Abstract
The plant growth-promoting rhizobacterium (PGPR) Pseudomonas simiae WCS417r stimulates lateral root formation and increases shoot growth in Arabidopsis thaliana (Arabidopsis). These plant growth-stimulating effects are partly caused by volatile organic compounds (VOCs) produced by the bacterium. Here, we performed a genome-wide association (GWA) study on natural genetic variation in Arabidopsis for the ability to profit from rhizobacteria-mediated plant growth-promotion. To this end, 302 Arabidopsis accessions were tested for root architecture characteristics and shoot fresh weight in response to exposure to WCS417r. Although virtually all Arabidopsis accessions tested responded positively to WCS417r, there was a large variation between accessions in the increase in shoot fresh weight, the extra number of lateral roots formed, and the effect on primary root length. Correlation analyses revealed that the bacterially-mediated increase in shoot fresh weight is related to alterations in root architecture. GWA mapping for WCS417r-stimulated changes in root and shoot growth characteristics revealed 10 genetic loci highly associated with the responsiveness of Arabidopsis to the plant growth-promoting activity of WCS417r. Several of the underlying candidate genes have been implicated in important plant growth-related processes. These results demonstrate that plants possess natural genetic variation for the capacity to profit from the plant growth-promoting function of a beneficial rhizobacterium in their rhizosphere. This knowledge is a promising starting point for sustainable breeding strategies for future crops that are better able to maximize profitable functions from their root microbiome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Beneficial microbes in the microbiome of plant roots provide important services to the plant as they improve plant nutrition and provide protection against plant pathogens (Berendsen et al. 2012; Bulgarelli et al. 2013; Philippot et al. 2013). In recent years, microarray and next-generation sequencing technologies provided detailed insight into the composition and assembly of microbiota communities on and in plants roots (Bai et al. 2015; Bulgarelli et al. 2012; Hardoim et al. 2015; Lareen et al. 2016; Lebeis et al. 2015; Lundberg et al. 2012; Mendes et al. 2011). Genotypic variations in host plants can influence the composition of the root microbiome with important adaptive consequences for plant health (Haney et al. 2015; Pieterse et al. 2016). Evidence is accumulating that domestication of crop species changed root microbiome characteristics in a plant genotype-dependent manner (Bulgarelli et al. 2015; Peiffer et al. 2013). It has been postulated that during the process of plant domestication, plant breeding for high yields under conditions of high fertilizer input in soils has resulted in the erosion of plant genotypic traits involved in beneficial plant–microbe interactions (Pérez-Jaramillo et al. 2016). Hence, the identification of loci associated with the capacity of plants to maximize profitable functions from their root microbiome will be highly instrumental for sustainable plant breeding strategies that are directed towards optimizing root microbiome functions in crops.
Plant root-colonizing bacteria that have a positive effect on plant growth are collectively known as plant growth-promoting rhizobacteria (PGPR). Mechanisms of growth promotion have been studied in detail and different PGPR may have different modes of action. Many PGPR stimulate plant growth through the inhibition of plant pathogens. For some PGPR this is due to direct inhibition of growth and activity of pathogens (Doornbos et al. 2012; Raaijmakers and Weller 1998), whereas others can suppress diseases indirectly by eliciting induced systemic resistance (ISR), a process that is characterized by priming plant-borne defense mechanisms to respond faster and stronger to pathogen attack (Conrath et al. 2015; Pieterse et al. 2014). Numerous rhizosphere inhabiting bacteria can elicit ISR in plants, including Pseudomonas spp. (Bakker et al. 2007; Pieterse et al. 2014) and Bacillus ssp. (Kloepper et al. 2004). Also abiotic stresses like salinity and drought stress can be alleviated by PGPR (Yang et al. 2009). Finally, PGPR can exhibit a direct influence on plant growth and development (Lugtenberg and Kamilova 2009) and this phenomenon is the focus of this study.
A common way by which PGPR directly promote plant growth is by aiding in the uptake of essential nutrients. For example, Rhizobium species stimulate the formation of root nodules in which atmospheric nitrogen is fixed and made available to the plant (Van Rhijn and Vanderleyden 1995). Other PGPR free up scarce nutrients in the soil, like phosphate (Browne et al. 2009), or induce plant uptake mechanisms for nitrate (Mantelin and Touraine 2004) or iron (Zamioudis et al. 2014, 2015; Zhang et al. 2009). PGPR can also influence plant growth by the production of plant hormones like auxin, abscisic acid, cytokinins, and gibberellins (Hayat et al. 2010), or influence plant ethylene concentrations by production of the enzyme 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase that breaks down ACC, the direct precursor of ethylene (Glick et al. 1998; Penrose et al. 2001). The latter process can effectively lower local ethylene concentrations resulting in increased root and shoot biomass (Glick et al. 2007).
More recently, volatile organic compounds (VOCs) have been reported to be involved in plant–microbe interactions, including rhizobacteria-mediated plant growth promotion (Blom et al. 2011; Junker and Tholl 2013). An early study reporting on VOCs and plant growth promotion deals with a Bacillus subtilis strain that promotes growth of Arabidopsis thaliana (Arabidopsis) via the bacterial VOCs acetoin and 2,3-butanediol (Ryu et al. 2003). Also root system architecture in Arabidopsis can be modulated by VOCs (Gutierrez-Luna et al. 2010). For instance, bacterial indole promotes root growth by interfering with auxin signaling in Arabidopsis (Bailly et al. 2014; Bhattacharyya et al. 2015). Moreover, Meldau et al. (2013) reported that the VOC dimethyl disulfide produced by Bacillus sp. B55 promotes growth in Nicotiana attenuata. Whereas most VOCs seem to act as inter-kingdom signaling molecules, dimethyl disulfide was suggested to be metabolized by the plant, thus reducing the need for energy costly sulfur uptake.
In Arabidopsis, colonization of the roots by the PGPR Pseudomonas simiae WCS417r (formerly known as Pseudomonas fluorescens WCS417r; Berendsen et al. 2015) can increase shoot fresh weight by approximately 30 % when co-cultivated in soil (Pieterse and Van Loon 1999). Using a split-plate system, it was shown that this growth-promoting effect is partly mediated by bacterial VOCs and unrelated to the ISR-inducing capacity of WCS417r (Zamioudis et al. 2013). Co-cultivating Arabidopsis accession Col-0 with WCS417r leads to inhibition of primary root elongation and promotion of lateral root and root hair formation, resulting in significant changes in the root architecture and increased shoot fresh weight (Zamioudis et al. 2013). This bacterially-induced process requires the action of the plant growth regulator auxin.
The involvement of microbial VOCs in plant growth promotion is typically studied by co-cultivating plants and PGPR in sealed Petri dishes in which microbially-produced CO2 can accumulate. Hence, it has been hypothesized that CO2 produced by the bacteria causes the growth promotion through the enhanced availability of this photosynthesis substrate. Whereas there is some evidence that CO2 can be partially involved (Kai and Piechulla 2009), the growth response stimulated by PGPR is far greater than can be explained by elevated CO2 alone (Blom et al. 2011). Elevated CO2 levels can increase plant biomass by up to 25 % (Sun et al. 2002; Ward and Strain 1997), but many PGPR easily surpass this as increases in plant biomass of over eightfold have been reported (Park et al. 2015). For WCS417r VOCs, Arabidopsis shoot fresh weight increases of up to fourfold have been reported (Zamioudis et al. 2013), indicating that also WCS417r VOCs have the capacity to stimulate plant growth beyond that caused solely by enhanced CO2 levels.
To investigate whether plant genotypic traits are involved in the plant’s capacity to profit from the plant growth-promoting activity of PGPR, we performed a genome-wide association (GWA) study. GWA mapping is a method initially utilized in human population studies to identify the genetic basis of complex traits (Hirschhorn and Daly 2005). GWA mapping has also been successfully utilized in plant studies (e.g. Aranzana et al. 2005; Atwell et al. 2010; Bac-Molenaar et al. 2015; Baxter et al. 2010; Chan et al. 2010; Kloth et al. 2012; Li et al. 2010). The underlying rationale of GWA studies is that natural variation in phenotypic traits in a population is caused by genetic differences that via single nucleotide polymorphisms (SNPs) in the genomes of the phenotyped genotypes can be linked to genetic loci and ideally candidate genes. Because plant growth promotion by PGPR is known to display host variation (Smith and Goodman 1999), GWA mapping may be a promising tool to investigate the genetic basis of PGPR-mediated plant growth promotion. In this study, we tested whether Arabidopsis has natural genetic variation in the ability to profit from the plant growth-promoting capacity of beneficial rhizosphere bacteria. If so, then this would hold promise for plant breeding strategies in crops that are aimed at maximizing profitable functions from the root microbiome. To test this, we used a global population of 302 Arabidopsis accessions that have been genotyped for 214 k SNPs (Baxter et al. 2010; Li et al. 2010) and performed a GWA study for growth-promotion characteristics in response to WCS417r. Virtually all accessions showed enhanced shoot fresh weight and increased lateral root formation upon exposure to WCS417r, while effects on primary root length were more variable. However, the magnitude of the plant growth-promoting effects was significantly different between Arabidopsis accessions and was independent of the intrinsic growth rate of the accessions. GWA mapping resulted in the identification of genetic loci associated with the plant’s capacity to benefit from growth-promoting effects of WCS417r rhizobacteria.
Materials and methods
Cultivation of bacteria
Pseudomonas simiae WCS417r (formerly known as Pseudomonas fluorescens WCS417r; Berendsen et al. 2015) was isolated in the 1980s from the rhizosphere of wheat (Lamers et al. 1988) and served since as a model Pseudomonas spp. strain for studying plant growth promotion (Van Peer and Schippers 1989; Zamioudis et al. 2013) and rhizobacteria-induced systemic resistance (Pieterse et al. 1996; Van Peer et al. 1991; Zamioudis and Pieterse 2012). WCS417r was grown for 24 h at 28 °C on King’s medium B agar medium supplemented with 50 µg mL−1 rifampicin as described (Van Wees et al. 2013). Subsequently, the WCS417r bacteria were suspended in 10 mL of 10 mM MgSO4 and centrifuged at 3200g for 5 min. The pellet was washed twice by resuspension in 10 mM MgSO4 and subsequent centrifuging at 3200g for 5 min after which the density was adjusted to 2 × 106 colony-forming units (cfu) mL−1 of 10 mM MgSO4 (OD600 = 0.002).
Plant growth conditions and PGPR treatment
A collection of 302 natural accessions of Arabidopsis thaliana (Supplemental Table S1) was used to investigate their responsiveness to growth-promoting effects of P. simiae WCS417r. Seeds were gas sterilized for 4 h in a desiccator as described (Van Wees et al. 2013). Sterilized seeds were sown on square Petri dishes with agar-solidified Murashige and Skoog (MS) medium supplemented with 0.5 % sucrose as described (Zamioudis et al. 2013). The seeds on MS agar were stratified for 2 days at 4 °C and subsequently incubated in a vertical position in a plant growth chamber at 21 °C with a 16-h day (100 µmol m−2 s−1) and 8-h night cycle. After 4 days of growth, seedlings of similar size were carefully transferred to 9 cm diameter Petri dishes containing MS agar medium. For the bacterial treatment, 240 µl of a WCS417r suspension containing 2 × 106 cfu mL−1 was applied on the agar medium 5 cm below the roots of the seedlings. For the mock treatment, 240 µl of 10 mM MgSO4 was applied in a similar manner. The Petri dishes were briefly dried in a laminar flow cabinet, sealed with a lid and two layers of Parafilm, and placed in a vertical position in the growth room.
Phenotyping responsiveness of Arabidopsis accessions to WCS417r
Per biological replicate, 4 seedlings of a tester Arabidopsis accession were grown on a MS agar plate alongside 4 seedlings of the reference accession Col-0. For each of the 302 tested accessions, 3 replicate plates with WCS417r-treated seedlings were compared to 3 replicate plates with mock-treated seedlings. After 8 days of growth, plates were photographed after which average primary root length per seedling and average number of lateral roots per seedling was determined for each plate. After 10 days of growth, average shoot fresh weight per seedling was determined for each plate of the mock- and WCS417r-treated seedlings. Data were obtained from three replicate plates per accession for both the control and the WCS417r treatment.
GWA mapping, SNP selection, and statistics
A collection of 302 accessions was used to investigate the genetic variation present within Arabidopsis (Baxter et al. 2010; Li et al. 2010; Platt et al. 2010). Each of these accessions were genotyped versus the Col-0 accession with ~214 k single nucleotide polymorphism markers (Kim et al. 2007). GWA mapping was performed on the WCS417r-mediated changes in shoot fresh weight (∆SFW), number of lateral roots formed (∆LRF), and primary root length (∆PRL). For all traits, means per seedling (n = 4) per biological replicate (n = 3) were used to calculate the mean per treatment per accession. Values obtained after subtracting the mean value of the mock treatment from that of the WCS417r treatment were used for the GWA analysis (∆SFW, ∆LRF, ∆PRL). The online tool GWAPP (http://gwas.gmi.oeaw.ac.at/) was used for GWA mapping as described (Seren et al. 2012). The data sets were normally distributed and the AMM algorithm was employed for mapping. This algorithm closely resembles the commonly used EMMAX (Kang et al. 2010). The GWAPP Geneviewer was used to zoom in on trait-associated SNPs and reveal their position in the genome to pinpoint candidate genes within 10 kb up- and down-stream of the identified SNP. For each of the candidate genes, the annotations were retrieved from TAIR10 (arabidopsis.org). With the exception of the GWA mapping, all statistics were carried out with IBM SPSS Statistics 20.
Results
Natural variation in Arabidopsis for responsiveness to PGPR-mediated growth promotion
In soil and plate assays, colonization of the roots of Arabidopsis accession Col-0 by P. simiae WCS417r has previously been shown to result in an increase in shoot fresh weight (Pieterse and Van Loon 1999; Zamioudis et al. 2013). On plates, it becomes visible that co-cultivation of Col-0 seedlings with WCS417r not only results in enhanced shoot growth, it also coincides with a faster development of lateral roots (Fig. 1), confirming previous findings (Zamioudis et al. 2013). In order to investigate the natural variation in responsiveness of Arabidopsis to the growth-promoting activity of WCS417r, we tested 302 Arabidopsis accessions for changes in shoot fresh weight and root architecture in response to exposure to WCS417r (Supplemental Table S1). PGPR-induced gain in shoot fresh weight (∆SFW), increase in the number of lateral roots formed (∆LRF), and effects on primary root length (∆PRL) were determined after an 8-day (root architecture) or 10-day (shoot fresh weight) period of co-cultivation with WCS417r by subtracting the average values of mock-treated plants from those of the WCS417r treatments. Figure 2a–c show that for all three parameters tested, the Arabidopsis accessions displayed extensive variation in the capacity to respond to WCS417r. Upon exposure to WCS417r, the reference accession Col-0 showed an average increase in shoot fresh weight of 10 mg, which is a 2.1-fold increase relative to the shoot fresh weight of mock-treated Col-0 plants. Among the 302 accessions, shoot fresh weight in response to WCS417r exposure changed between 0.9 and 4.2 fold, with Col-0 around the 65th percentile (Fig. 2a). For the increase in the number of lateral roots over the 8-day period of exposure to WCS417r, Col-0 performed in the upper range (Fig. 2b). While Col-0 formed on average 7 additional lateral roots in comparison to mock-treated seedlings, other accessions formed up to 14 additional lateral roots or none at all. The primary root length remained virtually unchanged in Col-0 in response to WCS417r, but in the other accessions it ranged from significant decreases to significant increases in length (Fig. 2c). Overall, these results demonstrate that Arabidopsis possesses extensive natural variation in responsiveness to the PGPR WCS417r.
Phenotyping of Arabidopsis accessions for responsiveness to the plant growth-promoting effect of P. simiae WCS417r. Shown are photographs of MS agar plates with 4 Col-0 seedlings (left half of the plate) and 4 seedlings of a tester accession (right half of the plate), 8 days after spotting 240 µl of 10 mM MgSO4 (Mock) or a WCS417r bacterial suspension (2 × 106 cfu mL−1) on the plate below the plant roots
Natural variation of 302 A. thaliana accessions in their responsiveness to the growth-promoting effect of P. simiae WCS417r. a Accessions sorted for increase in shoot fresh weight (∆SFW) in response to exposure to WCS417r for 10 days (Col-0 is indicated with arrow and black dot). b Accessions sorted for increase in the number of lateral roots formed (∆LRF) in response to exposure of seedlings to WCS417r for 8 days. c Accessions sorted for difference in the length of the primary root (∆PRL) after exposure of the seedlings to WCS417r for 8 days. d Average shoot fresh weight of mock-treated (grey dots) and WCS417r-treated (green dots) plants. e Number of lateral roots formed in mock-treated and WCS417r-treated (red dots) plants. f Primary root length of mock- and WCS417r-treated (purple dots) plants. Each dot represents the average of 3 biological replicates. Values in a–c were calculated by subtracting the mean of the mock treatment from the mean of the WCS417r treatment. In each panel, accessions were sorted for increasing values. The order of the accessions differs per panel. The Pearson correlation coefficient (R2) indicates the degree of correlation between values of control and WCS417r-treated plants
PGPR-mediated growth promotion capacity of Arabidopsis accessions is not linked to their intrinsic growth rates
To investigate whether the observed variation in the WCS417r-induced changes in the shoot and root growth parameters is related to intrinsic growth rates among the accessions, we plotted the absolute values of shoot fresh weight, number of lateral roots, and primary root length for both the mock and the WCS417r treatment and ranked the accessions ascendingly for the mock treatment (Fig. 2d–f). Figure 2d, e show that the values of the mock and the WCS417r treatment have a low correlation coefficient for the parameters shoot fresh weight and number of lateral roots (R2 = 0.313 and 0.025, respectively), indicating that the magnitude of these WCS417r-induced growth responses are, at the most, weakly related to the intrinsic growth capacity for the parameters tested under these conditions. Hence, faster growing accessions, or accessions that form more lateral roots in the experimental setup are not necessarily stronger responders to WCS417r. For the parameter primary root length, the correlation between mock and WCS417r treatment is stronger (R2 = 0.620; Fig. 2f).
PGPR-mediated enhancement of shoot fresh weight is correlated with root architecture parameters
In order to investigate whether the natural variation in the measured shoot and root growth characteristics of the 302 tested Arabidopsis accessions is correlated, we performed a correlation analysis for both the mock and the WCS417r data sets. Figure 3 shows that in mock-treated plants, shoot fresh weight only weakly correlates with the number of lateral roots (R2 = 0.228; Fig. 3a) and primary root length (R2 = 0.301; Fig. 3b), while primary root length and number of lateral roots do not correlate at all (R2 = 0.032; Fig. 3c). However, in WCS417r-treated plants, the correlation between number of lateral roots and shoot fresh weight (R2 = 0.515; Fig. 3a), primary root length and shoot fresh weight (R2 = 0.604; Fig. 3b), and primary root length and number of lateral roots (R2 = 0.385; Fig. 3c) is markedly higher than in the mock-treated plants. These results indicate that the PGPR-mediated increase in shoot fresh weight is at least partly related to PGPR-mediated changes in root architecture.
Correlation analysis of shoot fresh weight, number of lateral roots, and primary root length for 302 mock- and P. simiae WCS417r-treated Arabidopsis accessions. a Correlation between number of lateral roots and shoot fresh weight. b Correlation between primary root length and shoot fresh weight. c Correlation between primary root length and number of lateral roots. The correlation coefficient (R2) is presented in the graphs and is statistically significant at 0.01 (Pearson’s r, 2-tailed) unless stated otherwise (ns not significant)
GWA mapping reveals loci related to the capacity of Arabidopsis to respond to PGPR
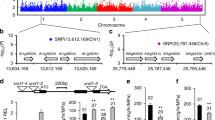
To study the genetic basis of PGPR-mediated plant growth promotion, we performed GWA mapping on the increase in shoot fresh weight data (∆SFW), the increase in number of lateral roots formed data (∆LRF), and the change in primary root length data (∆PRL), using the 214 k SNP set that is commonly used for GWA studies in Arabidopsis (Atwell et al. 2010; Bac-Molenaar et al. 2015; Horton et al. 2012; Kim et al. 2007; Li et al. 2010). Data for ∆SFW, ∆LRF, and ∆PRL (Supplementary Table S1) were uploaded to the online tool GWAPP (http://gwas.gmi.oeaw.ac.at/) (Seren et al. 2012) to analyze the quality of the data and perform GWA mapping. The (pseudo-)heritabilities for the recorded phenotypic variation in ∆SFW, ∆LRF, ∆PRL, were 0.28, 0.62, and 0.37, respectively. After confirming that all datasets were normally distributed, they were processed with the AMM algorithm. The GWA mapping results for all three parameters are depicted in Fig. 4. Of the loci with high SNP-trait associations [−log10(P) > 5.0], 5 loci were associated with PGPR-stimulated shoot fresh weight and 5 loci with PGPR-mediated increase in lateral root formation. No loci SNPs were highly associated with PGPR-mediated changes in primary root length (Table 1; Supplemental Table S2). When using a lower threshold level [−log10(P) > 4.0], 18 additional loci were detected for PGPR-induced increase in shoot fresh weight, 20 for the PGPR-mediated increase number of lateral roots, and 20 for the PGPR-induced changes in primary root length (Supplemental Table S2). Although we observed a clear level of correlation between WCS417r-mediated changes in shoot fresh weight and the root architecture parameters (Fig. 3), none of the identified trait-associated SNPs overlapped.
GWA mapping of P. simiae WCS417r-mediated effects on shoot fresh weight (∆SFW), lateral roots formation (∆LRF), and primary root length (∆PRL). Manhattan plots of the −log10(P) SNP marker–trait associations are shown. From left to right, different colors represent Arabidopsis chromosomes I–V. The dotted grey lines indicate the arbitrary thresholds of −log10(P) = 4 and −log10(P) = 5
The linkage disequilibrium of the natural population of Arabidopsis accessions used for GWA mapping is on average 10 kb (Kim et al. 2007). Therefore, all genes within 10 kb up- or downstream of the highly [−log10(P) > 5.0] and moderately [−log10(P) > 4.0] associated SNPs were identified. Their identities, gene annotation, and functional information are provided in Supplemental Table S2. In Table 1, the highly-associated SNPs are given along with their closest candidate gene. Among the candidate genes in the closest vicinity of the high SNP-trait associations for the PGPR-mediated changes in shoot and root growth are genes encoding proteins that are associated with important plant growth processes, such as the cytokinin transporter PUP19 (Cedzich et al. 2008), the photosynthesis-related protein Golden2-like 1 (GLK1) (Waters et al. 2009), the Rubisco biogenesis-related DEAD box protein RH39 (Nishimura et al. 2010), carbohydrate transmembrane transport protein At2G18480, Nitrilase-like protein 1 (NLP1) (Kusano et al. 2008), and the photorespiration-related mitochondrial serine hydroxymethyltransferase SHM2 (Engel et al. 2011; Voll et al. 2006) (Table 1). Overall, these results provide insight in the genetic components that may contribute to the plant’s capacity to benefit from growth-promoting rhizobacteria in the rhizosphere.
Discussion
Natural accessions of Arabidopsis have a high degree of variation in plant development, physiology, and adaptation to their biotic and abiotic environment (Alonso-Blanco et al. 2009). Using a small collection of Arabidopsis accessions, we previously explored the natural genetic variation in Arabidopsis for its capacity to develop PGPR-mediated ISR and identified quantitative trait loci related to this phenomenon (Ton et al. 1999, 2001). Haney et al. (2015) recently showed that genotypic variations in Arabidopsis accessions can influence the ability to associate with specific PGPR with consequences for plant fitness. Here we made use of the natural variation within 302 Arabidopsis accessions to investigate to what extent the different genotypes differ in their capacity to respond to growth-promoting effects of beneficial rhizosphere bacteria and to gain insight into the genetic basis of the profitable functions by which plants can benefit from their root microbiome. Analysis of the growth promotion parameters shoot fresh weight, lateral root formation, and primary root length in response to WCS417r treatment showed large natural variation among the tested accessions for all three parameters (Fig. 2). The capacity to respond to WCS417r only weakly correlated with the intrinsic growth rate of the accessions. Hence, accessions that gained more shoot fresh weight or produced more lateral roots in the mock-treatment, were not necessarily better responders to the PGPR treatment. By contrast, WCS417r-mediated changes in the growth parameters were moderately to highly correlated with each other. Especially the WCS417r-mediated increase in shoot fresh weight showed a high correlation with PGPR-induced changes in the root architecture parameters (Fig. 3), suggesting that changes in root architecture and shoot fresh weight are linked. Previously, Zamioudis et al. (2013) demonstrated that root colonization by WCS417r enhances the auxin response in the root of Arabidopsis and stimulates auxin-dependent developmental programs related to primary root, lateral root, and root hair development. Collectively, these root architectural changes enlarge the capacity of the root system to take up water and nutrients (Lopez-Bucio et al. 2003), which may contribute to the observed correlation between extra number of lateral roots formed and increased shoot fresh weight in WCS417r-treated Arabidopsis seedlings. It should, however, be noted that Arabidopsis seedlings were grown on plates in which nutrient availability was not limiting, hence we cannot rule out the possibility that the correlation between root architectural changes and increased shoot fresh weight is caused by another, so far unknown process.
In this study, we chose to treat the Arabidopsis seedlings with PGPR by co-cultivating them on the same MS agar plate while avoiding physical contact (Fig. 1). Previously, we showed that in this setup the effects on root and shoot growth are predominantly mediated by VOCs produced by the bacteria (Zamioudis et al. 2013). We realize that in this way we mainly capture the natural variation in VOCs-mediated plant growth effects in the Arabidopsis population, but because the magnitude of VOCs effects are very similar to those mediated by direct application of PGPR to the root system, we chose for this more simplified system. Also in other systems VOCs produced by PGPR have been shown to have major impacts on plant growth and development (Blom et al. 2011; Farag et al. 2013; Gutierrez-Luna et al. 2010; Park et al. 2015; Ryu et al. 2003). Microbial VOCs have been shown to stimulate auxin responses in the roots (Bailly et al. 2014; Bhattacharyya et al. 2015; Zamioudis et al. 2013), or are suggested to be taken up and utilized as e.g. sulfur nutrition (Meldau et al. 2013).
Although the PGPR-mediated increase in shoot fresh weight and changes in root architecture seem to be related in the population of natural Arabidopsis accessions tested (Fig. 3), the GWA analysis did not yield common candidate genes (Fig. 4, Table 1; Supplemental Table S2). Among the identified candidate genes for the PGPR-mediated increase in shoot fresh weight were genes associated with important plant growth processes. For instance, the GLK1 gene encoding transcription factor Golden2-like 1 (GLK1), which together with GLK2 plays a role in the regulation of photosynthetic genes to optimize photosynthetic capacity in varying environmental and developmental conditions (Waters et al. 2009). Or the gene encoding DEAD box protein RH39, which is required for maturation of chloroplast 23S rRNA and the biogenesis of ribulose 1,5-bisphosphate carboxylase/oxygenase (Rubisco) and other chloroplast-encoded, photosynthetic proteins (Nishimura et al. 2010). Also the cytokinin transporter gene PUP19 (Cedzich et al. 2008) is among the candidate genes associated with enhanced shoot fresh weight. Among the candidate genes related to the PGPR-mediated increase in lateral root formation is SHM2, which encodes the mitochondrial serine hydroxymethyltransferase SHM2 (Engel et al. 2011). The isozyme SHM1 catalyzes an essential step of the photorespiratory C2 cycle, but because SHM2 does not complement the mutant phenotype of shm1 and is relatively highly expressed in roots, its function is currently unknown (Voll et al. 2006). Another candidate gene related to increase in lateral root formation encodes Nitrilase-like protein 1 (NLP1). NLP1 is a N-carbamoylputrescine amidohydrolase involved in biosynthesis of polyamines that play an essential role in plant growth and survival (Kusano et al. 2008). Considering all candidate genes with SNP-trait associations in the GWA analysis, several have known or predicted functions that hold promise for being functional in mediating the plant growth-promoting effect of PGPR. Future analysis of knock-out mutants and ectopic over expressors of these genes and those with a currently unknown function should provide insight into the validity of the candidate genes and their role in the capacity of the plant to benefit from the plant growth-promoting activity of PGPR in their rhizosphere.
In summary, our survey of the natural genetic variation within Arabidopsis for the responsiveness to the plant growth-promoting activity of rhizosphere bacteria revealed that the capacity of plants to profit from their root-associated microbes has a clear genetic component. Hence, identification of genes and loci related to PGPR responsiveness in economically relevant crop species holds promise for breeding strategies that are aimed at introducing traits by which crops maximize profitable functions from their root microbiome.
References
Alonso-Blanco C, Aarts MGM, Bentsink L, Keurentjes JJB, Reymond M, Vreugdenhil D, Koornneef M (2009) What has natural variation taught us about plant development, physiology, and adaptation? Plant Cell 21:1877–1896
Aranzana MJ, Kim S, Zhao K, Bakker E, Horton M, Jakob K, Lister C, Molitor J, Shindo C, Tang C, Toomajian C, Traw B, Zheng H, Bergelson J, Dean C, Marjoram P, Nordborg M (2005) Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. PLoS Genet 1:531–539
Atwell S, Huang YS, Vilhjalmsson BJ, Willems G, Horton M, Li Y, Meng DZ, Platt A, Tarone AM, Hu TT, Jiang R, Muliyati NW, Zhang X, Amer MA, Baxter I, Brachi B, Chory J, Dean C, Debieu M, de Meaux J, Ecker JR, Faure N, Kniskern JM, Jones JDG, Michael T, Nemri A, Roux F, Salt DE, Tang CL, Todesco M, Traw MB, Weigel D, Marjoram P, Borevitz JO, Bergelson J, Nordborg M (2010) Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465:627–631
Bac-Molenaar JA, Fradin EF, Becker FFM, Rienstra JA, van der Schoot J, Vreugdenhil D, Keurentjes JJB (2015) Genome-wide association mapping of fertility reduction upon heat stress reveals developmental stage-specific QTLs in Arabidopsis thaliana. Plant Cell 27:1857–1874
Bai Y, Müller DB, Srinivas G, Garrido-Oter R, Potthoff E, Rott M, Dombrowski N, Münch PC, Spaepen S, Remus-Emsermann M, Hüttel B, McHardy AC, Vorholt JA, Schulze-Lefert P (2015) Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528:364–369
Bailly A, Groenhagen U, Schulz S, Geisler M, Eberl L, Weisskopf L (2014) The inter-kingdom volatile signal indole promotes root development by interfering with auxin signalling. Plant J 80:758–771
Bakker PAHM, Pieterse CMJ, Van Loon LC (2007) Induced systemic resistance by fluorescent Pseudomonas spp. Phytopathology 97:239–243
Baxter I, Brazelton JN, Yu DN, Huang YS, Lahner B, Yakubova E, Li Y, Bergelson J, Borevitz JO, Nordborg M, Vitek O, Salt DE (2010) A coastal cline in sodium accumulation in Arabidopsis thaliana is driven by natural variation of the sodium transporter AtHKT1;1. PLoS Genet 6:e1001193
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486
Berendsen RL, Van Verk MC, Stringlis IA, Zamioudis C, Tommassen J, Pieterse CMJ, Bakker PAHM (2015) Unearthing the genomes of plant-beneficial Pseudomonas model strains WCS358, WCS374 and WCS417. BMC Genom 16:539
Bhattacharyya P, Garladinne M, Lee YH (2015) Volatile indole produced by rhizobacterium Proteus vulgaris JBLS202 stimulates growth of Arabidopsis thaliana through auxin, cytokinin, and brassinosteroid pathways. J Plant Growth Regul 34:158–168
Blom D, Fabbri C, Connor EC, Schiestl FP, Klauser DR, Boller T, Eberl L, Weisskopf L (2011) Production of plant growth modulating volatiles is widespread among rhizosphere bacteria and strongly depends on culture conditions. Environ Microbiol 13:3047–3058
Browne P, Rice O, Miller SH, Burke J, Dowling DN, Morrissey JP, O’Gara F (2009) Superior inorganic phosphate solubilization is linked to phylogeny within the Pseudomonas fluorescens complex. Appl Soil Ecol 43:131–138
Bulgarelli D, Rott M, Schlaeppi K, Loren Ver, van Themaat E, Ahmadinejad N, Assenza F, Rauf P, Huettel B, Reinhardt R, Schmelzer E, Peplies J, Gloeckner FO, Amann R, Eickhorst T, Schulze-Lefert P (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488:91–95
Bulgarelli D, Schlaeppi K, Spaepen S, Van Themaat EVL, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838
Bulgarelli D, Garrido-Oter R, Munch PC, Weiman A, Droge J, Pan Y, McHardy AC, Schulze-Lefert P (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17:392–403
Cedzich A, Stransky H, Schulz B, Frommer WB (2008) Characterization of cytokinin and adenine transport in Arabidopsis cell cultures. Plant Physiol 148:1857–1867
Chan EKF, Rowe HC, Kliebenstein DJ (2010) Understanding the evolution of defense metabolites in Arabidopsis thaliana using genome-wide association mapping. Genetics 185:991–1007
Conrath U, Beckers GJM, Langenbach CJG, Jaskiewicz MR (2015) Priming for enhanced defense. Annu Rev Phytopathol 53:97–119
Doornbos RF, Van Loon LC, Bakker PAHM (2012) Impact of root exudates and plant defense signaling on bacterial communities in the rhizosphere. Rev Agron Sustain Dev 32:227–243
Engel N, Ewald R, Gupta KJ, Zrenner R, Hagemann M, Bauwe H (2011) The presequence of Arabidopsis serine hydroxymethyltransferase SHM2 selectively prevents import into mesophyll mitochondria. Plant Physiol 157:1711–1720
Farag MA, Zhang H, Ryu CM (2013) Dynamic chemical communication between plants and bacteria through airborne signals: induced resistance by bacterial volatiles. J Chem Ecol 39:1007–1018
Glick BR, Penrose DM, Li JP (1998) A model for the lowering of plant ethylene concentrations by plant growth-promoting bacteria. J Theor Biol 190:63–68
Glick BR, Todorovic B, Czarny J, Cheng ZY, Duan J, McConkey B (2007) Promotion of plant growth by bacterial ACC deaminase. Crit Rev Plant Sci 26:227–242
Gutierrez-Luna FM, Lopez-Bucio J, Altamirano-Hernandez J, Valencia-Cantero E, de la Cruz HR, Macias-Rodriguez L (2010) Plant growth-promoting rhizobacteria modulate root-system architecture in Arabidopsis thaliana through volatile organic compound emission. Symbiosis 51:75–83
Haney CH, Samuel BS, Bush J, Ausubel FM (2015) Associations with rhizosphere bacteria can confer an adaptive advantage to plants. Nat Plants 1:1–9
Hardoim PR, Van Overbeek LS, Berg G, Pirttila AM, Compant S, Campisano A, Doring M, Sessitsch A (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79:293–320
Hayat R, Ali S, Amara U, Khalid R, Ahmed I (2010) Soil beneficial bacteria and their role in plant growth promotion: a review. Ann Microbiol 60:579–598
Hirschhorn JN, Daly MJ (2005) Genome-wide association studies for common diseases and complex traits. Nat Rev Genet 6:95–108
Horton MW, Hancock AM, Huang YS, Toomajian C, Atwell S, Auton A, Muliyati NW, Platt A, Sperone FG, Vilhjalmsson BJ, Nordborg M, Borevitz JO, Bergelson J (2012) Genome-wide patterns of genetic variation in worldwide Arabidopsis thaliana accessions from the RegMap panel. Nat Genet 44:212–216
Junker RR, Tholl D (2013) Volatile organic compound mediated interactions at the plant-microbe interface. J Chem Ecol 39:810–825
Kai M, Piechulla B (2009) Plant growth promotion due to rhizobacterial volatiles—an effect of CO2? FEBS Lett 583:3473–3477
Kang HM, Sul JH, Service SK, Zaitlen NA, Kong SY, Freimer NB, Sabatti C, Eskin E (2010) Variance component model to account for sample structure in genome-wide association studies. Nat Genet 42:348–354
Kim S, Plagnol V, Hu TT, Toomajian C, Clark RM, Ossowski S, Ecker JR, Weigel D, Nordborg M (2007) Recombination and linkage disequilibrium in Arabidopsis thaliana. Nat Genet 39:1151–1155
Kloepper JW, Ryu C-M, Zhang SA (2004) Induced systemic resistance and promotion of plant growth by Bacillus spp. Phytopathology 94:1259–1266
Kloth KJ, Thoen MPM, Bouwmeester HJ, Jongsma MA, Dicke M (2012) Association mapping of plant resistance to insects. Trends Plant Sci 17:311–319
Kusano T, Berberich T, Tateda C, Takahashi Y (2008) Polyamines: essential factors for growth and survival. Planta 228:367–381
Lamers JG, Schippers B, Geels FP (1988) Soil-borne diseases of wheat in the Netherlands and results of seed bacterization with pseudomonads against Gaeumannomyces graminis var. tritici, associated with disease resistance. In: Jorna ML, Slootmaker LAJ (eds) Cereal breeding related to integrated cereal production. Pudoc, Wageningen, pp 134–139
Lareen A, Burton F, Schäfer P (2016) Plant root-microbe communication in shaping root microbiomes. Plant Mol Biol. doi:10.1007/s11103-015-0417-8
Lebeis SL, Paredes SH, Lundberg DS, Breakfield N, Gehring J, McDonald M, Malfatti S, del Rio TG, Jones CD, Tringe SG, Dangl JL (2015) Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349:860–864
Li Y, Huang Y, Bergelson J, Nordborg M, Borevitz JO (2010) Association mapping of local climate-sensitive quantitative trait loci in Arabidopsis thaliana. Proc Natl Acad Sci USA 107:21199–21204
Lopez-Bucio J, Cruz-Ramirez A, Herrera-Estrella L (2003) The role of nutrient availability in regulating root architecture. Curr Opin Plant Biol 6:280–287
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, del Rio TG, Edgar RC, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90
Mantelin S, Touraine B (2004) Plant growth-promoting bacteria and nitrate availability: impacts on root development and nitrate uptake. J Exp Bot 55:27–34
Meldau DG, Meldau S, Hoang LH, Underberg S, Wunsche H, Baldwin IT (2013) Dimethyl disulfide produced by the naturally associated bacterium Bacillus sp B55 promotes Nicotiana attenuata growth by enhancing sulfur nutrition. Plant Cell 25:2731–2747
Mendes R, Kruijt M, De Bruijn I, Dekkers E, Van der Voort M, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Nishimura K, Ashida H, Ogawa T, Yokota A (2010) A DEAD box protein is required for formation of a hidden break in Arabidopsis chloroplast 23S rRNA. Plant J 63:766–777
Park YS, Dutta S, Ann M, Raaijmakers JM, Park K (2015) Promotion of plant growth by Pseudomonas fluorescens strain SS101 via novel volatile organic compounds. Biochem Biophys Res Commun 461:361–365
Peiffer J, Spor A, Koren O, Jin Z, Tringe S, Dangl J, Buckler E, Ley R (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci USA 110:6548–6553
Penrose DM, Moffatt BA, Glick BR (2001) Determination of 1-aminocycopropane-1-carboxylic acid (ACC) to assess the effects of ACC deaminase-containing bacteria on roots of canola seedlings. Can J Microbiol 47:77–80
Pérez-Jaramillo JE, Mendes R, Raaijmakers JM (2016) Impact of plant domestication on rhizosphere microbiome assembly and functions. Plant Mol Biol. doi:10.1007/s11103-015-0337-7
Philippot L, Raaijmakers JM, Lemanceau P, van der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799
Pieterse CMJ, Van Loon LC (1999) Salicylic acid-independent plant defence pathways. Trends Plant Sci 4:52–58
Pieterse CMJ, Van Wees SCM, Hoffland E, Van Pelt JA, Van Loon LC (1996) Systemic resistance in Arabidopsis induced by biocontrol bacteria is independent of salicylic acid accumulation and pathogenesis-related gene expression. Plant Cell 8:1225–1237
Pieterse CMJ, Zamioudis C, Berendsen RL, Weller DM, Van Wees SCM, Bakker PAHM (2014) Induced systemic resistance by beneficial microbes. Annu Rev Phytopathol 52:347–375
Pieterse CMJ, de Jonge R, Berendsen RL (2016) The soil-borne supremacy. Trends Plant Sci. doi:10.1016/j.tplants.2016.1001.1018
Platt A, Horton M, Huang YS, Li Y, Anastasio AE, Mulyati NW, Agren J, Bossdorf O, Byers D, Donohue K, Dunning M, Holub EB, Hudson A, Le Corre V, Loudet O, Roux F, Warthmann N, Weigel D, Rivero L, Scholl R, Nordborg M, Bergelson J, Borevitz JO (2010) The scale of population structure in Arabidopsis thaliana. PLoS Genet 6:e1000843
Raaijmakers JM, Weller DM (1998) Natural plant protection by 2,4-diacetylphloroglucinol-producing Pseudomonas spp. in take-all decline soils. Mol Plant-Microbe Interact 11:144–152
Ryu C-M, Farag MA, Hu CH, Reddy MS, Wei HX, Pare PW, Kloepper JW (2003) Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci USA 100:4927–4932
Seren U, Vilhjalmsson BJ, Horton MW, Meng DZ, Forai P, Huang YS, Long Q, Segura V, Nordborg M (2012) GWAPP: a web application for genome-wide association mapping in Arabidopsis. Plant Cell 24:4793–4805
Smith KP, Goodman RM (1999) Host variation for interactions with beneficial plant-associated microbes. Annu Rev Phytopathol 37:473–491
Sun JD, Gibson KM, Kiirats O, Okita TW, Edwards GE (2002) Interactions of nitrate and CO2 enrichment on growth, carbohydrates, and rubisco in arabidopsis starch mutants. Significance of starch and hexose. Plant Physiol 130:1573–1583
Ton J, Pieterse CMJ, Van Loon LC (1999) Identification of a locus in Arabidopsis controlling both the expression of rhizobacteria-mediated induced systemic resistance (ISR) and basal resistance against Pseudomonas syringae pv. tomato. Mol. Plant-Microbe Interact 12:911–918
Ton J, Davison S, Van Wees SCM, Van Loon LC, Pieterse CMJ (2001) The Arabidopsis ISR1 locus controlling rhizobacteria-mediated induced systemic resistance is involved in ethylene signaling. Plant Physiol 125:652–661
Van Peer R, Schippers B (1989) Plant growth responses to bacterization and rhizosphere microbial development in hydroponic cultures. Can J Microbiol 35:456–463
Van Peer R, Niemann GJ, Schippers B (1991) Induced resistance and phytoalexin accumulation in biological control of fusarium wilt of carnation by Pseudomonas sp. strain WCS417r. Phytopathology 81:728–734
Van Rhijn P, Vanderleyden J (1995) The Rhizobium-plant symbiosis. Microbiol Rev 59:124–142
Van Wees SCM, Van Pelt JA, Bakker PAHM, Pieterse CMJ (2013) Bioassays for assessing jasmonate-dependent defenses triggered by pathogens, herbivorous insects, or beneficial rhizobacteria. Methods Mol Biol 1011:35–49
Voll LM, Jamai A, Renne P, Voll H, McClung CR, Weber APM (2006) The photorespiratory Arabidopsis shm1 mutant is deficient in SHM1. Plant Physiol 140:59–66
Ward JK, Strain BR (1997) Effects of low and elevated CO2 partial pressure on growth and reproduction of Arabidopsis thaliana from different elevations. Plant Cell Environ 20:254–260
Waters MT, Wang P, Korkaric M, Capper RG, Saunders NJ, Langdale JA (2009) GLK transcription factors coordinate expression of the photosynthetic apparatus in Arabidopsis. Plant Cell 21:1109–1128
Yang J, Kloepper JW, Ryu CM (2009) Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci 14:1–4
Zamioudis C, Pieterse CMJ (2012) Modulation of host immunity by beneficial microbes. Mol Plant-Microbe Interact 25:139–150
Zamioudis C, Mastranesti P, Dhonukshe P, Blilou I, Pieterse CMJ (2013) Unraveling root developmental programs initiated by beneficial Pseudomonas spp. bacteria. Plant Physiol 162:304–318
Zamioudis C, Hanson J, Pieterse CMJ (2014) β-Glucosidase BGLU42 is a MYB72-dependent key regulator of rhizobacteria-induced systemic resistance and modulates iron deficiency responses in Arabidopsis roots. New Phytol 204:368–379
Zamioudis C, Korteland J, Van Pelt JA, Van Hamersveld M, Dombrowski N, Bai Y, Hanson J, Van Verk JC, Ling HQ, Schulze-Lefert P, Pieterse CMJ (2015) Rhizobacterial volatiles and photosynthesis-related signals coordinate MYB72 in Arabidopsis roots during onset of induced systemic resistance and iron deficiency responses. Plant J. 84:308–322
Zhang H, Sun Y, Xie X, Kim M-S, Dowd SE, Paré PW (2009) A soil bacterium regulates plant acquisition of iron via deficiency-inducible mechanisms. Plant J 58:568–577
Acknowledgments
The authors thank Joel Boerefijn, Silvia Coolen, Silvia Proietti, and Christos Zamioudis for assisting in the GWA analysis and the growth promotion assays. This work was supported by ERC Advanced Grant 269072 of the European Research Council.
Authors contribution
P. C. A. W. designed and performed experiments, analyzed the data, and wrote the manuscript. P. A. H. M. B. designed experiments and wrote the manuscript. C. M. J. P. designed experiments and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wintermans, P.C.A., Bakker, P.A.H.M. & Pieterse, C.M.J. Natural genetic variation in Arabidopsis for responsiveness to plant growth-promoting rhizobacteria. Plant Mol Biol 90, 623–634 (2016). https://doi.org/10.1007/s11103-016-0442-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0442-2