Abstract
Plant recovery from viral infection is characterized by initial severe systemic symptoms which progressively decrease, leading to reduced symptoms or symptomless leaves at the apices. A key feature to plant recovery from invading nucleic acids such as viruses is the degree of the host’s initial basal immunity response. We review current links between RNA silencing, recovery and tolerance, and present a model in which, in addition to regulation of resistance (R) and other defence-related genes by RNA silencing, viral infections incite perturbations of the host physiological state that trigger reprogramming of host responses to by-pass severe symptom development, leading to partial or complete recovery. Recovery, in particular in perennial hosts, may trigger tolerance or virus accommodation. We discuss evidence suggesting that plant viruses can avoid total clearance but persistently replicate at low levels, thereby modulating the host transcriptome response which minimizes fitness cost and triggers recovery from viral-symptoms. In some cases a susceptible host may fail to recover from initial viral systemic symptoms, yet, accommodates the persistent virus throughout the life span, a phenomenon herein referred to as non-recovery accommodation, which differs from tolerance in that there is no distinct recovery phase, and differs from susceptibility in that the host is not killed. Recent advances in plant recovery from virus-induced symptoms involving host transcriptome reprogramming are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Symptom induction in infected plants is collectively caused by an accumulation of viral nucleic acids or proteins that alter functioning of the plant, resulting in hormonal, developmental and morphological disturbances manifested as symptoms. Plants are able to mount defence responses termed resistance, tolerance, immunity or recovery. For the purposes of this review we will differentiate between these terms that describe unique virus–host relationships. Resistance refers to the plant’s ability to limit virus multiplication (Fraile and García-Arenal 2010) by interfering with the disease cycle within the host plant. Complete immunity is when plants are unable to sustain virus replication and show no virus symptoms (Lecoq et al. 2004), while completely susceptible plants do not impair pathogen infection. Between these extremes, plants show different degrees of resistance or susceptibility. Because relative virus multiplication and yield are not always related, so-called tolerance to virus infection has been described. From a virologist point of view, tolerance describes a form of resistance where reduced symptom and virus levels persist throughout the lifespan of the plant (Fraile and García-Arenal 2010; Matthews 1991). Plants generally exhibit genetic variation for resistance, tolerance and susceptibility.
Recovery from virus pathogens is characterized by initial severe systemic symptoms which progressively decrease, resulting in a reduction or disappearance of symptoms in newly developed leaves (Ma et al. 2014; Ghoshal and Sanfaçon 2014; Nie and Molen 2015). In some cases the entire plant appears recovered at late stages of infection. To date, insights into the mechanism of plant tolerance to viral infection and recovery from viral-induced symptoms have been elusive. This is because the current literature describes recovery by targeting specific components in the RNA silencing pathway rather than considering the global changes occurring in a plant during the recovery process.
Symptom recovery in infected plants has been correlated with the accumulation of virus-targeting short interfering RNAs (vsRNAs) (Rodríquez-Negrete et al. 2009). Additionally, several disturbances in endogenous sRNA accumulation have been correlated with plant viral diseases, in particular miRNA-target interactions involving several miRNA families (Ramesh et al. 2014). Viral infections can modify miRNA-derived-trans-acting siRNAs or heterochromatic-siRNA production, which also leads to phenotypic changes during virus infection (Raja et al. 2008; Wang et al. 2010). Therefore, it is hardly surprising that virus infections cause global changes in the host transcriptome, either through direct interaction of virus proteins with host proteins, or indirectly via alterations in RNA silencing processes. More recently, transcriptomes data either covering partial or complete recovery processes in DNA- and RNA-viruses (Góngora-Castillo et al. 2012; Lu et al. 2012; Liu et al. 2014; Allie et al. 2014) are available. In this review we evaluate molecular interconnections based on previous literature regarding virus-persistence, adaptation, tolerance and recovery at the transcriptome level. Finally, we will discuss various elements of the recovery phenomenon that allows viral persistence, host transcriptome reprogramming and host tolerance.
Recovery and symptom reversion may share some similar genetic mechanisms
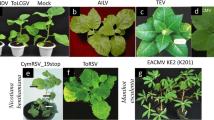
In all plant–virus interactions where recovery has been identified thus far, only emerging leaves at the top of the plant show recovery from systemic symptoms, such as in Manihot esculenta Crantz (cassava, Fig. 1), while older leaves or branches further down from the apex remain symptomatic (Góngora-Castillo et al. 2012; Nie and Molen 2015). Early pioneering field studies based on African cassava mosaic virus (ACMV), a geminivirus infecting cassava, observed that virus-free plants could be propagated from recovered plant portions via stem cuttings (Thresh et al. 1994; Fondong et al. 2000). This was termed reversion, and it was concluded that reversion was linked to the restricted distribution of the virus. The term reversion in cassava was described for the portion of a symptomless virus-free plant that resulted from stem cuttings taken from an infected mother-plant, while the production of a sequence of symptomless leaves (usually at the apex) by a plant that is known to be infected, and that had previously expressed symptoms, was referred to as ‘recovery’. However, Fargette et al. (1996) appeared to use the terms reversion and recovery interchangeably. The disease recovery phenomenon has been reported in many vegetatively propagated crops including Ipomoea batatas (sweet potato) (Gasura et al. 2008; Gasura and Mukasa 2010).
Despite the similar outcomes in recovery and reversion, there is no clear indication whether these two phenomena display analogous molecular mechanisms. In Fondong et al. (2000) it was concluded that reversion and recovery were genotype-dependent. Further support for genotype dependency comes from a study with Potato virus Y, where host recovery and reduced virus titers in the upper leaves occurred in Nicotiana tabacum L. (tobacco) and Solanum lycopersicum L. (tomato) but not in potato plants (Nie and Molen 2015). In a study on sweet potato in Uganda, recovery from Sweet potato virus disease (SPVD), caused by dual infection and synergistic interaction of Sweet potato feathery mottle potyvirus (SPFMV) and a whitefly transmitted Sweet potato chlorotic stunt crinivirus (SPCSV), was reported in three out of twenty graft-inoculated and naturally infected field cultivars (Gasura et al. 2009). Cultivar Munyeera was highly resistant followed by New Kawogo and Polyster (as reflected by the mean severity scores and mild symptoms observed), while the remaining cultivars ranged from moderately susceptible to susceptible, suggesting some level of genetically controlled resistance in these three cultivars. These three resistant cultivars displayed recovery from SPVD which was correlated with SPFMV reduced virus titers. Several cultivars were further selected for subsequent crosses based on their contrasting traits with respect to SPVD resistance. An interesting feature was that the level of segregation for SPVD resistance was high in the F1 progenies, and crosses between resistant and susceptible parents produced both resistant and susceptible progenies. From the bi-parental mating between resistant sweet potato cv. Munyeera and four maternal parents (Gasura 2008; Gasura et al. 2009), only some F1 progeny displayed recovery or reversion against Sweet potato feathery virus (SPFMV), further supporting a genetic basis for recovery. Thus, it is not unreasonable to suggest that reversion and recovery in plants may share a similar genetic basis.
Plants infected with viruses that counteract RNA silencing efficiently do not recover from infection
The process via which plants generate endogenous small interfering RNAs (sRNAs), microRNAs (miRNAs) and virus-derived small RNAs and amplify the silencing signals is well documented (Calo et al. 2012; Dunoyer et al. 2014) and reviewed in Tiwari et al. (2014). Mobile siRNA–mediated breakdown of RNA transcripts is referred to as post transcriptional gene silencing (PTGS). Recent literature indicates that translational repression mediated by siRNAs or miRNAs (Brodersen et al. 2008; Karran and Sanfaçon 2014; Ma et al. 2014) is an alternative to mRNA degradation. On the other hand, transcriptional gene silencing (TGS) is mainly induced through cytosine methylation of viral DNA targets and histone H3 methylation leading to repression of replication (Mette et al. 2000; Raja et al. 2008; Butterbach et al. 2014). Abundant data have shown that wild-type plants recover from viral infections by sequence-specific degradation of virus-encoded mRNA (Al-Kaff et al. 1998; Covey et al. 1997; Ratcliff et al. 1999). Links between PTGS and recovery (Chellappan et al. 2004), and TGS and recovery (Rodríquez-Negrete et al. 2009; Butterbach et al. 2014) are centred on the accumulation of 21–22 nt (PTGS) or 24 nt (TGS) classes of siRNAs, respectively. In principle, virus-derived small RNAs (vsRNAs) targeting the virus (by TGS or PTGS) increased with recovery of cassava and N. benthamiana infected with ACMV-[CM] (Chellappan et al. 2004) as well as in Pepper golden mosaic virus (PepGMV)—infecting Capsicum annuum L. (Rodríquez-Negrete et al. 2009), leading to lower virus titers. These studies indicate that RNA silencing pathways contributes to plant recovery from virus-induced symptoms.
Silencing of ARGONAUTE (AGO1) halts symptom recovery in N. benthamiana plants infected with Tomato ringspot virus (ToRSV) (Ghoshal and Sanfaçon 2014). This study provides evidence that when a component, AGO1, of RNA silencing is down-regulated, recovery is compromised. Although plant RNA silencing is a natural defense mechanism, viruses have devised strategies to thwart this layer of defense. Persuasive evidence comes from potyvirus Turnip mosaic virus (TuMV) and tobamovirus Turnip vein clearing virus (TVCV) that completely inhibit PTGS (Mourrain et al. 2000). The Rep protein in geminiviruses down-regulates methyltransferase MET1 and methyltransferase CMT3 to evade TGS (Rodriquez-Negrete et al. 2013). Furthermore, mixed infection of Tomato yellow leaf curl virus (TYLCV) with Cucumber mosaic virus (CMV) compromises genome hypermethylation and breaks down resistance conferred by Ty-1 resistance gene, encoding for an RNA-dependent RNA polymerase (Butterbach et al. 2014). Plant viruses that encode viral suppressors of RNA silencing often hinder the incorporation of siRNAs into RNA induced silencing complex (RISC) via destabilization, or inactivation of key enzymes at different levels of plant RNAi pathway (Hulo et al. 2011; Ghoshal and Sanfaçon 2015). Also, viruses that generate decoy RNAs that act to protect the viral RNAs from degradation (Blevins et al. 2011) and viruses with specialized replication mechanisms (Pooggin 2013) can overcome plant RNAi, thus no recovery occurs.
Abiotic factors that compromise RNA silencing could decrease the likelihood of plant recovery from viral infection. For instance low temperature (below 24 °C) inhibited RNA silencing—mediated defense by controlling siRNAs generation leading to no symptom recovery (Szittya et al. 2003). Also, low light intensities (<300 μE/m2/s) at a constant temperature of 25 °C reduced recovery from geminiviral infection despite strong systemic movement of silencing signals (Patil and Fauquet 2015). Another factor that can halt RNA silencing mediated recovery is removal of the first three leaves above the virus inoculated leaves as in the case of potyvirus Potato virus Y (PVY), an indication that the signals mediating recovery are probably generated within those leaves (Nie and Molen 2015). From current literature, it is tempting to conclude that the recovery process is multifactorial and can be compromised by several factors linked to plant RNA silencing defense. Hence, it is not unreasonable to suggest that plants may mobilize other immune pathways, such as pathogen-triggered immunity (PTI) involving pathogenesis related protein (PrP) genes or effector-triggered immunity (ETI) involving resistance (R) genes, to avoid succumbing to viral-induced symptoms when RNA silencing is prevented by the invading virus.
The NBS–LRR-encoding class of defence genes are targets of an extensive small RNA regulatory network, and it is noteworthy that a subset of R genes from the NBS–LRR family are regulated by miRNAs and secondary siRNAs (Shivaprasad et al. 2012; Zhai et al. 2011). Three highly abundant 22 nt miRNA families targeting conserved domains in NBS–LRR defense genes in Medicago truncatula were identified, and triggered the production of phased trans-acting (ta-siRNAs). Dicer-like 4 (DCL4) and SGS3 transcripts were also cleaved by 22 nt miRNAs leading to phasiRNA generation (Zhai et al. 2011). In cassava several ta-siRNAs were implicated in response to cassava bacterial blight, and 15 TAS loci were induced while 39 were repressed (Quintero et al. 2013). Virus-mediated suppression of RNA silencing can potentially lead to up-regulation of R genes (Shivaprasad et al. 2012). In tomato, miR482 and miR2118, which belong to a large superfamily, were found to target the p-loop motif in mRNA sequences for NBS–LRR proteins (Shivaprasad et al. 2012). Interestingly in SACMV-infected susceptible cassava T200 a significant number of R gene homologs were suppressed at 32 and 67 dpi, but not in tolerant TME3 (Allie et al. 2014). These and other studies demonstrate that silencing and pathogen defence pathways are highly linked and often synchronized.
Persistent viral titers do not always decrease during recovery
Persistent infection is a state of meta-stability in which the pathogen and the host co-exist (Goic and Saleh 2012). Metastability can be defined as a long term relationship where the equilibrium between viruses and hosts is maintained . A persistent infection could be considered as the most well-adapted or successful host–pathogen interaction (Goic and Saleh 2012). Decreased viral titers are associated with plants that undergo symptom recovery (Gasura and Mukasa 2010; Covey et al. 1997; Shaw et al. 2014; Ma et al. 2014; Nie and Molen 2015). Contrary to the general trend, N. benthamiana plants infected with Tomato ringspot virus (ToSRV) recovered without a commensurate reduction in the viral RNA titer (Jovel et al. 2007), which suggests recovery has variable outcomes in different virus–host interactions.
Interestingly, cassava (the natural host) and N. benthamiana, infected with a DNA begomovirus, East African cassava mosaic virus (EACMCV-[CM]), failed to recover (Chellappan et al. 2004). Chellappan et al. (2004) reported that the viral DNA and virus derived mRNA titers accumulated until the non-recovered plants succumbed to EACMCV-[CM] infection. Intriguing, with ACMV-[CM], a recovery phenotype geminivirus, there was an initial increase in viral DNA and viral-derived mRNA accumulation levels in N. benthamiana and cassava, which later on gradually decreased (Chellappan et al. 2004). Analogous results were observed in cantaloupe (Cucumis melo) and watermelon (Citrullus lunatus) recovering from a bipartite begomovirus Cucurbit leaf crumple virus (CuLCrV) (Hagen et al. 2008). Here, recovered leaves re-inoculated with CuLCrV failed to show symptoms and equally no increase in viral DNA was detected (Hagen et al. 2008). Re-infection of CuLCrV–recovered leaves with an RNA virus, CMV, resulted in severe disease symptoms (more severe than those induced by CMV or CuLCrV alone) and increased CuLCrV DNA levels (Hagen et al. 2008). On the basis of this study (Hagen et al. 2008), it was concluded that recovery is fully reversible in the presence of a new unrelated virus in recovered leaves. On the basis of the divergent pattern between recovery and viral load observed thus far, we conclude that viral titer often decreases during recovery but this is not a universal rule.
Persistent infection often leads to failed symptom recovery without killing the host
Viral infections can be acute, persistent (comprising of latent or chronic infections) and in some cases mutualistic. In animals, there is a high likelihood that in acute infections, viral titer might dramatically increase leading to severe infections and sudden death of the host (Goic and Saleh 2012; Swevers et al. 2013). In the case of cassava landraces TME3 and T200 infected with South African cassava mosaic virus (SACMV), susceptible T200 failed to recover or die from severe symptoms 67 days post inoculation (dpi), a condition we refer to as non-recovery viral accommodation (Fig. 2a). Viral-induced symptoms in T200 were manifested as curled and deformed leaves with a yellow mosaic, and plants were often stunted compared to non-infected controls. In contrast, tolerant TME3 leaves recovered from viral induced symptoms (Fig. 2b). Comparing the viral load in TME3 and T200, it was shown that SACMV replication was low in TME3 but high in T200 (Allie et al. 2014). Non-recovery viral accommodation would likely be more prevalent in perennial crops as they cannot afford to die prematurely, especially those that are vegetatively propagated. We propose that non-recovery accommodation differs from tolerance in that in the former severe symptoms and high virus titers persist yet the host continues to grow, while in tolerance virus titers and symptoms are lower compared to susceptible plants, and the host shows comparable growth to healthy plants. Tolerance may involve an initial symptomatic phase followed by recovery which then leads to accommodation, as in the case of cassava landrace TME3 (Allie et al. 2014). Non-recovery accommodation and susceptibility differ in that in susceptibility the host is eventually killed. Even when the host is cut back and new shoots emerge in susceptible cassava T200, the symptoms and virus load persist but the plant is able to grow. Hence, we propose that persistent viruses may not necessary kill a susceptible ‘non-recovery’ host, but persist to ensure survivability throughout the host life cycle.
Interaction of South African cassava mosaic virus (SACMV) with cassava landraces 100 days post inoculation. a T200 accommodates SACMV and fails to recover from viral-induced symptoms, showing curled and deformed leaves with a yellow mosaic. b Complete recovered TME3 hallmarked by symptomless leaves, which appear identical to normal uninfected leaves
Complete and partial recovery is associated with transcriptome alterations
Several lines of evidence in N. tabacum (Loebenstein et al. 1977) and N. benthamiana (Fuzawa et al. 2010) infected with the M-strain of Cucumber mosaic virus (M-CMV) have shown that disease development can involve initial and secondary pathogenicity processes that are interupted by a transient recovery phase. Furthermore, N. tabacum infected with M-CMV revealed that partial and complete recovery is followed by secondary symptoms induced by the same virus (Lu et al. 2012). Since recovery has only been studied over short periods of time, primarily in annual experimental hosts, this raises an intriguing question as to whether recovery is a transient phenomenon in annual hosts or cyclically intermittent in perennial plants, which could imply gene expression modulations. Transcriptome data indicated that complete recovery was marked by a low level of both up- and down- differentially expressed genes (DEGs) with respect to partial recovery (Fig. 3). KEGG pathways in partial and completely recovered leaves showed enrichment of carotenoid biosynthesis genes, genes involved in plant pathogen interactions, and several chitinases, protein kinases, EREP-like factors and ubiquitin-related enzymes. The above results (Lu et al. 2012) open new frontiers for investigation into the role of the transcriptome during and post recovery. For perennial plant crops such as cassava, recovery in tolerant phenotypes could be a cyclic process marked by recurrent appearance and disappearance of mild symptoms throughout their long life span.
Differentially expressed genes (DEGs) in different symptomatic stages following M-strain CMV infection of N. tabacum (Lu et al. 2012), permission was granted for the image. The profile reveals that recovery phenomenon is a transient process in some cases since recovered plants can succumb to secondary mosaic symptoms
Transcriptome reprograming and viral adaptation is vital for recovery
In plant–pathogen interactions, up to 20 % of the genome can undergo alteration in gene expression (Maleck et al. 2000; Tao et al. 2003) indicating the cell’s shift into a defensive mode of resistance or tolerance. Resistance is the ability to limit parasite burden to non-detectable levels of virus replication (Råberg et al. 2007; Fraile and García-Arenal 2010). Herein, tolerance to viruses is considered as a form of basal immune response that lowers virus replication to a detectable level, reduces crop damage and governs recovery from viral-induced symptoms.
An interesting perspective is that in animals, viral responsive host miRNAs, one regulatory aspect of transcriptome modulation, may promote persistence or tolerance (Mahajan et al. 2009). It is understood that host cells undergo significant reprogramming of their transcriptome during infection, which is conceivably a key requisite for mobilizing host defenses (Rodrigo et al. 2012). Host transcriptome reprogramming for a given virus–plant host interaction is genotype dependent (Rodrigo et al. 2012; Allie et al. 2014) and often leads to up- or down-regulation of genes (Lu et al. 2012). For instance, in SACMV–cassava T200 interaction manifested by non-recovery accommodation (Fig. 2a), transcriptome reprogramming led to a high number of differentially expressed genes (DEGS) and increase in viral titer (Fig. 4a). In contrast, the SACMV–cassava TME3 interaction, hallmarked by symptom recovery (Fig. 2b), was associated with a lower number of DEGs and decrease in viral replication (Fig. 4b) (Allie et al. 2014). Often, a severe symptom (Fig. 2a) is reflected by the magnitude of host transcriptome alterations (Fig. 4a) that leads to changes in cellular homeostasis and developmental processes (Hanley-Bowdoin 2013). It is tempting to suggest that different plant species/varieties reprogram gene expression differently which manifests either as symptoms, or symptom recovery to avoid succumbing to viral infection. Feasibly, transcriptome reprogramming in tolerant plant genotypes may prevent providing the virus with factors required for movement and replication as an energy-saving strategy priming recovery. This hypothesis encompasses the notion that tolerance is linked to levels of host-virus adaptation, attenuated genetic response to incoming virus and basal RNA silencing. Further support is provided by a study of Tobacco etch virus (TEV) infection of Arabidopsis thaliana (Agudelo-Romero et al. 2008). Wild virulent ancestral TEV (natural hosts belong to the Solanaceae) in non-host A. thaliana Ler causes mild symptoms and is poorly adapted. This ancestral TEV had lower fold changes in gene expression compared to the evolved TEV–At17 (responsive genes such as SAR and innate immunity were not activated) after 17 passages in A. thaliana Ler. During adaptation of a virus to a new host, natural selection may favour viral strategies/genomes to escape host defences. However, as the virus–host relationship evolves the dynamic interplay between virus and host shifts, determining levels of viral fitness and host adaptation which in the long term dictates the outcome i.e. susceptibility or tolerance and recovery.
Alterations in transcriptome to SACMV infection. a Cassava landrace T200 which exhibits non-recovering phenotype. b Tolerant cassava landrace TME3 exhibits recovery phenotype. The graphs are generated on the basis of the transcriptome data reported in Allie et al. (2014)
Transcriptome reprogramming from the plant and virus view point
A determining factor to symptom recovery is host tolerance. Therefore, it is important to consider transcriptome reprogramming and tolerance from the plant or virus view point. From the plant view point: (1) the plant may moderate viral replication via TGS in DNA viruses (Raja et al. 2008) and counteract the virus mediated inhibition of RNA silencing in RNA and DNA viruses (Pumplin and Voinnet 2013), (2) the plant host must co-evolve with the virus (Little et al. 2010), (3) the plant must minimize over–use of their energy resources so as to limit the synthesis of factors require for virus multiplication, and (4) the plant must avoid over-triggering its innate immune system. The latter is critical for tolerance as the host reprograms its transcriptome towards overall fitness (that is geared towards symptom recovery), down-regulating genes that may be involved in vital physiological processes such as photosynthesis, and reactivating the expression of genes involved in the stress response pathways.
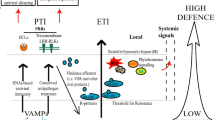
The proposed model (Fig. 5) gives credit to both RNA silencing and system-level transcriptome data reported in recovery studies (Hanssen et al. 2011; Lu et al. 2012; Liu et al. 2014). While the model depicts replication of geminiviruses (that encode 5–7 proteins) in the nucleus, all other host responses would be applicable to RNA or DNA viruses. Geminivirus proteins often redirect host machineries and processes to establish a productive infection (Hanley-Bowdoin 2013). Importantly, N. tabacum infected with the M-strain of CMV triggered transcriptome down-regulation of genes involved in photosynthesis and pigment metabolism (‘photosynthesis’, ‘carbon fixation in photosynthesis’, ‘porphyrin and chlorophyll metabolism’, ‘carotenoid biosynthesis’, and ‘anthocyanin biosynthesis’) at the initial and secondary phase of mosaic which flanked the partial recovery and complete recovery process (Fig. 3), respectively (Lu et al. 2012). It was equally observed that partial and complete recovery was accompanied with up-regulation of DEGs in KEGG pathways, namely: ‘monoterpenoid biosynthesis’, ‘flavonoids biosynthesis’, ‘cysteine and methionine metabolism’, and ‘plant-pathogen interaction’ pathway (Lu et al. 2012). This finding was interpreted as enhancement of innate immunity during the transient recovery process (Lu et al. 2012). Similarly, genes related to photosynthesis and pigment metabolism in tomato were suppressed by Pepino mosaic virus at the onset of infection (Hassen et al. 2011) and in ACMV–infected cassava (Liu et al. 2014). When rethinking the switch– off and –on of DEGs, it is not unreasonable to hypothesize that the initial host trade-off to accommodate a virus, the cause of transcriptome reprogramming, involves adjustment at the level of photosynthesis and pigment metabolism. A shift to over-expression of stress response genes has been reported during recovery in other transcriptome studies (Sahu et al. 2010; Góngora-Castillo et al. 2012).
The model for recovery from begomovirus-induced symptoms in plants, giving credit to links between RNA silencing and defence gene regulation. At the initial phase of infection, the host allow a begomovirus to recruit its replisome (host polymerase and associated replication factors) leading to an initial symptom development. As a cause (or consequence) of infection, the host tolerates the virus but reprograms its transcriptome by negatively modulating the interaction of either the replication protein (REP) or the replication enhancer protein (REN) with proliferating cell nuclear antigen (PCNA). As a result, the host reduces the amplitude and number of differentially expressed genes (DEGS) in an attempt to deprive the virus with valuable resources required for its proliferation, but triggers the expression of defense related genes (R and PrP). The host transcriptome reprogramming intends to reduce virus replication and synthesis of coat protein (CP), REN, REP, movement proteins (MP), anti-defense proteins (V2 and AV2). Consequently, a net decrease in viral load triggers transient symptom recovery hallmarked by an increase small RNAs and viral genome methylation. Full recovery with detectable but low viral titer is determined by abiotic factors such as temperature and light intensity. However, extreme variations in abiotic factors may lead to development of secondary symptoms due to increased fitness of the persistence virus. On the other hand, during viral infection, miRNAs specifically targeting loci for nucleotide binding site–Leu-rich repeat (NBS)–LRRs are decreased, priming upsurge in functional defense related proteins
Tolerance and symptom recovery from the virus viewpoint: (1) A DNA virus should allow for a certain level of methylation of its genome, (2) the virus should exhibit persistence in plant cells to circumvent clearance by the host defences, and (3) the virus should be transmitted vertically by seed or vegetatively propagated to allow adaptation and co-evolution with the host (Pagán et al. 2014). From the perspective of viral fitness, it was suggested that total host recovery might be highly catastrophic for the virus which can no longer replicate or be transmitted (Pagán et al. 2014). From the perspective of host fitness, partial or complete tolerance is an attempt to reduce virus replication which is often dictated by host type, such as perennial or annual crops. Tolerance of Arabidopsis to CMV is achieved via developmental reprogramming such that resources are reallocated from vegetative growth to reproduction (Pagán et al. 2008; Agudelo-Romero et al. 2008). Supposing a trade-off is struck in a virus–plant relationship, transcriptome reprograming and co-existence without severe collateral damage could be a beneficiary outcome. This benefit could be manifested as non-recovery accommodation (Fig. 2a) or recovery (Fig. 3b) in the long term. As previously suggested, it seems reasonable to suggest that host transcriptome reprograming is a key fine-tuning process for co-existence. This is supported by Little et al. (2010) definition of tolerance, in which the host minimizes virulence without necessarily (although it can as in the example of SACMV–cassava TME3) minimizing the viral load. Recovery and symptom re-emergence could fluctuate depending on concomitant host transcriptome reprograming and RNA silencing cycles during the plant lifespan, which to date has not been studied.
Conclusion
Attempts to determine factors governing recovery and reversion under field conditions have been reported (Gibson and Otim-Nape 1997), and other aspects of recovery not covered herein have been reviewed (Palukaitis 2011; Ghoshal and Sanfaçon 2015). The evolutionary advantage of breeding for tolerance over resistance against viral plant disease has been elaborated (Salomon 1999). Over the past decades the search for durable resistance in plant crops has grown within the circles of scientists as well as farmers. Plant tolerance to viruses is tightly linked to recovery and dependent on broader basal immunity rather than only mono- or oligo-genic resistance. Recovery does not appear to be governed by RNA silencing alone, but is accompanied by a broader change in the transcriptome that could lead to the expression of R and PrP genes, amongst other defence-related responses. Farmers are faced with the dilemma to choose between elite crops lines, recovery phenotypes (i.e. tolerant lines) or bred resistant lines. To date, most recovery experimental analyses are derived from annual hosts over a shorter infection period. This has limited our understanding of recovery and plant tolerance to viruses. In all reported recovery phenomena complete virus clearance does not occur, which is a hallmark of tolerance, virus persistence and host-virus adaptation. Tolerance could be effective in controlling disease caused by a single virus, but not effective in the case of mixed virus infections. Nonetheless, the benefits of tolerant plants with recovery phenotype and marginal transcriptome alterations attributes (such as cassava TME3) to viral infection have been neglected and should be further tested at the field level.
References
Agudelo-Romero IP, Carbonell P, De la Iglesia F, Carrera J, Rodrigo G, Jaramillo A, Amador MAP, Elena SF (2008) Changes in the gene expression profile of Arabidopsis thaliana after infection with Tobacco etch virus. Virol J 5:92
Al-Kaff NS, Covey SN, Kreike MM, Page AM, Pinder R, Dale PJ (1998) Transcriptional and post-transcriptional plant gene silencing in response to a pathogen. Science 279:2113–2115
Allie F, Pierce EJ, Okoniewski MJ, Rey MEC (2014) Transcriptional analysis of South African cassava mosaic virus-infected susceptible and tolerant landraces of cassava highlights differences in resistance, basal defense and cell wall associated genes during infection. BMC Genom 15:1006
Blevins T, Rajeswaran R, Aregger M, Borah BK, Schepetilnikov M, Baerlocher L, Farinelli L, Meins Jr-F, Hohn T, Pooggin MM (2011) Massive production of small RNAs from a non-coding region of Cauliflower mosaic virus in plant defense and viral counter-defense. Nucleic Acids Res 39:5003–5014
Brodersen P, Sakvarelidze-Achard L, Bruun-Rasmussen M, Dunoyer P, Yamamoto YY, Sieburth L, Voinnet O (2008) Widespread translational inhibition by plant miRNAs and siRNAs. Science 320:1185–1190
Butterbach P, Verlaan MG, Dullemans A, Lohuis D, Visser RGF, Bai Y, Kormelink R (2014) Tomato yellow leaf curl virus resistance by Ty-1 involves increased cytosine methylation of viral genomes and is compromised by Cucumber mosaic virus infection. Proc Natl Acad Sci USA 111(35):12942–12947. doi:10.1073/pnas.1400894111
Calo S, Nicolás FE, Vila A, Torres-Martínez S, Ruiz-Vázquez RM (2012) Two distinct RNA-dependent RNA polymerases are required for initiation and amplification of RNA silencing in the basal fungus Mucor circinelloides. Mol Microbiol 83(2):379–394
Chellappan R, Vanitharani R, Fauquet CM (2004) Short interfering RNA accumulation correlates with host recovery in DNA virus-infected hosts, and gene silencing targets specific viral sequences. J Virol 78:7465–7477
Covey SN, Al-Kaff NS, Langara A, Turner DS (1997) Plants combat infection by gene silencing. Nature 385:781–782
Dunoyer P, Schoot G, Himber C, Meyer D, Takeda A, Carrington JC, Voinnet O (2014) Small duplexes function as mobile silencing signals between plant cells. Science 328:912
Fargette D, Colon LT, Bouveau D, Fauquet C (1996) Components of resistance of cassava to African cassava mosaic virus. Eur J Plant Pathol 102:645–654
Fondong VN, Thresh JM, Fauquet C (2000) Field experiments in Cameroon on cassava mosaic virus disease and the reversion phenomenon in susceptible and resistant cultivars. Int J Pest Manag 4:211–217
Fraile A, García-Arenal F (2010) The coevolution of plants and viruses: resistance and pathogenicity. Adv Virus Res 76:1–32
Fukuzawa N, Itchoda N, Goto K, Masuta C, Matsumura T (2010) HC-pro, a potyvirus RNA silencing suppressor, cancels cycling of Cucumber mosaic virus in Nicotiana benthamiana plants. Virus Gene 40(3):440–2446
Gasura E (2008) Mechanisms associated with sweet potato virus disease resistance in Ugandan sweet potato genotypes. MSc. Thesis. Makerere University. Kampala, Uganda. P.58
Gasura E, Mukasa SB (2009) Prevalence and implications of sweet potato recovery from Sweet potato virus disease in Uganda. Afr Crop Sci J 18:195–205
Gasura E, Mashingaidze AB, Mukasa SB (2010) Genetic variability for tuber yield, quality, and virus disease complex traits in Uganda sweet potato germplasm. Afr Crop Sci J 16(2):147–160
Ghoshal B, Sanfaçon H (2014) Temperature-dependent symptom recovery in Nicotiana benthamiana plants infected with tomato ringspot virus is associated with reduced translation of viral RNA2 and requires ARGONAUTE 1. Virology 456–457:188–197
Ghoshal B, Sanfaçon H (2015) Symptom recovery in virus-infected plants: revisiting the role of RNA silencing mechanisms. Virology 479–480:167–179
Gibson RW, Otim-Nape GW (1997) Factors determining recovery and reversion in mosaic-affected affecting cassava mosaic virus resistant cassava. Ann Appl Biol 131:259–271
Goic B, Saleh M-C (2012) Living with the enemy: viral persistent infections from a friendly viewpoint. Curr Opin Microbiol 15:531–537
Góngora-Castillo E, Ibarra-Laclette E, Trejo-Saavedra DL, Rivera-Bustamante RF (2012) Transcriptome analysis of symptomatic and recovered leaves of geminivirus-infected pepper (Capsicum annuum). Virol J 9:295
Hagen C, Rojas MR, Kon T, Gilbertson RL (2008) Recovery from Cucurbit leaf crumple virus (Family Geminiviridae, Genus Begomovirus) infection is an adaptive antiviral response associated with changes in viral small RNAs. Phytopathology 98:1029–1037
Hanley-Bowdoin L, Bejarano ER, Robertson D, Mansoor H (2013) Geminiviruses: masters at redirecting and reprogramming plant processes. Nature Rev Microbiol 11:777–788
Hanssen IM, Peter van Esse H, Ballester AR, Hogewoning SW, Parra NO, Lievens A, Bovy AG, Thomma BP (2011) Differential tomato transcriptomics responses induced by pepino mosaic virus isolates with differential aggressiveness. Plant Physiol 156:301–318
Hulo C, de Castro E, Masson P, Bougueleret L, Bairoch A, Xenarios I, Mercier LP (2011) ViralZone: acknowledge resource understand virus diversity. Nucleic Acids Res . doi:10.1093/nar/gkq901 (Database issue)
Jovel J, Walker M, Sanfaçon H (2007) Recovery of Nicotiana bethamiana plants from a necrotic response induced by a nepovirus is associated with RNA silencing but not with reduced virus titer. J Virol 81:12285
Karran RA, Sanfacon H (2014) Tomato ringspot virus coat protein binds to ARGONAUTE 1 and suppresses the translation repression of a reporter gene. Mol Plant Microbe Interact 27:933–943
Lecoq H, Moury B, Desbiez C, Palloix A, Pitrat M (2004) Durable virus resistance in plants through conventional approaches: a challenge. Virus Res 100:31–39
Little TJ, Shuker DM, Colegrave N, Day N, Graham AL (2010) The coevolution of virulence: tolerance in perspective. PLoS Pathog 6:e1001006
Liu J, Yang J, Bi H, Zhang P (2014) Why mosaic? Gene expression profiling of African cassava mosaic virus-infected cassava reveals the effect of chlorophyll degradation on symptom development. JIPB 56(2):122–132
Loebenstein G, Cohen J, Shabtai S, Coutts RHA, Wood KR (1977) Distribution of Cucumber mosaic virus in systemically infected tobacco leaves. Virology 81:117–125
Lu J, Du Z-X, Kong J, Chen L-N, Qiu Y-H, Li G-F, Meng X-H, Zhu S-F (2012) Transcriptome analysis of Nicotiana tabacum infected by Cucumber mosaic virus during systemic symptom development. PLoS ONE 7:e43447
Ma X, Nicole M-C, Meteignier L-V, Hong N, Wang G, Moffett P (2014) Different roles for RNA silencing and RNA processing components in of virus recovery and virus-induced gene silencing in plants. J Exp Bot 65(1):311–322. doi:10.1093/jxb/eru447
Mahajan VS, Drake A, Chen J (2009) Virus-specific host miRNAs: antiviral defenses or promoters of persistent infection? Trends Immunol 30:1–7
Maleck K, Levine A, Eulgem T, Morgan A, Schmid J, Lawton KA, Dangl JL, Dietrich RA (2000) The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nature Genet 26:403–410
Matthews REF (1991) Plant Virology, 3rd edn. Academic Press Inc., Harcourt Brace Jovanovich Publishers, San Diego
Mette MF, Aufsatz W, Van der Winden J, Matzke MA, Matzke AJM (2000) Transcriptional silencing and promoter methylation triggered by double stranded RNA. EMBO J 19:5194–5201
Mourrain P, Beclin C, Elmayan T, Feuerbach F, Godon C, Morel J, Jouette D, Lacombe A, Nikic S, Picault N, Remoue K, Sanial M, Vo T, Vaucheret H (2000) Arabidopsis SGS2 and SGS3 genes are required for post-transcriptional gene silencing and natural virus resistance. Cell 101:533–542
Nie X, Molen TA (2015) Host recovery and reduced virus level in the upper leaves after Potato virus y infection occur in tobacco and tomato but not in potato plants. Viruses 7:680–698
Pagán I, Alonso-Blanco C, García-Arenal F (2008) Host responses in life-history traits and tolerance to virus infection in Arabidopsis thaliana. PLoS Pathog 4:e1000124
Pagán I, Montes N, Milgroom MG, García-Arenal F (2014) Vertical transmission selects for reduced virulence in a plant virus and increased resistance in the host. PloS Pathog 10:e1004293
Palukaitis P (2011) The road to RNA silencing is paved with plant-virus interactions. Plant Pathol J 27(3):197–206
Patil BL, Fauquet CM (2015) Light intensity and temperature affect systemic spread of silencing signal in transient agroinfiltration studies. Mol Plant Pathol 16(5):48–494. doi:10.1111/mpp.12205
Pooggin MM (2013) How can plant DNA viruses evade siRNA-directed DNA methylation and silencing? Int J Mol Sci 14:15233–15259
Pumplin N, Voinnet O (2013) RNA silencing suppression by plant pathogens: defence, counter-defence and counter-counter-defence. Nature Rev Microbiol 11:745–760
Quintero A, Perez-Quintero AL, Lopez C (2013) Identification of ta-siRNAs and cis-nat-siRNAs in cassava and their roles in response to cassava bacterial blight. GPB 11(3):172–181. doi:10.1016/j.gpb.2013.03.001
Råberg L, Sim D, Read AF (2007) Disentangling genetic variation for resistance and tolerance to infectious diseases in animals. Science 318:812–814
Raja P, Sanville BC, Buchmann Bisaro DM (2008) Viral genome methylation as an epigenetic defense against geminiviruses. J Virol 82:8997–9007
Ramesh SV, Ratnaparkhe MB, Gupta GK, Husain SM (2014) Plant miRNAome and antiviral resistance: a retrospective view and prospective challenges. Virus Genes 48:1–14
Ratcliff FG, MacFarlane SG, Baulcombe DC (1999) Gene silencing without DNA: RNA mediated cross-protection between viruses. Plant Cell 11:1207–1215
Rodrigo G, Carrera J, Ruiz-Ferrer V, del Toro FJ, Llave C, Voinnet O, Elena SF (2012) A meta-analysis reveals the commonalities and differences in Arabidopsis thaliana response to different viral pathogens. PLoS ONE 7(7):e40526
Rodriquez-Negrete E, Lozano-Duran R, Piedra-Aguilera A, Cruzado L, Bejarano ER, Castillo AG (2013) Geminivirus Rep protein interferes with the plant DNA methylation machinery and suppresses transcriptional gene silencing. New Phytol 199:464–475
Rodríquez-Negrete EA, Carrillo-Tripp J, Rivera-Bustamante RF (2009) RNA silencing against geminivirus: complementary action of posttranscriptional gene silencing and transcriptional gene silencing in host recovery. J Virol 83:1332–1340
Sahu PP, Rai NK, Chakraborty S, Singh M, Chandrappa PH, Ramesh B, Chattopadhyay D, Prasad M (2010) Tomato cultivar tolerant to Tomato leaf curl New Delhi virus infection induces virus-specific short interfering RNA accumulation and defense-associated host gene expression. Mol Plant Pathol 11:531–544
Salomon R (1999) The evolutionary advantage of breeding for tolerance over resistance against viral plant disease. Isr J Plant Sci 47:I35–I39
Shaw J, Love AJ, Makarova SS, Kalinima NO, Harrison BD, Taliansky ME (2014) Coilin, the signature protein of cajal bodies, differentially modulates the interactions of plants with viruses in widely different taxa. Nucleus 5(1):85–94
Shivaprasad PV, Chen HM, Patel K, Bond DM, Santos ACM, Baulcombe DC (2012) A microRNA superfamily regulates nucleotide binding site-leucine rich repeats and other mRNAs. Plant Cell 24:859–874
Swevers L, Broeck JV, Smagghe G (2013) The possibible impact of persistent virus infection on the function of the RNAi machinery in insects: hypothesis. Front Physiol 4 319:1–15
Szittya G, Silhavy D, Molnár A, Havelda Z, Lovas A, Lakatos L, Bánfalvi Z, Burgyán J (2003) Low temperature inhibits RNA silencing-mediated defence by the control of siRNA generation. EMBO J 22(3):633–640
Tao Y, Xie Z, Chen W, Glazebrook J, Chang H-S, Han B, Zhu T, Zou G, Katagiri F (2003) Quantitative nature of Arabidopsis responses during compatible and incompatible interactions with the bacterial pathogen Pseudomonas syringae. Plant Cell 15:317–330
Thresh JM, Otim-Nape GW, Jennings DL (1994) Exploiting resistance to African cassava mosaic virus. Asp Appl Biol 39:51–60
Tiwari M, Sharma D, Trivedi PK (2014) Artificial microRNA mediated gene silencing in plants: progress and perspectives. Plant Mol Biol 86:1–18
Wang XB, Wu Q, Ito T, Cillo F, Li WX, Chen X, Yu JL, Ding SW (2010) RNAi-mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proc Natl Acad Sci USA 107(1):484–489
Zhai J, Jeong D-H, De Paoli E, Park S, Rosen BD, Li Y, González JA, Yan Z, Kitto LS, Grusak AM, Jackson SA, Stacey G, Cook DR, Green JP, Sherrier JD, Meyers CB (2011) MicroRNAs as master regulators of the plant NB-LRR defense gene family via the production of phased trans-acting siRNAs. Genes Dev 25(23):2540–2553
Acknowledgments
The authors wish to apologize to all authors whose valuable work was not mentioned within this review article due to space constraint. We thank the peer-reviewers for all the comments they made leading to the final version of the article. L. Bengyella was supported by URC post-doctoral funding from the School of Molecular and Cell Biology (CSM: 13203), University of the Witwatersrand, Johannesburg, South Africa.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Bengyella, L., Waikhom, S.D., Allie, F. et al. Virus tolerance and recovery from viral induced-symptoms in plants are associated with transcriptome reprograming. Plant Mol Biol 89, 243–252 (2015). https://doi.org/10.1007/s11103-015-0362-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0362-6