beijing, China). First-strand cDNA was synthesised using a PrimeScript RT reagent kit (Takara, Kusatsu, Japan), and qRT-PCR was performed on a CFX Connect Real-Time PCR Detection System (Bio-Rad) using AceQ SYBR qPCR Master Mix (Vazyme, Nanjing, China). The Actin01 gene was used for normalisation, and each experiment included three biological replicates. The primers used are listed in Table S1, and were tested in our previous study (Fan et al. 2019).
Western blotting
Rice flour from mature grains was weighted and suspended in total protein extraction buffer comprising 125 mM Tris-HCl pH 6.8, 4 M urea, 4 % sodium dodecyl sulphate (SDS) and 5 % 2-mercaptoethanol at 37 °C at a ratio of 1:15 (1 mg flour to 15 µL buffer). The buffer volume for glutinous rice was doubled due to its high stickiness. The extracted total proteins were mixed with SDS loading buffer (4×), denatured at 99 °C for 10 min, separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequently transferred to a polyvinylidene difluoride (PVDF) membrane. The antibody specifically recognizing GBSSI (Clontech) was used for examining the accumulation of GBSSI, and the acid subunit of glutelin was set as loading control.
Measuring rice grain physicochemical properties
Air-dried mature seeds were dehusked using a SY88-TH instrument (Sangyong, Korea), polished (Pearlest, Kett, Japan), ground into powder using a FOSS 1093 Cyclotec Sample Mill (Sweden), and passed through a 100-mesh screen (pore size 0.15 mm). The starch content of the milled rice flour was analysed using a K-TSTA total starch assay kit (Megazyme, Ireland). The crude protein content was calculated from the nitrogen content of the rice flour using a FOSS TECATOR Kjeltec2300 instrument (according to AOAC standard method 990.03) (Zhang et al. 2013). AAC, GC, GT and RVA were determined according to our previously reported method (Zhang et al. 2016). In brief, AAC was calculated using an iodine staining method, GC was determined by measuring the length of rice gel after gelatinisation and cooling, and GT was measured using a DSC 200 F3 differential scanning calorimeter (Netzsch, Germany). The obtained DSC curves included the temperature of onset (To), peak (Tp) and end (Te), and the enthalpy of gelatinisation (△H). RVA values were measured using a Techmaster RVA instrument (Newport Scientific, Warriewood, Australia). The primary RVA parameters include peak viscosity (PKV), hot paste viscosity (HPV), cool paste viscosity (CPV), setback viscosity (SBV), breakdown viscosity (BDV), peak time (PeT) and pasting temperature (PaT). All tests were performed in duplicate or triplicate.
Gel permeation chromatography (GPC) and fluorophore-assisted carbohydrate electrophoresis (FACE) analyses
Starch was extracted from milled rice endosperm using the neutral protease method (Wang and Wang 2001) with slight modifications. In brief, purified rice starch was debranched with isoamylase (EC3.2.1.68, E-ISAMY, Megazyme), dissolved in dimethyl sulphoxide (DMSO), and used to determine the relative molecular weight distribution by GPC using a PL-GPC 220 instrument (Agilent, USA). The three well-resolved fractions from GPC curves represent A and B1 chains of amylopectin (Ap1), B2 to B4 chains of amylopectin (Ap2), and amylose (Am), respectively.
Debranched starch was also quantitatively analysed using FACE with a PA-800 Plus instrument (Beckman, USA) and APTS-labelled linear glucans to determine the chain-length distribution (CLD) of amylopectin, denoted Nde(X) (Gu et al. 2019). The ΔNde (X) value, representing the change in CLD, was then calculated from the Nde(X) value for transgenic lines minus the Nde(X) value for the corresponding WT, where, de indicates linear glucans and X indicates the degree of polymerisation (DP).
Statistical analysis
At least two replicates were performed for each experiment. All data are presented as means ± standard deviations (means ± SD). One-way analysis of variance (ANOVA) was used to determine the level of significance (* and ** indicate significant differences at p < 0.05 and p < 0.01, respectively). Different lower-case letters indicate statistically significant differences at p < 0.05.
Results
SSII-2 RNAi simultaneously regulates amylose content and amylopectin structure to improve the quality of non-waxy rice
In our previous study, we successfully generated SSII-2 RNAi transgenic rice in Nip (Wxb) (Li et al. 2018; Xu et al. 2020b). Both AAC and GT were decreased in these transgenic rice, while GC was increased, which led to significantly improved rice ECQ without any defects in endosperm transparency. This novel soft rice has a far better appearance than traditional soft rice. To determine whether the effect of SSII-2 is dependent on the specific allele type of the Wx gene, a key determinant of rice ECQ, we further introduced the same SSII-2 RNAi construct into NILs Nip(Wxa) carrying the representative Wxa alleles in non-waxy rice, as shown in Table 1. The main agronomic traits of all transgenic lines were similar to those of their WT counterparts (Fig. S2), consistent with our previous studies (Li et al. 2018; Xu et al. 2020b). Therefore, neither suppression of SSII-2 expression nor changes in Wx alleles affected rice growth and development.
In the high AC Nip(Wxa) background, knockdown of SSII-2 expression had no effect on the total starch or protein content (Fig. S3A, B). However, both AAC and GT data from DSC, including To, Tp and Te, were dramatically decreased in transgenic lines (Fig. 1 A and C and Fig. S3C, D), while GC values were significantly increased (Fig. 1B). RVA profile of rice flours revealed an overall downward trend for transgenic lines compared with WT (Fig. 1D, E). In more detail, PKV, HPV, CPV, SBV, PeT and PaT were significantly decreased, while only BDV was increased (Table S2). Low AAC, soft GC, high BDV and low SBV are all representative characteristics favored by Chinese rice consumers. These results indicate that suppression of SSII-2 in the Wxa background had the same effect on grain physicochemical properties as those in Wxb rice, suggesting that knockdown of SSII-2 expression could be a universal strategy for improving rice quality.
Comparison of physicochemical properties and starch fine structures of mature seeds between SSII-2 RNAi transgenic rice and their corresponding Wx NIL wild types. A Apparent amylose content (AAC) of rice flours. B Gel consistency (GC) of rice flours. C Peak gelatinisation temperature (GT) of rice flours determined by DSC. D−F Rapid visco-analyser (RVA) profile of rice flours in Nip(Wxa), Nip(Wxb) and Nip(wx) backgrounds, respectively. (G−I) Fine structures of starches determined by gel permeation chromatography (GPC) in Nip(Wxa), Nip(Wxb) and Nip(wx) backgrounds, respectively. J Changes in chain length distribution of amylopectin determined by fluorophore-assisted carbohydrate electrophoresis (FACE). Error bars represent SD (*p < 0.05 and **p < 0.01; ns no significant difference)
Next, GPC and FACE were used to examine the starch fine structure and CLD of SSII-2 RNAi transgenic rice in both Nip(Wxa) and Nip(Wxb) backgrounds. The GPC results indicated that the amylose proportion (Am) in isolated grain starch from transgenic lines was significantly decreased, while the amylopectin proportion (Ap), especially Ap1, was increased (Fig. 1G, H). The FACE data showed that the proportion of amylopectin A chains with a DP value from 7 to 11 was significantly increased, and the proportion of B1 chains with a DP value from 13 to 20 was slightly decreased in SSII-2 RNAi transgenic rice (Fig. 1J). These results implied that suppression of SSII-2 expression in non-waxy rice could simultaneously modulate both AC and amylopectin structure, thereby leading to improved rice ECQ.
SSII-2 RNAi improves the amylopectin structure of glutinous rice
To explore the specific effect of SSII-2 on the amylopectin structure, we also generated SSII-2 RNAi/Nip(wx) transgenic rice, and grains contained similar amounts of starch and crude protein to those in the WT Nip(wx) counterpart (Fig. S3A, B). GT, estimated from To, Tp and Te using DSC analysis, was dramatically decreased in both transgenic lines. The above results are consistent with those for the Nip(Wxb) and Nip(Wxa) backgrounds. Regarding AAC and GC, no remarkable changes were observed (Fig. 1 A−C and Fig. S3E). In addition, the overall RVA profile of SSII-2 RNAi/Nip(wx) transgenic rice showed a general downward trend (Fig. 1F). The only difference was that the PKV of SSII-2 RNAi/Nip(wx) rice was slightly increased, unlike that of the Nip(Wxa) and Nip(Wxb) backgrounds (Table S2).
From the GPC data, no amylose peak was observed either of the SSII-2 RNAi transgenic lines or the Nip(wx) control (Fig. 1I). This is a typical feature of waxy rice due to the absence of the GBSSI enzyme. However, the Ap1 fraction of amylopectin was significantly increased in SSII-2 RNAi/Nip(wx) transgenic rice compared with the WT counterpart, while the Ap2 fraction was decreased. As expected, consistent changes in amylopectin CLD were observed for the SSII-2 RNAi/Nip(wx) transgenic lines (Fig. 1J). The results indicate that RNA interference of SSII-2 only influenced the amylopectin structure of waxy rice.
Interestingly, when we compared the altered amylopectin CLD in SSII-2 RNAi transgenic lines with different Wx alleles and AAC, the amylopectin structure in SSII-2 RNAi/Nip(Wxb) showed greater change than that of SSII-2 RNAi/Nip(Wxa), while the change in SSII-2 RNAi/Nip(wx) was greater than that of SSII-2 RNAi/Nip(Wxb), indicating that the lower the AC, the greater the change in amylopectin structure (Fig. 1J). These results further confirmed that downregulating SSII-2 in rice endosperm could help to coordinate the synthesis and structure of amylose and amylopectin.
SSII and Wx are downregulated in the endosperm of SSII-2 RNAi transgenic rice
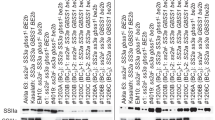
To clarify the mechanism by which SSII-2 affects the physicochemical properties and structure of starch, we examined the expression of SSII subfamily genes and Wx gene, especially the two major genes (Wx and SSII-3) respectively controlling AC and GT. The qRT-PCR data showed that expression of SSII-2 was dramatically decreased in the developing seeds of SSII-2 RNAi lines with different Wx alleles, demonstrating that suppressing SSII-2 expression via RNAi is a promising strategy (Fig. 2A). In addition, both the transcript and protein abundance of Wx were significantly reduced in the caryopses of SSII-2 RNAi non-waxy rice, while no Wx expression was detected in any of the Nip(wx)-related lines (Fig. 2B, C). Therefore, interfering with the expression of SSII-2 simultaneously reduced the expression of Wx, which may largely account for the decline in AC in non-waxy SSII-2 RNAi transgenic rice. Regarding the other two SSII members, expression of SSII-3 was also markedly reduced in all SSII-2 RNAi lines (Fig. 2D), while SSII-1 expression was almost unchanged (Fig. 2E). Given that SSII-3 is the major gene controlling rice GT, the reduced SSII-3 expression could explain the decrease in GT and modified amylopectin CLD in the SSII-2 RNAi transgenic lines.
Expression analysis of SSII and Wx genes in SSII-2 RNAi transgenic rice and the wild type controls. A–E qRT-PCR expression analysis of SSII-2, Wx, SSII-1 and SSII-3. C Western blotting analysis of the GBSSI protein. Rabbit anti-GBSSI antibody was used to detect GBSSI. The acid subunit of glutelin served as a loading control. NA, not applicable. Error bars represent SD (*p < 0.05 and **p < 0.01; ns, no significant difference)
Generation of null ssii-2 and ssii-3 mutants by CRISPR/Cas9
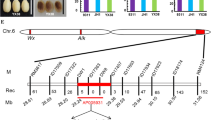
Since the expression of SSII-3, the close isoform of SSII-2, was also decreased, the CRISPR/Cas9 system was used to further confirm the effects of SSII-2, and avoid the potential off-target effect caused by the RNAi method. Thus, ssii-2 and ssii-3 single mutants and the ssii-2ssii-3 double mutant were generated via CRISPR/Cas9 (Fig. S1). Ten mutants without T-DNA insertions were selected after several generations of screening, including five ssii-2 single mutants (s2), two ssii-3 single mutants (s3) and three ssii-2ssii-3 double mutants (s23). All these mutants had undergone one insertion or deletion event in the target site that caused a frameshift mutation in the target gene (Fig. 3A). Except for a slight decrease in kernel weight, none of the other agronomic traits of the mutants were altered (Fig. S4). Therefore, two typical mutants of each type were selected for subsequent analyses.
Generation of s2, s3 and s23 mutants, and expression analysis of SSII and Wx genes in s2, s3 and s23 mutants. A Mutation information for the target sites of SSII-2 and SSII-3 genes. Red nucleotide sequences and red dashes in different lines indicate sequence changes. PAM, proto-spacer-motif; Var., variants of identified mutations; WT, wild type. B−D Relative expression of SSII-2, SSII-3 and Wx. E The accumulation of GBSSI in s2, s3 and s23 mutants. Different lower-case letters indicate statistically significant differences at p < 0.05
Effects of SSII-2 and SSII-3 mutations on SSII and Wx expression in endosperm
The abundance of SSII and Wx transcripts in developing endosperm of mutants and WT plants was analysed. The qRT-PCR results showed that expression of both SSII-2 and SSII-3 was dramatically decreased in s2 mutants (Fig. 3B, C). In s3 mutants, expression of both SSII-2 and SSII-3 was slightly lower than in controls. In s23 double mutants, both SSII-2 and SSII-3 expression was also greatly decreased (Fig. 3B, C). Furthermore, the Wx expression was significantly decreased in s2 mutants, slightly decreased in s23 double mutants but a little bit increased in s3 mutants (Fig. 3D), while only the amount of GBSSI protein was decreased in s2 mutants and no significant change was observed in s3 and s23 mutants (Fig. 3E). These results indicate that knockout of SSII-2 can markedly reduce the expression of SSII-2, SSII-3 and Wx genes, while knockout of SSII-3 alone or both SSII-2 and SSII-3 mainly affected the expression of SSII-2 and SSII-3, with minimal or no effect on Wx expression.
Improvements in grain quality and starch structure in ssii-2 and ssii-3 null mutants are similar to those in SSII-2 RNAi transgenic lines
The grain physicochemical properties of the mutants were analysed and compared in detail. First, AAC analysis revealed a significant reduction in AAC for s2 single mutants, while s3 and s23 mutants exhibited minimal change (Fig. 4A). In the case of GT, the To, Tp and Te values of all mutant grains were significantly decreased (Fig. 4B and Table S3). However, the decrease in GT in s3 mutants was greater than that in s2 mutants. Interestingly, the s23 double mutants displayed the lowest grain GT values among the mutants, suggesting an additive effect for SSII-2 and SSII-3 on grain GT. Regarding starch viscosity properties, the RVA profiles of s2 mutants showed an overall downward trend compared with those of WT samples, reflected by the lower PKV, HPV, BDV, CPV, SBV and PaT values. The RVA profiles of s3 mutants also displayed a decrease in PKV, BDV, CPV and PaT, but HPV and SBV were increased (Fig. 4C). Moreover, the overall RVA profile of s23 double mutants were lower than those of s2 and s3 single mutants, and similar to those of SSII-2 RNAi lines (Fig. 1D).
Physicochemical properties and starch fine structure of mature seeds from s2, s3 and s23 mutants. A Apparent amylose content (AAC) of rice flours. B Peak gelatinisation temperature (GT) of rice flours determined by DSC. C Rapid visco-analyser (RVA) profile of rice flours. D Fine structures of starches determined by gel permeation chromatography (GPC). E Changes in chain length distribution of amylopectin determined by fluorophore-assisted carbohydrate electrophoresis (FACE). Different lower-case letters indicate statistically significant differences at p < 0.05
The GPC results showed that the AC of s2 mutants was significantly decreased, while that of s3 and s23 mutants was not altered (Fig. 4D). Regarding amylopectin, only Ap1 of s2 mutants was significantly increased, while Ap1 of s3 and s23 mutants and Ap2 of all mutants were not changed (Table S5). Furthermore, data from FACE analysis indicated that the absence of SSII-2 alone caused a minor change in amylopectin fine structure, including a slight increase in the proportion of chains with DP 10−12 and > 34, and a decrease in chains with DP 5−9 and 13−33 (Fig. 4E). The chain-length distributions of amylopectin from s3 and s23 mutants were similar, both showing a significant increase in DP 5−12 and a significant decrease in DP 13−24 chains (Fig. 4E). Compared with SSII-2 RNAi lines, the amylopectin structure of s2 grains showed a consistent change in chains with DP 13−24, while that of s3 and s23 grains showed a consistent change in chains with DP 7−11.
Taken together, the above results revealed that mutating SSII-2 can indeed decrease rice AC and GT, albeit to a lesser extent than that caused by SSII-2 RNAi. Meanwhile, ssii-3 single mutants and ssii-2ssii-3 double mutants exhibited similar amylopectin CLD and GT levels to those in SSII-2 RNAi rice, but there was no change in AC. In conclusion, SSII-2 is a promising biotechnological target for improving rice ECQ via either RNAi or CRISPR/Cas9 strategies during rice breeding programs.
Discussion
Synergetic downregulation of Wx, SSII-2 and SSII-3 expression in the endosperm of SSII-2 RNAi lines
Three NILs within the Wx locus were generated in the Nipponbare background, among which non-waxy Nip(Wxa) and Nip(Wxb) rice contained different amounts of mature Wx mRNA and GBSSI, while no mature Wx mRNA or GBSSI protein were detected in the glutinous Nip(wx) rice. The SSII-2 RNAi construct was subsequently introduced into the three NILs, and qRT-PCR and western blotting results indicated that expression of the Wx gene was markedly lower in the resulting SSII-2 RNAi lines. It is intriguing how suppression of SSII-2 could trigger such a striking decrease in expression of the Wx gene. Gene sequence analysis indicated that the SSII-2 fragment used for RNAi shared low sequence identity with the Wx gene, with a maximum of only 9 consecutive identical bases. Additionally, the Wx gene was not in the list of potential off-target genes predicted by dsCheck (http://dsCheck.RNAi.jp/) (Naito et al. 2005), suggesting that the dramatic decrease in Wx expression was not caused by off-target effects (Supplemental Table 6). To further confirm this notion, we analysed the expression of Wx in ssii-2 mutants, which were generated by the CRISPR/Cas9 editing system. The results showed that both the expression of Wx transcripts and GBSSI proteins were significantly reduced in the mutants, suggesting that the decrease in Wx expression was indeed caused by SSII-2 RNAi. Therefore, we speculate that either knockdown or knockout of SSII-2 expression could decrease Wx expression, and consequently AC. Note that the reduction in Wx transcripts and GBSSI proteins in ssii-2 mutants was lower than in SSII-2 RNAi lines (Figs. 2B, C and 3D, E), indicating that other factors might be involved in the grain quality formation in SSII-2 RNAi transgenic lines. For SSII-3 gene, mutation of SSII-3, the closest isoform of SSII-2, caused slightly increased expression of Wx transcripts but no remarkable change in GBSSI protein accumulation (Fig. 3 C and E).
Moreover, expression levels of both SSII-2 and SSII-3 were dramatically decreased in SSII-2 RNAi transgenic lines, but SSII-1 expression was not altered, suggesting that SSII-2 and SSII-3 both contributed to the improved grain quality of SSII-2 RNAi transgenic rice. Mutation of either SSII-2 or SSII-3 alone via CRISPR/Cas9 could lead to a decrease in expression of the other, indicating interdependency and reciprocal regulation of these two SSII isoforms.
Simultaneous modulation of amylose content and amylopectin structure in SSII-2 RNAi lines
Analysis of grain physicochemical properties revealed that both AC and amylopectin fine structure were improved in SSII-2 RNAi/Nip(Wxa) and SSII-2 RNAi/Nip(Wxb) transgenic rice, resulting in increased GC, decreased GT and RVA profile, and hence enhanced rice ECQ. Meanwhile in SSII-2 RNAi/Nip(wx) rice, only amylopectin synthesis was affected, also leading to decreased GT and improved stickiness, hardness and cooking properties.
The absence of SSII-2 resulted in minor changes in amylopectin CLD, consistent with its low expression in rice endosperm (Ohdan et al. 2005a), and this distinguished it from SSII-3 knockout lines. The proportion of amylopectin with DP 5−8 was decreased while the portion with DP 9−12 was increased in short chains, and the proportion with DP 13−30 was decreased while the proportion with DP ≥ 31 was increased in intermediate chains for ssii-2 mutants, compared with WT (Figs. 4E and S5). We speculate that the increase in amylopectin A chain with DP 9−12 and the decrease in amylopectin A chain with DP 5−8 and B chain with DP 13−30 together lowered GT in ssii-2 rice. Previous studies indicate that short-chain amylopectin with DP ≤ 12 can only form short double helices, whereas long parallel chains of amylopectin can form longer double helices (Miura et al. 2018).
Both ssii-3 and ssii-2ssii-3 mutants displayed similar GT and CLD for amylopectin to those of SSII-2 RNAi lines, but for RVA profiles, a consistent shape was only observed for ssii-2ssii-3 double mutants and SSII-2 RNAi lines, suggesting that both SSII-2 and SSII-3 were responsible for improving ECQ in SSII-2 RNAi rice. Thus, the formation of grain quality in SSII-2 RNAi lines and ssii-2 mutants was likely caused by a moderate decrease in Am and Ap-B1, and an increase in Ap-A via coordinated downregulation of the expression of Wx, SSII-2 and SSII-3 (Fig. 5). We speculated that the downregulation of SSII-3 expression was not only caused by suppressing of SSII-2, but also the possible off-target effect of RNAi to SSII-3 expression, even though SSII-3 was not in the list of potential off-target genes predicted by dsCheck (Supplemental Table 6).
Proposed mechanism by which SSII-2, SSII-3 and Wx regulate rice grain quality. Either knockdown or knockout of SSII-2 may suppress the expression of SSII-2, SSII-3 and Wx, resulting in decreased amylose (Am) and amylopectin B1 chains (Ap-B1), and increased amylopectin A chains (Ap-A), and a consequent improvement in rice ECQ
By specific knockout of SSII-2 and SSII-3 using CRISPR/Cas9, the ssii-2ssii-3 (s23) mutants were created trying to reproduce the moderate AC reduction as in SSII-2 RNAi lines. But the GBSSI accumulation as well as the AC showed no significant change in ssii-2ssii-3 mutants. Enzyme complexes were observed during amylopectin synthesis and SSII-3 was reported to form complexes with SSI and SBEs in rice, maize and wheat (Crofts et al. 2015; Hennen-Bierwagen et al. 2008; Tetlow et al. 2008). SSII-3 is the core of the SSI/SSII-3/SBEII (SBE3/SBEIIb or SBE4/SBEIIa) trimer, located between SSI and SBEII. In addition to extending the chain length of amylopectin, SSII-3 also transports SSI and SBEII to starch granules (Liu et al. 2012). The ssii-3 mutant in maize, typically known as sugary 2 (su2), showed decreased SSI and SSIIb proteins on starch granules. Besides, new enzymes complexes were observed after mutation of one component of the original enzyme complex (Liu et al. 2011). In present study, the expression of SSII-3 was knockdown in SSII-2 RNAi lines, while the SSII-3 protein was fully absence in ssii-3 mutants. This could be a possible reason for the AC difference between these lines. By using the genome editing system, the AC reduction in ssii-2ssii-3 mutants can be further achieved by application of natural Wx allelic variants, as well as the novel edited Wx alleles (Huang et al. 2020a; Xu et al. 2020a; Zeng et al. 2020; Zhang et al. 2019a).
Moreover, the SSII-3 was successfully knockout in ssii-3 mutants resulting from the CLD and dramatically decreased GT, while slight AC changes were observed in ssii-3 mutants created by CRISPR/Cas9. It distinguishes from the increased AC in ssii-3 mutants in previous studies (Chen et al. 2020; Miura et al. 2018). The reason need to be further explained.
SSII-2 is a novel biotechnological target for improving rice ECQ
The present work revealed that the SSII-2 gene is involved in the synthesis of both rice amylose and amylopectin by modulating the expression of Wx and SSII-3 genes. SSII-2 knockdown or knockout rice displayed decreased AC and GT, but there were almost no negative effects on other agronomic traits, including grain appearance, which may be beneficial for high-quality rice breeding applications. In the past few decades, tremendous effort has been expended to improve rice grain quality without lowering the yield, and regulation of rice starch biosynthesis appears to be a promising approach. However, most previous studies focused on modulating SSRGs with high or seed-specific expression, including Wx, SSI, SSII-3, SSIII-2, SBE1, SBE3, ISA1 and Pull (Jeon et al. 2010; Nishi et al. 2001; Zhou et al. 2016; Zhu et al. 2012). Other SSRGs with low expression abundance, including SSII-2, have received less attention, and their functions typically remain unknown. For example, SSII-2 is believed to be involved in the biosynthesis of transient starch in plant leaves, based on its expression pattern in maize and rice (Harn et al. 1998; Jiang et al. 2004). Various studies on rice ECQ improvement have successfully identified regulators, including transcription factors and starch/enzyme-binding proteins such as RSR1, OsbZIP58, OsMADS7, and the NF-YB1-YC12-bHLH144 complex (Bello et al. 2019; Fu and Xue 2010; Wang et al. 2013; Zhang et al. 2018), as well as PTST1, PTST2 (flo6) and OsGBP (Peng et al. 2014; Seung et al. 2015; Wang et al. 2019). These transcription factors mainly affect AC, GC and GT in rice endosperm, but accompanying defects in rice yield or grain appearance greatly limits their application in rice breeding programs. At present, application of natural allelic variation in Wx and SSII-3 remains the major approach for improving rice ECQ (Phing Lau et al. 2016). Excitingly, our current study suggests that SSII-2, a neglected SSII isoform, could be a novel beneficial target for improving rice ECQ during breeding programs.
Conclusions
In summary, our study revealed the underlying mechanism of SSII-2 RNAi-mediated improvements in rice ECQ, and highlighted the key role of the coordinated regulation of amylose and amylopectin synthesis via suppression of the expression of SSII-2, SSII-3 and Wx. Furthermore, we also uncovered the function of SSII-2 in the regulation of both amylose and amylopectin synthesis by studying ssii-2 mutants. The findings provide new strategies and germplasm resources for breeding high-ECQ rice.
Data availability
All data supporting the findings of this study are available within the paper and within its supplementary materials published online.
References
Anacleto R, Badoni S, Parween S, Butardo VM Jr., Misra G, Cuevas RP et al (2019) Integrating a genome-wide association study with a large-scale transcriptome analysis to predict genetic regions influencing the glycaemic index and texture in rice. Plant Biotechnol J 17:1261–1275
Bao JS, Corke H, Sun M (2006) Nucleotide diversity in starch synthase IIa and validation of single nucleotide polymorphisms in relation to starch gelatinization temperature and other physicochemical properties in rice (Oryza sativa L.). Theor Appl Genet 113:1171–1183
Bello BK, Hou Y, Zhao J, Jiao G, Wu Y, Li Z et al (2019) NF-YB1-YC12-bHLH144 complex directly activates Wx to regulate grain quality in rice (Oryza sativa L.). Plant Biotechnol J 17:1222–1235
Cai X, Wang Z, Xing Y, Zhang J, Hong M (1998) Aberrant splicing of intron 1 leads to the heterogeneous 5′ UTR and decreased expression of Waxy gene in rice cultivars of intermediate amylose content. Plant J 14:459–465
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697
Chen Z, Lu Y, Feng L, Hao W, Li C, Yang Y et al (2020) Genetic dissection and functional differentiation of ALKa and ALKb, two natural alleles of the ALK/SSIIa gene, responding to low gelatinization temperature in rice. Rice 13:39
Commuri PD, Keeling PL (2001) Chain-length specificities of maize starch synthase I enzyme: studies of glucan affinity and catalytic properties. Plant J 25:475–486
Crofts N, Abe N, Oitome NF, Matsushima R, Hayashi M, Tetlow IJ et al (2015) Amylopectin biosynthetic enzymes from developing rice seed form enzymatically active protein complexes. J Exp Bot 66:4469–4482
Donohoue PD, Barrangou R, May AP (2018) Advances in industrial biotechnology using CRISPR-Cas systems. Trends Biotechnol 36:134–146
Fan X, Li Y, Lu Y, Zhang C, Li E, Li Q et al (2019) The interaction between amylose and amylopectin synthesis in rice endosperm grown at high temperature. Food Chem 301:125258
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Fu FF, Xue HW (2010) Coexpression analysis identifies rice starch regulator1, a rice AP2/EREBP family transcription factor, as a novel rice starch biosynthesis regulator. Plant Physiol 154:927–938
Fujita N, Yoshida M, Asakura N, Ohdan T, Miyao A, Hirochika H, Nakamura Y (2006) Function and characterization of starch synthase I using mutants in rice. Plant Physiol 140:1070–1084
Furukawa K, Tagaya M, Inouye M, Preiss J, Fukui T (1990) Identification of lysine 15 at the active site in Escherichia coli glycogen synthase. Conservation of Lys-X-Gly-Gly sequence in the bacterial and mammalian enzymes. J Biol Chem 265:2086–2090
Gao Z, Zeng D, Cui X, Zhou Y, Yan M, Huang D et al (2003) Map-based cloning of the ALK gene, which controls the gelatinization temperature of rice. Sci China C Life Sci 46:661–668
Gu F, Gong B, Gilbert RG, Yu W, Li E, Li C (2019) Relations between changes in starch molecular fine structure and in thermal properties during rice grain storage. Food Chem 295:484–492
Harn C, Knight M, Ramakrishnan A, Guan H, Keeling PL, Wasserman BP (1998) Isolation and characterization of the zSSIIa and zSSIIb starch synthase cDNA clones from maize endosperm. Plant Mol Biol 37:639–649
Hennen-Bierwagen TA, Liu F, Marsh RS, Kim S, Gan Q, Tetlow IJ et al (2008) Starch biosynthetic enzymes from developing maize endosperm associate in multisubunit complexes. Plant Physiol 146:1892–1908
Hirano HY, Eiguchi M, Sano Y (1998) A single base change altered the regulation of the Waxy gene at the posttranscriptional level during the domestication of rice. Mol Biol Evol 15:978–987
Huang L, Li Q, Zhang C, Chu R, Gu Z, Tan H et al (2020a) Creating novel Wx alleles with fine-tuned amylose levels and improved grain quality in rice by promoter editing using CRISPR/Cas9 system. Plant Biotechnol J 18:2164–2166
Huang L, Sreenivasulu N, Liu Q (2020b) Waxy editing: old meets new. Trends Plant Sci 25:963–966
Jeon JS, Ryoo N, Hahn TR, Walia H, Nakamura Y (2010) Starch biosynthesis in cereal endosperm. Plant Physiol Biochem 48:383–392
Jiang H, Dian W, Liu F, Wu P (2004) Molecular cloning and expression analysis of three genes encoding starch synthase II in rice. Planta 218:1062–1070
Li Z, Chu X, Mouille G, Yan L, Kosar-Hashemi B, Hey S et al (1999) The localization and expression of the class II starch synthases of wheat. Plant Physiol 120:1147–1156
Li H, Prakash S, Nicholson TM, Fitzgerald MA, Gilbert RG (2016) Instrumental measurement of cooked rice texture by dynamic rheological testing and its relation to the fine structure of rice starch. Carbohydr Polym 146:253–263
Li QF, Huang LC, Chu R, Li J, Jiang MY, Zhang CQ et al (2018) Down-regulation of SSSII-2 gene expression results in novel low-amylose rice with soft, transparent grains. J Agric Food Chem 66:9750–9760
Li C, Wu A, Yu W, Hu Y, Li E, Zhang C, Liu Q (2020) Parameterizing starch chain-length distributions for structure-property relations. Carbohydr Polym 241:116390
Liu L, Ma X, Liu S, Zhu C, Jiang L, Wang Y et al (2009) Identification and characterization of a novel Waxy allele from a Yunnan rice landrace. Plant Mol Biol 71:609–626
Liu F, Ahmed Z, Lee EA, Donner E, Liu Q, Ahmed R et al (2011) Allelic variants of the amylose extender mutation of maize demonstrate phenotypic variation in starch structure resulting from modified protein–protein interactions. J Exp Bot 63:1167–1183
Liu F, Romanova N, Lee Elizabeth A, Ahmed R, Evans M, Gilbert Elliot P et al (2012) Glucan affinity of starch synthase IIa determines binding of starch synthase I and starch-branching enzyme IIb to starch granules. Biochem J 448:373–387
Liu W, Xie X, Ma X, Li J, Chen J, Liu Y (2015) DSDecode: a web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations. Mol Plant 8:1431–1433
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X et al (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23:1233–1236
Mikami I, Aikawa M, Hirano HY, Sano Y (1999) Altered tissue-specific expression at the Wx gene of the opaque mutants in rice. Euphytica 105:91–97
Mikami I, Uwatoko N, Ikeda Y, Yamaguchi J, Hirano HY, Suzuki Y, Sano Y (2008) Allelic diversification at the Wx locus in landraces of Asian rice. Theor Appl Genet 116:979–989
Miura S, Crofts N, Saito Y, Hosaka Y, Oitome NF, Watanabe T et al (2018) Starch synthase IIa-deficient mutant rice line produces endosperm starch with lower gelatinization temperature than japonica rice cultivars. Front Plant Sci 9:645–645
Morell MK, Kosar-Hashemi B, Cmiel M, Samuel MS, Chandler P, Rahman S et al (2003) Barley sex6 mutants lack starch synthase IIa activity and contain a starch with novel properties. Plant J 34:173–185
Naito Y, Yamada T, Matsumiya T, Ui-Tei K, Saigo K, Morishita S (2005) dsCheck: highly sensitive off-target search software for double-stranded RNA-mediated RNA interference. Nucleic Acids Res 33:W589–W591
Nakamura Y, Francisco PB, Hosaka Y, Sato A, Sawada T, Kubo A, Fujita N (2005) Essential amino acids of starch synthase IIa differentiate amylopectin structure and starch quality between japonica and indica rice varieties. Plant Mol Biol 58:213–227
Nishi A, Nakamura Y, Tanaka N, Satoh H (2001) Biochemical and genetic analysis of the effects of amylose-extender mutation in rice endosperm. Plant Physiol 127:459–472
Ohdan T, Francisco PB Jr, Sawada T, Hirose T, Terao T, Satoh H, Nakamura Y (2005a) Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J Exp Bot 56:3229–3244
Ohdan T, Francisco PJT, Hirose T, Terao T, Satoh H, Nakamura Y (2005b) Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J Exp Bot 56:3229–3244
Peng C, Wang Y, Liu F, Ren Y, Zhou K, Lv J et al (2014) FLOURY ENDOSPERM6 encodes a CBM48 domain-containing protein involved in compound granule formation and starch synthesis in rice endosperm. Plant J 77:917–930
Phing Lau WC, Latif MA, Rafii MY, Ismail MR, Puteh A (2016) Advances to improve the eating and cooking qualities of rice by marker-assisted breeding. Crit Rev Biotechnol 36:87–98
Sano Y (1984) Differential regulation of Waxy gene expression in rice endosperm. Theor Appl Genet 68:467–473
Sato H, Suzuki Y, Sakai M, Imbe T (2002) Molecular characterization of Wx-mq, a novel mutant gene for low-amylose content in endosperm of rice (Oryza sativa L.). Breed Sci 52:131–135
Seung D, Soyk S, Coiro M, Maier BA, Eicke S, Zeeman SC (2015) PROTEIN TARGETING TO STARCH is required for localising GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis. PLoS Biol 13:e1002080
Su Y, Rao Y, Hu S, Yang Y, Gao Z, Zhang G et al (2011) Map-based cloning proves qGC-6, a major QTL for gel consistency of japonica/indica cross, responds by Waxy in rice (Oryza sativa L.). Theor Appl Genet 123:859
Tan Y, Yu S, Xing Y, Xu C, Zhang Q, Li J (1999) The three important traits for cooking and eating quality of rice grains are controlled by a single locus in an elite rice hybrid, Shanyou 63. Theor Appl Genet 99:642–648
Tetlow IJ, Beisel KG, Cameron S, Makhmoudova A, Liu F, Bresolin NS et al (2008) Analysis of protein complexes in wheat amyloplasts reveals functional interactions among starch biosynthetic enzymes. Plant Physiol 146:1878–1891
Tian Z, Qian Q, Liu Q, Yan M, Liu X, Yan C et al (2009) Allelic diversities in rice starch biosynthesis lead to a diverse array of rice eating and cooking qualities. Proc Natl Acad Sci USA 106:21760–21765
Umemoto T, Yano M, Satoh H, Shomura A, Nakamura Y (2002) Mapping of a gene responsible for the difference in amylopectin structure between japonica-type and indica-type rice varieties. Theor Appl Genet 104:1–8
Wang L, Wang YJ (2001) Comparison of protease digestion at neutral pH with alkaline steeping method for rice starch isolation. Cereal Chem 78:690–692
Wang Z, Wu Z, Xing Y, Zheng F, Guo X, Zhang W, Hong M (1990) Nucleotide sequence of rice Waxy gene. Nucleic Acids Res 18:5898–5898
Wang JC, Xu H, Zhu Y, Liu QQ, Cai XL (2013) OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. J Exp Bot 64:3453–3466
Wang C, Shen L, Fu Y, Yan C, Wang K (2015) A simple CRISPR/Cas9 system for multiplex genome editing in rice. J Genet Genom 42:703–706
Wang X, Pang Y, Wang C, Chen K, Zhu Y, Shen C et al (2016) New candidate genes affecting rice grain appearance and milling quality detected by genome-wide and gene-based association analyses. Front Plant Sci 7:1998
Wang W, Wei X, Jiao G, Chen W, Wu Y, Sheng Z et al (2019) GBSS-BINDING PROTEIN, encoding a CBM48 domain-containing protein, affects rice quality and yield. J Integr Plant Biol 62:948–966
Xie X, Ma X, Zhu Q, Zeng D, Li G, Liu Y (2017) CRISPR-GE: a convenient software toolkit for CRISPR-based genome editing. Mol Plant 10:1246–1249
Xu Y, Lin Q, Li X, Wang F, Chen Z, Wang J et al (2020a) Fine-tuning the amylose content of rice by precise base editing of the Wx gene. Plant Biotechnol J 19:11–13
Xu Z, Yu M, Yin Y, Zhu C, Ji W, Zhang C et al (2020b) Generation of selectable marker-free soft transgenic rice with transparent kernels by downregulation of SSSII-2. Crop J 8:53–61
Yang Y, Guo M, Sun S, Zou Y, Yin S, Liu Y et al (2019) Natural variation of OsGluA2 is involved in grain protein content regulation in rice. Nature Commun 10:1949
Zeng D, Liu T, Ma X, Wang B, Zheng Z, Zhang Y et al (2020) Quantitative regulation of Waxy expression by CRISPR/Cas9-based promoter and 5’UTR-intron editing improves grain quality in rice. Plant Biotechnol J 18:2385–2387
Zhang C, Zhu L, Shao K, Gu M, Liu Q (2013) Toward underlying reasons for rice starches having low viscosity and high amylose: physiochemical and structural characteristics. J Sci Food Agric 93:1543–1551
Zhang H, Zhang J, Wei P, Zhang B, Gou F, Feng Z et al (2014) The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol J 12:797–807
Zhang C, Zhou L, Zhu Z, Lu H, Zhou X, Qian Y et al (2016) Characterization of grain quality and starch fine structure of two japonica rice (Oryza Sativa) cultivars with good sensory properties at different temperatures during the filling stage. J Agric Food Chem 64:4048–4057
Zhang H, Xu H, Feng M, Zhu Y (2018) Suppression of OsMADS7 in rice endosperm stabilizes amylose content under high temperature stress. Plant Biotechnol J 16:18–26
Zhang C, Zhu J, Chen S, Fan X, Li Q, Lu Y et al (2019a) Wxlv, the ancestral allele of rice Waxy gene. Mol Plant 12:1157–1166
Zhang H, Zhou L, Xu H, Wang L, Liu H, Zhang C et al (2019b) The qSAC3 locus from indica rice effectively increases amylose content under a variety of conditions. BMC Plant Biol 19:275
Zhao DS, Li QF, Zhang CQ, Zhang C, Yang QQ, Pan LX et al (2018) GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat Commun 9:1240
Zhou H, Wang L, Liu G, Meng X, Jing Y, Shu X et al (2016) Critical roles of soluble starch synthase SSIIIa and granule-bound starch synthase Waxy in synthesizing resistant starch in rice. Proc Natl Acad Sci USA 113:12844–12849
Zhou H, Xia D, He Y (2020) Rice grain quality—traditional traits for high quality rice and health-plus substances. Mol Breed 40:1
Zhu F (2018) Relationships between amylopectin internal molecular structure and physicochemical properties of starch. Trends Food Sci Technol 78:234–242
Zhu L, Gu M, Meng X, Cheung SC, Yu H, Huang J et al (2012) High-amylose rice improves indices of animal health in normal and diabetic rats. Plant Biotechnol J 10:353–362
Zhu Q, Zeng D, Yu S, Cui C, Li J, Li H et al (2018) From golden rice to aSTARice: bioengineering astaxanthin biosynthesis in rice endosperm. Mol Plant 11:1440–1448
Acknowledgements
This work was supported by National Natural Science Foundation of China (31825019 and 31901517), the Ministry of Agriculture of China (2016ZX08001006-005), the programs from Jiangsu Province Government (CX(20)3004 and PAPD), and Young Elite Scientists Sponsorship Program by China Association for Science and Technology (2018QNRC001).
Author information
Authors and Affiliations
Contributions
QQL, QFL and HL conceived the project; LH, ZG, ZC, JY, RC, HT, DZ and XF performed the experiments and analysed the data; LH, QQL and QFL wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, L., Gu, Z., Chen, Z. et al. Improving rice eating and cooking quality by coordinated expression of the major starch synthesis-related genes, SSII and Wx, in endosperm. Plant Mol Biol 106, 419–432 (2021). https://doi.org/10.1007/s11103-021-01162-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-021-01162-8