Abstract
The Enhancer of Zeste Polycomb group proteins, which are encoded by a small gene family in Arabidopsis thaliana, participate to the control of plant development. In the tomato (Solanum lycopersicum), these proteins are encoded by three genes (SlEZ1, SlEZ2 and SlEZ3) that display specific expression profiles. Using a gene specific RNAi strategy, we demonstrate that repression of SlEZ2 correlates with a general reduction of H3K27me3 levels, indicating that SlEZ2 is part of an active PRC2 complex. Reduction of SlEZ2 gene expression impacts the vegetative development of tomato plants, consistent with SlEZ2 having retained at least some of the functions of the Arabidopsis CURLY LEAF (CLF) protein. Notwithstanding, we observed significant differences between transgenic SlEZ2 RNAi tomato plants and Arabidopsis clf mutants. First, we found that reduced SlEZ2 expression has dramatic effects on tomato fruit development and ripening, functions not described in Arabidopsis for the CLF protein. In addition, repression of SlEZ2 has no significant effect on the flowering time or the control of flower organ identity, in contrast to the Arabidopsis clf mutation. Taken together, our results are consistent with a diversification of the function of CLF orthologues in plants, and indicate that although partly conserved amongst plants, the function of EZ proteins need to be newly investigated for non-model plants because they might have been recruited to specific developmental processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polycomb group (PcG) proteins were initially identified in Drosophila for their role in repressing the expression of homeotic genes (Orlando and Paro 1995; Grimaud et al. 2006). Since then PcG proteins have been identified in several organisms from Drosophila to plants and mammals, where they are involved in the long-term repression of gene expression (Köhler and Villar 2008). They contribute to specifying cell-specific expression patterns that are necessary for proper development, thereby determining a type of cell memory through consecutive cell divisions (Köhler and Hennig 2010).
PcG proteins function as multi-proteins complexes, known as Polycomb Repressive Complex 1 and 2 (PRC1 and PRC2). Although the precise molecular mechanisms responsible for gene repression by the PRCs are not fully understood, the classical view describes the following sequential events: (1) recruitment of the PRC2 at a given locus; (2) trimethylation of histone H3 on lysine 27 (H3K27me3) at this locus by the PRC2; (3) recognition of H3K27me3 by the PRC1; (4) monoubiquitination of the histone H2A by the PRC1 leading to a stable repression state (Wang et al. 2004). Recent studies suggest that this classical view may not correspond to a general mechanism. For example, the PRC1 can be recruited to some target genes in the absence of any H3K27me3 (Schoeftner et al. 2006; Tavares et al. 2012) and recruits the PRC2 itself (Yang et al. 2013; Molitor et al. 2014; Blackledge et al. 2014).
Contrary to PRC1 components (Molitor and Shen 2013; Kim and Sung 2014), proteins constituting PRC2s are highly conserved from animal to plant systems and have also been identified in unicellular algae (Shaver et al. 2010). In Drosophila, the PRC2 complex is composed of four core proteins encoded by unique genes: the Suppressor of Zeste (12) [Suz(12)], the Enhancer of Zeste [E(z)], the Extra Sex Comb [Esc] and the P55 protein. The methylation activity of the PRC2 complex is carried out by the E(z) protein which contains the SET domain characteristic of histone methyl transferases (Cao et al. 2002; Czermin et al. 2002; Kuzmichev et al. 2002; Müller et al. 2002). In A. thaliana, genetic and biochemical evidence suggest that at least three PRC2 complexes are present that differ by their composition in E(z) and Suz(12) paralogs (Luo et al. 2000; Spillane et al. 2000; Yadegari et al. 2000; Köhler et al. 2003; Chanvivattana et al. 2004; Katz et al. 2004; Wang et al. 2006; Wood et al. 2006; De Lucia et al. 2008).
The A. thaliana genome contains three genes encoding E(z) proteins, namely MEDEA, CURLY LEAF and SWINGER (Goodrich et al. 1997; Grossniklaus et al. 1998; Kiyosue et al. 1999; Luo et al. 1999; Chanvivattana et al. 2004), and three other genes, EMRYONIC FLOWER 2 (EMF2), VERNALIZATION 2 (VRN2), and FERTILISATION-INDEPENDENT SEED 2 (FIS 2) encode Suz(12) proteins (Luo et al. 1999; Gendall et al. 2001; Yoshida et al. 2001). Five genes encode p55 homologues, namely MULTISUPPRESSOR of IRA 1 to 5 (MSI1 to 5), only one of which has been shown to be part of PRC2, MSI1 (Derkacheva et al. 2013). On the other hand, ESC is encoded by a unique gene, FERTILIZATION INDEPENDENT ENDOSPERM (FIE) (Ach et al. 1997; Ohad et al. 1999; Hennig et al. 2003, 2005). Specific PRC2s control particular aspects of plant development (Mozgova et al. 2015; Xiao and Wagner 2015). For example, seed development is controlled by the MEDEA-FIS2-PRC2 (Chaudhury et al. 1997; Grossniklaus et al. 1998; Kiyosue et al. 1999; Ohad et al. 1999; Spillane et al. 2000; Yadegari et al. 2000; Köhler et al. 2003), transition from vegetative growth to flowering by the CLF/SWN-EMF2-PRC2 (Kinoshita et al. 2001; Yoshida et al. 2001; Chanvivattana et al. 2004; Jiang et al. 2008), and the induction of flowering in response to vernalization by a third PRC2, that contains CLF/SWN and VRN2 (Wood et al. 2006; De Lucia et al. 2008).
Although the understanding of PRC2 function is much less advanced in other plants, recent studies have outlined the conservation of PRC2 protagonists among the plant kingdom, suggesting a strong functional conservation during evolution. This has been evidenced by the partial complementation of fie mutations between A. thaliana and mosses (Mosquna et al. 2009). Components of PRC2 complexes, such as E(z), Suz(12) or Esc, have however been duplicated in many plants therefore generating paralogous genes (Danilevskaya et al. 2003; Butenko and Ohad 2011). The diversification of PRC2s in plants seems to be dependent on the species under study, as is their recruitment for the control of specific developmental processes. For example, the AtMEDEA and AtFIS2 genes are specific to A. thaliana and related species, where they are essential for the inhibition of central cell proliferation in the absence of fertilization (Spillane et al. 2007; Miyake et al. 2009). Other PcG proteins may fulfill this function in some other plant species like rice, as suggested by the analysis of rice transgenic RNAi lines where both OsFIE1 and OsFIE2 were down-regulated (Li et al. 2013). But this may be not a general fact, as in Hieracium pilosella there is no evidence for autonomous central cell proliferation after down regulation of the unique HFIE gene in the embryo sac (Rodrigues et al. 2008). In this context, the tomato plant, which originates from the Andean region in South America, forms fleshy fruits, a developmental process not found in the plant model A. thaliana (Moyle 2008). Hence, tomato has become a recognized model system to study fleshy fruit development and a wide range of genomic and genetic resources have been developed (Fei et al. 2011) that led to the recent release of the tomato genome (Consortium 2012). Although most studies on tomato plant and fruit development have relied on molecular genetics and metabolomics approaches (Giovannoni 2007), epigenetic mechanisms are emerging as important regulators of tomato fruit development, as illustrated by the identification of the Cnr epiallele responsible for the inhibition of fruit ripening caused by a high DNA methylation level in the CNR gene promoter region (Manning et al. 2006; Chen et al. 2015). Analysis of DNA methylation in tomato fruits showed tissue-specific changes in the overall DNA methylation levels and patterns during fruit development (Teyssier et al. 2008) and locus specific loss of DNA methylation during ripening (Zhong et al. 2013, Liu et al. 2015). Furthermore it has been recently shown that a DNA demethylase gene governs tomato fruit ripening (Liu et al. 2015). Beside DNA methylation, genes involved in histone regulation, such as the DE-ETIOLATED1 (DET1) gene, were also shown to play a role during fruit ripening (Benvenuto et al. 2002; Davuluri et al. 2004). To further characterize the contribution of histone modifications to the control of fruit development and quality, we have initiated the study of PRC2s in the tomato plant (How Kit et al. 2010). The tomato genome contains only one ESC and one Suz(12), but three E(z) genes. Two of them, SlEZ1 and SlEZ2, respectively orthologous to the A. thaliana SWINGER and CURLY LEAF genes, encode proteins targeted to the nucleus whereas the third gene, SlEZ3, was suggested to encode a truncated protein lacking the SET domain. Functional analysis of the SlEZ1 gene provided evidence that the corresponding protein is specifically involved in determining the carpel number of tomato flowers and in stamen development (How Kit et al. 2010).
In the present work we now show that in addition to SlEZ1 and SlEZ2, SlEZ3 is also likely to encode a functional E(z) protein. All three SlEZ genes are expressed in all organs analyzed, albeit with specific expression profiles. Since both SlEZ2 and SlEZ3 appear likely to have been generated following the duplication of a CLF-like ancestor gene, we have analyzed whether SlEZ2, which presents the highest homology level with AtCLF, is its functional homologue. Reduction of SlEZ2 gene expression resulted in a decrease in the H3K27me3 mark, consistent with SlEZ2 being involved in an active tomato PRC2 complex. RNAi plants displayed various phenotypes consistent with SlEZ2 having a pleiotropic role in this plant species. However, contrary to AtCLF in A. thaliana, SlEZ2 function is not restricted to plant growth, leaf formation, and flower development, but seems to have been recruited to new functions in fruit development and ripening.
Materials and methods
Plant materials and growth conditions
Germination of tomato seeds (Solanum lycopersicum, cv WVA106) was performed in vitro in a growth chamber with a day/night temperature of 25/21 °C and a 12 h/12 h day/night photoperiod. After germination in vitro, tomato plants were grown in glasshouse conditions in spring or autumn. Leaves at selected developmental stages were collected and immediately frozen in liquid nitrogen. Fruits were harvested at 5, 10, 20 days post anthesis (dpa), breaker (35dpa), orange and red ripe stages of fruit ripening. All fruits were measured and hand-dissected immediately after harvest in order to separate seeds and locular tissue from pericarp and columella. Pericarp and columella were frozen in liquid nitrogen and stored at −80 °C until processed. Seeds were separated from the locular tissue by using a nylon mesh, dried and stored at 4 °C. Stamens, petals, sepals, styles, and ovary structures, collected from open and closed flowers, were hand dissected and observed using a Leica binocular MZFLIII microscope equipped with a Leica DC300F Camera.
Transgenic plant regeneration
To specifically repress SlEZ2 gene expression, an RNAi construct was generated using the 3’ untranslated region (UTR) of the SlEZ2 gene cloned in sense and antisense orientations in plasmid pK7GWIWG2 (I) (http://gateway.psb.ugent.be/vector/show/pK7GWIWG2). For SlEZ2 promoter analysis, a 2633-bp fragment of the SlEZ2 gene promoter region was inserted upstream of the EGFP and GUS reporter genes carried by the binary vector pKGWFS7,0. These two recombinant plasmids named, respectively, pK7GWEZ2 and pKGWFSEZ2, were independently introduced in the Agrobacterium tumefaciens strain GV3101.
Subsequently, tomato cotyledon transformation was performed as described in Gonzalez et al. (2007). Eight in vitro kanamycin resistant SlEZ2 RNAi T0 plants were selected from independent calli and cultured as described in How Kit et al. (2010). After transfer to greenhouse, six transgenic plants could grow (namely 2–4, 2–5, 2–6, 2–9, 2–11 and 2–14) and generated T1 seeds. Genomic PCR was used to confirm that the six remaining lines were transgenic. We classified the T1 plants based on the level of SlEZ2 expression, as determined by semi-quantitative RT-PCR on total leaf RNA. Plants from each group that displayed a 3:1 ratio on kanamycin, indicative of a single locus insertion, were selected for further study. Ten T1 plants were grown to obtain seeds. Homozygous T1 plants were identified on kanamycin plates (100 % of kanamycin resistant T2 seeds) for lines 2–9 and 2–5. Analyses were performed on T2 seeds obtained from a given homozygous T1 plant selected based on their residual SlEZ2 expression levels. In the case of line 2–11 and 2–14, T1 hemizygous plants were obtained, and T2 plants were therefore analyzed to determine whether they were hemizygous or homozygous.
Two independent lines transformed with pKGWFSEZ2 were selected and T2 plants were used for GUS staining.
Histochemical localization of GUS activity in tomato plants
Localization of GUS activity was performed essentially as described in Gallusci et al. (1994). Freshly harvested plant material (leaf, flower, fruits at various developmental stages) were used un-dissected (leaf, flower, fruit) or hand-sectioned with a razor blade to expose internal tissues (flowers, fruits). Tomato samples were immerged in staining buffer (Gallusci et al. 1994) and vacuum infiltrated for 3 min (3 × 1 min). The reactions were carried out at 37 °C for 12 h. Samples were washed and cleared in 70 % (v/v) ethanol at room temperature to remove pigments. Observations were performed using a Leica binocular MZFLIII microscope equipped with a Leica DC300F Camera.
RNA extraction and RT-PCR analysis
Frozen leaf and fruit tissues were ground to powder in liquid nitrogen. Total RNA were isolated from 100 mg of powder using Trizol reagent (Invitrogen) followed by a turbo DNase (Ambion) treatment according to manufacturer’s recommendations. RNA quality was controlled by electrophoresis on 1 % agarose gels and RNA was quantified using a Nanovue (GE Healthcare) spectrophotometer. To control for genomic DNA contamination, quantitative PCR was performed on total RNA using Ef1α primers (Table S2). One µg of total RNA were reverse transcribed with Improm II enzyme (Promega) using oligo d(T) primers according to manufacturer’s instructions.
Semi-quantitative RT-PCR analyses were performed as described in How Kit et al. (2010), and PCR products were analyzed on 1.5 % agarose gels.
Real time RT-PCR analysis was performed using the Biorad CFX C1000TM Thermal Cycler (CFX96™ Real-Time system). Primers were designed to amplify fragments of approximately 100 bp (Table S2). PCR reactions were performed using IQ™ SYBR Green Supermix 2X according to manufacturer’s instructions (Biorad) in 96 well plates (BIORAD). Typical reactions contained 4 pmol of each primer, 4 µL of ten times diluted RT reactions (corresponding to 20 ng of total RNA), and 10 µL of SYBR Green Supermix (2X) in a final volume of 20 µL. Amplifications of EF1 and ACTIN fragments were used as controls for normalization. For each gene, differences between samples were calculated using the ΔΔCt method. Each PCR analysis was performed in triplicate using cDNA synthesized from two to four independent RNA extractions each prepared from a minimum of three independent biological replicates.
Western blot analysis
Equal amounts of nuclear protein extracts proteins (10 µg) (Gendrel et al. 2005), were separated on a 12 % SDS/PAGE and transferred to a nitrocellulose membrane (Millipore). Histone levels were determined using anti-H3 (Millipore 07-690), anti-H3K27me3 (Millipore 07-449) and HRP conjugated goat anti-rabbit IgG antibodies (Agrisera AS09602). The chemiluminescent signals were detected with the ECL Prime Western Blotting Detection Reagent (Amersham) using the ChemiDoc™ MP Imaging System (Bio-Rad). The quantification was performed with the Image Lab Software (Bio-Rad). H3K27me3, H3 and secondary antibodies were diluted, respectively, 1/2500th, 1/50,000th, and 1/2500th fold.
Cutin monomer and wax analysis
Cuticular waxes were extracted by immersing fruits for 30 s in chloroform containing docosane as internal standard. Extracts were dried under nitrogen flux and lipids were derivatized and analyzed as previously described (Bourdenx et al. 2011).
For the cutin monomer analysis, 1 cm diameter fruit epidermis discs were cut off, carefully scratched with a scalpel blade in order to remove exocarp cells, and incubated 30 min in hot isopropanol at 85 °C. After cooling, samples were extensively delipidated by extracting the soluble lipids, then dried and depolymerized as described (Domergue et al. 2010). Extraction, derivatization and analysis were perfomed as previously described (Domergue et al. 2010).
Results
The tomato genome contains three E(z) genes that display specific expression patterns
In a previous study, we demonstrated that in addition to SlEZ1 and SlEZ2, respectively orthologous to the SWN and CLF genes of A. thaliana, the tomato genome contains SlEZ3, a third E(z) gene also orthologous to CLF, with a stop codon interrupting the coding sequence at position 1104 (How Kit et al. 2010). Screening of an S. peruvianum EST library (http://solgenomics.net/) allowed the identification of a full length SlEZ3 mRNA (VO2SLM0055024.2) that lacks this stop codon. Re-cloning of SlEZ3 cDNAs from S. lycopsersicum (WVA106) fruits and leaves RNA revealed that this gene generates three distinct mRNAs, one of them potentially encoding a protein with all characteristics of functional E(z) proteins (Fig S1).
To detect possible overlaps between the expression patterns of all tomato SlEZ genes, the expression levels of each gene was analyzed in tomato leaves, flowers, apex, stem and fruits using real time RT-PCR. As shown in Fig. 1, SlEZ1, SlEZ2 and SlEZ3 are expressed in all tomato plant organs tested, albeit with different expression profiles. Consistent with previous analyses, SlEZ1 mRNA accumulates in all organs and is most abundant in flowers and in expanded leaves (How Kit et al. 2010). Low expression levels were detected in stems and stem apices, with a 4 fold reduction as compared to closed flowers. It is noteworthy that SlEZ1 expression remains stable in the pericarp of developing fruits, with expression levels in the same range as found in leaves and open flowers (Fig. 1A). This contrasts with SlEZ2 mRNA abundance, which decreases during organ development (Fig. 1B). SlEZ2 mRNA is therefore abundant in young fruit pericarp but becomes barely detectable during fruit ripening. A similar observation was made during leaf development. Significant variations were also observed between organs with a very high expression level, observed in closed and open flowers, as compared to fruits. Finally, SlEZ3 mRNA displays a pattern of accumulation in leaves similar to SlEZ1 mRNA, characterized by an increase during leaf expansion (Fig. 1C). SlEZ3 transcript abundance decreases during the period of fruit growth corresponding to the cell expansion phase before increasing again as fruits ripen.
Abundance of SlEZ mRNA in different tomato plant organs. RT-QPCR analysis performed as described in the methods using primers for SlEZ1 (A), SlEZ2 (B), SlEZ3 (the longest transcript shown in Fig. S1), (C) and ACTIN and EF1α as control genes (table S1). Relative quantification has been performed using the ΔΔCt method. Leaves are numbered relative to the plant apex: leaves 3 and 4 (L3-4) were pooled and correspond to young expanding leaves, leaves 8 (L8) were fully expanded and leaves 16 (L16) were old leaves but not senescing. CF: Closed Flowers 2 days before anthesis; OF: Open Flowers at anthesis; A: cauline Apex; S: Stem; 5dpa, 10dpa, 20dpa, B, O and RR: fruit pericarp (P) hand dissected respectively at 5, 10 and 20 days post anthesis (dpa), breaker (B), orange (O) and red ripe (RR) stages. For each gene, values have been reported to gene expression level in leaves 3–4 that represents the 100 %. Data are means of RT-QPCR experiments performed on three independent biological replicates analyzed as described in the methods. For each gene, student tests were performed using as reference either 3–4 leaves (black stars) or 5dpa fruits (grey stars), *p < 0.05, **p < 0.01, ***p < 0.001
Taken together these results indicate that SlEZ genes follow contrasting expression patterns, consistent with the hypothesis that tomato E(z) proteins perform specific functions during tomato plant and organ development. The low expression level of SlEZ3, together with the numerous changes observed in the structure of this gene as compared to SlEZ2 (Fig S1) would suggest that the latter is the functional homologue to the Arabidopsis CLF gene.
In planta SlEZ2 gene expression patterns
To analyse SlEZ2 tissue-specific expression pattern, a transcriptional fusion between a 2633 bp SlEZ2 promoter (pSlEZ2) fragment and the GUS reporter gene was generated in the vector pKGWFS7. Among the transgenic lines obtained, two were shown to have a single transgene insertion and were selected for further study. Consistent with RT-PCR analysis (Fig. 1 and How Kit et al. 2010), the SlEZ2 promoter region was found to direct GUS expression in all tested organs (Fig. 2). In open flowers, the GUS staining was distributed uniformly in the sepals but was restricted to the petal margins (Fig. 2C, D) and to the adaxial part of stamens (Fig. 2A). The different parts of the pistil were intensely stained: the whole ovary (Fig. 2E), the stigmata, and the style (basal and apical parts together with the transmitting tract) (Fig. 2B). Such a pattern is consistent with in situ hybridization results described in How Kit el al (2010) showing the accumulation of SlEZ2 mRNA in most part of the flower and indicate that the promoter fragment used here is sufficient for proper SlEZ2 gene expression. The pSlEZ2-driven GUS staining was also found in fruits at all developmental stages, with a progressive reduction of staining intensity as fruit developed (Fig. 2F, G), consistent with the reduction in SlEZ2 mRNA accumulation observed in the pericarp of developing tomato fruits (Fig. 1). All fruit tissues were stained, including the seeds. Finally in young developing leaves (Fig. 2H), the blue color was not evenly distributed but was mainly located at leaf margins corresponding to the sites of lobe formation.
Histochemical localization of GUS activity in T2 tomato plants transformed with a pSlEZ2::GUS construct. Tomato tissues have been harvested and treated as described in “Materials and methods”. Fully opened flowers were harvested at anthesis and hand dissected after GUS staining except for carpel coloration: stamen (A), style (B), sepal (C), petal (D) and carpel (E). Immature green fruits (F, 20 dpa), ripening fruits (G, orange stage) and young leaf (H) are shown. Carpel and fruits were cut before staining to allow efficient diffusion of the staining solution
In conclusion, SlEZ2 is expressed in most plant organs, but displays cell-type specific expression in leaves, petals and style. On the other hand, young fruits and sepals were uniformly stained.
The tomato SlEZ2 protein belongs to an active PRC2 complex leading to trimethylation of H3K27
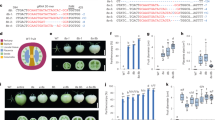
To study the function of SlEZ2 in tomato, we used a gene specific RNAi approach designed to specifically target the SlEZ2 mRNA (“Materials and methods”). Eight primary transformants (T0) were selected from independent calli, among which only six were viable. Indeed, two T0 plants were lost after transfer to the greenhouse following arrest of the shoot apical meristem. Residual SlEZ2 expression was subsequently measured in T0 and T1 plants, and lines 2–9, 2–11 and 2–14, that displayed the lowest SlEZ2 mRNA levels, were selected for further studies. All subsequent experiments were performed on T2 plants derived from single T1 parents that were either homozygous (T1 plants 2–11, 2–9) or hemizygous (T1 plants 2-14) for the transgene (“Materials and methods”). Non transgenic plants were used as controls. In addition RNAi plants affected in the expression of SlEZ1 only (How kit et al. 2010) were grown at the same time as SlEZ2 RNAi plants, and were used to verify that developmental alterations of SlEZ2 RNAi plants were specific to these plants. Expression levels of SlEZ2 were reduced in leaves of all T2 transgenic plants with residual expression levels ranging from 47 to 52 %, as compared to WT leaves of the same age (Fig. 3A). SlEZ1 and SlEZ3 gene expression remained essentially unchanged (Fig. 3B, C).
SlEZ gene expression in SlEZ2 RNAi transgenic tomato plants (A–C) and H3K27me3 mark abundance (D). (A–C) RT QPCR analysis were performed as detailed in the methods using RNA samples prepared from fully expanded leaves (12th leaf from apex) of WT and transgenic plants of lines 2–9, 2–11 and 2–14 and primers specific to SlEZ2 (A), SlEZ1 (B) and SlEZ3 (C) genes (Table S1 supplementary data) as indicated. Relative quantification was performed using the ΔΔCt method. For each gene, expression values are normalized to null transformed wild-type (WT) control plants (100 %). Data ± sd are means of three independent biological replicates (three different T2 plants for each line) performed using two independent RNA preparations obtained from three different cultures (A–C). Student’s t-tests indicate significant difference at 0.05 (*), 0.01 (**) or 0.001 (***). (D) Abundance of trimethylation of lysine 27 of histone 3 (H3K27me3) and histone 3 (H3) are quantified by western blot analysis using the same plant samples as in (A–C). Values represent the ratio between signals obtained with H3 and H3K27me3 antibodies quantitated with Chemi-Capt software using the Western blot shown here. An independant Western blot with equal amounts of nuclear porteins isolated from leaf samples of WT and transgenic plants of lines 2–9, 2–11 and 2–14 are shown in Fig S2
In Arabidopsis leaves, it has been established that the partial impairment of PRC2 function results in a global decrease in the abundance of the histone H3 lysine 27 trimethylation (H3K27me3) mark (Katz et al. 2004; Lafos et al. 2011). Western blot analyses of leaf nuclear proteins showed a reduction ranging from 30 to 50 % in H3K27me3 abundance in the three tested tomato T2 transgenic plants compared to WT (Fig. 3D, Fig S2), consistent with a partial loss of PRC2 activity in the SlEZ2 RNAi tomato plants.
Plants with reduced SlEZ2 gene expression are affected in several aspects of vegetative development
Although limited, the reduction in SlEZ2 gene expression (Fig. 3) resulted in several alterations during tomato plant vegetative development (Fig. 4). As mentioned above, strong effects were already observed in T0 plants, some of them developing only a few leaves prior to meristem arrest. A similar phenotype was found in T2 plants from line 2–11. Out of 20 germinated T2 plants from line 2–11, half developed two to three leaves before meristem arrest (Fig. 4A), which in most cases resulted in plant death shortly after germination. Occasionally, a secondary meristem was initiated (Fig. 4B) that led to normal plant development. In addition, all T2 RNAi plants were characterized by a significant size reduction that ranged between 40 and 74 % (Fig. 4C and Table S1). Size reduction was due to a decreased number and shorter internodes (Table S1). Occasionally, fasciated stems formed on secondary axes (Fig. 4D). Leaves of transgenic plants displayed a reduction in the number of secondary and intercalary leaflets (Fig. 4E, F and Fig. S2a (Jasinski et al. 2008)). Variability in leaf phenotype intensity was observed between lines and occasionally between plants of the same line, the most severe phenotypes being observed on lines 2–11 and 2–14 (Fig. S2b). Leaflet shape was also affected. Primary leaflet serration was reduced although not abolished and secondary leaflets, characterized by an asymmetrical growth of the abaxial versus adaxial surface, became curly and crinkled and eventually fused to the leaf axe (Fig. 4I–N) Although such modified leaflets were occasionally observed in WT plants, their frequency increased significantly in SlEZ2 silenced lines (Fig S3). Altogether, these results demonstrate that SlEZ2 knock-down leads to pleiotropic effects on tomato plant vegetative development.
Abnormal stem and growth phenotypes displayed by SlEZ2 silenced plants. (A) Three weeks old plantlet of line 2–11 showing meristem arrest. (B) Five weeks old plantlet of line 2–11 that initiates a secondary meristem (white arrow) after primary meristem arrest. (C) Representative plant of line 2–11 (left) displaying a reduced growth 52 days after germination in comparison to control plant (right). (D) Fasciated stem on a T2 plant from line 2–9. Leaves from WT (E) and T2 transgenic (F) plants. Primary leaflets from WT (G) and a T2 transgenic (line 2–11) (H) plant showing reduced serration. Secondary leaflets from WT (I) and transgenic lines (J–N). Secondary leaflets from the transgenic plants show asymmetric growth between the adaxial and abaxial surfaces leading to crinkled surfaces (J–N) which are sometimes fused to the axe (N). Bars: 5 cm (E–F), 1 cm (G–N)
Studies in A. thaliana described that a lack of PCR2 activity leads to upregulation of two MADS-box (AtAG and AtSHP1) and two class I KNOX genes (AtKNAT2 and AtSTM) in leaves (Katz et al. 2004; Lafos et al. 2011). The tomato genes SlTAG1, SlTAGL1, SlTKN4 and LeT6, respectively orthologous to AtAG, AtSHP1, AtKNAT2 and AtSTM were analyzed in a gene targeted approach to measure their expression level in fully expanded tomato leaves when these genes are normally weakly expressed. None of these genes were consistently upregulated in the transgenic lines under study. In contrast to the expected results, SlTAG1 was consistently down regulated in transgenic leaves as compared to WT leaves of the same age (Fig S4) which may suggest an indirect regulation of this gene by SlEZ2.
SlEZ2 down regulation affects flower and fruit development
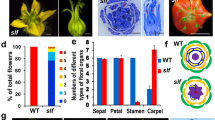
Contrary to WT flowers characterized by a style enclosed within the stamen cone (Fig. 5A), transgenic plants from all generations tested (T0, T1 and T2) developed abnormally shaped flowers with twisted stamens leading to style exposition (Fig. 5B, E–H), outgrowths on petals and stamens (Fig. 5B–D), style enlargement (Fig. 5I, J), altered sepal shape (Fig. 5K, L), and occasionally sepals with extra lobes (Fig. 5M). However, neither stamen number and identity nor pollen production and fertility were modified. Although regularly observed the frequency of these phenotypes varied between lines (Fig. S5).
Flower phenotypes. (A) Wild type (WT) open flower with closed staminal cone; (B) abnormal flower from line 2–9 showing slightly twisted stamens (arrow head) and serrated petals with ectopic organs (white arrows); ectopic outgrowth formed on petals (left WT, right transgenic) (C) or stamens (D). Stamens from WT (left) and transgenic flower (right) (E); compared to WT control flowers (F), transgenic stamens are slightly (G) or strongly (H) modified. (I) Styles of T2 RNAi plant flowers (right) are larger than WT styles (left) and (J) eventually fasciated. Transgenic sepals are larger (left) than wild type sepal (right) (K), have modified shape (left and middle: transgenic; right: wild type,) (L), or present extra lobes (M)
Fruit set from transgenic lines 2–9, 2–11 and 2–14, was not as high as in the WT, as revealed by the analysis of flower abortion rate (Fig. S4). Moreover the increased frequency of flower abortion in the SlEZ2 RNAi plants was strictly correlated with the increase in the percentage of abnormal flowers most likely due to a decrease in self-pollination efficiency leading to fruit development abortion (Fig. S4). Alternatively the decrease in transgenic fruit set could reveal a function for SlEZ2 in fruit set and/or during early fruit development. In order to discriminate between these two possibilities, manual pollination of transgenic flowers with WT pollen was performed, leading to transgenic fruits with increased seed content that grew similarly to the WT controls (Fig. S5). Thus, providing WT pollen restored seed set and thereby fruit development indicating that the transgene per se did not affect fruit set and growth kinetic.
Fruits from lines 2–9, 2–11 and 2–14 either selfed or back-crossed, were different from WT fruits in several other aspects, including their shape, texture, color, and surface aspect (Fig. 6). Transgenic fruits were flat and much softer when fully ripe than the spherical WT fruits (Fig. 6A, B). In some cases, fruit resistance could not even be measured, such as those produced by plants from line 2–11. The color of transgenic fruits was also different from WT fruits of the same age. Hence, immature green fruits from the transgenic plants were light green to white, with some irregular green dots whereas those from WT plants were largely green with some regular light slightly yellow streaks. Also the mature fruits from both WT and transgenic plants were largely red, although red color intensity was often reduced in transgenic fruits consistent with their slightly reduced lycopene content (Fig S5). In addition, as transgenic fruits appeared brighter than WT (Fig. 7A, Fig S7), showed orange dots on their surface and shriveled more rapidly than WT when left overripe, we also analyzed their surface (Fig. 6, Fig S7). Contrary to WT fruits, transgenic SlEZ2 RNAi fruits are sticky and present a higher trichome density at maturity as compared to WT fruits, as revealed by binocular microscopy observations (Fig. 7B). In addition, fresh pealed outer epidermis observed under optical microscope appeared smoother than that of wild-type fruits. Surface irregularities due to the cutinized epidermal cell walls which separates two adjacent epidermal cells, were large and clearly visible on wild type fruits, but were thinner and hardly visible on transgenic fruits (Fig. 7C). consistent with an alteration of their cutin content (Petit et al. 2014). This was confirmed by measuring major components of epicuticular waxes and cutin monomers in transgenic fruits. We found a significant decrease in four cutin acid monomers (p-coumaric acid, hexadecane-1,16-dioic acid, 16-OH hexadecanoic acidand 10,16-diOH hexeadecanoic acid) in transgenic fruits as compared to WT fruits (Fig. 7D). In contrast, transgenic fruits contain significantly more C32 alkane and C31 iso-alkane, two major components of the epicuticular waxes, than the wild type (Fig. 7D). Finally all lines occasionally developed fruits with extra carpels (Fig. 6C–H) that developed internally to the sepals. In addition to carpels, ectopic flowers and/or leaves were observed (Fig. 6G, H).
A Atypical fruits from SlEZ2RNAi plants. 1 Transgenic fruits obtained after self-pollination of plants from line 2–9 (upper) and WT (lower) harvested at 20dpa. 2 Fruits from WT plants (lower) and fruits from line 2–9 (upper) at the red ripe stage obtained after backcrossing transgenic flowers with WT pollen. Bars: 1 cm; (B) Measurement of fruit pericarp softening with a penetrometer. Student statistic test shows significant difference at 0.05 (*), 0.01 (**) or 0.001 (***). (C) Ectopic organ development on SlEZ2 RNAi plants. Fruits with additional carpel from line 2–9 (1–3, 5), 2–11 (4) and 2–14 (6). Extra carpel number varied from two (2) to more than 10 carpels (1, 3, 4) that developed internally giving a “fruit in a fruit” phenotype (1), were fused to fruits (2) or remained separated from each other (3) or from the main fruit (4). Extra carpels developed concomitantly to the main fruit (2–3) or their formation was delayed, which resulted in young developing carpels when fruits were already mature (4). Ectopic leaves (5) and flowers (6) development in addition to abnormal carpels
Surface analysis of red ripe fruits. (A–C) and quantification of the major cutin acid monomers and wax aliphatic components (D). (A) Fruits from WT (upper) and from line 2–9 (lower) showing differences in color and brightness, (B) fruit surface viewed at a shallow angle with a stereomicroscope, and (C) fresh pealed outer epidermis observed under optical microscope. Scale bar 1 cm in (A), 2 mm in (B) and 30 µm in (C). (D) Quantifications (µg of components by cm2 of fruit fresh weight) were performed with 3 fruits of 3 transgenic plants for each line. All these fruits presented altered color and texture. Major components of cutin and epicuticular waxes are respectively 10, 16-diOH hexadecanoic acid (10,16-diOH C16:0), p-coumaric acid (Coumaric acid), hexadecane-1,16-dioic acid (C16:0 DCA), 16-OH hexadecanoic acid (16-OH C16:0), 9,10,18-triOH octadecanoic acid (9,10,18-triOH C18:0) and C31 iso-alkane (iso-ALK31), C31 alkane (ALK31), C32 alkane (ALK32). Student statistic test shows significant difference at 0.05 (*), 0.01 (**) or 0.001 (***)
In summary, SlEZ2 gene repression leads to modifications in flower morphology, as well as various alteration of fruit development, including control of carpel initiation, fruit development and ripening, and modification of fruit cuticle formation.
Discussion
E(z) genes in tomato
Polycomb proteins have been intensively studied in the model plant A. thaliana, leading to the demonstration that they constitute three different PRC2 complexes which are essential for developmental transitions during plant development (Köhler and Villar 2008; Holec and Berger Holec 2012). Notwithstanding, study of the specific function of Arabidopsis E(z) proteins has proven to be challenging because they are encoded by three closely related genes with partly redundant functions: SWN, CLF and MEA (Hennig and Derkacheva 2009). MEDEA is specifically involved in controlling the central cell division in the female gametophyte and early seed development (Grossniklaus et al. 1998) whereas the CLF protein is essential for Arabidopsis vegetative development (Goodrich et al. 1997). However, the functional redundancy between SWN and CLF, revealed by the severe defects of clf swn double mutants, masks their specific functions (Chanvivattana et al. 2004; Schubert et al. 2005).
As in A. thaliana (Butenko and Ohad 2011), tomato E(z) proteins are encoded by a multigenic family composed of three genes (How Kit et al. 2010). Notwithstanding, the situation differs markedly between these two plant species. Firstly, in tomato the CLF gene has been duplicated to give SlEZ2/SlCLF1 and SlEZ3/SlCLF2, whereas SWN has not. Hence, SlEZ1/SlSWN is unique in tomato and there is no gene orthologous to MEDEA, a gene paralogous to SWN in Arabidopsis (Spillane et al. 2007). Finally, whereas Arabidopsis swn mutants have no phenotype, SlEZ1 knock-down in tomato impacts stamen development and carpel number indicating that the functional balance of individual E(z) proteins has diverged between these two plants (How Kit et al. 2010).
SlEZ2 is a functional E(z) protein
To determine to what extent SlEZ2 has functions in tomato plants similar to AtCLF in Arabidopsis, RNAi plants that are specifically affected in the expression of the SlEZ2 gene have been generated. Surprisingly we failed to obtain plants with a residual SlEZ2 gene expression below 40 %. This is significantly higher than the remaining SlEZ1 mRNA level observed in SlEZ1 RNAi lines that ranged between 20 and 40 % (How Kit et al. 2010) and suggests that SlEZ2 may have essential functions in tomato. Indeed, the global level of the H3K27me3 mark was reduced in SlEZ2 RNAi plants compared to WT, to an extent comparable to the reduction of H3K27me3 observed in the Arabidopsis clf-28 loss of function mutant (Lafos et al. 2011). This result indicates that SlEZ2 is a functional E(z) protein which cannot be fully complemented by either of the two other SlEZ proteins. This implies that SlEZ3 has rapidly evolved since the gene duplication event that generated SlEZ2 and SlEZ3, consistent with the significant modification of the SlEZ3 gene structure compared to SlEZ2 and AtCLF and the identification both in leaves and fruits of multiple SlEZ3 RNA forms (Fig S1) as was described for its petunia counterpart PhCLF1 (Mayama et al. 2003). The absence of full complementation of SlEZ2 functions by other SlEZ genes is also consistent with the observation that SlEZ2 displays an expression pattern distinct from SlEZ1 and SlEZ3, being characterized by a high expression level in young developing tissues that decreases in mature organs. GUS staining further indicated that SlEZ2 gene expression in leaves is restricted to margins, and is highly expressed in flowers and young fruits, as previously observed using in situ hybridization (How Kit et al. 2010), both of which are tissues containing actively dividing cells (Hagemann and Gleissberg 1996; Berger et al. 2009). This is consistent with the gene being involved in the early reprogramming of chromatin states during organogenesis.
SlEZ2 is the tomato functional ortholog to CLF
Similar to the clf mutation in Arabidopsis, SlEZ2 knock down leads to phenotypes that affect several aspects of plant development. These include plant size reduction and shorter internodes, flower development and leaf shape alterations, all of which are consistent with the idea that SlEZ2 is the functional ortholog to AtCLF. Yet there are many phenotypic differences between tomato SlEZ2 RNAi plants and Arabidopsis clf mutants. For example, leaves from SlEZ2 RNAi plants display decreased complexity and crinkled leaflets but are never curly contrary to the leaves of Arabidopsis clf mutants. In a similar way, SlEZ2 RNAi flowers are characterized by twisted stamens and outgrowths on petals whereas flowers from Arabidopsis clf mutants lack petals and have staminoid petals. Another difference corresponds to the absence of effect on flowering time contrary to Arabidopsis clf mutants that flower 3 weeks earlier than WT plants under short days conditions (Goodrich et al. 1997).
Although such phenotypic differences may reflect a diversification of SlEZ2 function as compared to AtCLF, other reasons could account for these observations. Firstly, SlEZ2 RNAi plants still accumulate SlEZ2 mRNA (60 % of WT level) and most likely SlEZ2 proteins, whereas null clf mutants produce no functional CLF protein and are therefore likely to display more pronounced phenotypes (Goodrich et al. 1997). Secondly, the E(z) gene family composition differs between Arabidopsis and tomato. As a consequence, eventual functional complementation between SlEZ genes may lead to situations distinct from Arabidopsis. Indeed, our previous results suggest that the functional balance between SlEZ1/SlSWN and SlEZ2/SlCLF has diverged between tomato and Arabidopsis (How Kit et al. 2010). However, we cannot formally rule out a partial complementation of SlEZ2 by SlEZ1, which would be reminiscent of the situation already described in Arabidopsis between SWN and CLF (Chanvivattana et al. 2004). Finally SlEZ3, a protein not found in Arabidopsis, might also be able to partly complement SlEZ2 function, thereby also contributing to the phenotypic differences between Arabidopsis clf mutants and tomato SlEZ2 RNAi plants.
SlEZ2 has acquired new functions in tomato
We also found that SlEZ2 RNAi plants produced fruits with developmental alterations. Such phenotypes were never observed in Arabidopis clf mutants or in Arabidopsis plants impaired for other PRC2s (Katz et al. 2004). Interestingly there has been no clear description of the role of PcG in Arabidopsis fruits with the exception of FIE suppressed plants, which are characterized by multi-carpel gynoecia (Katz et al. 2004) and clf mutants which were reported to occasionally produce flowers with unfused carpels (Goodrich et al. 1997). Given the very specific alterations of fleshy fruit phenotypes, these results suggest a specific recruitment of SlEZ2 in fruit development, a function not identified for the Arabidopsis CLF protein. Most notably, tomato fruits from SlEZ2 RNAi plants showed modified shapes, texture, and color. Interestingly fruit shape and texture depend on events occurring early during fruit development (Chaïb et al. 2007; van der Knaap et al. 2014), concomitant to the highest expression level of SlEZ2 (Figs. 1 and 2, How Kit et al. 2010). In addition, changes in the surface of SlEZ2 RNAi fruits were correlated to low cutin content and a high trichome density, suggesting a role for SlEZ2 in the control of tomato fruit epidermis cell identity. This is reminiscent to a recently identified function of Arabidopsis PRC2s that were suggested to control guard cell fate stability by repressing stomatal stem cell genes in cotyledons (Lee et al. 2014).
However, effects on fruits trichome could also be indirect due to the altered cuticule composition. In plants, cuticle formation is an important epidermal property and all aerial epidermal cells produce cuticle (Javelle et al., 2011). Although no direct link between epidermal cell fate specification and cuticle biosynthesis has been made, several Arabidopsis lines with mutations in genes involved in cutin biogenesis show severe defects in cuticle composition, together with abnormalities in the development of the epidermis. For instance, knock-down mutations in genes of the fatty elongation complex (AtKCR-RNAi lines or pas2-1 mutant) display strong developmental defects, such as spontaneous organ fusions and abnormal epidermal cell morphology (Faure et al. 1998; Beaudoin et al. 2009). Mutations in the ABCG11 transporter which is involved in cutin precursor export lead to numerous organ fusion events, development of asymmetric stomata, shorter trichomes with irregular branching or collapsed trichomes (Bird et al. 2007; Panikashvili et al. 2007). These data together with others hint at a complex cross-talk between cuticle formation and epidermis differentiation, but also underlie the difficulty of functionally separating both.
Contrary to this study, the functional analysis of SlEZ1 had not revealed any role for this protein in tomato fruit. Knocking down FIE, a partner of the EZ proteins in PRC2 complexes, resulted in tomato lines characterized by increased sepal and petal numbers in flowers, fused ovules and pistils, and parthenocarpic fruit formation (Liu et al. 2012). Although Liu et al. (2012) demonstrated that SlFIE could interact with SlEZ2, the phenotypes described in that study are different from those of SlEZ2 RNAi plants. As FIE is encoded by a unique gene in tomato, knockdown of this gene may have a stronger effect than SlEZ2 as the corresponding protein is expected to be present in all PRC2 complexes.
Noteworthy, although several genes orthologous to CLF target genes in Arabidopsis have been analyzed, none of them was found to be consistently upregulated in the tomato lines analyzed in this work (Fig. S4). Indeed, SlEZ2 gene repression was limited to 40 % and had a significant but limited impact on the global level of H3K27me3 abundance. This in turn may results in transient and/or weak effects on gene expression. Consistent with this view, the most affected transgenic plants were not viable in our conditions, and all studies were therefore performed on plants presenting rather mild phenotypes. Several other reasons could explain such a result: the morphology of SlEZ2 RNAi leaves was not observed in any tomato single mutant described in the literature. Hence the identification of the genes whose expression is deregulated in the transgenic tomato plant leaves and in a more general way flowers and fruits is not simple, and will require transcriptomic studies and genome wide analysis of H3K27me3 distribution. Furthermore, the gene deregulation may be restricted to a small number of cells during a short developmental window. It has now been shown that tomato leaf morphogenesis involves complex regulatory genes networks implying dynamic spatial and temporal gene activity (Bar and Ori 2014). Finally, only a limited number of H3K27me3 targets are mis-expressed in Arabidopsis PcG mutants (Weinhofer et al. 2010; Bouyer et al. 2011; Farrona et al. 2011; Lafos et al. 2011), suggesting that in most cases their deregulation also depends on the presence of other regulators, and/or epigenetic marks.
Conclusion
Altogether the functional analysis of SlEZ2 indicates that this gene has retained most of the ancestral functions of the tomato CLF like gene that generated SlEZ2 and SlEZ3. However, our results also demonstrate that SlEZ2 is likely to have additional functions in fleshly fruits and support the idea that E(z) proteins have been recruited to specific processes during the evolution of land plants (Butenko and Ohad 2011). So far, the results described here are consistent with SlEZ2 participating to a functional PRC2 complex. Indeed other tomato PRC2 components need to be identified, but as the tomato genome contains only one ESC like and one Su(z)12 like encoding gene, it is likely that both proteins will contribute to all tomato PRC2 complexes. Tomato PRC2s could be defined by their E(z) component, contrary to A. thaliana where the specificity of PRC2s relies on the combination between the E(z) and the Su(z)12 protagonists. Hence, this work together with previous results (How Kit et al. 2010) is consistent with at least two tomato PRC2 complexes, respectively containing SlEZ1 or SlEZ2, with partially overlapping functions in plant, flower and early fruit development. It is not clear at this time whether a third complex containing the SlEZ3 protein also exists, although this gene is likely to encode a functional E(z) protein.
Transcriptomic together with ChIP-SEQ analysis of tomato plants impaired in PRC2 function will provide a more precise view of the functions of PRC2s in this plant and will probably highlight further differences with Arabidopsis and other plant species.
References
Ach RA, Taranto P, Gruissem W (1997) A conserved family of WD-40 proteins binds to the retinoblastoma protein in both plants and animals. The Plant Cell Online 9:1595–1606
Bar M, Ori N (2014) Leaf development and morphogenesis. Development 141:4219–4230
Beaudoin F, Wu X, Li F et al (2009) Functional characterization of the arabidopsis β-ketoacyl-coenzyme a reductase candidates of the fatty acid elongase. plant Physiol 150:1174–1191
Benvenuto G, Formiggini F, Laflamme P, Malakhov M, Bowler C (2002) The photomorphogenesis regulator DET1 binds the amino-terminal tail of histone H2B in a nucleosome context. Curr Biol 12:1529–1534
Berger Y, Harpaz-Saad S, Brand A, Melnik H, Sirding N, Alvarez JP, Zinder M, Samach A, Eshed Y, Ori N (2009) The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development 136:823–832
Bird D, Beisson F, Brigham A et al (2007) Characterization of Arabidopsis ABCG11/WBC11, an ATP binding cassette (ABC) transporter that is required for cuticular lipid secretion. Plant J 52:485–498
Blackledge Neil P, Farcas Anca M, Kondo T, King Hamish W, McGouran Joanna F, Hanssen Lars L, Ito S, Cooper S, Kondo K, Koseki Y, Ishikura T, Long Hannah K, Sheahan Thomas W, Brockdorff N, Kessler Benedikt M, Koseki H, Klose Robert J (2014) Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell 157:1445–1459
Bourdenx B, Bernard A, Domergue F, Pascal S, Léger A, Roby D, Pervent M, Vile D, Haslam RP, Napier JA, Lessire R, Joubés J (2011) Overexpression of Arabidopsis ECERIFERUM1 promotes wax very-long-chain alkane biosynthesis and influences plant response to biotic and abiotic stresses. Plant Physiol 156:29–45
Bouyer D, Roudier F, Heese M, Andersen ED, Gey D, Nowack MK, Goodrich J, Renou J-P, Grini PE, Colot V, Schnittger A (2011) Polycomb repressive complex 2 controls the embryo-to-seedling phase transition. PLoS Genet 7:e1002014
Butenko Y, Ohad N (2011) Polycomb-group mediated epigenetic mechanisms through plant evolution. Biochim Biophys Acta 1809:395–406
Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P, Jones RS, Zhang Y (2002) Role of histone H3 Lysine 27 methylation in polycomb-group silencing. Science 298:1039–1043
Chaïb J, Devaux M-F, Grotte M-G, Robini K, Causse M, Lahaye M, Marty I (2007) Physiological relationships among physical, sensory, and morphological attributes of texture in tomato fruits. J Exp Bot 58:1915–1925
Chanvivattana Y, Bishopp A, Schubert D, Stock C, Moon YH, Sung ZR, Goodrich J (2004) Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 131:5263–5276
Chaudhury AM, Ming L, Miller C, Craig S, Dennis ES, Peacock WJ (1997) Fertilization-independent seed development in Arabidopsis thaliana. Proc Natl Acad Sci USA 94:4223–4228
Chen W, Kong J, Qin C, Sheng Y, Tan J, Chen Y-r WuC, Wang H, Shi Y, Li C, Li B, Zhang P, Wang Y, Lai T, Yu Z, Zhang X, Shi N, Wang H, Osman T, Liu Y, Manning K, Jackson S, Rolin D, Zhong S, Seymour GB, Gallusci P, Hong Y (2015) Requirement of CHROMOMETHYLASE3 for somatic inheritance of spontaneous tomato epimutation Colourless non-ripening. Sci Rep 5:9192
Consortium TTG (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Czermin B, Melfi R, McCabe D, Seitz V, Imhof A, Pirrotta V (2002) Drosophila enhancer of zeste/ESC complexes have a histone H3 methyltransferase activity that marks chromosomal polycomb sites. Cell 111:185–196
Danilevskaya ON, Hermon P, Hantke S, Muszynski MG, Kollipara K, Ananiev EV (2003) Duplicated fie genes in maize: expression pattern and imprinting suggest distinct functions. Plant Cell 15:425–438
Davuluri GR, van Tuinen A, Mustilli AC, Manfredonia A, Newman R, Burgess D, Brummell DA, King SR, Palys J, Uhlig J, Pennings HMJ, Bowler C (2004) Manipulation of DET1 expression in tomato results in photomorphogenic phenotypes caused by post-transcriptional gene silencing. Plant J 40:344–354
De Lucia F, Crevillen P, Jones AME, Greb T, Dean C (2008) A PHD-polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc Natl Acad Sci USA 105:16831–16836
Derkacheva M, Steinbach Y, Wildhaber T, Mozgová I, Mahrez W, Nanni P, Bischof S, Gruissem W, Hennig L (2013) Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J 32:2073–2085
Domergue F, Vishwanath SJ, Joubés J, Ono J, Lee JA, Bourdon M, Alhattab R, Lowe C, Pascal S, Lessire R, Rowland O (2010) Three Arabidopsis fatty acyl-coenzyme a reductases, FAR1, FAR4, and FAR5, generate primary fatty alcohols associated with suberin deposition. Plant Physiol 153:1539–1554
Farrona S, Thorpe FL, Engelhorn J, Adrian J, Dong X, Sarid-Krebs L, Goodrich J, Turck F (2011) Tissue-specific expression of flowering locus T in arabidopsis is maintained independently of polycomb group protein repression. Plant Cell 23:3204–3214
Faure JD, Vittorioso P, Santoni V et al (1998) The PASTICCINO genes of Arabidopsis thaliana are involved in the control of cell division and differentiation. Development 125:909–918
Fei Z, Joung J-G, Tang X, Zheng Y, Huang M, Lee JM, McQuinn R, Tieman DM, Alba R, Klee HJ, Giovannoni JJ (2011) Tomato functional genomics database: a comprehensive resource and analysis package for tomato functional genomics. Nucl Acids Res 39:D1156–D1163
Gallusci P, Salamini F, Thompson RD (1994) Differences in cell type-specific expression of the gene Opaque 2 in maize and transgenic tobacco. Mol Gen Genet 244:391–400
Gendall AR, Levy YY, Wilson A, Dean C (2001) The vernalization 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 107:525–535
Gendrel A-V, Lippman Z, Martienssen R, Colot V (2005) Profiling histone modification patterns in plants using genomic tiling microarrays. Nat Methods 2:213–218
Giovannoni JJ (2007) Fruit ripening mutants yield insights into ripening control. Curr Opin Plant Biol 10:283–289
Gonzalez N, Gévaudant F, Hernould M, Chevalier C, Mouras A (2007) The cell cycle-associated protein kinase WEE1 regulates cell size in relation to endoreduplication in developing tomato fruit. Plant J 51:642–655
Goodrich J, Puangsomlee P, Martin M, Long D, Meyerowitz EM, Coupland G (1997) A polycomb-group gene regulates homeotic gene expression in Arabidopsis. Nature 386:44–51
Grimaud C, Negre N, Cavalli G (2006) From genetics to epigenetics: the tale of Polycomb group and trithorax group genes. Chromosome Res 14:363–375
Grossniklaus U, Vielle-Cazalda JP, Hoepner MA, Gagliano W (1998) maternal control of embryogenesis by MEDEA, a polycomb group gene in Arabidopsis thaliana. Science 280:448–449
Hagemann W, Gleissberg S (1996) Organogenetic capacity of leaves: the significance of marginal blastozones in angiosperms. Plant Syst Evol 199:121–152
Hennig L, Derkacheva M (2009) Diversity of Polycomb group complexes in plants: same rules, different players? Trends Genet 25:414–423
Hennig L, Taranto P, Walser M, Schonrock N, Gruissem W (2003) Arabidopsis MSI1 is required for epigenetic maintenance of reproductive development. Development 130:2555–2565
Hennig L, Bouveret R, Gruissem W (2005) MSI1-like proteins: an escort service for chromatin assembly and remodeling complexes. Trends Cell Biol 15:295–302
Holec S, Berger F (2012) Polycomb group complexes mediate developmental transitions in plants. Plant Physiol 158:35–43
How Kit A, Boureau L, Stammitti-Bert L, Rolin D, Teyssier E, Gallusci P (2010) Functional analysis of SlEZ1 a tomato enhancer of zeste (E(z)) gene demonstrates a role in flower development. Plant Mol Biol 74:201–213
Jasinski S, Tattersall A, Piazza P, Hay A, Martinez-Garcia JF, Schmitz G, Theres K, McCormick S, Tsiantis M (2008) PROCERA encodes a DELLA protein that mediates control of dissected leaf form in tomato. Plant j 56:603–612
Javelle M, Vernoud V, Rogowsky PM et al (2011) Epidermis: the formation and functions of a fundamental plant tissue. New Phytol 189:17–39
Jiang D, Wang Y, Wang Y, He Y (2008) Repression of flowering locus C and flowering locus T by the Arabidopsis polycomb repressive complex 2 components. PLoS ONE 3:e3404
Katz A, Oliva M, Mosquna A, Hakim O, Ohad N (2004) FIE and curly leaf polycomb proteins interact in the regulation of homeobox gene expression during sporophyte development. Plant J 37:707–719
Kim D-H, Sung S (2014) Polycomb-mediated gene silencing in Arabidopsis thaliana. Mol Cells 37:841–850
Kinoshita T, Harada JJ, Goldberg RB, Fischer RL (2001) Polycomb repression of flowering during early plant development. Proc Natl Acad Sci USA 98:14156–14161
Kiyosue T, Ohad N, Yadegari R, Hannon M, Dinneny J, Wells D, Katz A, Margossian L, Harada JJ, Goldberg RB, Fisher RL (1999) Control of fertilization independent endosperm development by the MEDEA polycomb gene in Arabidopsis. Proc Natl Acad Sci USA 96:4186–4191
Köhler C, Hennig L (2010) Regulation of cell identity by plant Polycomb and trithorax group proteins. Curr Opin Genet Dev 20:541–547
Köhler C, Villar CBR (2008) Programming of gene expression by Polycomb group proteins. Trends Cell Biol 18:236–243
Köhler C, Hennig L, Bouveret R, Gheyselinck J, Grossniklaus U, Gruissem W (2003) Arabidopsis MSI1 is a component of the MEA/FIE polycomb group complex and required for seed development. EMBO J 22:4804–4814
Kuzmichev ARD, Nishioka K, Erdjument-Bromage H, Tempst P, Reinberg D (2002) Histone methyltransferase activity associated with a human multiprotein complex containing the enhancer of Zeste protein. Genes Dev 16:2893–2905
Lafos M, Kroll P, Hohenstatt ML, Thorpe FL, Clarenz O, Schubert D (2011) Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet 7:e1002040
Lee E, Lucas JR, Goodrich J, Sack FD (2014) Arabidopsis guard cell integrity involves the epigenetic stabilization of the FLP and FAMA transcription factor genes. Plant J 78:566–577
Li S, Zhou B, Peng X, Kuang Q, Huang X, Yao J, Du B, Sun M-X (2013) OsFIE2 plays an essential role in the regulation of rice vegetative and reproductive development. New Phytol 201:66–79
Liu D-D, Dong Q-L, Fang M-J, Chen K-Q, Hao Y-J (2012) Ectopic expression of an apple apomixis-related gene MhFIE induces co-suppression and results in abnormal vegetative and reproductive development in tomato. J Plant Physiol 169:1866–1873
Liu R, How Kit A, Stammitti L et al (2015) A demeter-like DNA demethylase governs tomato fruit ripening. Proc Natl Acad Sci USA 112:10804–10809
Luo M, Bilodeau P, Koltunow A, Dennis ES, Peacock WJ, Chaudhury AM (1999) Genes controlling fertilization-independent seed development in Arabidopsis thaliana. Proc Natl Acad Sci USA 96:296–301
Luo M, Bilodeu P, Dennis ES, Peacock JW, Chaudhury AML (2000) Expression and parent-of-origin effects for FIS2, E(Z), and FIE in the endosperm and embryo of developing Arabidopsis seeds. Proc Natl Acad Sci USA 97:10637–10642
Manning K, Tör M, Poole M, Hong Y, Thompson AJ, King GJ, Giovannoni JJ, Seymour GB (2006) A naturally occuring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nature genet 38:948–952
Mayama T, Ohtsubo E, Tsuchimoto S (2003) Isolation and expression analysis of petunia curly leaf-like genes. Plant Cell Physiol 44:811–819
Miyake T, Takebayashi N, Wolf DE (2009) Possible diversifying selection in the imprinted gene, medea, in Arabidopsis. Mol Biol Evol 26:843–857
Molitor A, Shen W-H (2013) The polycomb complex PRC1: composition and function in plants. J Genet Genomics 40:231–238
Molitor AM, Bu Z, Yu Y, Shen W-H (2014) Arabidopsis AL PHD-PRC1 complexes promote seed germination through H3K4me3-to-H3K27me3 chromatin state switch in repression of seed developmental genes. PLoS Genet 10:e1004091
Mosquna A, Katz A, Decker EL, Rensing SA, Reski R, Ohad N (2009) Regulation of stem cell maintenance by the Polycomb protein FIE has been conserved during land plant evolution. Development 136:2433–2444
Moyle LC (2008) Ecological and evolutionary genomics in the wild tomatoes (Solanum sect. Lycopersicon). Evolution 62:2995–3013
Mozgova I, Köhler C, Hennig L (2015) Keeping the gate closed: functions of the polycomb repressive complex PRC2 in development. Plant J. doi:10.1111/tpj.12828
Müller J, Hart CM, Francis NJ, Vargas ML, Sengupta A, Wild B, Miller EL, O’Connor MB, Kingston RE, Simon JA (2002) Histone methyltransferase activity of a drosophila polycomb group repressor complex. Cell 111:197–208
Ohad N, Yadegari R, Margossian L, Hannon M, Michaeli D, Harada JJ, Goldberg RB, Fischer RL (1999) Mutations in FIE, a WD polycomb group gene, allow endosperm development without fertilization. Plant Cell 11:407–416
Orlando V, Paro R (1995) Chromatin multiprotein complexes involved in the maintenance of transcription patterns. Curr Opin Genet Dev 5:174–179
Panikashvili D, Savaldi-Goldstein S, Mandel T et al (2007) The Arabidopsis desperado/AtWBC11 transporter is required for cutin and wax secretion. Plant Physiol 145:1345–1360
Petit J, Bres C, Just D, Garcia V, Mauxion J-P, Marion D, Bakan B, Joubés J, Domergue F, Rothan C (2014) Analyses of tomato fruit brightness mutants uncover both cutin-deficient and cutin-abundant mutants and a new hypomorphic allele of GDSL lipase. Plant Physiol 164:888–906
Rodrigues JCM, Tucker MR, Johnson SD, Hrmova M, Koltunow AMG (2008) Sexual and apomictic seed formation in hieracium requires the plant polycomb-group gene fertilization independent endosperm. Plant cell 20:2372–2386
Schoeftner S, Sengupta AK, Kubicek S, Mechtler K, Spahn L, Koseki H, Jenuwein T, Wutz A (2006) Recruitment of PRC1 function at the initiation of X inactivation independent of PRC2 and silencing. EMBO J 25:3110–3122
Schubert D, Clarenz O, Goodrich J (2005) Epigenetic control of plant development by Polycomb-group proteins. Curr Opin Plant Biol 8:553–561
Shaver S, Casas-Mollano JA, Cerny RL, Cerutti H (2010) Origin of the polycomb repressive complex 2 and gene silencing by an E(z) homolog in the unicellular alga Chlamydomonas. Epigenetics 5:301–312
Spillane C, MacDougall C, Stock C, Kohler C, Vielle-Gazalda JP, Nunes SM, Grossniklaus U, Goodrich J (2000) Interaction of the Arabidopsis polycomb group proteins FIE and E(Z) mediates their common phenotypes. Curr Biol 10:1535–1538
Spillane C, Schmid KJ, Laoueille-Duprat S, Pien S, Escobar-Restrepo JM, Baroux C, Gagliardini V, Page DR, Wolfe KH, Grossniklaus U (2007) Positive darwinian selection at the imprinted MEDEA locus in plants. Nature 448:349–352
Tavares L, Dimitrova E, Oxley D, Webster J, Poot R, Demmers J, Bezstarosti K, Taylor S, Ura H, Koide H, Wutz A, Vidal M, Elderkin S, Brockdorff N (2012) RYBP-PRC1 complexes mediate H2A ubiquitylation at polycomb target sites independently of PRC2 and H3K27me3. Cell 148:664–678
Teyssier E, Bernacchia G, Maury S, How Kit A, Stammitti-Bert L, Rolin D, Gallusci P (2008) Tissue dependent variations of DNA methylation and endoreduplication levels during tomato fruit development and ripening. Planta 228:391–399
van der Knaap E, Chakrabarti M, Chu YH, Clevenger JP, Illa-Berenguer E, Huang Z, Keyhaninejad N, Mu Q, Sun L, Wang Y, Wu S (2014) What lies beyond the eye: the molecular mechanisms regulating tomato fruit weight and shape. Front Plant Sci 5:227
Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, Zhang Y (2004) Role of histone H2A ubiquitination in polycomb silencing. Nature 431:873–878
Wang D, Tyson MD, Jackson SS, Yadegari R (2006) Partially redundant functions of two SET-domain polycomb-group proteins in controlling initiation of seed development in Arabidopsis. Proc Natl Acad Sci USA 103:13244–13249
Weinhofer I, Hehenberger E, Roszak P, Hennig L, Köhler C (2010) H3K27me3 profiling of the endosperm implies exclusion of polycomb group protein targeting by DNA methylation. PLoS Genet 6:e1001152
Wood CC, Robertson M, Tanner G, Peacock WJ, Dennis ES, Helliwell CA (2006) The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes vernalization insensitive 3. Proc Natl Acad Sci USA 103:14631–14636
Xiao J, Wagner D (2015) Polycomb repression in the regulation of growth and development in Arabidopsis. Curr Opin Plant Biol 23:15–24
Yadegari R, Kinoshita T, Lotan O, Cohen G, Katz A, Choi Y, Nakashima K, Harada JJ, Goldberg RB, Fischer RL, Ohad N (2000) Mutations in the FIE and E(Z) genes that encode interacting polycomb proteins cause parent-of-origin effects on seed development by distinct mechanisms. Plant Cell 12:2367–2382
Yang C, Bratzel F, Hohmann N, Koch M, Turck F, Calonje M (2013) AL- and AtBMI1-mediated H2Aub initiate the switch from embryonic to postgerminative growth in Arabidopsis. Curr Biol 23:1324–1329
Yoshida N, Yanai Y, Chen L, Kato Y, Hiratsuka J, Miwa T, Sung ZR, Takahashi S (2001) Embryonic flower2, a novel polycomb group protein homolog, mediates shoot development and flowering in arabidopsis. Plant Cell Online 13:2471–2481
Zhong S, Fei Z, Chen Y-R, Zheng Y, Huang M, Vrebalov J, McQuinn R, Gapper N, Liu B, Xiang J, Shao Y, Giovannoni JJ (2013) Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat Biotech 31:154–159
Acknowledgments
LB and AHK were in receipt of a grant from the French Ministry of Research and Higher Education and MR from the Italian Ministry of Agriculture. Research work was in part funded by the French National Research Agency in the Frame of the ENDOREPIGENE project, by the Research Federation, Integrative Biology and Environment (FR BIE) and by the National Transgenic Program of China (2016ZX08009-001).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Boureau, L., How-Kit, A., Teyssier, E. et al. A CURLY LEAF homologue controls both vegetative and reproductive development of tomato plants. Plant Mol Biol 90, 485–501 (2016). https://doi.org/10.1007/s11103-016-0436-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0436-0