Abstract
Recent reports have shown that the molecular mechanisms involved in root stem-cell niche development in Arabidopsis thaliana are complex and contain several feedback loops and non-additive interactions that need to be analyzed using computational and formal approaches. Complex systems cannot be understood in terms of the behavior of their isolated components, but they emerge as a consequence of largely non-linear interactions among their components. The study of complex systems has provided a useful approach for the exploration of system-level characteristics and behaviors of the molecular networks involved in cell differentiation and morphogenesis during development. We analyzed the complex molecular networks underlying stem-cell niche patterning in the A. thaliana root in terms of some of the key dynamic traits of complex systems: self-organization, modularity and structural properties. We use these analyses to integrate the available root stem-cell niche molecular mechanisms data and postulate novel hypotheses, missing components and interactions and explain apparent contradictions in the literature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Located at the tip of the root, the root stem-cell niche (RSCN) sustains the development and growth of all below-ground tissues. Given its anatomical simplicity and accessibility, the Arabidopsis thaliana RSCN has become an excellent model system. The RSCN has been amenable to cellular and molecular genetic analyses unraveling a plethora of molecular regulatory mechanisms (MRMs) involved in maintaining its pattern and functionality. Partially due to the lack of data, until recently, most of the RSCN MRMs were understood as fragmented and isolated processes that were many times assumed to exhibit a linear relationship between genotype and phenotype. For example, currently, the identity and location of the RSCN is explained by the intersection of the expression patterns of a small set of transcription factors (Aida et al. 2004). However, recent findings reveal that a complex network composed of many interacting elements underlie RSCN patterning.
In her great introductory book to complexity, Mitchell (2009) described a complex system as “…a system that exhibits nontrivial emergent and self-organizing behaviors”. Indeed, complex systems comprise feedback loops and other non-linear interactions that produce the emergence of often non-intuitive behaviors that without the use of theoretical approaches seem impenetrable and many times preclude clear interpretations of the experimental data. The RSCN regulatory network involves several components interacting in non-linear ways. This does not mean that actual approaches are not useful; instead, systematic and integrative approaches can complement detailed analyses of particular molecular components, improving our understanding of the system. Such integrative approaches become more relevant if we consider that complex networks have systemic key structural and dynamic properties, such as self-organization and modularity, which cannot be understood by characterizations of isolated components.
Theoretical and computational approaches are necessary to study complex systems, like biological systems, affording the verification, prediction, and deeper understanding of experimental data with a more integral and systemic view (Strogatz 2001; Kitano 2002). While we will not review the many different tools available (But see: Alvarez-Buylla et al. 2007; Ay and Arnosti 2011), we will focus on the conceptual integration of experimental and theoretical approaches while analyzing the MRMs of RSCN patterning. To do this, we will describe some key properties, namely self-organization, modularity, and some structural and dynamic network properties, to guide the integrative description of the RSCN MRMs, detect missing components and interactions and provide novel hypotheses and plausible explanations for apparently contradictory data. Importantly, apart from the functional modularity and auxin-transport self-organization properties (Azpeitia et al. 2010; Mironova et al. 2010; Leyser 2011), the properties that we will describe have not been explicitly tested in the RSCN. However, the available data suggest that a robust complex molecular network with certain structural and dynamic characteristics, which are typical of complex networks, underlies RSCN patterning. Moreover, some of these properties appear to be generic to previously characterized MRMs (Barabási and Oltvai 2004; Kitano 2007).
In this review, first, we briefly describe the RSCN and the current explanation of how the RSCN is maintained. We then analyze the structural and dynamic characteristics of the RSCN MRMs with regard to complex systems approaches with a particular focus on network theory. Our analysis enabled us to propose novel explanations and propose experimentally verifiable predictions. For example, this approach is useful for uncovering and understanding the specific mechanisms of cell patterning, regenerative capacity and the maintenance of stem cells (SC) in the system under study. Finally, we discuss the implications and future directions of the ideas presented here.
The RSCN
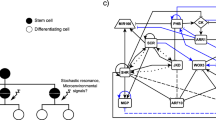
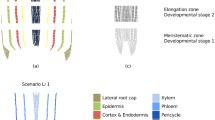
All primary tissues of the root develop from the RSCN. The RSCN consists of a small group of cells with low division rates called the quiescent center (QC), which are surrounded by a cell layer composed of four different cell types of initial or SCs (Fig. 1; Dolan et al. 1993). The QC is necessary for SC maintenance because its ablation or malfunction produces premature SC differentiation, RSCN consumption, and, finally, if not reestablished, root determinacy (van den Berg et al. 1995; van den Berg et al. 1997; Xu et al. 2006; Sarkar et al. 2007).
Multiple MRMs are involved in RSCN maintenance, and the most important MRMs identified thus far include the following: (1) The regulatory interactions sustained among the GRAS family transcription factors SHORTROOT (SHR) and SCARECROW (SCR) and a few additional genes, (2) the interplay between the redundant transcription factors PLETHORA1 (PLT1), 2, 3, BBM, and the auxin signaling pathway, (3) the CLAVATA LIKE40 (CLE40) and WUSCHEL RELATED HOMEOBOX5 (WOX5) MRM, (4) many hormonal signaling pathways in addition to the auxin signaling pathway and (5) epigenetic mechanisms (reviewed in Scheres 2007; Benková and Hejátko 2009; Shen and Xu 2009; Sablowski 2011).
Molecular genetic approaches have suggested that the identity and location of the RSCN depends on the intersection of the SHR, SCR and PLT protein domains (Aida et al. 2004) and the negative regulation of the WOX5 QC identity marker by CLE40 (Stahl et al. 2009). Because the PLT transcriptional and protein domains depend on the auxin concentration (Aida et al. 2004; Zhou et al. 2010), the maximum auxin concentration coincides with the QC location (Brunoud et al. 2012), and auxin signaling, transport and metabolism modifications alter the RSCN (Ding and Friml 2010), auxin is assumed to have a fundamental role in RSCN specification. Finally, epigenetic mechanisms modulate the expression location and level of, at least, the SCR and PLT genes (Shen and Xu 2009). We recently published a model that demonstrated that the concerted action of at least the first three MRMs mentioned above, and not the isolated activity of any such MRMs, is necessary to understand how the RSCN is specified and maintained (Azpeitia et al. 2010). Importantly, our results suggested that the characterized RSCN regulatory network is incomplete because the model was not capable of reproducing important processes observed in the RSCN such as its robustness. We believe that a complex systems perspective such as the one used here may be used to propose the missing components and interactions necessary for the production of the RSCN observed systemic behaviors and aid in the achievement of a better understanding of the properties of the MRMs underlying the RSCN.
Complex system approaches to RSCN patterning
We now use a complex systems-based approach to analyze the integrated action of the above mentioned MRMs during RSCN patterning and study some systems-level traits and behaviors of the integrated network. We also discuss whether such a systematic and integrative approach reveals novel predictions to be tested experimentally or innovative approaches towards understanding how the cellular patterns and organization of the RSCN emerge.
Structural network-based study of the RSCN MRM
A network is composed of components called nodes that are connected through edges. In molecular regulatory networks, the nodes usually represent genes, proteins or molecules (e.g., hormones), while the edges represent regulatory interactions (reviewed in Barabási and Oltvai 2004; Albert 2007).
The most basic structural features of networks are their degree (also called connectivity) and degree distribution. The connectivity or degree k refers to the number of direct links that one node has with the other nodes of the network, while the degree distribution P(k) refers to the probability that a randomly selected node has a specific degree k. Many biological networks follow a power law degree distribution (Babu et al. 2004) or a similar long-tail distribution. A power law degree distribution means that P(k) ≈ Ak −λ, where A is a normalization constant and λ is the degree exponent. Networks with a power law degree distribution are also known as scale-free networks. As observed with the degree distribution, in scale-free networks, there are many low degree nodes, while a few of the nodes, known as hubs, have high degrees (Barabási and Oltvai 2004; Albert 2007). Because of their high connectivity, hubs have been proposed as important nodes for network functionality, connecting nodes that can participate in different processes and bringing together the network as a whole (Barabási and Oltvai 2004).
Although we lack a large enough network structure or architecture for RSCN patterning to allow for statistical analyses of the degree distribution, more than one of the genes involved in RSCN maintenance are probably hubs. For example, SHR and SCR regulate hundreds of genes (Sozzani et al. 2010), a fact that is reflected by the many processes in which they are involved apart from RSCN maintenance such as root regeneration (Xu et al. 2006; Sena et al. 2009), lateral root development (Lucas et al. 2011), cell cycle (Sozzani et al. 2010), root radial patterning (Helariutta et al. 2000), middle cortex formation (Cui and Benfey 2009), vascular development (Carlsbecker et al. 2010; Cui et al. 2011) and stress response (Cui et al. 2012).
Scale-free networks present the small-world propertiy. The small-world property refers to the shortest possible path to travel from a node to any other node using only directly linked network nodes (Watts and Strogatz 1998). In small-world networks, nodes are connected to each other through a short path. Importantly, the small-world property has been reported in biological networks (Wagner and Fell 2001). Most of the MRMs involved in RSCN maintenance were initially characterized as independent of each other; however, recent work has discovered some links among them, creating short communication paths. For example, the SHR/SCR and PLT/auxin MRM were first described as independent MRMs (Aida et al. 2004). However, Lucas et al. (2011) recently reported that shr single mutants have an excessive accumulation and synthesis of auxin during the first 6 days after germination and have a progressive reduction in the auxin transport facilitators PINFORMED (PIN) expression in the root tip, likely regulating PIN abundance at the posttranscriptional level or indirectly regulating their expression, as suggested by Levesque et al. (2006). Moreover, SHR and SCR up-regulate the expression of miR165a, miRNA166a and miR166b (collectively referred as miR165/6). miR165/6 can diffuse from its site of expression and negatively regulate the post transcriptional expression of the HD-ZIP III gene PHABULOSA (PHB) (Carlsbecker et al. 2010; Miyashima et al. 2011). HD-ZIP III genes apparently act by antagonizing the PLT genes in the RSCN (Smith and Long 2010). Moreover, during embryogenesis, HD-ZIP III genes regulate auxin flow (Izhaki and Bowman 2007), a fact that needs to be tested in the case of the root. Importantly, the PLT/auxin MRM feeds back to the SHR/SCR MRM. An analysis of whole seedlings revealed that HD-ZIP III genes expression is induced by auxin (Zhou et al. 2007), and the inhibition of PHB and its redundant gene PHAVOLUTA in the basal pole during embryogenesis is necessary for SCR and WOX5 expression and thus proper RSCN development (Grigg et al. 2009; Fig. 2).
Shortest paths connecting the SHR/SCR (purple) and the PLT/Auxin (orange) modules as described in the main text. Simplified versions of these modules are depicted. Blue edges highlight the short paths that connect both modules. As observed, even when these were characterized as independent pathways or modules, they have multiple short communication paths. Arrowheads represent positive interactions, T arrowheads represent negative interactions, doted arrowhead auxin transport facilitation and diamond arrowhead antagonistic probably non-regulatory interactions. ARFa ARF activator, ARFr ARF repressor, CK cytokinin
The last example is not the only example of the communication of MRMs through short paths. Many hormones are important for the RSCN including auxin, cytokinins (CK), ethylene, brassinosteroids (BR), jasmonate and abscisic acid, all of which alter the RSCN pattern, functionality or development (Ortega-Martínez et al. 2007; Müller and Sheen 2008; Ding and Friml 2010; Zhang et al. 2010; Chen et al. 2011a; González-García et al. 2011). However, hormones do not act trough isolated pathways or MRMs; they instead regulate each other at the biosynthesis, signal transduction and transport levels. For example, ethylene, CK and auxin regulate the synthesis of each other (Nordström et al. 2004; Tsuchisaka and Theologis 2004; Stepanova et al. 2005; Swarup et al. 2007; Stepanova et al. 2008; Jones et al. 2010; Zhou et al. 2011), PIN auxin transporter expression is affected by CK, BR, ethylene and auxin itself (Blilou et al. 2005; Vieten et al. 2005; Ruzicka et al. 2007; Dello Ioio et al. 2008; Ruzicka et al. 2009; Zhang et al. 2011), and the effects of ethylene on cell elongation are dependent on the auxin signaling pathway (Swarup et al. 2007; Stepanova et al. 2007; Fig. 3). Indeed, hormonal cross-talk is important for root patterning (reviewed in Benková and Hejátko 2009). The evidence reviewed here suggests that the small-world property is present in the whole RSCN network and demonstrates that the SHR/SCR, the PLT/auxin MRMs, and the hormone signaling pathways, which were originally reported as independent MRMs, are interconnected through short and most likely multiple pathways.
Cross-talk among auxin, cytokinin and ethylene pathways involved in RSCN patterning as described in the main text. As observed, these hormones pathways have multiple interactions or crosstalk connections at the signaling, synthesis and transport levels demonstrating that they are part of an integrated complex network with short communicating paths
Importantly, due to the presence of short communication paths, the small-world property proposes that modifications in one MRM can have unexpected effects in other MRMs (Watts and Strogatz 1998), while, at the same time, allowing for simpler and direct explanations of such effects. For example, the PIN genes and WOX5 expression are affected in scr and shr single mutant backgrounds, even though neither PIN nor WOX5 appear to be direct target genes of SHR or SCR (Levesque et al. 2006; Sarkar et al. 2007; Sozzani et al. 2010). Based on the fact that SHR directly up-regulates the expression of cytokinin oxidase 3 (CKX3; Cui et al. 2011), a CK catabolism enzyme, and that CK represses PIN expression (Ruzicka et al. 2009; Zhang et al. 2011), one possible explanation is that SHR indirectly regulates PIN expression through its down-regulation of CK synthesis. Other possible explanations are (1) that SHR and SCR regulate PIN expression through its effect on auxin, as shr mutants accumulate auxin and high concentrations of auxin reduce the PIN protein levels (Vieten et al. 2005; Lucas et al. 2011), or (2) through an effect of SHR and SCR on PHB (Carlsbecker et al. 2010) given that the HD-ZIP III genes regulate auxin flow (Izhaki and Bowman 2007; Fig. 2). SHR and SCR could modulate WOX5 through its regulation of the HD-ZIP III genes or through another indirect mechanism. For example, CLE10 was recently reported as a putative SHR target gene (Cui et al. 2011). However, CLE10 has been described as a peptide involved in protoxylem vessel formation and not in RSCN maintenance (Kondo et al. 2011).
Interestingly, neither hub importance nor small-world properties are rules in biological networks. The deletion of the more connected genes does not necessarily lead to the most drastic phenotypes (e.g., Espinosa-Soto et al. 2004), and the perturbation of a MRM does not alter all other MRMs. Why does this happen in biological networks? High connectivity is not necessarily directly related to functionality in a network. Other measurements, such as betweenness (i.e., the number of shortest paths that pass through a node), may also determine the functionality of a node (Goh et al. 2002). In addition, positive feedback loops appear to make biological networks more robust against mutations in highly connected nodes (Espinosa-Soto et al. 2004). However, understanding how structure and function are related is a difficult task and entails different approaches. Theoretical biology has proposed other properties of biomolecular networks, such as modularity, which may help explain why neither hub importance nor small-world properties are rules in biological networks.
The RSCN as a modular system
Recent research suggests that biological networks usually have modular organization. At a structural level, network modules are usually defined as a subset of network components that are more connected with each other than with other components of the network (Fig. 4; Wagner et al. 2007; Espinosa-Soto and Wagner 2010). Modular organization may reduce the pleiotropic effects of perturbations (such as mutants) in the network (von Dassow and Munro 1999; Wagner et al. 2007) because modules have a relatively autonomous behavior with respect to the rest of the network. Hence, such modularity may help explain why, in biological networks, mutations of highly interconnected nodes may not alter the phenotype, or they do not necessarily behave as expected for small-world networks.
As previously mentioned, some interactions occur between the SHR/SCR MRM and the PLT/auxin MRM. However, there are multiple interactions within PLT/auxin and SHR/SCR MRMs (Figs. 2, 5, 6). PLT genes expression patterns are altered by auxin addition, transport inhibition and signaling pathway mutants (Aida et al. 2004; Blilou et al. 2005; Galinha et al. 2007). The PLTs response to auxin is partially dependent on tyrosylprotein sulfotransferase (TPST). TPST controls the activity of the secreted peptide portion of the root meristem growth factors 1 (RGF1), 2 and 3 (herein RGFs) by Tyr sulfation (Matsuzaki et al. 2010). RGFs expression is auxin-independent, while TPST expression is auxin-dependent. Matsuzaki et al. (2010) proposed that RGFs probably stabilize PLT proteins based on the fact that wild-type seedlings treated with RGF1 expand PLT1 and 2 protein domains but not PLT1 or PLT2 transcriptional domains. However, other results demonstrated that tpst mutants reduce PLT at the transcriptional and protein levels, demonstrating that TPST can control PLT expression at both levels (Zhou et al. 2010). Interestingly, PLT genes control auxin transport, which is indispensable for the observed auxin graded concentration in the root, creating a loop in which PLT genes simultaneously control and are controlled by auxin distribution (Aida et al. 2004; Blilou et al. 2005; Galinha et al. 2007). The RopGEF7 gene is positively regulated by auxin and acts as a positive regulator of PLT expression. Interestingly, RopGEF7 also affects the auxin transport and response in the RSCN (Chen et al. 2011b). Moreover, as described below, there are multiple feedback loops within the auxin signaling pathway, greatly increasing the connectivity of the MRM (Fig. 5).
The PLT/Auxin (orange) and the SHR/SCR (purple) structural modules. The whole RSCN network is probably divided into several structural modules, based on their relative intra-module versus inter-module interactions: in these two modules there are many more within-module interactions than between-module connections (see also Fig. 2). Arrowheads as in Fig. 2
On the other hand, SCR and SHR form a dimer, and they together directly up-regulate MAGPIE (MGP) expression and JACKDAW (JKD) postembryonic expression (Levesque et al. 2006; Welch et al. 2007; Cui et al. 2011). Mutations in JKD diminish SCR expression in the QC and cortex-endodermis initials (CEI), causing a misspecification of the QC and ectopic periclinal divisions of the CEI. The double mutant jkd mgp restores SCR expression in the QC and CEI, suggesting that MGP is a negative regulator of SCR. Yeast two-hybrid and transient assays have shown that SHR, SCR, JKD, and MGP can physically interact and modulate the expression of and transcriptional activity of each other (Welch et al. 2007; Ogasawara et al. 2011). SCR also acts as a negative regulator of MGP when it forms a dimer with like heterochromatin protein1 (LHP1) (Cui and Benfey 2009). LHP1 is a candidate protein for the histone trimethylation that is necessary for PcG repression activity (Exner et al. 2009). As mentioned above, SHR and SCR negatively regulate PHB through miR165/6. Recently, Miyashima et al. (2011) demonstrated that the restriction of PHB to the stele is necessary for maintaining JKD expression. Consistent with this result, the gain-of-function allele phd-1d has a similar ground tissue phenotype as the jkd single mutant with low SCR expression levels, which may explain why PHB repressed SCR expression in a previous study (Grigg et al. 2009; Fig. 5).
Until now, we have considered a structural definition of modularity. However, dynamic regulatory network models may be used to test whether a set of interacting genes or other molecular components constitute a functional module that is sufficient and necessary to recover an observed gene-state configuration and its spatial pattern. Importantly, structural and functional modules may not coincide (Ten Tusscher and Hogeweg 2011). For example, in Azpeitia et al. (2010), we aimed to analyze whether PLT/auxin, SHR/SCR, and CLE/WOX5 MRMs were sufficient to reproduce the stable gene configurations that characterize each cell type within the RSCN and their observed spatial distribution. We found that some important interactions are missing, but by adding a few predictions, our RSCN regulatory network model constitutes a functional module that incorporates the necessary and sufficient MRMs required to recover the genetic configurations that have been described for the main cell types of the RSCN and their observed qualitative spatial patterning. However, as mentioned above, each of the MRMs considered in the proposed RSCN network may constitute a different structural module (Fig. 6). Thus, dynamic analyses are fundamental to the understanding of how the MRM structure and function are related.
Once a dynamic model is developed, the model can be validated with experimental evidence. For example, our model is capable of reproducing most of the mutant phenotypes observed, but it is not as robust as expected from experimental observations and from comparison with other biomolecular networks (Azpeitia et al. 2010). This indicates that the uncovered regulatory module is incomplete. Additional components, interactions or complete modules may be necessary to recover the observed robustness and overall RSCN behavior. For example, epigenetic MRMs were not considered in our model and are fundamental to RSCN maintenance because SCR or PLT have an altered expression in mutants of the TONSOKU (TSK), TEBICHI (TEB), FASCIATA1 (FAS1) and FAS2, NAP1-related protein1 (NRP1) and NRP2, Polycomb group (PcG), general control nonderepressible protein5 (GCN5) and Pickle (PKL) genes, which are all involved in epigenetic regulation (reviewed in Shen and Xu 2009). Moreover, all mutants of these genes have alterations in RSCN patterning, the expression of QC markers, and/or columella SC differentiation (Shen and Xu 2009). These epigenetic MRMs, which are part of structural modules that are different than the ones incorporated thus far, may be required to recover observed dynamic behaviors in the RSCN patterning and the regulatory networks involved within it. Thus, such epigenetic mechanisms should be an important part of the RSCN functional module even when they are likely a part of different structural modules. However, we believe that one of the most interesting and important properties of the dynamic analysis of complex networks is self-organization. This may also provide important biological insights into such networks, e.g., in terms of uncovering missing components or interactions.
The RSCN as a self-organized system
Self-organization refers to the emergence of patterns or behaviors at the global system level as a consequence of non-linear interactions among system components without depending upon the action of a central controller (Seeley 2002). For example, the stable gene-state configurations (attractors) that characterize each cell type within a RSCN constitute a self-organized property of the complex regulatory module. These attractors emerge as a consequence of the concerted action of all of the molecular interactions considered in the particular network under consideration. At a different level of organization, the spatial cellular arrangement that characterizes the RSCN may also be considered as a self-organized pattern resulting from the coupled dynamics of intracellular networks in a multicellular spatial domain. In the case under consideration, the coupling of the dynamics of the intracellular networks occurs via the intercellular movement of some of the network components (Azpeitia et al. 2010).
The lack of a central controller means that self-organization arises from the local interactions of the system components and not from an individual component at any level of organization that works as a “guiding” unit. In this sense, it is relevant to uncover the structure and dynamics of intracellular biological networks and their coupling mechanisms among cells, rather than only concentrating on the role of so-called “key” genes. In contrast, we should understand, in terms of integrated regulatory networks, what makes a “key” gene key and why the mutation of such genes are sometimes sufficient to take the system from one multigenic and multicellular configuration to another one in contrast to mutations in other genes that are not “key”.
In the root, there are many traits and behaviors that suggest the existence of self-organizing processes at the macro and micro levels. The regeneration process is perhaps one of the most obvious and best examples of self-organization in the RSCN. Normally, the RSCN develops from early embryonic stages (Dolan et al. 1993); however, when the root tip is excised or the QC ablated, a new RSCN is formed (Xu et al. 2006; Sena et al. 2009). The process of root and RSCN regeneration follows an ordered sequence of events. It begins with the formation of a new auxin maximum, which induces PLT expression and, later, SHR nuclear localization and SCR expression where the new RSCN will be relocated. SHR, SCR, PLT genes and the auxin maximum are all necessary for the RSCN regeneration process (Xu et al. 2006). After the root tip is excised, the RSCN is completely eliminated, and the root tip pattern is severely affected. However, even in the absence of the original pattern, RSCN regeneration proceeds (Sena et al. 2009). Moreover, callus regeneration from root, cotyledon, and petals, all of which have completely different morphologies, resemble the root regeneration process (Sugimoto et al. 2010). The fact that the RSCN can regenerate without a specific pre-established pattern and employ multiple molecular components strongly suggests that no single molecular component, module, external agent or pre-pattern directs the RSCN regeneration and patterning process. The process is instead due to the self-organizing capability of the molecular regulatory network involved, and it is likely that additional coupling constraints such as physical and chemical (e.g., hormone) fields are also involved.
The root auxin gradient is another excellent example of a self-organizing process involved in RSCN maintenance (Leyser 2011). Theoretical and experimental studies have suggested that the graded distribution of auxin along the root longitudinal axis depends on the polar localization of the PIN proteins (Blilou et al. 2005; Vieten et al. 2005; Grieneisen et al. 2007; Mironova et al. 2010) and on auxin metabolism (synthesis and degradation; Stepanova et al. 2008; Petersson et al. 2009). However, the polar localization of the PIN transporters is, in turn, regulated by the auxin-signaling pathway (Vieten et al. 2005; Sauer et al. 2006), which partly depends on the auxin cell concentration that at the same time, depends on the auxin gradient (Tiwari et al. 2001; Kepinski and Leyser 2005; Fig. 5). Hence, the auxin gradient is at the same time a cause and effect through its interdependences with the auxin signaling and transport mechanisms and not a process directed by any specific fixed force.
Self-organizing systems have a dissipative structure behavior, meaning that they are far from thermodynamic equilibrium. Interestingly, dissipative structures are maintained in steady states or attractors (Prigogine 1978), which can adapt and adjust to internal and external changes, allowing them to maintain coherent patterns or functions under a wide range of conditions and several types of perturbations (Heylighen 2001) that could include genetic loss or gain of function mutations. The latter behaviors are observed in the above examples of RSCN regeneration because it occurs in different types of RSCN lesions and different organs (Xu et al. 2006; Sena et al. 2009; Sugimoto et al. 2010). The auxin gradient is maintained even under some PIN and auxin signaling mutant backgrounds and during auxin addition or overproduction (Blilou et al. 2005; Vieten et al. 2005; Grieneisen et al. 2007). Finally, the overall multigenic RSCN configurations and cellular patterns are maintained in the presence of several mutations. However, without understanding how such apparently dispensable nodes are connected to the “key” nodes, we cannot understand the molecular regulatory basis of the overall behavior of the system.
Hormone signaling pathways are probably better examples of self-organizing, dissipative, and adaptable systems, i.e., they return to a basal state or attractor after perturbation. Such attractors are usually the inactive state of the pathway when the hormone is absent or present at low concentrations. Such systems also modulate and stabilize (adapt) their response when their inputs, usually hormone concentrations, change. There are multiple ways in which hormone pathways can reach this adaptable behavior (e.g., Dreher and Callis 2007).
As previously mentioned, one of the best-studied pathways during RSCN patterning is the auxin pathway. Auxin signaling can be modulated by adjusting the auxin concentration. As stated above, the auxin concentration depends on its transport, which is a self-organizing process, but it is also dependent on auxin metabolism. Auxin metabolism is regulated by multiple signals. Interestingly, auxin signaling also regulates some of these signals. For example, auxin inhibits CK synthesis (Nordström et al. 2004) and promotes ethylene synthesis (Rahman et al. 2001), while both CK and ethylene promote auxin synthesis (Stepanova et al. 2008; Jones et al. 2010; Zhou et al. 2011; Fig. 3). The internal components of the auxin pathway are also regulated to maintain a coherent adaptive response. Receptors of the transport inhibitor response1/auxin signaling f-box protein1–5 (TIR1/AFB) detect auxin. TIR/AFB are components of the SKP1/Cullin/F-box protein (SCF)TIR1/AFB ubiquitin ligase complex (Mockaitis and Estelle 2008) and are negatively regulated post-transcriptionally by miR393. The SCFTIR1/AFB complex promotes Aux/IAA degradation in the presence of auxin (Tiwari et al. 2001; Kepinski and Leyser 2005). Interestingly, Aux/IAAs act as an auxin co-receptor because TIR/AFB cannot readily bind auxin in the absence of Aux/IAA proteins (Calderón Villalobos et al. 2012). The Aux/IAA proteins form heterodimers with the Auxin Response Factor (ARF) proteins that mediate auxin transcriptional regulation (Tiwari et al. 2001). Some ARFs act as transcriptional activators (ARFa), while others act as transcriptional repressors (ARFr; Guilfoyle and Hagen 2007), and all compete for the regulation of the same target genes (Ulmasov et al. 1999); thus, the ARFa/ARFr ratio also modulates the auxin signaling response (Vernoux et al. 2011). Through its transcriptional activity, ARFa generates many feedback loops inducing the induction of Aux/IAA expression, and probably miR393 and some ARF family members (Wang et al. 2005; Paponov et al. 2008; Parry et al. 2009; Chen et al. 2011c; Fig. 3).
Importantly, the auxin signaling pathway can differentially modulate its response, depending on the Aux/IAA, ARF and TIR/AFB members involved in each specific process, because the members of these gene families have partially redundant functions, but also have specific characteristics that are involved in different responses (Hardtke et al. 2004; Parry et al. 2009; Calderón Villalobos et al. 2012; Rademacher et al. 2012). For example, the members of the TIR/AFB family have different strengths of interaction with Aux/IAAs, and different Aux/IAA-TIR/AFB complexes have different sensitivities to the auxin concentration (Parry et al. 2009; Calderón Villalobos et al. 2012). Thus, auxin signaling can adapt to several conditions, modulating the auxin concentration, adjusting the expression level and activity of the different receptors and proteins involved in the pathway through many feedback loops, adjusting the ARFa/ARFr ratio and selecting specific members of the Aux/IAA, ARF and TIR/AFB gene families that are involved in each response and condition.
Explicitly considering the self-organizing properties of a complex dynamic system could help resolve some apparently conflicting points regarding the MRMs involved in RSCN patterning. For example, some reports have shown that auxin promotes the expression of WOX5 (Gonzali et al. 2005; Sarkar et al. 2007; Sena et al. 2009; Sugimoto et al. 2010). In contrast, a recent manuscript by Ding and Friml (2010) demonstrated that elevated auxin concentrations in the root tip lead to the consumption of the RSCN and WOX5 expression inhibition. It is likely that some of these apparent contradictions can be explained by invoking the different responses and roles of the members of the ARF, Aux/IAA and TIR1/AFB gene families. However, another complementary hypothesis that may help reconcile such apparently contradictory results is that different responses to auxin signaling are explained by the self-organizing and self-regulating properties of the pathway. For example, self-organizing mechanisms generally involve, as has been uncovered for biological networks, many regulatory motifs such as feedback loops, which provide several properties to self-organizing processes. Positive feedback loops amplify a received signal or stimulus, allowing switch-like behavior, bistability, and hysteresis (Kitano 2004; Mitrophanov and Groisman 2008). On the other hand, negative feedback loops play an important role by dampening fluctuations, providing stability and limiting the fluctuating range of the components (Kitano 2004; Becskei and Serrano 2000). As mentioned above, positive and negative feedback loops are present in the auxin signaling pathway self-organizing process. The expression of ARF16, which is a putative ARFr, and ARF19, which is an ARFa, is promoted by auxin, thus creating a negative and a positive feedback loop, respectively (Wang et al. 2005; Paponov et al. 2008). The presence of these feedback loops could generate different auxin responses under different conditions, which could help explain why, under some circumstances, auxin inhibits WOX5 expression, while in other conditions auxin promotes WOX5 expression. For example, ARFa and ARFr compete for the same target genes (Ulmasov et al. 1999), and the expression patterns of ARF16 and ARF19 in the root tip are similar (Rademacher et al. 2011). Thus, if ARF16 expression is promoted to a greater extent by auxin than ARF19 expression, we expect that auxin addition would repress WOX5 expression as observed by Ding and Friml (2010). However, if ARF19 is induced by auxin to a greater extent than ARF16, we could expect an induction of WOX5 under auxin addition conditions as observed by the research of Gonzali et al. (2005) and Sugimoto et al. (2010). Hence, we may observe auxin inductive and repressive responses over WOX5 under different conditions, and this would not imply contradictory experimental observations.
Missing links in the RSCN network
The analyses of structural and dynamic MRM properties also allow for the detection of missing links and non–intuitive behaviors in the MRM that may guide future experimental studies that may otherwise be obviated. For example, Miyashima et al. (2011) suggested that miR165/6 acts as a morphogen. Theoretical research has demonstrated that the morphogen graded distribution alone is not capable of generating such highly precise patterns as the ones observed in the root vasculature. Theoretical analyses of complex systems have demonstrated that feedback loops have a critical role in the generation of robust and precise patterns (e.g., Jaeger et al. 2008). HD-ZIP III genes form a negative feedback loop with ZPR proteins in the shoot (Kim et al. 2008). It will be interesting to investigate if HD-ZIP III genes also create this or similar feedback loops in the root as suggested by theoretical studies and the observed patterns (Jaeger et al. 2008).
CLE40/WOX5 MRM is fundamental for RSCN maintenance (Sarkar et al. 2007; Stahl et al. 2009). However, until now, little information has been gathered regarding this MRM in the RSCN. Stahl et al. (2009) reported that CLE40 reduces WOX5 expression. A similar MRM acts in the shoot stem cell niche, where the expression of WUS, a WOX5 homolog, is repressed by CLV3 (Sablowski 2011). The WOX5/CLE40 and WUS/CLV3 MRMs are likely similar in structure and dynamic behavior. This is supported by the fact that WOX5 can be substituted by WUS, that CLV3 can be partially substituted by CLE40 in the shoot, and that CLV3 and CLE40 over expression and peptide addition produce similar root phenotypes (Hobe et al. 2003; Fiers et al. 2005; Sarkar et al. 2007). Moreover, the protein phosphatases POLTERGEITS (POL) and PLL1 act downstream of the WOX5/CLE40 and WUS/CLV3 MRMs. In both cases, the CLV pathway inhibits POL and PLL1 activity, which is necessary for WOX5 and WUS expression in the root and shoot, respectively (Song et al. 2008; Gagne et al. 2010). However, unlike the WOX5/CLE40 MRM, the WUS/CLV3 MRM has been thoroughly studied, revealing the presence of many feedback loops (Gordon et al. 2009; Chickarmane et al. 2012). The similarity between the WOX5/CLE40 and the WUS/CLV3 MRMs and the key importance of feedback loops in network behavior and emerging pattern robustness may guide researchers to search for feedback motifs that are similar to the ones that have been uncovered in the shoot in the WOX5/CLE40 MRM. Our own studies suggest a few yet uncovered feedback loops in the root MRMs that should be experimentally documented: a negative feedback loop between CLE40 and WOX5 and a positive self-regulation of WOX5, which likely occurs via the auxin signaling pathway (Azpeitia et al. 2010, and unpublished data).
Conclusions
Complex systems approaches are becoming fundamental to the understanding of regulatory systems that result from non-linear interactions among multiple components that act in concert during a particular cell differentiation or morphogenetic process, rather than by being directed by single and isolated genes or any type of central controller. In this review, we have used structural and dynamic properties of complex networks, like modularity and self-organization, to integrate and provide novel explanations for the plethora of molecular genetic involved in the RSCN of A. thaliana. Due to space limitations, we only focused on a few structural and dynamic properties of complex networks. However, there are other properties, such as robustness, which are pervasive among complex systems. Robustness refers to the ability of systems to retain their functionality despite perturbations. For the interested reader, excellent reviews have focused on this aspect (Kitano 2007; Whitacre 2012) and other important properties of complex systems (Mitchell 2009).
These properties constitute useful conceptual frameworks for analysis at the systemic level, and importantly, they yield complementary information to achieve a better and more profound mechanistic understanding of complex systems such as the regulatory networks involved in the patterning of SC niches and other biological structures. For example, structural analyses allowed for a global picture of the involved network topology, which may be useful for uncovering missing components or interactions and identifying novel behaviors or roles for particular nodes. Modularity at the structural level also helps us understand why relying on only the simplest structural traits, such as degree and degree distribution, may be misleading with regards to the role of particular nodes in overall network dynamics. However, importantly, structural modularity does not necessarily coincide with functional modularity; hence, when trying to uncover the structure–function relationship, additional dynamic approaches are required to verify the necessity and sufficiency of the components that are being considered in the overall system behavior or dynamics. Hence, structural and dynamic analyses complement each other, which also allows for the prediction of novel components and interactions and the evaluation of the functional role of characterized components or the proposal of innovative systemic explanations. As we observed, all of these properties are interconnected and together provide a powerful vision for the study of complex systems.
In this review, we have shown that enough molecular genetic information has been uncovered for the MRMs involved in RSCN patterning to allow for an integrative approach that explores the structural and dynamic properties characteristic of complex systems. This approach has allowed us to postulate a novel, non-intuitive hypothesis, which may guide future experimental studies, and provide explanations for apparently contradictory experimental evidence. Therefore, this type of analysis may guide research and then feedback to further the understanding of RSCN patterning. A dynamic interplay among theoretical, experimental and comparative analyses will greatly contribute to the further understanding of the complex nature of the regulatory systems involved in cell differentiation and morphogenesis during plant and animal development.
References
Aida M, Beis D, Heidstra R, Willemsen V, Blilou I, Galinha C, Nussaume L, Noh YS, Amasino R, Scheres B (2004) The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell 119:109–120
Albert R (2007) Network inference, analysis, and modeling in systems biology. Plant Cell 19:3327–3338
Alvarez-Buylla ER, Benítez M, Dávila EB, Chaos A, Espinosa-Soto C, Padilla-Longoria P (2007) Gene regulatory network models for plant development. Curr Opin Plant Biol 10:83–91
Ay A, Arnosti DN (2011) Mathematical modeling of gene expression: a guide for the perplexed biologist. Crit Rev Biochem Mol Biol 46:137–151
Azpeitia E, Benítez M, Vega I, Villarreal C, Alvarez-Buylla ER (2010) Single-cell and coupled GRN models of cell patterning in the Arabidopsis thaliana root stem cell niche. BMC Syst Biol 4:134
Babu MM, Luscombe NM, Aravind L, Gerstein M, Teichmann SA (2004) Structure and evolution of transcriptional regulatory networks. Curr Opin Struct Biol 14:283–291
Barabási AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–113
Becskei A, Serrano L (2000) Engineering stability in gene networks by autoregulation. Nature 405:590–593
Benková E, Hejátko J (2009) Hormone interactions at the root apical meristem. Plant Mol Biol 69:383–396
Blilou I, Xu J, Wildwater M, Willemsen V, Paponov I, Friml J, Heidstra R, Aida M, Palme K, Scheres B (2005) The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433:39–44
Brunoud G, Wells DM, Oliva M, Larrieu A, Mirabet V, Burrow AH, Beeckman T, Kepinski S, Traas J, Bennett MJ, Vernoux T (2012) A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482:103–106
Calderón Villalobos LI, Lee S, De Oliveira C, Ivetac A, Brandt W, Armitage L, Sheard LB, Tan X, Parry G, Mao H, Zheng N, Napier R, Kepinski S, Estelle M (2012) A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nat Chem Biol 8:477–485
Carlsbecker A, Lee JY, Roberts CJ, Dettmer J, Lehesranta S, Zhou J, Lindgren O, Moreno-Risueno MA, Vatén A, Thitamadee S, Campilho A, Sebastian J, Bowman JL, Helariutta Y, Benfey PN (2010) Cell signalling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465:316–321
Chen Q, Sun J, Zhai Q, Zhou W, Qi L, Xu L, Wang B, Chen R, Jiang H, Qi J, Li X, Palme K, Li C (2011a) The basic helix-loop-helix transcription factor MYC2 directly represses PLETHORA expression during jasmonate-mediated modulation of the root stem cell niche in Arabidopsis. Plant Cell 23:3335–3352
Chen M, Liu H, Kong J, Yang Y, Zhang N, Li R, Yue J, Huang J, Li C, Cheung AY, Tao LZ (2011b) RopGEF7 regulates PLETHORA-dependent maintenance of the root stem cell niche in Arabidopsis. Plant Cell 23:2880–2894
Chen ZH, Bao ML, Sun YZ, Yang YJ, Xu XH, Wang JH, Han N, Bian HW, Zhu MY (2011c) Regulation of auxin response by miR393-targeted transport inhibitor response protein 1 is involved in normal development in Arabidopsis. Plant Mol Biol 77:619–629
Chickarmane VS, Gordon SP, Tarr PT, Heisler MG, Meyerowitz EM (2012) Cytokinin signaling as a positional cue for patterning the apical-basal axis of the growing Arabidopsis shoot meristem. Proc Natl Acad Sci USA 109:4002–4007
Cui H, Benfey PN (2009) Interplay between SCARECROW, GA and LIKE HETEROCHROMATIN PROTEIN 1 in ground tissue patterning in the Arabidopsis root. Plant J 58:1016–1027
Cui H, Hao Y, Kovtun M, Stolc V, Deng XW, Sakakibara H, Kojima M (2011) Genome-wide direct target analysis reveals a role for SHORT-ROOT in root vascular patterning through cytokinin homeostasis. Plant Physiol 157:1221–1231
Cui H, Hao Y, Kong D (2012) SCARECROW has a SHORT-ROOT independent role in modulating sugar response. Plant Physiol 158:1769–1778
Dello Ioio R, Nakamura K, Moubayidin L, Perilli S, Taniguchi M, Morita MT, Aoyama T, Costantino P, Sabatini S (2008) A genetic framework for the control of cell division and differentiation in the root meristem. Science 322:1380–1384
Ding Z, Friml J (2010) Auxin regulates distal stem cell differentiation in Arabidopsis roots. Proc Natl Acad Sci USA 107:12046–12051
Dolan L, Janmaat K, Willemsen V, Linstead P, Poethig S, Roberts K, Scheres B (1993) Cellular organisation of the Arabidopsis thaliana root. Development 119:71–84
Dreher K, Callis J (2007) Ubiquitin, hormones and biotic stress in plants. Ann Bot 99:787–822
Espinosa-Soto C, Wagner A (2010) Specialization can drive the evolution of modularity. PLoS Comput Biol 6:e1000719
Espinosa-Soto C, Padilla-Longoria P, Alvarez-Buylla ER (2004) A gene regulatory network model for cell-fate determination during Arabidopsis thaliana flower development that is robust and recovers experimental gene expression profiles. Plant Cell 16:2923–2939
Exner V, Aichinger E, Shu H, Wildhaber T, Alfarano P, Caflisch A, Gruissem W, Köhler C, Hennig L (2009) The chromodomain of LIKE HETEROCHROMATIN PROTEIN 1 is essential for H3K27me3 binding and function during Arabidopsis development. PLoS ONE 4:e5335
Fiers M, Golemiec E, Xu J, van der Geest L, Heidstra R, Stiekema W, Liu CM (2005) The 14-amino acid CLV3, CLE19, and CLE40 peptides trigger consumption of the root meristem in Arabidopsis through a CLAVATA2-dependent pathway. Plant Cell 17:2542–2553
Gagne JM, Gish LA, Clark SE (2010) The role of the acyl modification, palmitoylation, in Arabidopsis stem cell regulation. Plant Signal Behav 5:1048–1051
Galinha C, Hofhuis H, Luijten M, Willemsen V, Blilou I, Heidstra R, Scheres B (2007) PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 449:1053–1057
Goh KI, Oh E, Jeong H, Kahng B, Kim D (2002) Classification of scale-free networks. Proc Natl Acad Sci USA 99:12583–12588
González-García MP, Vilarrasa-Blasi J, Zhiponova M, Divol F, Mora-García S, Russinova E, Caño-Delgado AI (2011) Brassinosteroids control meristem size by promoting cell cycle progression in Arabidopsis roots. Development 138:849–859
Gonzali S, Novi G, Loreti E, Paolicchi F, Poggi A, Alpi A, Perata P (2005) A turanose-insensitive mutant suggests a role for WOX5 in auxin homeostasis in Arabidopsis thaliana. Plant J. 44:633–645
Gordon SP, Chickarmane VS, Ohno C, Meyerowitz EM (2009) Multiple feedback loops through cytokinin signaling control stem cell number within the Arabidopsis shoot meristem. Proc Natl Acad Sci USA 106:16529–16534
Grieneisen VA, Xu J, Marée AF, Hogeweg P, Scheres B (2007) Auxin transport is sufficient to generate a maximum and gradient guiding root growth. Nature 449:1008–1013
Grigg SP, Galinha C, Kornet N, Canales C, Scheres B, Tsiantis M (2009) Repression of apical homeobox genes is required for embryonic root development in Arabidopsis. Curr Biol 19:1485–1490
Guilfoyle TJ, Hagen G (2007) Auxin response factors. Curr Opin Plant Biol 10:453–460
Hardtke CS, Ckurshumova W, Vidaurre DP, Singh SA, Stamatiou G, Tiwari SB, Hagen G, Guilfoyle TJ, Berleth T (2004) Overlapping and non-redundant functions of the Arabidopsis auxin response factors MONOPTEROS and NONPHOTOTROPIC HYPOCOTYL 4. Development 131:1089–1100
Helariutta Y, Fukaki H, Wysocka-Diller J, Nakajima K, Jung J, Sena G, Hauser MT, Benfey PN (2000) The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 101:555–567
Heylighen F (2001) The science of self-organization and adaptivity. In: Kiel LD (ed) Knowledge management, organizational intelligence and learning, and complexity. The encyclopedia of life support systems. Eolss Publishers, Oxford
Hobe M, Müller R, Grünewald M, Brand U, Simon R (2003) Loss of CLE40, a protein functionally equivalent to the stem cell restricting signal CLV3, enhances root waving in Arabidopsis. Dev Genes Evol 213:371–381
Izhaki A, Bowman JL (2007) KANADI and class III HD-Zip gene families regulate embryo patterning and modulate auxin flow during embryogenesis in Arabidopsis. Plant Cell 19:495–508
Jaeger J, Irons D, Monk N (2008) Regulative feedback in pattern formation: towards a general relativistic theory of positional information. Development 135:3175–3183
Jones B, Gunnerås SA, Petersson SV, Tarkowski P, Graham N, May S, Dolezal K, Sandberg G, Ljung K (2010) Cytokinin regulation of auxin synthesis in Arabidopsis involves a homeostatic feedback loop regulated via auxin and cytokinin signal transduction. Plant Cell 22:2956–2969
Kepinski S, Leyser O (2005) The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435:446–451
Kim YS, Kim SG, Lee M, Lee I, Park HY, Seo PJ, Jung JH, Kwon EJ, Suh SW, Paek KH, Park CM (2008) HD-ZIP III activity is modulated by competitive inhibitors via a feedback loop in Arabidopsis shoot apical meristem development. Plant Cell 20:920–933
Kitano H (2002) Computational systems biology. Nature 420:206–210
Kitano H (2004) Biological robustness. Nat Rev Genet 5:826–837
Kitano H (2007) Towards a theory of biological robustness. Mol Syst Biol 3:137
Kondo Y, Hirakawa Y, Kieber JJ, Fukuda H (2011) CLE peptides can negatively regulate protoxylem vessel formation via cytokinin signaling. Plant Cell Physiol 52:37–48
Levesque MP, Vernoux T, Busch W, Cui H, Wang JY, Blilou I, Hassan H, Nakajima K, Matsumoto N, Lohmann JU, Scheres B, Benfey PN (2006) Whole-genome analysis of the SHORT-ROOT developmental pathway in Arabidopsis. PLoS Biol 4:e143
Leyser O (2011) Auxin, self-organisation, and the colonial nature of plants. Curr Biol 21:R331–R337
Lucas M, Swarup R, Paponov IA, Swarup K, Casimiro I, Lake D, Peret B, Zappala S, Mairhofer S, Whitworth M, Wang J, Ljung K, Marchant A, Sandberg G, Holdsworth MJ, Palme K, Pridmore T, Mooney S, Bennett MJ (2011) Short-Root regulates primary, lateral, and adventitious root development in Arabidopsis. Plant Physiol 155:384–398
Matsuzaki Y, Ogawa-Ohnishi M, Mori A, Matsubayashi Y (2010) Secreted peptide signals required for maintenance of root stem cell niche in Arabidopsis. Science 329:1065–1067
Mironova VV, Omelyanchuk NA, Yosiphon G, Fadeev SI, Kolchanov NA, Mjolsness E, Likhoshvai VA (2010) A plausible mechanism for auxin patterning along the developing root. BMC Syst Biol 4:98
Mitchell M (2009) Complexity: a guided tour. Oxford University Press, Oxford
Mitrophanov AY, Groisman EA (2008) Positive feedback in cellular control systems. BioEssays 30:542–555
Miyashima S, Koi S, Hashimoto T, Nakajima K (2011) Non-cell-autonomous microRNA165 acts in a dose-dependent manner to regulate multiple differentiation status in the Arabidopsis root. Development 138:2303–2313
Mockaitis K, Estelle M (2008) Auxin receptors and plant development: a new signaling paradigm. Annu Rev Cell Dev Biol 24:55–80
Müller B, Sheen J (2008) Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 453:1094–1097
Nordström A, Tarkowski P, Tarkowska D, Norbaek R, Astot C, Dolezal K, Sandberg G (2004) Auxin regulation of cytokinin biosynthesis in Arabidopsis thaliana: a factor of potential importance for auxin-cytokinin-regulated development. Proc Natl Acad Sci USA 101:8039–8044
Ogasawara H, Kaimi R, Colasanti J, Kozaki A (2011) Activity of transcription factor JACKDAW is essential for SHR/SCR-dependent activation of SCARECROW and MAGPIE and is modulated by reciprocal interactions with MAGPIE, SCARECROW and SHORT ROOT. Plant Mol Biol 77:489–499
Ortega-Martínez O, Pernas M, Carol RJ, Dolan L (2007) Ethylene modulates stem cell division in the Arabidopsis thaliana root. Science 317:507–510
Paponov IA, Paponov M, Menges M, Teale W, Chakrabortee S, Murray JA, Palme K (2008) Comprehensive transcriptome analysis of auxin responses in Arabidopsis. Mol Plant 1:321–337
Parry G, Calderon-Villalobos LI, Prigge M, Peret B, Dharmasiri S, Itoh H, Lechner E, Gray WM, Bennett M, Estelle M (2009) Complex regulation of the TIR1/AFB family of auxin receptors. Proc Natl Acad Sci USA 106:22540–22545
Petersson SV, Johansson AI, Kowalczyk M, Makoveychuk A, Wang JY, Moritz T, Grebe M, Benfey PN, Sandberg G, Ljung K (2009) An auxin gradient and maximum in the Arabidopsis root apex shown by high-resolution cell-specific analysis of IAA distribution and synthesis. Plant Cell 21:1659–1668
Prigogine I (1978) Time, structure, and fluctuations. Science 201:777–785
Rademacher EH, Möller B, Lokerse AS, Llavata-Peris CI, van den Berg W, Weijers D (2011) A cellular expression map of the Arabidopsis AUXIN RESPONSE FACTOR gene family. Plant J 68:597–606
Rademacher EH, Lokerse AS, Schlereth A, Llavata-Peris CI, Bayer M, Kientz M, Freire Rios A, Borst JW, Lukowitz W, Jürgens G, Weijers D (2012) Different auxin response machineries control distinct cell fates in the early plant embryo. Dev Cell 22:211–222
Rahman A, Amakawa T, Goto N, Tsurumi S (2001) Auxin is a positive regulator for ethylene-mediated response in the growth of Arabidopsis roots. Plant Cell Physiol 42:301–307
Ruzicka K, Ljung K, Vanneste S, Podhorská R, Beeckman T, Friml J, Benková E (2007) Ethylene regulates root growth through effects on auxin biosynthesis and transport-dependent auxin distribution. Plant Cell 19:2197–2212
Ruzicka K, Simásková M, Duclercq J, Petrásek J, Zazímalová E, Simon S, Friml J, Van Montagu MC, Benková E (2009) Cytokinin regulates root meristem activity via modulation of the polar auxin transport. Proc Natl Acad Sci USA 106:4284–4289
Sablowski R (2011) Plant stem cell niches: from signalling to execution. Curr Opin Plant Biol 14:4–9
Sarkar AK, Luijten M, Miyashima S, Lenhard M, Hashimoto T, Nakajima K, Scheres B, Heidstra R, Laux T (2007) Conserved factors regulate signalling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 446:811–814
Sauer M, Balla J, Luschnig C, Wisniewska J, Reinöhl V, Friml J, Benková E (2006) Canalization of auxin flow by Aux/IAA-ARF-dependent feedback regulation of PIN polarity. Genes Dev 20:2902–2911
Scheres B (2007) Stem-cell niches: nursery rhymes across kingdoms. Nat Rev Mol Cell Biol 8:345–354
Seeley TD (2002) When is self-organization used in biological systems? Biol Bull 202:314–318
Sena G, Wang X, Liu HY, Hofhuis H, Birnbaum KD (2009) Organ regeneration does not require a functional stem cell niche in plants. Nature 457:1150–1153
Shen WH, Xu L (2009) Chromatin remodeling in stem cell maintenance in Arabidopsis thaliana. Mol Plant 2:600–609
Smith ZR, Long JA (2010) Control of Arabidopsis apical-basal embryo polarity by antagonistic transcription factors. Nature 464:423–426
Song SK, Hofhuis H, Lee MM, Clark SE (2008) Key divisions in the early Arabidopsis embryo require POL and PLL1 phosphatases to establish the root stem cell organizer and vascular axis. Dev Cell 15:98–109
Sozzani R, Cui H, Moreno-Risueno MA, Busch W, Van Norman JM, Vernoux T, Brady SM, Dewitte W, Murray JA, Benfey PN (2010) Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 466:128–132
Stahl Y, Wink RH, Ingram GC, Simon R (2009) A signaling module controlling the stem cell niche in Arabidopsis root meristems. Curr Biol 19:909–914
Stepanova AN, Hoyt JM, Hamilton AA, Alonso JM (2005) A Link between ethylene and auxin uncovered by the characterization of two root-specific ethylene-insensitive mutants in Arabidopsis. Plant Cell 17:2230–2242
Stepanova AN, Yun J, Likhacheva AV, Alonso JM (2007) Multilevel interactions between ethylene and auxin in Arabidopsis roots. Plant Cell 19:2169–2185
Stepanova AN, Robertson-Hoyt J, Yun J, Benavente LM, Xie DY, Dolezal K, Schlereth A, Jürgens G, Alonso JM (2008) TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 133:177–191
Strogatz SH (2001) Exploring complex networks. Nature 410:268–276
Sugimoto K, Jiao Y, Meyerowitz EM (2010) Arabidopsis regeneration from multiple tissues occurs via a root development pathway. Dev Cell 18:463–471
Swarup R, Perry P, Hagenbeek D, Van Der Straeten D, Beemster GT, Sandberg G, Bhalerao R, Ljung K, Bennett MJ (2007) Ethylene upregulates auxin biosynthesis in Arabidopsis seedlings to enhance inhibition of root cell elongation. Plant Cell 19:2186–2196
Ten Tusscher KH, Hogeweg P (2011) Evolution of networks for body plan patterning; interplay of modularity, robustness and evolvability. PLoS Comput Biol 7:e1002208
Tiwari SB, Wang XJ, Hagen G, Guilfoyle TJ (2001) AUX/IAA proteins are active repressors, and their stability and activity are modulated by auxin. Plant Cell 13:2809–2822
Tsuchisaka A, Theologis A (2004) Unique and overlapping expression patterns among the Arabidopsis 1-amino-cyclopropane-1-carboxylate synthase gene family members. Plant Physiol 136:2982–3000
Ulmasov T, Hagen G, Guilfoyle TJ (1999) Dimerization and DNA binding of auxin response factors. Plant J 19:309–319
van den Berg C, Willemsen V, Hage W, Weisbeek P, Scheres B (1995) Cell fate in the Arabidopsis root meristem determined by directional signalling. Nature 378:62–65
van den Berg C, Willemsen V, Hendriks G, Weisbeek P, Scheres B (1997) Short-range control of cell differentiation in the Arabidopsis root meristem. Nature 390:287–289
Vernoux T, Brunoud G, Farcot E, Morin V, Van den Daele H, Legrand J, Oliva M, Das P, Larrieu A, Wells D, Guédon Y, Armitage L, Picard F, Guyomarc’h S, Cellier C, Parry G, Koumproglou R, Doonan JH, Estelle M, Godin C, Kepinski S, Bennett M, De Veylder L, Traas J (2011) The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol Syst Biol 7:508
Vieten A, Vanneste S, Wisniewska J, Benková E, Benjamins R, Beeckman T, Luschnig C, Friml J (2005) Functional redundancy of PIN proteins is accompanied by auxin-dependent cross-regulation of PIN expression. Development 132:4521–4531
von Dassow G, Munro E (1999) Modularity in animal development and evolution: elements of a conceptual framework for EvoDevo. J Exp Zool 285:307–325
Wagner A, Fell DA (2001) The small world inside large metabolic networks. Proc Biol Sci 268:1803–1810
Wagner GP, Pavlicev M, Cheverud JM (2007) The road to modularity. Nat Rev Genet 8:921–931
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17:2204–2216
Watts DJ, Strogatz SH (1998) Collective dynamics of ‘small-world’ networks. Nature 393:440–442
Welch D, Hassan H, Blilou I, Immink R, Heidstra R, Scheres B (2007) Arabidopsis JACKDAW and MAGPIE zinc finger proteins delimit asymmetric cell division and stabilize tissue boundaries by restricting SHORT-ROOT action. Genes Dev 21:2196–2204
Whitacre JM (2012) Biological robustness: paradigms, mechanisms, and systems principles. Front Genet 3:67
Xu J, Hofhuis H, Heidstra R, Sauer M, Friml J, Scheres B (2006) A molecular framework for plant regeneration. Science 311:385–388
Zhang H, Han W, De Smet I, Talboys P, Loya R, Hassan A, Rong H, Jürgens G, Paul Knox J, Wang MH (2010) ABA promotes quiescence of the quiescent centre and suppresses stem cell differentiation in the Arabidopsis primary root meristem. Plant J 64:764–774
Zhang W, To JP, Cheng CY, Eric Schaller G, Kieber JJ (2011) Type-A response regulators are required for proper root apical meristem function through post-transcriptional regulation of PIN auxin efflux carriers. Plant J 68:1–10
Zhou GK, Kubo M, Zhong R, Demura T, Ye ZH (2007) Overexpression of miR165 affects apical meristem formation, organ polarity establishment and vascular development in Arabidopsis. Plant Cell Physiol 48:391–404
Zhou W, Wei L, Xu J, Zhai Q, Jiang H, Chen R, Chen Q, Sun J, Chu J, Zhu L, Liu CM, Li C (2010) Arabidopsis Tyrosylprotein sulfotransferase acts in the auxin/PLETHORA pathway in regulating postembryonic maintenance of the root stem cell niche. Plant Cell 22:3692–3709
Zhou ZY, Zhang CG, Wu L, Zhang CG, Chai J, Wang M, Jha A, Jia PF, Cui SJ, Yang M, Chen R, Guo GQ (2011) Functional characterization of the CKRC1/TAA1 gene and dissection of hormonal actions in the Arabidopsis root. Plant J 66:516–527
Acknowledgments
This paper constitutes a partial fulfillment of the Graduate Program in Biological Sciences of the National Autonomous University of México (UNAM). E.A. acknowledges the scholarship and financial support provided by the National Council of Science and Technology (CONACyT), and UNAM. E.A.B. thanks financial support from Conacyt (81433, 81542, 1667705,180098,180380) and PAPIIT (IN229003-3, IN226510-3, IN204011-3, IB201212-2). E.A.B. is currently sponsored by the Miller Institute for Basic Research in Science, University of California, Berkeley, USA. We thank Rigoberto V. Perez-Ruız and Diana Romo for technical and logistical assistance. We thank Lynna Kiere for the detailed revision of the manuscript. Finally, we also thank Dr. Joseph Dubrovsky, Dr. Carlos Villarreal, Dr. Mariana Benítez and M.S. Úrsula Abad for their thoughtful comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Azpeitia, E., Alvarez-Buylla, E.R. A complex systems approach to Arabidopsis root stem-cell niche developmental mechanisms: from molecules, to networks, to morphogenesis. Plant Mol Biol 80, 351–363 (2012). https://doi.org/10.1007/s11103-012-9954-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-012-9954-6