Abstract
Sumoylation is an essential post-translational regulator of plant development and the response to environmental stimuli. SUMO conjugation occurs via an E1-E2-E3 cascade, and can be removed by SUMO proteases (ULPs). ULPs are numerous and likely to function as sources of specificity within the pathway, yet most ULPs remain functionally unresolved. In this report we used loss-of-function reverse genetics and transcriptomics to functionally characterize Arabidopsis thaliana ULP1c and ULP1d SUMO proteases. GUS reporter assays implicated ULP1c/d in various developmental stages, and subsequent defects in growth and germination were uncovered using loss-of-function mutants. Microarray analysis evidenced not only a deregulation of genes involved in development, but also in genes controlled by various drought-associated transcriptional regulators. We demonstrated that ulp1c ulp1d displayed diminished in vitro root growth under low water potential and higher stomatal aperture, yet leaf transpirational water loss and whole drought tolerance were not significantly altered. Generation of a triple siz1 ulp1c ulp1d mutant suggests that ULP1c/d and the SUMO E3 ligase SIZ1 may display separate functions in development yet operate epistatically in response to water deficit. We provide experimental evidence that Arabidopsis ULP1c and ULP1d proteases act redundantly as positive regulators of growth, and operate mainly as isopeptidases downstream of SIZ1 in the control of water deficit responses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To cope with a changing environment, plants developed a number of molecular, biochemical and morphological strategies. Tactics to specifically overcome low water availability include the control of stomata opening, changes in root morphology, modulation of photosynthesis, cell wall modification and the accumulation of osmotically compatible metabolites (Aroca et al. 2012; Setter 2012). To implement these strategies, plants carry out physiological adjustments and gene expression reprogramming that partially require phytohormone signalling circuits (Golldack et al. 2014; Kilian et al. 2012). The most important stress hormone is abscisic acid (ABA), a key regulator of dehydration avoidance and drought tolerance, controlling the biosynthesis of protective components, stomata movement, and seed maturation and germination (Cutler et al. 2010; Raghavendra et al. 2010; Sreenivasulu et al. 2012).
Post-translational modifications (PTMs) are essential regulators of plant stress responses, rapidly modulating protein function. Among PTMs, modification by ubiquitin and ubiquitin-like small peptides (UBLs) has been found to be essential to the control of key components in abiotic stress responses (Lyzenga and Stone 2012; Miura and Hasegawa 2010). Small Ubiquitin-like Modifier (SUMO) is a UBL that gained importance over the past decade, after functional studies implicated this modifier in the fast and reversible modulation of protein activity, without the necessity for degradation or de novo synthesis. SUMO may exert different effects depending on the target protein, controlling its conformation, or creating or blocking interacting interfaces (Wilkinson and Henley 2010). Generally, SUMO conjugates accumulate drastically during stress, a feature that seems characteristic of all eukaryotes (Kurepa et al. 2003; Lallemand-Breitenbach et al. 2008; Zhou et al. 2004). In plants, SUMO conjugation has been associated with extreme temperatures, drought and salinity tolerance, oxidative stress modulation, control of nutritional homeostasis, and control of sugar signalling and germination time (Castro et al. 2012, 2015). Many of these responses involve the regulation of salicylic acid (SA), ABA and auxin hormones (Miura et al. 2009, 2010, 2011). SUMO modulation of cellular processes occurs primarily at the nuclear level, as SUMO pathway components and most known SUMO targets are nuclear-located (Budhiraja et al. 2009; Miller et al. 2010). Sumoylation is normally considered to have a repressor effect on transcription, targeting key regulators of nuclear mechanisms, such as transcription factors (TFs) and chromatin remodelling components (Garcia-Dominguez and Reyes 2009; van den Burg and Takken 2009).
A cyclic pathway mediates the conjugation/deconjugation of SUMO to target proteins. Pre-SUMO peptides are initially maturated by SUMO proteases of the ULP/SENP type (Ubiquitin-Like Proteases/SENtrin-specific Proteases), which display endopeptidase activity, cleaving the C-terminal end of the pre-SUMO to expose a di-glycine motif. Sumoylation, the covalent attachment of SUMO to a target, requires the sequential activity of three enzymes, E1, E2, and E3 (Gareau and Lima 2010). In vivo, sumoylation is greatly enhanced by SUMO E3 ligases that aid in the reaction, and promote specificity (Gareau and Lima 2010). The SUMO modifier can be removed from the target by ULPs presenting isopeptidase activity, allowing SUMO to re-enter the conjugation pathway. SUMO seems to be essential for plant development, as disruption of SUMO conjugation components, namely SAE2, SCE1 and the SUM1/SUM2 modifiers, results in developmental arrest at the early stages of embryogenesis (Saracco et al. 2007). Mutants for the SUMO E3 ligases SIZ1 and HPY2/MMS21 display pleiotropic phenotypes, while their double mutant is lethal (Catala et al. 2007; Huang et al. 2009; Ishida et al. 2009, 2012; Jin et al. 2008; Miura et al. 2010). Recently, E4-type SUMO ligases have been identified and implicated in SUMO chain formation (Tomanov et al. 2014). In contrast to the low number of SUMO-conjugating components, Arabidopsis thaliana ULPs comprise a family of at least seven elements (ESD4, ULP1a/ELS1, ULP1b, ULP1c/OTS2, ULP1d/OTS1, ULP2a and ULP2b), which may result in both specificity and redundancy within the SUMO pathway (Chosed et al. 2006; Colby et al. 2006; Lois 2010). ESD4 and ULP1a were previously associated with the control of flowering time and plant development (Hermkes et al. 2011; Murtas et al. 2003). ULP1c and ULP1d (herein ULP1c/d) have been implicated in salt stress responses, in DELLA-dependent regulation of growth, in modulation of SA signalling and the desumoylation of phytochrome-B (Bailey et al. 2016; Conti et al. 2008, 2014; Sadanandom et al. 2015).
In the present work, we demonstrate that ULP1c/d are highly expressed and display unequal redundancy in the control of important developmental traits, including rosette growth and seed germination. Genome-wide transcriptome analysis of ulp1c/d indicates that a surprisingly large set of differentially expressed genes (DEGs) are associated with drought and ABA responses. These results led us to uncover that ULP1c and ULP1d act as modulators of osmotic stress, and control stomatal aperture independently of stress-related stimuli. We genetically demonstrate that ULP1c and ULP1d are hypostatic to the E3 ligase SIZ1 with regards to these stress responses, and propose that these ULPs control specific stress regulons, by mediating desumoylation of known SUMO targets such as ABI5 and ICE1.
Materials and methods
Plant material and growth conditions
T-DNA insertion mutants for A. thaliana ULP1c (At1g10570) and ULP1d (At1g60220) were identified using SIGnAL (signal.salk.edu), and consisted of SALK lines SALK_050441 (ulp1c-2), SALK_151423 (ulp1c-3), SALK_029340 (ulp1d-2) and SALK_065397 (siz1-2). Ecotype Columbia-0 (Col) was used as the wild-type. Genotypes were ordered through NASC (arabidopsis.info) and ABRC (www.biosci.ohio-state) seed stock centres. Homozygous insertion mutants were genotyped based on SIGnAL T-DNA Primer Design (signal.salk.edu/tdnaprimers.2.html), using the primers in Table S1.
Synchronized seeds were stratified as previously reported (Silva-Correia et al. 2014). Seeds were sown onto 1.2 % agar-solidified MS medium containing 1.5 % sucrose, 0.5 g L−1 MES, pH 5.7, and grown vertically in culture rooms with a 16/8 h light/dark cycle under cool white light (80 µE m−2 s−1) at 23 °C. For standard growth, 7-day-old in vitro-grown seedlings were transferred to a 4:1 soil to vermiculite mixture, and maintained under identical growth conditions, with regular watering. Developmental characterization of the mutants was based on the developmental map of Boyes et al. (2001). Morphological traits were measured using ImageJ (imagej.nih.gov/ij/). For germination assays, seeds were sterilized as previously detailed, sown onto 0.8 % agar MS medium and grown horizontally under identical conditions. Each replica plate contained >30 seeds per genotype. Germination was scored as the emergence of two green cotyledon leaves, over time.
GUS histochemical staining of transgenic Arabidopsis (Col background) plants containing proULP1c::GUS and proULP1d::GUS constructs was performed as previously described (Posé et al. 2009) (Methods S1).
Osmotic stress and ABA-related experiments
For rapid dehydration, the rosette of 4-week-old soil-grown plants was detached from roots and air-dried at room temperature. Fresh weight was measured at different time points. When required, seedlings were grown in vitro for 7 days, and subsequently transferred to 0.5× MS 1.2 % agar plates. Plates were supplemented with either 0, 5, 10, 20 or 30 µM ABA, 100 mM NaCl, 200 mM mannitol or PEG-infusion (Methods S2). Vertical root growth was measured every 2 days for up to 10 days. Mesophyll and stomata cell morphology was analysed as previously detailed (Miura et al. 2010). Quantification of real-time stomatal aperture was adapted from Chitrakar and Melotto (2010) (Methods S3), and performed in 4-week-old plants or 2-week-old seedlings. Quantification and ABA inhibition of stomatal opening was performed on isolated epidermal strips from rosette leaves of 4-week-old plants, as previously described (Lozano-Duran et al. 2011) (Methods S4). Stomata size, aperture and density were measured using ImageJ. ABA and mannitol germination assays were performed as previously detailed for germination, using medium supplemented with 1 µM ABA, or 400 mM mannitol.
Plasmid construction and plant transformation
Plasmids were constructed using standard DNA cloning techniques, and confirmed by DNA sequencing. For promoter::GUS constructs, ULP1c and ULP1d promoter regions were selected as the intergenic region (ULP1c) or the −2000 bp span (ULPd) encompassing significant cis-regulatory elements as detected by AthaMap (www.athamap.de/; Steffens et al. 2004). Regions were amplified by PCR from Arabidopsis genomic DNA (Edwards et al. 1991). Incorporated restriction sites (EcoRI and NcoI) were used to clone fragments into the pCAMBIA 1303 vector (www.cambia.org/daisy/cambia/585). For complementation purposes, the ULP1d open reading frame was amplified from cDNA by PCR, with incorporated restriction sites EcoRI and ClaI. The amplification product was sub-cloned into the pGEM-T Easy Vector (Promega) and subsequently cloned into the pHANNIBAL vector (Wesley et al. 2001) to create a pro35S::ULP1d-NOS terminator fusion. The construct was excised using NotI, and subsequently cloned into the plant expression vector pGREEN II 0229 (www.pgreen.ac.uk). Agrobacterium tumefaciens strain EHA105 was used for plant transformation. A. thaliana (Col ecotype) plants were transformed by the floral dip method (Clough and Bent 1998). A resistance marker (Kanamycin) selection strategy was employed to establish homozygous transformants.
Microarray analysis and quantitative RT-PCR
Genome-wide transcription studies were performed using the ATH1 microarray chip (Affymetrix; www.affymetrix.com) with three independent pools per genotype, each representing RNA from nine separate 5-week-old plants. Plants were grown in culture chambers, as previously stated. Three rosette leaves were sampled from each plant. RNA was extracted using a standard Trizol protocol (Invitrogen), including a treatment with Recombinant DNase I (Takara Biotechnology), followed by RNeasy Plant Mini Kit (QIAGEN) column cleaning. Microarray hybridisations and differential expression analysis were conducted at Unité de Recherche en Génomique Végétale (Université d’Evry Val d’Essonne, France), and data was deposited in ArrayExpress (www.ebi.ac.uk/arrayexpress). Gene Ontology (GO) term functional categorization was performed in VirtualPlant 1.2 (virtualplant.bio.nyu.edu/cgi-bin/vpweb), using the BioMaps function with a 0.05 P value cut-off (Katari et al. 2010). Redundancy exclusion and scatterplot analysis were performed using REVIGO (revigo.irb.hr/), with a 0.9 C-value. Venn diagrams were obtained using Venn Diagram Generator (www.pangloss.com/seidel/Protocols/venn.cgi).
For quantitative Real-Time PCR (qPCR) analysis, RNA was extracted from plant tissues using an RNeasy Plant Mini Kit (QIAGEN), and RNA quantity and quality were assessed using both a Nanodrop ND-1000 spectrophotometer, and standard agarose-gel electrophoretic analysis. The RNA samples were treated with Recombinant DNase I (Takara Biotechnology), and cDNA was subsequently generated using a Superscript II Reverse Transcriptase Kit (Invitrogen). Ssofast Evagreen Supermix (Bio-Rad) was used in the qPCR reaction mixture, as per the manufacturer’s indications. The reaction was performed in a Rotor Gene Q System (QIAGEN) or a MyIQ Single-Color Real-Time PCR Detection System (Bio-Rad). Primers for qPCR (Table S2) were designed using NCBI Primer-BLAST (www.ncbi.nlm.nih.gov/tools/primer-blast) (Ye et al. 2012). ACT2 (At3g18780) was used as the reference gene (Lozano-Duran et al. 2011).
Protein extraction and immunoblotting
For stress imposition, 10-day-old seedlings were submerged into 200 mM mannitol, 100 mM NaCl or water (control condition), and infiltration was promoted by applying vacuum for 3 × 5 min, at 50 kPa. Seedlings were then incubated for 3 h in the growth chamber under standard conditions, and subsequently snap frozen in liquid nitrogen. To induce rapid dehydration, 10-day-old seedlings were exposed to constant air flux in a horizontal flow chamber for 4 h. Plant tissue was grinded in a microtube in liquid nitrogen, with the help of polypropylene pestles. Protein extracts were obtained by adding extraction buffer (50 mM Tris; 150 mM NaCl; 0.2 % (v/v) Triton X-100) supplemented with Complete Protease Inhibitor Cocktail (Roche), as per the manufacturer’s instructions. Following incubation for 1 h at 4 °C with agitation, microtubes were centrifuged for 2 × 30 min at 16,000g. The supernatant was recovered and stored at −80 °C. Protein was spectrophotometrically quantified using Bradford Reagent (Sigma-Aldrich) (Bradford 1976). SUMO-conjugate levels were monitored by standard SDS-PAGE and Western-blot analysis (Methods S5).
In silico analysis
See Methods S6.
Results
ULP1c and ULP1d show a similar expression pattern
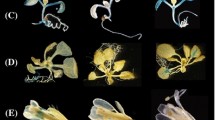
The fairly large number of Arabidopsis ULPs, and the high phylogenetic proximity of several family members, suggests the existence of various redundant gene pairs, one of which comprises Arabidopsis SUMO protease genes ULP1c/OTS2 (At1g10570) and ULP1d/OTS1 (At1g60220) (Chosed et al. 2006; Colby et al. 2006; Lois 2010). ULP1c/d have been implicated in salt stress responses as well as in DELLA-dependent regulation of growth (Conti et al. 2008, 2014), yet little is known regarding their additional involvement in plant development or other abiotic stress responses. To gain new insight on ULP1c/d function, we first established their spatial and developmental expression pattern using a GUS reporter system (Fig. 1). Analysis of several independent lines showed similar expression patterns between ULP1c and ULP1d. Both genes seemed to be expressed in early germination stages, while in plate-grown seedlings, expression could be detected in the entire leaves and cotyledons, with prevalence in vascular tissues and petioles. In roots, expression was restricted to the vascular tissue, with a slight signal increase in emerging lateral roots. In leaves of adult soil-grown plants, expression of both genes was reduced. For most tissues, expression was stronger for ULP1d. Conversely, in flowers and siliques, proULP1c::GUS lines showed stronger expression, although the pattern remained similar: staining was observed at the top and at the base of developing siliques, in the vascular tissues of petals and sepals, in the stamen filament and at the base of the stigma. The observed expression patterns were consistent with systematic gene expression maps of Arabidopsis development (Fig. S1), and both indicate the prevalent expression of ULP1d over ULP1c. Subsequent phylogenetic analysis placed ULP1c and ULP1d as a likely redundant pair, closer to the ULP2-type of SUMO proteases (Fig. S2a). Additional in silico analysis also demonstrated that both genes (1) are highly co-expressed during development and other experimental conditions, (2) are highly co-expressed within the Arabidopsis ULP gene family, and (3) are syntenically located within a large genomic block (Fig. S2). Collectively, the present data supports the existence of functional redundancy between ULP1c and ULP1d.
Expression profile of proULP1c::GUS and proULP1d::GUS by histochemical β-glucoronidase (GUS) staining. a, b Schematic representation of the ULP1c (a) and ULP1d (b) promoter regions, used for promoter::GUS fusions. c–e GUS staining during different stages of in vitro seed germination. f–h 10-day-old (f), 15-day-old (g) and 21-day-old (h) in vitro-grown shoots. i Cotyledons of 10-day-old seedlings. j, k 10-day-old root tips (j) and emerging lateral roots (k). l–o Expression in different adult-stage tissues: 5-week-old leaves (l), developing silique (m), flowers (n), and flowering structures (o). Scale bar indicates 1 mm unless stated
ULP1c and ULP1d are determinants of important developmental traits
ULP1c and ULP1d expression patterns suggested a potential involvement in various aspects of development. We investigated this hypothesis using reverse genetics, and previously unreported T-DNA lines for ULP1c (ulp1c-2; SALK_050441) and ULP1d (ulp1d-2; SALK_029340) (Fig. 2a, b). These lines are located upstream from earlier employed SALK lines ots1-1/ulp1d-1 and ots2-1/ulp1c-1 (Conti et al. 2008). To validate the efficiency of the present insertion mutants, gene knockout was confirmed by quantitative RT-PCR (qPCR) (Fig. 2c). The double mutant ulp1c-2 ulp1d-2 (herein ulp1c/d) also displayed the same late flowering and in vitro salt stress sensitivity phenotypes that were previously reported by Conti et al. (2008) (Figs. S3a, b; S4a). Subsequently, we performed a comprehensive morphological characterization of the single and double mutants (Fig. 2d–g). Morphology of 4-week-old plants is depicted in Fig. 2d. Quantification of shoot fresh weight, rosette radius and number of leaves, indicate that ulp1c/d plants have reduced growth. In fact, ulp1d-2 also showed smaller shoot growth (Fig. 2d), although statistical significance was only observed for rosette radius. As previously reported (Conti et al. 2008), in vitro-grown seedlings were not significantly altered in morphology or root growth rate (Fig. S3c, d). Since lethality of SUM1/2, E1 and E2 knockouts in Arabidopsis have strongly implicated sumoylation in embryo formation (Saracco et al. 2007), we investigated whether seed development or germination were also compromised in ulp1c/d. While siliques did not show differences in morphology or seed number (Fig. S3e, f), seeds displayed a delay of approximately one day in the formation of green cotyledons (Fig. 2h, i). Complementation of ulp1c/d by ectopic expression of a pro35S::ULP1d construct in the mutant background (C-ulp1c/d) reverted the rosette radius and delayed germination phenotypes, providing genetic confirmation (Fig. 2i; Fig. S3g). The present characterization implicates ULP1c/d in the control of leaf development and seed germination, and establishes the existence of unequal redundancy, with a more significant role in development being played by ULP1d.
Developmental characterization of ulp1c, ulp1d and ulp1c/d mutants. a, b Schematic representation of ULP1c (At1g10570) and ULP1d (At1g60220), with indication of T-DNA insertion sites (inverted triangles) and primer locations for diagnostic PCR genotyping (arrows); selected T-DNA lines are highlighted; exons and UTRs are represented by grey and black boxes, respectively. c Quantitative RT-PCR analysis of ULP1c and ULP1d relative expression levels in the wild-type (Col) and ulp1c/d backgrounds. d Morphology of soil-grown 4-week-old plants; scale bar indicates 1 cm. e Fresh weight of the shoot of 4-week-old plants (n = 9). f Maximum radius of the rosette of 4-week-old plants (n ≥ 12). g Leaf number in 4-week-old plants (n = 9). h Seed germination rate (formation of green cotyledons) (n ≥ 4). i Seedling morphology 4 days after germination; scale bar indicates 1 mm. Error bars represent standard error of the means (SEM). Asterisks represent statistically significant differences of mutants in relation to the wild-type (unpaired t test; *P < 0.05, **P < 0.01)
Microarray transcript profiling implicates ULP1c/d in drought stress responses
We performed a microarray analysis of adult wild-type and ulp1c/d plants, observing that a total of 59 genes were up-regulated and 53 were down-regulated by at least twofold in ulp1c/d (File S1). Gene Ontology analysis was then used for functional inference on the differential transcriptome of the double mutant. We identified an overrepresentation of genes related to shoot development, including organ morphogenesis, which is consistent with ulp1c/d developmental defects (Fig. 3a; Table 1). Interestingly, a significant number of genes, particularly up-regulated genes, correlated with the plant response to both pathogens and various abiotic stresses (Fig. 3a; Table 1). Considering that (1) genes with identical expression patterns will be controlled by the same TF, thus sharing common cis-elements in their promoters, and (2) sumoylation is a known modulator of transcriptional regulators, we subsequently searched for statistically over-represented cis-elements in the promoters of ulp1c/d DEGs (Table 2). Significantly, all the identified TF-binding site motifs (DREB1A/CBF3, ABRE-like, G-box and ATHB6) were associated with drought-dependent transcriptional regulation. In support, we observed that approximately 38 and 23 % of ulp1c/d DEGs were similarly de-regulated after drought and ABA treatments, respectively (Fig. 3b, c) (Catala et al. 2007; Nemhauser et al. 2006). Results strongly suggest that ulp1c/d displays a transcriptional signature of plants responding to drought stress. This hypothesis was validated by qPCR confirmation of microarray data on several up- and down-regulated genes, including drought-responsive genes RD20, GOLS1 and CIPK11 (Fig. 3d).
Microarray analysis of ulp1c/d in standard growth conditions. a Redundancy exclusion and scatterplot analysis of enriched Gene Ontology (GO) terms for ulp1c/d differentially expressed genes. The scatterplot represents the cluster representatives in a two dimensional space (x- and y-axis) derived by applying multidimensional scaling to a matrix of the GO terms’ semantic similarities (Supek et al. 2011); bubble size indicates the frequency of the GO term. b, c Venn diagram comparison of differentially expressed genes in ulp1c/d against differential expression following drought stress (b) and ABA (c) treatment (Catala et al. 2007; Nemhauser et al. 2006). d Quantitative RT-PCR (qPCR) analysis of ULP1c/d regulation of gene expression. Relative expression levels (fold difference) in ulp1c/d vs wild-type (Col) plants were compared with microarray data for the following genes: QQS (At3g30720); RD20 (At2g33380); GOLS1 (At2g47180); CIPK11 (At2g30360); RPT2 (At2g30520); CalB (for Calcineurin B subunit-related; At2g45670). As a loading control, ACT2 (At3g18780) mRNA was amplified within each sample. Error bars represent three independent biological replicates
ULP1c/d are positive regulators of osmotic stress tolerance and stomatal closure
Studies on plant adaptation to low water potential (ψw) stress use osmotic assays to lower ψw in the growth media in a precise fashion (Verslues et al. 2006), therefore we analysed ulp1c/d growth in MS medium containing PEG or 200 mM mannitol. Seven-day-old Col and ulp1c/d seedlings grown in standard MS agar plates were transferred to osmoticum-containing plates and root growth was subsequently monitored (Fig. 4a–d). In both experiments, ulp1c/d plants were sensitive to the presence of osmotic stress, a phenotype that was reversed by genetic complementation. Results are consistent with the reported sensitivity of ULP1c/d loss-of-function mutants to salt stress (present work, Conti et al. 2008), and implicate these proteases in resistance to the osmotic, rather than the ionic component of salt stress.
Characterization of the ulp1c/d mutant concerning water-deficit response phenotypes. a Morphology of seedlings grown in standard conditions for 7 days and transferred to media supplemented with 200 mM mannitol for 10 days; scale bar indicates 1 cm. b Relative root growth of seedlings subjected to 200 mM mannitol-induced osmotic stress; (n ≥ 10). c Morphology of seedlings grown in standard conditions for 7 days and transferred to PEG-infused MS plates (−0.70 MPa) for 10 days; scale bar indicates 1 cm. d Relative root growth of seedlings subjected to PEG-induced osmotic stress; (n ≥ 24). e Quantification of the stomatal aperture in the abaxial surface of 4-week-old wild-type, ulp1c/d and C-ulp1c/d plants; (n = 5 independent plants with >15 measurements per plant). f Morphology of abaxial stomata in 4-week-old wild-type, ulp1c/d and C-ulp1c/d plants; scale bar indicates 5 µm. g Rapid dehydration assay by quantification of water loss in percentage of fresh weight lost over time, in 4-week-old wild-type and ulp1c/d plants; (n = 6). h Morphology of mesophyll cells belonging to the fifth rosette leaves of 4-week-old wild-type, ulp1c/d and C-ulp1c/d plants; scale bar indicates 100 µm. Error bars represent SEM. Asterisks represent statistically significant differences between genotypes (unpaired t test; *P < 0.05, **P < 0.01, ***P < 0.001)
Since water-deficit responses are mainly determined by stomatal conductance (Schroeder et al. 2001), we analysed whether stomatal aperture was affected in the ulp1c/d mutant. We observed that ulp1c/d displayed increased stomatal aperture in optimally-grown adult plants, while the complementation mutant was able to suppress this phenotype, placing ULP1c/d as positive regulators of stomatal closure (Fig. 4e, f). We used a rapid dehydration assay to check if the higher stomatal aperture translated into faster water loss rates in ulp1c/d. The aerial part of each plant was detached from roots and exposed to dehydration while the decline in fresh weight was monitored. We observed that the rate of water loss was marginally higher in ulp1c/d mutants, though differences were not statistically significant (Fig. 4g). Interestingly, when subjected to long-term drought, adult ulp1c/d mutants displayed a moderate resistance phenotype (Fig. S4). To rule out the possibility that these observations were affected by leaf cellular density, we demonstrated that mesophyll cells were similar in volume and density between ulp1c/d and the wild-type, and no differences in stomata density or morphology at the abaxial leaf surface were observed (Fig. 4h; Fig. S5a–d), thus suggesting an identical number of cells per unit area. Additionally, we observed that the significant increase in stomatal aperture in ulp1c/d could also be replicated in in vitro-grown seedlings (Fig. S5e, f), strongly implicating ULP1c/d in stomatal aperture control.
Exogenous ABA triggers a similar response between Col and ulp1c/d
ABA is a key regulator of plant water status, regulating various dehydration responses, therefore stomatal closure was investigated in the ulp1c/d double mutant, after application of exogenous ABA (Fig. 5a, b). Under light and stomata-opening solution, aperture was significantly higher in ulp1c/d than in wild-type plants. Addition of ABA proportionally closed the stomata in both genotypes, maintaining the higher aperture in ulp1c/d. ABA inhibition of root growth was also measured in in vitro-grown seedlings (Fig. 5c, d). Once more, no differences were observed towards hypo- or hyper-sensitivity of ulp1c/d to exogenous ABA application. Since ABA levels are also fundamental for seed dormancy and maintenance (Finkelstein et al. 2008), and the ulp1c/d mutant previously displayed a delay in germination, we analysed its phenotype in the presence of ABA (Fig. 5e, f). This hormone induced a 6-day delay in germination for both genotypes, and the 1-day-late germination phenotype of ulp1c/d, previously observed in ABA-free medium (Fig. 2h, i), was maintained in this assay. The overall results indicate that the response to exogenous ABA is similar between ulp1c/d and wild-type plants.
Characterization of the ulp1c/d response to ABA. a Representative images of stomatal aperture in the presence or absence of 5 µM ABA. Scale bars indicate 5 µm. b Stomatal aperture in response to ABA; error bars represent SEM (n ≥ 130). c Morphology of seedlings grown in standard conditions for 7 days and transferred to media supplemented with 0 µM (control) or 10 µM of ABA, and grown for 8 days; scale bar indicates 1 cm. d Root growth rate of seedlings (n ≥ 18) transferred to media supplemented with different ABA concentrations. e Seedling morphology 10 days after germination in the presence of 1 µM ABA; scale bar indicates 1 mm. f Seed germination rate (formation of green cotyledons) (n = 6) in the presence of 1 µM ABA. Error bars represent SEM. Asterisks represent statistically significant differences between genotypes (unpaired t test; ***P < 0.001)
The ulp1c/d mutant displays altered SUMO-conjugate levels
In plants, an increase in SUMO conjugates appears to be an early and common event following numerous abiotic stress challenges including rapid dehydration (Castro et al. 2012; Catala et al. 2007). We investigated the role of ULP1c and ULP1d in the SUMO conjugation profile after exposure of 10-day-old seedlings to rapid dehydration. The immunoblot was performed using an antibody raised against the main SUMO modifiers SUM1 and -2 from Arabidopsis (Conti et al. 2008). As shown in Fig. 6, wild-type plants displayed a low level of high molecular weight SUM1/2 conjugates (HMWC), while heat shock (HS) treatment induced the massive accumulation of HMWC, as previously reported (Kurepa et al. 2003; Miller et al. 2010; Saracco et al. 2007). HMWC in the E3 ligase mutant siz1 were considerably lower but still increased following stress imposition. Most significantly, ulp1c/d plants accumulated HMWC in control conditions, indicating an important role for these proteases as SUMO-deconjugating enzymes. While exposure to rapid dehydration induced the accumulation of HMWC in the wild-type, this accumulation was not significant in ulp1c/d due to the endogenously higher levels of HMWC.
Western blot analysis of high molecular weight SUM1-conjugates (HMWC) following rapid dehydration. Ten-day-old plants were subjected to rapid dehydration for 4 h. As a positive control, similar plants were subjected to a 37 °C heat shock for 1 h (HS). Protein extracts (50 µg per lane) were analysed by protein gel blots using anti-AtSUM1 polyclonal antibodies. As a loading control, Ponceau S staining of the large subunit (LS) of Rubisco (55 kD) is displayed
SIZ1 operates upstream of ULP1c/d in the control of stress responses
SUMO conjugate accumulation in ulp1c/d suggests that ULP1c/d act downstream of E3 ligases to promote deconjugation of sumoylated SUMO targets. Thus we investigated the epistatic relationship between ULP1c/d and the major stress-associated E3 ligase SIZ1, by generating the triple siz1 ulp1c/d mutant. As shown in Fig. 7a, b, the triple mutant showed enhanced developmental defects in comparison with siz1. Likewise, it displayed a stronger delay in seed germination than siz1 or the double ulp1c/d mutant (Fig. 7c, d). A similar germination phenotype was observed in the presence of osmoticum-suplemented medium (Fig. S6). The additive phenotypes of loss-of-function mutants suggest that ULP1c/d are, at least partially, independent of SIZ1 in the control of key developmental traits. This is supported by the lack of significant overlap in DEGs between the siz1 mutant (Catala et al. 2007) and the ulp1c/d double mutant (Fig. 7e).
Characterization of the triple mutant siz1 ulp1c/d. a Morphology of soil-grown 4-week-old plants; scale bar indicates 1 cm. b Maximum radius of the rosette of 4-week-old plants (n ≥ 6). c Seed germination rate inferred by the formation of green cotyledons (n = 6). d Seedling morphology 4 days after germination; scale bar indicates 1 mm. e Comparison of differentially expressed genes between ulp1c/d and previously published siz1-3 microarray data (Catala et al. 2007). f Relative root growth of seedlings grown in standard conditions for 7 days and transferred to media supplemented with 200 mM mannitol for 10 days; error bars represent SEM (n ≥ 15). g Quantification of stomata length/width ratio in 2-week-old seedlings grown in standard conditions, 100 mM NaCl or 200 mM mannitol (n ≥ 46). h Western blot analysis of high molecular weight SUM1-conjugates (HMWC) in 10-day-old seedlings grown in 100 mM NaCl or 200 mM mannitol. i Proposed model for the regulation of development and water-deficit responses by Arabidopsis SUMO proteases ULP1c and ULP1d. Asterisks represent statistically significant differences between genotypes (unpaired t test; ***P < 0.001; a–d represent statistically different populations)
Interestingly, with regards to the stress response, we demonstrated that siz1 was resistant whereas ulp1c/d was sensitive to osmotic stress set by the presence of mannitol in the medium, while the triple mutant displayed a siz1-like phenotype (Fig. 7f). A similar behaviour was observed for stomatal aperture. Seedlings grown in vitro in the presence of mannitol or NaCl constitutively displayed increased stomatal aperture in ulp1c/d (Fig. 7g). Meanwhile, stomata were more closed in both siz1 and the triple siz1 ulp1c/d mutants. These phenotypes were confirmed at the transcriptional level (Fig. S7), by monitoring the expression level of drought marker genes previously known to be differentially expressed in siz1 (Catala et al. 2007). Subsequent analysis of high molecular weight SUMO conjugate levels allowed us to (1) observe that ulp1c/d accumulates HMWC even under different environmental stimuli, (2) establish that NaCl is capable of triggering sumoylation to a higher extent than the equivalent osmotic stress, (3) genetically confirm that ULP1c/d acts hypostatically to SIZ1 during stress-induced sumoylation (Fig. 7h).
Discussion
ULP1c/d are implicated in the control of development
In this study we demonstrate, for the first time, that ULP1c/d are expressed in numerous developmental stages and their loss-of-function generates leaf growth defects and late germination, also confirming previous evidence for the existence of an early flowering phenotype (Conti et al. 2008). In previous studies, overexpression of ULP1c/d already suggested a role for these proteases in rosette development (Conti et al. 2009), and, more recently, ULP1c/d were shown to prevent accumulation of sumoylated DELLA proteins, which themselves lead to growth restraint (Conti et al. 2014). Our current phenotypes are consistent with a model in which ulp1c/d growth defects can be attributed to SUMO-dependent stabilization of DELLAs (Fig. 7i). Also, microarray analysis of the ulp1c/d mutant identified a GO term enrichment in DEGs associated with the positive regulation of organ morphogenesis. Genes under positive transcriptional control of ULP1c/d include the AS1/MYB91 TF, which is associated with leaf development (Byrne et al. 2002), and the TCP3/TCP4 gene pair that is essential for correct morphogenesis of several shoot organs (Koyama et al. 2007). Interestingly, data from Conti et al. (2014) suggests that control of flowering time is independent of DELLA proteins. Here, we observed that the negative flowering time regulator FLC was down-regulated in ulp1c/d (Table 1; Fig. 7i). FLC is equally down-regulated in SUMO pathway mutants siz1 and esd4, and was recently shown to be sumoylated by the Arabidopsis E3 ligase HIGH PLOIDY 2 (HPY2) (Catala et al. 2007; Kwak et al. 2016; Reeves et al. 2002). All four mutants (siz1, hpy2, esd4 and ulp1c/d) display early-flowering phenotypes. Moreover, FLC is transcriptionally repressed by FLOWERING LOCUS D (FLD), and FLD is rendered inactive by SIZ1-dependent SUMO conjugation (Jin et al. 2008), collectively suggesting that FLD and FLC are key components of SUMO control over flowering time.
ULP1c/d influence stomatal aperture and responses to osmotic stress
Overall results support opposing roles for ULP1c/d and SIZ1 in the control of physiological traits associated with water usage, including root-level responses to osmotic stress and the control of stomatal aperture. Since ulp1c/d mutants were previously shown to be hypersensitive to salt (Conti et al. 2008), the present data suggests that osmotic rather than the ionic component of salt stress is responsible for the phenotype previously observed by Conti and co-workers. Results also establish ULP1c/d as positive regulators of stomatal closure, while confirming previous reports that siz1 mutants have reduced stomatal aperture (Li et al. 2012; Miura et al. 2012). Stomatal responses are strongly associated with ABA-mediated signalling (Schroeder et al. 2001). Even though ulp1c/d stomatal and seed germination responses did not differ from wild-type responses following exogenous application of ABA, alterations in endogenous ABA levels or ABA-dependent signalling (as suggested by the ulp1c/d transcriptional signature) are not to be excluded, and mounting evidence suggests that both ABA-dependent and -independent mechanisms are involved in SUMO-abiotic stress associations (reviewed by Castro et al. 2012). Significantly, the siz1 closed stomata phenotype was reported to involve SA-dependent reactive oxygen species (ROS) production, rather than ABA-dependent ROS production (Miura et al. 2012). A similar SA-dependent and ABA-independent association was reported for WRKY46 TF. Overexpression of WRKY46 diminished starch levels in guard cells by positively regulating QUA-QUINE STARCH (QQS) expression, and as a consequence, stomatal closure was impaired and more water loss was observed (Ding et al. 2014; Li et al. 2009). Here, not only did ulp1c/d display stomatal aperture defects, but we also confirmed that QQS was amongst the most up-regulated genes in this mutant. Moreover, ULP1c/d were recently implicated in the modulation of SA biosynthesis and signalling (Bailey et al. 2016), further supporting the hypothesis that modulation of stomatal responses by ULP1c/d may involve SA.
Stress-induced SUMO-conjugate levels are constitutively modulated by ULP1c/d
SUMO conjugate accumulation during stress imposition is ubiquitous in eukaryotes (Castro et al. 2012; Golebiowski et al. 2009; Kurepa et al. 2003; Zhou et al. 2004), and tends to match the duration and intensity of the stress (Kurepa et al. 2003; Miller and Vierstra 2011), suggesting an important role for ULP isopeptidase activity in the recovery period following stress challenges. In the present report, SUMO conjugate levels were constitutively increased in the ulp1c/d double mutant, therefore stress imposition did not render significant differences in SUMO conjugate levels in comparison with non-stressed mutant plants. Because under standard growth the ulp1c/d SUMO conjugate profile mimics that of drought-stressed plants, this may lead to the constitutive activation of SUMO-dependent stress signalling mechanisms. In support, we showed that, under normal growth, ulp1c/d mutants display a drought stress transcriptional signature. We also confirmed that the ulp1c/d mutations up-regulate effector genes involved in different drought stress responses, including the drought marker gene RD20 (Aubert et al. 2010), GOLS1 (important in the biosynthesis of the osmoprotectant raffinose; Taji et al. 2002), and CIPK11, a protein kinase associated with stomatal movement (Fuglsang et al. 2007). Finally, we observed that adult ulp1c/d plants are resistant to limiting water conditions, suggesting that later in development they are more adaptable to drought, after a lifetime of priming for drought resistance responses.
ULP1c/d act downstream of SIZ1 in the control of osmotic stress and stomatal aperture
A key issue in the characterization of SUMO dynamics concerns the dual mode-of-action of ULPs as either SUMO maturases/endopeptidases (placing them at the beginning of the pathway), and deconjugases/isopeptidases, operating downstream of the SUMO conjugation machinery. Previously, both SUMO E3 ligases SIZ1 and MMS21/HPY2 were associated with drought stress. MMS21/HPY2 was proposed to be a negative regulator of drought tolerance (Zhang et al. 2012), since the knockout mutant mms21 showed hypersensitivity to exogenous ABA in the control of stomatal aperture and seed germination/cotyledon greening. The mms21 phenotypes, however, do not seem to correlate with ULP1c/d function, since ulp1c/d responses to exogenous ABA did not differ from the wild-type’s responses. Conversely, SIZ1 has been more consistently associated with various abiotic stress responses (reviewed by Castro et al. 2012). Miura et al. (2012) demonstrated that the reduced stomatal aperture of siz1 was ABA-independent and the stomatal index (no. stomata/no. epidermal cells) was not affected, which supports a functional correlation between SIZ1 and ULP1c/d at phenotype level. Here, we used the siz1 mutant to provide the first genetic evidence of Arabidopsis ULPs operating downstream of SUMO E3 ligases, suggesting that SUMO deconjugation/isopeptidase activity is the primary role of ULP1c/d. We demonstrate that SIZ1 is epistatic to ULP1c/d in the control of osmotic stress responses and stomatal aperture. Since siz1 drought-related phenotypes oppose those of ulp1/d, ULP1c/d are likely to operate by directly counteracting SIZ1-driven SUMO modification of drought-responsive targets.
Genes which were differentially expressed in ulp1c/d did not include known components of drought-associated signalling pathways, suggesting that ULP1c/d operates on targets that are located upstream of the deregulated stress responsive genes. We observed that cis-element enrichment analysis specifically implicated ULP1c/d in the repression of three drought/ABA-related regulons, controlled by DREB1A/CBF3, ABF/AREB, and ATHB6. Despite the existence of hundreds of putative SUMO targets, mechanistic links between SUMO and abiotic stress have been extremely scarce, yet they include the SIZ1-dependent modulation of ABI5 and ICE1 activities (Miura et al. 2007, 2009). ABI5 physically interacts with ABRE (ABA-responsive element) cis regulatory elements to control the ABF regulon, while ICE1 sumoylation regulates the expression of DREB1A and its regulon. Thus, a strong correlation can be established between ulp1c/d DEGs and known mechanisms involving SIZ1-drought stress interplay. We propose a model in which ULP1c/d act downstream of SIZ1 in the maintenance of SUMO-target homeostatic levels, including the modulation of ABI5 and ICE1 activities (Fig. 7i). Analysis also seems to implicate ATHB6, a protein that (1) belongs to class I of the homeodomain-leucine zipper (HD-Zip) family of plant TFs, (2) is transcriptionally regulated by ABA levels, and (3) acts as a negative regulator of several ABA responses, including seed germination and stomatal closure (Ariel et al. 2007; Henriksson et al. 2005; Himmelbach et al. 2002). Like ABI5, ATHB6 interacts with components of the primary ABA signalling module, and seems to be post-translationally regulated by ubiquitin E3s (Lechner et al. 2011). Most significantly, ATHB6 has been repeatedly singled out as a SUMO target in high-throughput SUMO proteomic surveys (Miller et al. 2010, 2013), which suggests that ATHB6 sumoylation is likely to constitute a third molecular mechanism implicated in ULP1c/d-drought stress interplay (Fig. 7i).
Future perspectives
In recent years, data from non-plant models has revealed an increasing complexity in SUMO pathway components and in the dynamics of SUMO conjugation/deconjugation cycles. These findings include the identification of new SUMO proteases, and the growing importance of poly-SUMO dynamics (Hickey et al. 2012; Vertegaal 2010). Molecular characterization of plant ULPs must specifically account for the fact that ULPs (1) may discriminate between SUMO isoforms, (2) are likely to contribute differently to total isopeptidase and endopeptidase activities, (3) present different expression patterns, and (4) display different subcellular/subnuclear localizations (Chosed et al. 2006; Colby et al. 2006; Conti et al. 2008; Hermkes et al. 2011; Lois 2010; Murtas et al. 2003). In addition, biological redundancy between SUMO proteases above the canonical redundant pairs is not to be excluded. This complexity will certainly be prompting future research efforts on SUMO protease function, particularly with regards to the resolution of ULP-target specificities.
References
Ariel FD, Manavella PA, Dezar CA, Chan RL (2007) The true story of the HD-Zip family. Trends Plant Sci 12:419–426. doi:10.1016/j.tplants.2007.08.003
Aroca R, Porcel R, Ruiz-Lozano JM (2012) Regulation of root water uptake under abiotic stress conditions. J Exp Bot 63:43–57. doi:10.1093/jxb/err266
Aubert Y et al (2010) RD20, a stress-inducible caleosin, participates in stomatal control, transpiration and drought tolerance in Arabidopsis thaliana. Plant Cell Physiol 51:1975–1987. doi:10.1093/pcp/pcq155
Bailey M, Srivastava A, Conti L, Nelis S, Zhang C, Florance H, Love A, Milner J, Napier R, Grant M, Ari Sadanandom A (2016) Stability of small ubiquitin-like modifier (SUMO) proteases OVERLY TOLERANT TO SALT1 and -2 modulates salicylic acid signalling and SUMO1/2 conjugation in Arabidopsis thaliana. J Exp Bot 67:353–363. doi:10.1093/jxb/erv468
Boyes DC, Zayed AM, Ascenzi R, McCaskill AJ, Hoffman NE, Davis KR, Görlach J (2001) Growth stage-based phenotypic analysis of Arabidopsis: a model for high throughput functional genomics in plants. Plant Cell 13:1499–1510. doi:10.1105/TPC.010011
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Budhiraja R et al (2009) Substrates related to chromatin and to RNA-dependent processes are modified by Arabidopsis SUMO isoforms that differ in a conserved residue with influence on desumoylation. Plant Physiol 149:1529–1540. doi:10.1104/pp.108.135053
Byrne ME, Simorowski J, Martienssen RA (2002) ASYMMETRIC LEAVES1 reveals knox gene redundancy in Arabidopsis. Development 129:1957–1965
Castro PH, Tavares RM, Bejarano ER, Azevedo H (2012) SUMO, a heavyweight player in plant abiotic stress responses. Cell Mol Life Sci 69:3269–3283. doi:10.1007/s00018-012-1094-2
Castro PH, Verde N, Lourenço T, Magalhães AP, Tavares RM, Bejarano ER, Azevedo H (2015) SIZ1-dependent post-translational modification by SUMO modulates sugar signalling and metabolism in Arabidopsis thaliana. Plant Cell Physiol 56(12):2297–2311. doi:10.1093/pcp/pcv149
Catala R, Ouyang J, Abreu IA, Hu Y, Seo H, Zhang X, Chua NH (2007) The Arabidopsis E3 SUMO ligase SIZ1 regulates plant growth and drought responses. Plant Cell 19:2952–2966. doi:10.1105/tpc.106.049981
Chitrakar R, Melotto M (2010) Assessing stomatal response to live bacterial cells using whole leaf imaging. J Vis Exp. doi:10.3791/2185
Chosed R, Mukherjee S, Lois LM, Orth K (2006) Evolution of a signalling system that incorporates both redundancy and diversity: Arabidopsis SUMOylation. Biochem J 398:521–529. doi:10.1042/BJ20060426
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
Colby T, Matthai A, Boeckelmann A, Stuible HP (2006) SUMO-conjugating and SUMO-deconjugating enzymes from Arabidopsis. Plant Physiol 142:318–332. doi:10.1104/pp.106.085415
Conti L, Price G, O’Donnell E, Schwessinger B, Dominy P, Sadanandom A (2008) Small ubiquitin-like modifier proteases OVERLY TOLERANT TO SALT1 and -2 regulate salt stress responses in Arabidopsis. Plant Cell 20:2894–2908. doi:10.1105/tpc.108.058669
Conti L, Kioumourtzoglou D, O’Donnell E, Dominy P, Sadanandom A (2009) OTS1 and OTS2 SUMO proteases link plant development and survival under salt stress. Plant Signal Behav 4:225–227. doi:10.4161/psb.4.3.7867
Conti L et al (2014) Small Ubiquitin-like Modifier protein SUMO enables plants to control growth independently of the phytohormone gibberellin. Dev Cell 28:102–110. doi:10.1016/j.devcel.2013.12.004
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679. doi:10.1146/annurev-arplant-042809-112122
Ding ZJ, Yan JY, Xu XY, Yu DQ, Li GX, Zhang SQ, Zheng SJ (2014) Transcription factor WRKY46 regulates osmotic stress responses and stomatal movement independently in Arabidopsis. Plant J 79:13–27. doi:10.1111/tpj.12538
Edwards K, Johnstone C, Thompson C (1991) A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res 19:1349. doi:10.1093/nar/19.6.1349
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415. doi:10.1146/annurev.arplant.59.032607.092740
Fuglsang AT et al (2007) Arabidopsis protein kinase PKS5 inhibits the plasma membrane H+-ATPase by preventing interaction with 14-3-3 protein. Plant Cell 19:1617–1634. doi:10.1105/tpc.105.035626
Fujita Y, Yoshida T, Yamaguchi-Shinozaki K (2013) Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol Plant 147:15–27. doi:10.1111/j.1399-3054.2012.01635.x
Garcia-Dominguez M, Reyes JC (2009) SUMO association with repressor complexes, emerging routes for transcriptional control. Biochim Biophys Acta 1789:451–459. doi:10.1016/j.bbagrm.2009.07.001
Gareau JR, Lima CD (2010) The SUMO pathway: emerging mechanisms that shape specificity, conjugation and recognition. Nat Rev Mol Cell Biol 11:861–871. doi:10.1038/nrm3011
Golebiowski F et al (2009) System-wide changes to SUMO modifications in response to heat shock. Sci Signal 2:ra24. doi:10.1126/scisignal.2000282
Golldack D, Li C, Mohan H, Probst N (2014) Tolerance to drought and salt stress in plants: unraveling the signaling networks. Front Plant Sci 5:151. doi:10.3389/fpls.2014.00151
Henriksson E, Olsson AS, Johannesson H, Johansson H, Hanson J, Engstrom P, Soderman E (2005) Homeodomain leucine zipper class I genes in Arabidopsis. Expression patterns and phylogenetic relationships. Plant Physiol 139:509–518. doi:10.1104/pp.105.063461
Hermkes R et al (2011) Distinct roles for Arabidopsis SUMO protease ESD4 and its closest homolog ELS1. Planta 233:63–73. doi:10.1007/s00425-010-1281-z
Hickey CM, Wilson NR, Hochstrasser M (2012) Function and regulation of SUMO proteases. Nat Rev Mol Cell Biol 13:755–766. doi:10.1038/nrm3478
Himmelbach A, Hoffmann T, Leube M, Hohener B, Grill E (2002) Homeodomain protein ATHB6 is a target of the protein phosphatase ABI1 and regulates hormone responses in Arabidopsis. EMBO J 21:3029–3038. doi:10.1093/emboj/cdf316
Huang L et al (2009) The Arabidopsis SUMO E3 ligase AtMMS21, a homologue of NSE2/MMS21, regulates cell proliferation in the root. Plant J 60:666–678. doi:10.1111/j.1365-313X.2009.03992.x
Ishida T et al (2009) SUMO E3 ligase HIGH PLOIDY2 regulates endocycle onset and meristem maintenance in Arabidopsis. Plant Cell 21:2284–2297. doi:10.1105/tpc.109.068072
Ishida T, Yoshimura M, Miura K, Sugimoto K (2012) MMS21/HPY2 and SIZ1, two Arabidopsis SUMO E3 ligases, have distinct functions in development. PLoS One 7:e46897. doi:10.1371/journal.pone.0046897
Jin JB et al (2008) The SUMO E3 ligase, AtSIZ1, regulates flowering by controlling a salicylic acid-mediated floral promotion pathway and through affects on FLC chromatin structure. Plant J 53:530–540. doi:10.1111/j.1365-313X.2007.03359.x
Katari MS et al (2010) VirtualPlant: a software platform to support systems biology research. Plant Physiol 152:500–515. doi:10.1104/pp.109.147025
Kilian J, Peschke F, Berendzen KW, Harter K, Wanke D (2012) Prerequisites, performance and profits of transcriptional profiling the abiotic stress response. Biochim Biophys Acta 1819:166–175. doi:10.1016/j.bbagrm.2011.09.005
Koyama T, Furutani M, Tasaka M, Ohme-Takagi M (2007) TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis. Plant Cell 19:473–484. doi:10.1105/tpc.106.044792
Kurepa J et al (2003) The small ubiquitin-like modifier (SUMO) protein modification system in Arabidopsis. Accumulation of SUMO1 and -2 conjugates is increased by stress. J Biol Chem 278:6862–6872. doi:10.1074/jbc.M209694200
Kwak JS, Son GH, Kim S-I, Song JT, Seo HS (2016) Arabidopsis HIGH PLOIDY2 sumoylates and stabilizes Flowering Locus C through its E3 Ligase activity. Front Plant Sci 7:530. doi:10.3389/fpls.2016.00530
Lallemand-Breitenbach V et al (2008) Arsenic degrades PML or PML-RARalpha through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat Cell Biol 10:547–555. doi:10.1038/ncb1717
Lechner E et al (2011) MATH/BTB CRL3 receptors target the homeodomain-leucine zipper ATHB6 to modulate abscisic acid signaling. Dev Cell 21:1116–1128. doi:10.1016/j.devcel.2011.10.018
Li L, Foster CM, Gan Q, Nettleton D, James MG, Myers AM, Wurtele ES (2009) Identification of the novel protein QQS as a component of the starch metabolic network in Arabidopsis leaves. Plant J 58:485–498. doi:10.1111/j.1365-313X.2009.03793.x
Li S, Pandey S, Gookin TE, Zhao Z, Wilson L, Assmann SM (2012) Gene-sharing networks reveal organizing principles of transcriptomes in Arabidopsis and other multicellular organisms. Plant Cell 24:1362–1378. doi:10.1105/tpc.111.094748
Lois LM (2010) Diversity of the SUMOylation machinery in plants. Biochem Soc Trans 38:60–64. doi:10.1042/BST0380060
Lozano-Duran R, Rosas-Diaz T, Gusmaroli G, Luna AP, Taconnat L, Deng XW, Bejarano ER (2011) Geminiviruses subvert ubiquitination by altering CSN-mediated derubylation of SCF E3 ligase complexes and inhibit jasmonate signaling in Arabidopsis thaliana. Plant Cell 23:1014–1032. doi:10.1105/tpc.110.080267
Lyzenga WJ, Stone SL (2012) Abiotic stress tolerance mediated by protein ubiquitination. J Exp Bot 63:599–616. doi:10.1093/jxb/err310
Maruyama K et al (2004) Identification of cold-inducible downstream genes of the Arabidopsis DREB1A/CBF3 transcriptional factor using two microarray systems. Plant J 38:982–993. doi:10.1111/j.1365-313X.2004.02100.x
Miller MJ, Vierstra RD (2011) Mass spectrometric identification of SUMO substrates provides insights into heat stress-induced SUMOylation in plants. Plant Signal Behav 6:130–133. doi:10.4161/psb.6.1.14256
Miller MJ, Barrett-Wilt GA, Hua Z, Vierstra RD (2010) Proteomic analyses identify a diverse array of nuclear processes affected by small ubiquitin-like modifier conjugation in Arabidopsis. Proc Natl Acad Sci USA 107:16512–16517. doi:10.1073/pnas.1004181107
Miller MJ, Scalf M, Rytz TC, Hubler SL, Smith LM, Vierstra RD (2013) Quantitative proteomics reveals factors regulating RNA biology as dynamic targets of stress-induced SUMOylation in Arabidopsis. Mol Cell Proteomics 12:449–463. doi:10.1074/mcp.M112.025056
Miura K, Hasegawa PM (2010) Sumoylation and other ubiquitin-like post-translational modifications in plants. Trends Cell Biol 20:223–232. doi:10.1016/j.tcb.2010.01.007
Miura K et al (2007) SIZ1-mediated sumoylation of ICE1 controls CBF3/DREB1A expression and freezing tolerance in Arabidopsis. Plant Cell 19:1403–1414. doi:10.1105/tpc.106.048397
Miura K, Lee J, Jin JB, Yoo CY, Miura T, Hasegawa PM (2009) Sumoylation of ABI5 by the Arabidopsis SUMO E3 ligase SIZ1 negatively regulates abscisic acid signaling. Proc Natl Acad Sci USA 106:5418–5423. doi:10.1073/pnas.0811088106
Miura K, Lee J, Miura T, Hasegawa PM (2010) SIZ1 controls cell growth and plant development in Arabidopsis through salicylic acid. Plant Cell Physiol 51:103–113. doi:10.1093/pcp/pcp171
Miura K et al (2011) SIZ1 regulation of phosphate starvation-induced root architecture remodeling involves the control of auxin accumulation. Plant Physiol 155:1000–1012. doi:10.1104/pp.110.165191
Miura K, Okamoto H, Okuma E, Shiba H, Kamada H, Hasegawa PM, Murata Y (2012) SIZ1 deficiency causes reduced stomatal aperture and enhanced drought tolerance via controlling salicylic acid-induced accumulation of reactive oxygen species in Arabidopsis. Plant J 73:91–104. doi:10.1111/tpj.12014
Murtas G, Reeves PH, Fu YF, Bancroft I, Dean C, Coupland G (2003) A nuclear protease required for flowering-time regulation in Arabidopsis reduces the abundance of SMALL UBIQUITIN-RELATED MODIFIER conjugates. Plant Cell 15:2308–2319. doi:10.1105/tpc.015487
Nemhauser JL, Hong F, Chory J (2006) Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses. Cell 126:467–475. doi:10.1016/j.cell.2006.05.050
O’Connor TR, Dyreson C, Wyrick JJ (2005) Athena: a resource for rapid visualization and systematic analysis of Arabidopsis promoter sequences. Bioinformatics 21:4411–4413. doi:10.1093/bioinformatics/bti714
Posé D et al (2009) Identification of the Arabidopsis dry2/sqe1-5 mutant reveals a central role for sterols in drought tolerance and regulation of reactive oxygen species. Plant J 59:63–76. doi:10.1111/j.1365-313X.2009.03849.x
Raghavendra AS, Gonugunta VK, Christmann A, Grill E (2010) ABA perception and signalling. Trends Plant Sci 15:395–401. doi:10.1016/j.tplants.2010.04.006
Reeves PH, Murtas G, Dash S, Coupland G (2002) Early in short days 4, a mutation in Arabidopsis that causes early flowering and reduces the mRNA abundance of the floral repressor FLC. Development 129:5349–5361. doi:10.1242/dev.00113
Sadanandom A, Ádámb E, Orosaa B, Vicziánb A, Klosec C, Zhanga C, Jossed E-M, Kozma-Bognárb L, Nagyb F (2015) SUMOylation of phytochrome-B negatively regulates light-induced signaling in Arabidopsis thaliana. Proc Natl Acad Sci USA 11108–11113 doi:10.1073/pnas.1415260112
Saracco SA, Miller MJ, Kurepa J, Vierstra RD (2007) Genetic analysis of SUMOylation in Arabidopsis: conjugation of SUMO1 and SUMO2 to nuclear proteins is essential. Plant Physiol 145:119–134. doi:10.1104/pp.107.102285
Schroeder JI, Kwak JM, Allen GJ (2001) Guard cell abscisic acid signalling and engineering drought hardiness in plants. Nature 410:327–330. doi:10.1038/35066500
Setter TL (2012) Analysis of constituents for phenotyping drought tolerance in crop improvement. Front Physiol 3:180. doi:10.3389/fphys.2012.00180
Shen Q, Ho TH (1995) Functional dissection of an abscisic acid (ABA)-inducible gene reveals two independent ABA-responsive complexes each containing a G-box and a novel cis-acting element. Plant Cell 7:295–307. doi:10.1105/tpc.7.3.295
Silva-Correia J, Freitas S, Tavares RM, Lino-Neto T, Azevedo H (2014) Phenotypic analysis of the Arabidopsis heat stress response during germination and early seedling development. Plant Methods 10:7. doi:10.1186/1746-4811-10-7
Sreenivasulu N, Harshavardhan VT, Govind G, Seiler C, Kohli A (2012) Contrapuntal role of ABA: does it mediate stress tolerance or plant growth retardation under long-term drought stress? Gene 506:265–273. doi:10.1016/j.gene.2012.06.076
Steffens NO, Galuschka C, Schindler M, Bülow L, Hehl R (2004) AthaMap: an online resource for in silico transcription factor binding sites in the Arabidopsis thaliana genome. Nucleic Acids Res 32:D368–D372. doi:10.1093/nar/gkh017
Supek F, Bosnjak M, Skunca N, Smuc T (2011) REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6:e21800. doi:10.1371/journal.pone.0021800
Taji T et al (2002) Important roles of drought- and cold-inducible genes for galactinol synthase in stress tolerance in Arabidopsis thaliana. Plant J 29:417–426. doi:10.1046/j.0960-7412.2001.01227.x
Toledo-Ortiz G, Huq E, Quail PH (2003) The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15:1749–1770. doi:10.1105/tpc.013839
Tomanov K, Zeschmann A, Hermkes R, Eifler K, Ziba I, Grieco M, Novatchkova M, Hofmann K, Hesse H, Bachmair A (2014) Arabidopsis PIAL1 and 2 promote SUMO chain formation as E4-Type SUMO ligases and are involved in stress responses and sulfur metabolism. Plant Cell 26:4547–4560. doi:10.1105/tpc.114.131300
van den Burg HA, Takken FL (2009) Does chromatin remodeling mark systemic acquired resistance? Trends Plant Sci 14:286–294. doi:10.1016/j.tplants.2009.02.003
Verslues PE, Agarwal M, Katiyar-Agarwal S, Zhu J, Zhu JK (2006) Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status. Plant J 45:523–539. doi:10.1111/j.1365-313X.2005.02593.x
Vertegaal AC (2010) SUMO chains: polymeric signals. Biochem Soc Trans 38:46–49. doi:10.1042/BST0380046
Wesley SV, Helliwell CA, Smith NA, Wang M, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA, Robinson SP, Gleave AP, Green AG, Waterhouse PM (2001) Construct design for efficient, effective and high-throughput gene silencing in plants. Plant J 27:581–590. doi:10.1046/j.1365-313X.2001.01105.x
Wilkinson KA, Henley JM (2010) Mechanisms, regulation and consequences of protein SUMOylation. Biochem J 428:133–145. doi:10.1042/BJ20100158
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13:134. doi:10.1186/1471-2105-13-134
Zhang S, Qi Y, Liu M, Yang C (2012) SUMO E3 ligase AtMMS21 regulates drought tolerance in Arabidopsis thaliana. J Integr Plant Biol 55:83–95. doi:10.1111/jipb.12024
Zhou W, Ryan JJ, Zhou H (2004) Global analyses of sumoylated proteins in Saccharomyces cerevisiae. Induction of protein sumoylation by cellular stresses. J Biol Chem 279:32262–32268. doi:10.1074/jbc.M404173200
Acknowledgments
This work was supported by FEDER through the Operational Competitiveness Program—COMPETE—and by national funds through the Foundation for Science and Technology—FCT—within the scope of project “SUMOdulator” [Refs. FCOMP-01-0124-FEDER-028459 and PTDC/BIA-PLA/3850/2012]. PHC was supported by FCT [grant ref. SFRH/BD/44484/2008 and PTDC/BIA-PLA/3850/2012]. HA was supported by the “Genomics and Evolutionary Biology” project, co-financed by North Portugal Regional Operational Programme 2007/2013 (ON.2—O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF), and by FEDER (COMPETE) and FCT, for Rede de Investigação em Biodiversidade e Biologia Evolutiva [UID/BIA/50027/2013 and POCI-01-0145-FEDER-006821].
Author contributions
P.H.C. and D.C. performed most of the experiments; S.F., N.V. and A.P.M. performed or helped perform specific experiments; S.H. performed microarray experimentation; H.A., E.R.B., R.M.T., J. R.-A. and M.A.B. supervised the experiments. H.A., P.H.C. and D.C. conceived the project and analyzed the data. H.A. and P.H.C. wrote the article with contributions from all the authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Pedro Humberto Castro and Daniel Couto have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Castro, P.H., Couto, D., Freitas, S. et al. SUMO proteases ULP1c and ULP1d are required for development and osmotic stress responses in Arabidopsis thaliana . Plant Mol Biol 92, 143–159 (2016). https://doi.org/10.1007/s11103-016-0500-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0500-9