Abstract
Abscisic acid (ABA) and sugars have been well established to be crucial factors controlling seed germination of Arabidopsis. Here we demonstrate that AtMKK1 and AtMPK6 are both critical signals involved in ABA and sugar-regulated seed germination. Wild type plants depended on stratification and after-ripening for seed germination, whereas this dependence on either stratification or after-ripening was not required for mutants of mkk1 and mpk6 as well as their double mutant mkk1 mpk6. While seed germination of wild type plants was sensitively inhibited by ABA and glucose, mkk1, mpk6 and mkk1 mpk6 were all strongly resistant to ABA or glucose treatments, and in contrast, plants overexpressing MKK1 or MPK6 were super-sensitive to ABA and glucose. Glucose treatment significantly induced increases in MKK1 and MPK6 activities. These results clearly indicate that MKK1 and MPK6 are involved in the ABA and sugar signaling in the process of seed germination. Further experiments showed that glucose was capable of inducing ABA biosynthesis by up-regulating NCED3 and ABA2, and furthermore, this up-regulation of NCED3 and ABA2 was arrested in the mkk1 mpk6 double mutant, indicating that the inhibition of seed germination by glucose is potentially resulted from sugar-induced up-regulation of the ABA level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Seed germination has been increasingly become a model system for researches on cellular signaling in plants (Koornneef et al. 2002). ABA is well known to play extensive roles in plant physiology, such as stomatal movement, root growth, embryo development, stress-tolerant responses as well as seed germination. To date many genes or proteins involved in ABA signaling have been characterized in Arabidopsis (Smeekens 2000; Coruzzi and Zhou 2001; Rolland et al. 2001, 2002; Schroeder et al. 2001; Bray 2002; Finkelestein et al. 2002; Finkelstein and Gibson 2002; Zhu 2002; Assmann 2003; Chen et al. 2004; Chow and McCourt 2004; Yamaguchi-Shinozaki and Shinozaki 2006; Fujii et al. 2007; Xing et al. 2008), and among these genes or proteins, some of them are suggested to play critical roles in the seed germination (Koornneef et al. 1984; Giraudat et al. 1992; Finkelstein et al. 1998; Finkelstein and Lynch 2000). ABI1 and ABI2, characterized as protein phosphatases, are demonstrated to be involved in ABA signaling in various physiological processes, whereas ABI3, ABI4 and ABI5, characterized as transcription factors, are suggested to show ABA insensitivity only in seed germination and early seedlings (Koornneef et al. 1984; Finkelstein and Somerville 1990; Ooms et al. 1993; Parcy et al. 1994). Although the screening of ABA-responsive mutants has identified some proteins that play crucial roles in seed germination, a relatively complete profile outlining the ABA signaling network, especially the upstream signaling cascades, still remains unclear.
Besides ABA, sugars have been recently documented to be a critical factor controlling seed germination and early seedling development (Koornneef et al. 2002; Rolland et al. 2001, 2002, 2006). Surprisingly, it has been increasingly suggested that the sugar-associated signaling cascades overlap very much with the ABA signaling in various processes especially the seed germination (Rolland et al. 2002; Leon and Sheen 2003; Gibson 2005), and moreover, the ABA and sugar signaling frequently interacts in a rather elusive pattern. For example, while sugars have been demonstrated to be able to inhibit Arabidopsis seed germination, there are evidences that exogenous sugars were capable of relieving the inhibitory effect of ABA on seed germination (Dekkers et al. 2004; Price et al. 2003). It appears that the signaling interactions between sugar and ABA vary depending on different concentrations and different stages, and positive interactions between sugar and ABA signaling are more obvious during early seedling development (Leon and Sheen 2003; Gibson 2005). There is little doubt that sugar signaling and ABA signaling is closely related under some conditions, but the exact mechanism for this remains elusive (Zhou et al. 1998; Arenas-Huertero et al. 2000; Gazzarrini and McCourt 2001; Finkelstein and Gibson 2002; Rolland et al. 2006).
In as early as 70’s the last century, it was suggested that catalase might play a crucial role in seed germination since inhibition of catalase was capable of promoting the seed germination of lettuce and pigweed (Hendricks and Taylorson 1975). In keeping with this proposition, it has increasingly demonstrated that reactive oxygen species (ROS), such as NO and H2O2, is able to promote seed germination (Kwak et al. 2003; Apel and Hirt 2004; Laloi et al. 2004). In recent works we have demonstrated that a MAPK kinse, i.e. AtMKK1, is able to mediate ABA and stress-induced gene expression of calase via AtMPK6-coupled signaling in vegetative tissues of Arabidopsis (Xing et al. 2007, 2008). MAPK cascades minimally consist of a MAPKKK-MAPKK-MAPK module that is linked in various ways to upstream and downstream signaling events (Nakagami et al. 2005). While it has been extensively shown that MAPK cascades is involved in variety of processes from plant development to environmental responses in vegetative tissues, much less is known about the roles of the MAPK cascades in seed germination (Hirt 1997; Ligterink et al. 1997; Mizoguchi et al. 1997; Hirt and Asard 2000; Nakagami et al. 2005). In view of the potential roles of catalse in seed germination and the important roles of the AtMKK1-AtMPK6 module in ABA-induced gene expression of catalase, and also in view of the fact that ABA-associated signals in vegetative tissues are liable to function in seed germination, we have investigated whether the module of AtMKK1-AtMPK6 might be involved in the seed germination signaling. The results strongly indicate that AtMKK1 and AtMPK6 are critical mediators in the signaling processes of both ABA and sugar-regulated seed germination, and that the seed germination inhibition of glucose is potentially resulted from sugar-induced expressions of genes encoding key enzymes in ABA biosynthesis pathway.
Materials and methods
Identification and isolation T-DNA insertion lines
The seeds of Arabidopsis mkk1 and mpk6 T-DNA insertion lines (SALK_015914 and SALK_127507) were obtained from the ABRC (Alonso et al. 2003). The method of identification and isolation T-DNA insertion lines of homozygous was described in Xing et al. (2008).
Total RNA extraction, RT-PCR and quantitative PCR analysis
Total RNA was extracted using the RNeasy Plant Mini Kit (Qiagen) according to the manufacturer’s instructions and then stored at −80°C after DNase treatment. Reverse transcription reactions were performed using 0.5 μg of total RNA and SuperScript II first-strand synthesis system (Invitrogen). PCR reactions were performed using Taq DNA polymerase (Invitrogen).
Real-time quantitative PCR analysis was performed with the iQ5 Real-time PCR detection system (Bio-Rad) according to the manufacturer’s recommendations. Real-time quantitative PCR reaction contained 10 μl 2 × SYBR Green Supermix (Bio-Rad), 2 μl primer mix, 4 μl cDNA and 4 μl deionized water to make a total volume of 20 μl. To quantify the copy number of each RNA, the threshold cycle value (Ct) was compared with a standard curve generated using PCR products for each gene that was purified and quantified by UV light absorbance. Primers that used for both RT-PCR and quantitative RT-PCR are given in Supplementary Table 1. Three replicates were performed for each experiment.
Sugar and ABA treatments
The effects of different sugars on seed germination were studied by determining the germination rates of 100 seeds pretreated with deionized water or 100 μM fluridone and then planted in triplicate on medium containing glucose and mannitol (Sigma). The effects of ABA were studied in the similar manner. Sterilized seeds were sown on plates containing different concentrations of sugars (d-glucose and d-mannitol) after stratification at 4°C. For comparisons of germination rates, each plate was subdivided and all seed lines were sown on the same plate.
Germination assay
Germination rates were compared between seed lots that were produced, harvested and stored under identical conditions. Light and chilling (stratification) are typically required for germination and, as such, must also be controlled in comparative studies on germination. Before sowing, seeds were surface-sterilized with 70% ethanol for 1 min, then with 2% hypochlorite for 5 min and rinsed six times with sterile deionized water. One hundred of seeds from wild-type, mkk1 mpk6, mkk1, and mpk6 were stratified at 4°C for 4 days and planted in half-strength Murashige and Skoog (MS) medium without sucrose under continuous white light (~80 μmolm−2 s−1) at 22°C. Seeds germination rate were scored when the radicals completely penetrated the seed coat.
ABA analysis
After various treatments, samples were immediately frozen in liquid nitrogen, and then homogenized in water at 4°C. The samples were then centrifuged for 25 min at 20000×g and the supernatants were used for the ABA assay. ABA analyses were carried out using the radioimmunoassay (RIA) method described by Quarrie et al. (1988). The highly specific monoclonal antibody (Mac 252) was provided by Dr. S. A. Quarrie (John Innes Centre, UK). Aqueous extracts of root tissues were used for the assay without purification. 50 μl of crude extracts was mixed with 200 μl of phosphate-buffered saline (pH 6.0), 100 μl diluted antibody solution and 100 μl 3H-ABA (about 8,000 cpm) solution. The reaction mixture was incubated at 4°C for 45 min and the bound radioactivity was measured in 50%-saturated (NH4)2SO4 precipitated pellets with a liquid scintillation counter.
In vitro kinase assays
Protein kinase assays were performed basically according to the protocol by Xing et al. (2008). Kinase-inactive MPK6-GST fusion protein was generated by exchanging a conserved lysine residue in the ATP binding domains to methionine and arginine using the Quick-Change kit from Stratagene (LA Jolla, CA). The point mutations for MPK6 were K92M and K93R. MKK1 was immunoprecipitated from Arabidopsis seedlings using anti-MKK1 and incubated with the kinase inactive MPK6-GST protein in the kinase reaction mixture [20 μl of kinase buffer containing 50 mM Tris (pH 7.5), 1 mM DTT, 10 mM MgCl2, 0.1 mM ATP and 6 μCi of γ-32P-ATP] for 30 min at room temperature. Kinase reactions were stopped after 30 min by adding 4 μl SDS loading buffer and heating for 5 min at 95°C. Reaction products were analyzed by SDS-PAGE, autoradiography and Coomassie brilliant R250 staining. Prestained size markers (Bio-Rad) were used to calculate the size of the kinases. The protein concentration was determined using the protein assay kit (Bio-Rad) with BSA as a standard. Kinase activity of immunoprecipitated MPK6 from seedlings using anti-MPK6 was measured with MBP as artificial substrate in vitro kinase assays described as above. Loading levels of MKK1 and MPK6 proteins were detected with western blot. The polyclonal antibodies of MKK1 and MPK6 were produced from rabbits ordered from Strategic Diagnostics Inc.
Generation of MKK1 and MPK6 over-expressing plants
The MKK1 and MPK6 over-expressing lines were obtained as the method described in Xing et al. (2007).
Accession numbers
Sequence data from the article can be found in the GenBank data libraries or TIGR database (Arabidopsis thaliana Genome Project) under the following accession numbers: MKK1, At4g26070; MPK6, At2g43790; NCED3, At3g14440; ABA2, At1g52340; AAO3, At2g27150; NCED9, At1g78390.
Results
After-ripening and stratification are not required for seed germination of mkk1 and mpk6 mutants
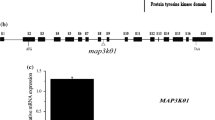
The T-DNA insertion mutants of mkk1 (SALK_015914) and mpk6 (SALK_127507) were obtained from the Arabidopsis Biological Resource Center’s (ABRC) collection of T-DNA transformed Arabidopsis lines (Alonso et al. 2003). T-DNA in mkk1 was located in the 5’ untranslated region (UTR) near the start codon and the T-DNA in mpk6 was located in the intron between exon 2 and exon 3. Determination of the T-DNA effect on mRNA levels for both AtMKK1 and AtMPK6 was conducted in our previous works, where it was shown that no message of AtMKK1 or AtMPK6 was detected while mRNA was detected in wild-type Col-0 plants, and furthermore, the complementation experiment excluded the possibility that other site mutation might be responsible for the mutant phenotypes (Xing et al. 2007, 2008). To obtain the double mutant of mkk1 and mpk6, single mutants of mkk1 and mpk6 were crossed and further RT-PCR analysis confirmed that expression of both MKK1 and MPK6 was abolished in the double mutant mkk1 mpk6 (Fig. 1a).
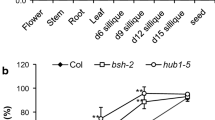
Mutant verification and after-ripening effect on the seed germination of different mutants. a Mutant verification. RT-PCR analysis with MKK1, MPK6 and ACT2 primers using total RNA extracted from seedlings of the wild-type (Col-0) and mkk1 mpk6, mkk1 and mpk6 as the template. b Seed germination of mkk1 mpk6, mkk1, mpk6 and wild-type in absence of stratification. Seeds were sterilized and sowed on Petri dishes after storage of different times. Values presented are the mean germination rate from three separate repeats and the error bar indicates ± SE (n = 3)
Generally ABA-related seed dormancy in Arabidopsis requires either after-ripening process and/or cold stratification (moist pre-chilling at 4°C) for facilitating germination (Bewley 1997). For dormancy experiment, plants of each genotype were grown in different sections of the same tray and seeds were harvested at the same time to minimize the difference in seed maturation and storage period. Figure 1b shows the different germination rates of wild-type and mutants seeds after different times of harvesting. Within the first 3 days after harvest, while less than only 15% of the seeds germinated for wild type, more than 70% of the seeds germinated for the mutant of mkk1 and double mutant of mkk1 mpk6. Two weeks after harvest, a germination rate of more than 90% could be recorded for all mutants, in contrast, only about 50% for the wild-type (Fig. 1b). In the absence of stratification, the seeds of mkk1 mpk6 showed a much higher germination rate than that of wild-type (Fig. 2a). Stratification treatment greatly promoted the seed germination for wild type, although it also had a positive influence on the seed germination for all the mutants (Fig. 2b). These results indicate that after-opening and stratification ware basically not required for seed germination of all the mutants while they ware strong factors limiting the seed germination for the wild type plants.
Effect of stratification on seed germination. a Germination of each genotype in the absence of stratification after storage of 2 weeks. b Germination of each genotype in the presence of stratification (4°C, 4 days) after storage of 2 weeks. Values presented are the mean germination rate from three separate repeats and the error bar indicates ± SE (n = 3)
Sensitivity of seed germination to either glucose or ABA is impaired in mkk1, mkk6 and mkk1 mpk6
Glucose treatment greatly inhibited the seed germination of wild type, as was shown from a decrease of more than 80% in the seed germination rate at 6% glucose (Fig. 3a). Compared to wild type, the seed germination of all the mutants was much less sensitive to glucose especially for the double mutant of mkk1 mpk6 in which the germination rate was arrested by only less than 20% at 6% glucose. To investigate whether glucose-induced germination inhibition is purely due to its osmotic effect, we also tested the effect of mannitol on seed germination. Indeed, mannitol treatment was able to inhibit the seed germination to some extent for both wild type and mutants, but unlike the case of glucose, no difference was observed in the seed germination sensitivity to mannitol between wild type and mutants, indicating that the inhibitory effect of glucose on seed germination is achieved through a glucose-specific signaling pathway (Fig. 3b).
Effects of glucose on seed germination of different genotypes. The seeds of each genotype were sterilized, stratified and sown on the Petri dish containing the indicated concentrations of glucose a and mannitol b under continuous white light at 22°C. The germination rate of each genotype was scored at 3 days after the end of stratification. Values are mean of three separate repeats ± SE (n = 3)
Seed germination of wild type plants was very sensitive to ABA, and it was completely arrested when 2 μM ABA was applied. Compared to that for wild type, the inhibitory effects of ABA on the seed germination was much less for all the mutants especially for the double mutant of mkk1 mpk6 that showed a decrease in seed germination rate of only less than 10% when 2 μM ABA was applied (Fig. 4a), and in terms of green cotyledons for statistics, the ABA effect was basically similar (Fig. 4b).
Effects of ABA on seed germination of mkk1 mpk6, mkk1, mpk6 and wild-type. a Quantification of radicle emergence as the germination rate of each genotype at 3 days after the end of stratification on the indicated of ABA. Each measurement consisted of at least 100 seeds. b Quantification of the percentage of seedlings with green cotyledons after 7 days on the indicated of ABA. Values are mean of three separate repeats ± SE (n = 3)
Overexpression of MKK1 and MPK6 confer hypersensitivity to glucose and ABA
To provide further evidences for the roles of AtMKK1 and AtMPK6 in seed germination, the overexpression lines of AtMKK1 and AtMPK6 were examined. The full-length AtMKK1 and AtMPK6 sequences were placed under the control of the 35S CaMV promoter and transformed into the wild type. A total of 22 and 28 independent transgenic lines (T2) were generated and screened for AtMKK1 and AtMPK6, and the transgenic lines with highest mRNA were chosen for the present research (Xing et al. 2008). The detailed information about the construction and analysis of the transgenic plants were described in our previous work, so the related date is not shown here.
As shown in Fig. 5a, seeds of different genotypes all germinated well on the control medium with the germination rate nearly reaching 100%. While glucose treatment strongly inhibited seed germination of the wild type, the inhibitory effect of glucose on the seed germination was relatively much less in all the mutants, especially in the mkk1mpk6 double mutant (Fig. 5b). To more exactly estimate the effects of ABA and glucose on the seed germination, we had performed statistical analysis of the seed germination of all the related mutants as well as the plants with gene over-expression (Fig. 5c, d). As shown in Fig. 5c, while mkk1, mpk6 and mkk1 mpk6 double mutant had much higher germination rate than that of the wild type in the presence of either glucose or ABA, the seed germination of MKK1-OE and MPK6-OE were almost completely inhibited especially for MKK1-OE. ABA at a concentration of only 0.5 μM resulted in a decrease of more than 60% in the germination rate for MKK1-OE and MPK6-OE, whereas the same concentration of ABA only led to a decrease of less than 20% for the wild type, indicating that plants overexpressing AtMKK1 and AtMPK6 are hypersensitive to ABA (Fig. 5d).
Seed germination in response to ABA and glucose treatments. a and b Pictures of seed germination of different genotype without or with glucose treatment respectively. The seeds of each genotype were sterilized, stratified and sown on the medium with no glucose or containing 6% glucose, and pictures were taken at the 7th day after sown. c and d Statistical analysis of the see germination of different genotype in response to ABA and glucose treatments. The seeds of each genotype were sterilized, stratified and sown on the medium containing 1 μM ABA or 6% glucose. c Germination was scored at 3 days after the end of stratification. d Seeds of each genotype were sterilized, stratified and sown on the medium containing the indicated concentrations of ABA. The germination was scored at 3 days after the end of stratification. Each measurement consisted of at least 100 seeds. Values presented are the mean germination rate from three separate repeats ± SE (n = 3)
ABA is required for seedling establishment (Lopez-Molina et al. 2001). Low concentration of ABA (<1 μM) is known to stimulate root growth (Ephritikhine et al. 1999). To further investigate the roles of AtMKK1 and AtMPK6 in postgermination development, we transferred 4-days-old seedlings germinated on agar plates to the same medium with or without ABA, and the length of the primary root and seedling fresh weight were measured 2 weeks later. Slight stimulation of root elongation in the wild-type in response to 0.5 μM ABA was observed in our experiments (Fig. 6a), whereas this stimulation by 0.5 μM ABA was not observed in the double mutant of mkk1 mpk6 and all the single mutants (mkk1 and mpk6). Under high ABA concentration (5–50 μM), root elongation and seedling biomass accumulation were generally inhibited by ABA (Fig. 6b, c). Double mutant of mkk1 mpk6 grew the most and single mutants of mkk1 and mpk6 grew more than the wild-type under the high ABA treatment. Results suggest that the mutants had lost their sensitivity to the regulation by ABA.
Effect of ABA on post-germination growth of different genotypes. a Quantification of root length for seedlings at 14 days after transfer to the medium with or without 0.5 μM ABA. b and c Quantification of root length and seedling fresh weight for seedlings at 14 days after transfer to the control medium (MS medium with 3% sucrose) or containing the indicated concentrations of ABA. For fresh weight determination, ten seedlings were weighted at one time and the result was divided by ten. Seedlings were 4-days-old at the time of transfer and had equal root lengths at that time. Data are means ± SE (n = 30 for root length and n = 10 for fresh weight)
Effect of glucose on seed germination is associated with regulation of ABA biosynthesis
Glucose and ABA signaling in seed germination is well known to be frequently overlapped very much (Rolland et al. 2002; Leon and Sheen 2003; Price et al. 2003; Dekkers et al. 2004; Gibson 2005; Rolland et al. 2006). To clarify whether glucose may function directly through endogenous ABA, further experiment with a pharmacological method was performed. Seeds pretreated with the ABA biosynthesis inhibitor fluridone were germinated with the presence of glucose, and it was found that the inhibitory effect of glucose on seed germination and the difference among genotypes largely diminished (Fig. 7a), suggesting that the glucose inhibition to seed germination is possibly associated with a regulation of the endogenous ABA level. In order to confirm this hypothesis, changes in the endogenous ABA content were determined. Floridans pretreatment significantly reduced the endogenous ABA levels in both wild type and all the mutants, and more importantly, glucose treatment indeed significantly increased the ABA levels in wild type and mutants. Further more, compared to that in wild type plants, the glucose-induced ABA increase was much less in mutants especially the double mutant mkk1 mpk6 (Fig. 7b).
Effect of fluridone on seed germination and ABA content of different genotypes. a Effect of fluridone on seed germiantion. For glucose treatment, seeds were sown on the medium containing 6% glucose after pretreatment with or without 100 μM fluridone (Glc and Flu + Glc). b Effects of fluridone and glucose treatments on endogenous ABA contents in germination seeds. Seeds were pretreated with water or 100 μM fluridone at 4°C for 2 days, sown in the medium with or without 6% glucose at 22°C for 3 days, and analyzed ABA content. Values presented are the mean ± SE (n = 3)
AtMKK1 and AtMPK6 mediate glucose-induced ABA biosynthesis
In view of the discovery that glucose was able to induce different increases in ABA content in wild type and all the mutants, we further investigated whether the glucose-induced ABA increase was possibly mediated by AtMKK1 and AtMPK6. The expressions of several genes encoding key enzymes in ABA biosynthesis pathway were determined. As shown in Fig. 8a, while glucose treatment induced a dramatic increase in NCED3 expression in both wild type and MKK1-OE and MPK6-OE plants, the increase in NCED3 expression was higher in over-expression plants than that in wild type plants. More interestingly, the glucose-induced NCED3 expression largely diminished in all the mutants, especially the double mutant of mkk1 mpk6 in which the NCED3 expression was nearly completely arrested. The expressing patterns of ABA2 were basically similar with that of NCED3 (Fig. 8b). Compared with NCED3 and ABA2, the transcription levels of other related genes, such as NCED9 and AAO3, in ABA biosynthesis pathway were relatively lower, and also, little difference was found among the different genotypes treated with glucose, indicating that the glucose-induced increase in ABA biosynthesis is mediated by AtMKK1 and AtMPK6 signaling in seed germination.
Expression of ABA biosynthesis genes in wild-type, double mutant (mkk1 mpk6), single mutants (mkk1 and mpk6) and transgenic lines (MKK1-OE and MPK6-OE). Expressions of the genes were analyzed by quantitative RT-PCR in different genotype. The seedlings were under control conditions or after 4 h of exposure to 6% glucose. Values presented are the mean ± SE (n = 3)
To further investigate the roles of AtMKK1 and AtMPK6 in seed germination, AtMKK1 and AtMPK6 activities in response to glucose treatment were determined. As shown in Fig. 9, glucose treatment significantly induced the increase in both AtMKK1 and AtMPK6 activity, which further demonstrates that AtMKK1 and AtMPK6 are downstream regulators of the glucose signal during the process of seed germination.
MKK1 and MPK6 kinase activities response to glucose. The MKK1 and MPK6 kinase activities were assayed in wild-type after treatment with different concentration of glucose (1, 3 and 6%). The proteins of MKK1 or MPK6 were immunoprecipitated using antibody of anti-MKK1 or anti-MPK6 from seedlings after treatments. MKK1 kinase activity was determined by in vitro kinase assays using kinase-inactive GST-MPK6 as a substrate. Kinase activity of immunoprecipitated MPK6 was measured with MBP as artificial substrate in vitro kinase assays. Loading levels of MKK1 and MPK6 proteins were detected in western blot
Discussion
A MAPK cascade minimally consists of a MAPKKK-MAPKK-MAPK module that is linked in various ways to upstream and downstream signaling. A surprisingly large number of genes encoding MAPK pathway components have been uncovered by analyzing model plant genomes, suggesting that MAPK cascades are abundant players of signal transduction. Numerous studies demonstrated that have confirmed major roles of defined MAPK pathways in development, cell proliferation and hormone physiology, as well as in biotic and abiotic stress signaling. Nevertheless, information about MAPKs signaling in seed germination is relatively much less. Very recently, we have demonstrated that AtMKK1, a MAPK kinase, was able to mediate stress and ABA-induced catalase CAT1 expression via AtMPK6, a MAPK, coupled signaling in vegetative tissues (Xing et al. 2007, 2008). It is well known that catalase plays a central role in H2O2 scavenging. It has been increasing suggested that H2O2 and other reactive oxygen species (ROS) may be crucial factors controlling seed germination (Apel and Hirt 2004; Kwak et al. 2003; Laloi et al. 2004). It is hence reasonable to confer that AtMKK1-AtMPK6 signaling may play a role in seed germination. In consistent with this hypothesis, the loss of function mutants of AtMKK1 and AtMPK6 showed insensitivity to glucose and ABA, and in contrast, over-expression of AtMKK1 and AtMPK6 exhibited hyper-sensitivity to both ABA and glucose in the seed germination (Fig. 5). The results clearly demonstrated that both AtMKK1 and AtMPK6 were involved in the glucose and ABA signaling in seed germination. Compared to single mutant of either mkk1 or mpk6, the double mutant mkk1 mpk6 was shown to be more insensitive to glucose and ABA in seed germination (Figs. 4, 5), and also, more insensitive in glucose-induced gene expression (Fig. 8), which means that AtMPK6 dose not likely act as a sole target of AtMKK1 and there may be some other MAPKs associated with the signaling in seed germination. In consistent with this proposition, AtMPK3 was also reported to be involved in the regulation of seed germination (Lu et al. 2002; Kuhn and Schroeder 2003; Himmelbach et al. 2003; Laloi et al. 2004), but how AtMKK1 interacts with the downstream MAPKs remains to be further investigated.
Seed germination is a complicated process dominated by a combination of environmental and endogenous signals with both synergistic and competing effects (Sheen et al. 1999; Ullah et al. 2002; Chen and Jones 2004; Chen et al. 2004; Gibson 2005). Because of this, seed germination has been increasingly a model system for researches on plant signaling. As regards the endogenous signals, sugars and some hormones, i.e. ABA, GA and ethylene, have been well demonstrated to play crucial roles in the seed germination (Smeekens 2000; Coruzzi and Zhou 2001; Finkelestein et al. 2002; Rolland et al. 2002). Genetic screening has isolated a number of sugar-insensitive and sugar hypersensitive mutants in Arabidopsis. Surprisingly, many sugar-responsive signals tune out to be ABA signaling-associated proteins. Examples for this are the characterization of glucose insensitive5 (gin5) and gin6/sucrose uncoupling6 (sun6)/sugar insensitive5 (sis5) as mutant alleles of ABA3 and the transcription factorABI4, respectively (Arenas-Huertero et al. 2000; Leon and Sheen 2003; Gibson 2005), the glucose insensitive sis4/gin1 mutants are allelic to aba2, a mutant deficient in SDR1 required for ABA biosynthesis (Cheng et al. 2002; Leon and Sheen 2003; Gibson 2005), and also, two Glc and Suc insensitive mutants, sis7 and sis10, were found to lie in NCED3 and ABI3 respectively (Huang et al. 2008). Theses findings strongly suggested a central role of ABA in sugar signaling, but how ABA exhibits its roles in the glucose signaling remains unclear. It appears that the interaction between ABA and sugar is quite complicate and may vary when their concentrations change. Arenas-Huertero et al. (2000) reported that sugar inhibition of seed germination is due to the increase in the active endogenous ABA level. ABA biosynthetic mutant seeds are insensitive to glucose (Huijser et al. 2000; Laby et al. 2000). Nevertheless, it has been proposed that the glucose-inhibition on seed germination is ABA dependent but not caused by an increase in cellular ABA concentrations, and rather it is associated with a slowing down of the decline in endogenous ABA (Rolland et al. 2006). In the present study, we provide the genetic evidence that AtMKK1/AtMPK6 is capable of mediating the sugar signaling through regulation of ABA biosynthesis. Active endogenous ABA contents of mkk1 mpk6, mkk1 and mpk6 mutants were all lower than that of wild-type in germinating seeds. Fluridone, an ABA biosynthetic inhibitor, reduced the active endogenous ABA level and also alleviated the inhibitory effects of glucose on seed germination. Fluridone treatment diminished the differences between the mutants and wild-type in their responses to glucose on seed germination (Fig. 7). These results has not only provided evidences that glucose-inhibition on seed germination was due to an increase in endogenous ABA level, but also more importantly, demonstrated that the AtMKK1/AtMPK6 signaling play a critical role in the cross-talking between ABA and sugar signaling.
The pathways of ABA biosynthesis and catabolism have been extensively documented, and AtNCED3 has been well characterized as the key enzyme in ABA biosynthesis pathway (Finkelestein et al. 2002; Nambara and Marion-Poll 2003; Xiong and Zhu 2003). Overexpression of NCED genes increases seed ABA content and seed dormancy, and delays seed germination (Thompson et al. 2000; Qin and Zeevaart 2002). In vegetative tissues, it was reported that glucose was not able to induce AtNCED3 expression although it was able to induce the expression of ABA2, a gene encoding the enzyme required for ABA biosynthesis below the AtNCED3-catalyzed step (Cheng et al. 2002; Leon and Sheen 2003). In the present study we have found that glucose was not only able to induce ABA2 expression but also induce AtNCED3 expression, indicating a distinctive sugar signaling pathway in seed germination (Fig. 8). The findings of the AtMKK1/AtMPK6 mediated AtNCED3/AtABA2 expression further confirmed the central role of ABA signaling in seed germination. Lefebvre et al. (2006) reported that AtNCED6 and AtNCED9 are required for ABA biosynthesis during seed development and reduced dormancy was observed in the nced6 nced9 double-mutant seeds. Seo et al. (2006) revealed that ABA metabolism was phytochrome-regulated during photoreversible seed germination and nced6-1, aba2-2 and aao3-4 exhibited an enhanced ability to germinate relative to wild type when imbibed in the dark after irradiation with an FR light pulse. More recently, Toh et al. (2008) showed that ABA levels in imbibed seeds are elevated at high temperature and NCED9 plays a major role and NCED5 and NCED2 play a relatively minor role in high temperature-induced ABA synthesis and germination inhibition. Our previous research showed that the transcriptional level of NCED3 is relatively higher than other NCEDs, such as NCED9 and 6, during seed germination although these genes might be also involved in the seed germination. These results suggest that the expressional regulation of the genes in ABA biosynthesis and catabolism is very complicated, and the roles of the genes in seed germination may be different at different conditions.
To date, many proteins associated with the ABA-regulated seed germination have been isolated, but notably, most of these proteins were identified as transcription factors or enzyme responsible for ABA biosynthesis, such as ABI3, ABI4, ABI5, ABF2,ABF3, ABF4, ABA1 and ABA2 (Rolland et al. 2006). Although ABI1 and ABI2, two members of protein phosphatase 2C family, were identified as downstream signals in the ABA signaling pathway, they play major roles in vegetative stress responses (Rolland et al. 2006). Clearly, up today we still know much less about signaling mechanism for the ABA-regulated seed germination. As mentioned above, catalase and ROS have been suggested to be implicated in seed germination (Hendricks and Taylorson 1975; Kwak et al. 2003; Apel and Hirt 2004; Laloi et al. 2004). In vegetative tissues, we have demonstrated that AtMKK1/AtMPK6 mediate ABA-induced CAT1 expression and H2O2 production (Xing et al. 2007, 2008). Relationships among catalase, ROS and seed germination are quite complex, we don know whether the AtMKK1/AtMPK6-regualted seed germination is associated with the regulation of theses factors.
Recently, Yoshida et al. (2006) identified and characterized several mutants which showed hypersensitivity to ABA during germination and early growth. Among these mutants ahg3 shows the strongest ABA hypersensitivity, and AHG3 was afterwards clarified to be a gene encoding a protein phosphatase 2C. In addition, Nishimura et al. (2007) also identified ABA hypersensitive mutants, ahg1-1, which showed hypersensitivity to ABA, NaCl, KCl, mannitol, glucose and sucrose during germination and post-germination growth. Likely, AHG1-1 also encodes a protein phosphatase 2C. It is well known that the reversible phosphorylation is catalyzed by both protein phosphatase and kinase. Our results demonstrate that the MAPKs signaling pathway is involved in ABA and glucose signaling during seed germination and the reversible phosphorylation should play the crucial roles in seed germination.
As summarized in Fig. 10, the data presented here demonstrate that AtMKK1 and AtMPK6 are crucial signals in seed germination of Arabidopsis. AtMKK1 and AtMPK6 can both mediate the sugar and ABA signaling in the seed germination. Sugar-regulated seed germination is closely correlated with regulation of the endogenous ABA level in seed, and AtMKK1 and AtMPK6 are just the mediators for the glucose-induced regulation of ABA biosynthesis (Fig. 10).
AtMKK1 and AtMPK6 are crucial signals in seed germination of Arabidopsis. AtMKK1 and AtMPK6 can both mediate the sugar and ABA signaling in the seed germination. Sugar-regulated seed germination is closely correlated with regulation of the endogenous ABA level in seed, and AtMKK1 and AtMPK6 are just the mediators for the glucose-induced regulation of ABA biosynthesis
References
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H et al (2003) Genome-wide insertional mutagensis of Arabidopsis thaliana. Science 301:653–657. doi:10.1126/science.1086391
Apel K, Hirt H (2004) Metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399. doi:10.1146/annurev.arplant.55.031903.141701
Arenas-Huertero F, Arroyo A, Zhou L, Sheen J, Leon P (2000) Analysis of Arabidopsis glucose insensitive mutants, gin5 and gin6, reveals a central role of the plant hormone ABA in the regulation of plant vegetative development by sugar. Genet Dev 14:2085–2096
Assmann SM (2003) OPEN STOMATA1 opens the door to ABA signaling in Arabidopsis guard cells. Trends Plant Sci 8:151–153. doi:10.1016/S1360-1385(03)00052-9
Bewley JD (1997) Seed germination and dormancy. Plant Cell 9:1055–1066. doi:10.1105/tpc.9.7.1055
Bray EA (2002) Abscisic acid regulation of gene expression during water-deficit stress in the ear of the Arabidopsis genome. Plant Cell Environ 25:153–161. doi:10.1046/j.1365-3040.2002.00746.x
Chen JG, Jones AM (2004) AtRGS1 function in Arabidopsis thaliana. Methods Enzymol 389:338–350. doi:10.1016/S0076-6879(04)89020-7
Chen JG, Pandey S, Huang J, Alonso JM, Ecker JR, Assmann SM, Jones AM (2004) GCR1 can act independently of heterotrimeric G-protein in response to brassinosteroids and gibberellins in Arabidopsis seed germination. Plant Physiol 135:907–915. doi:10.1104/pp.104.038992
Cheng WH, Endo A, Zhou L, Penney J, Chen HC, Arroyo A, Leon P, Nambara E, Asami T, Seo M, Koshiba T, Sheen J (2002) A unique short-chain dehydrogenase/reductase in Arabidopsis glucose signaling and abscisic acid biosynthesis and functions. Plant Cell 14:2723–2743. doi:10.1105/tpc.006494
Chow B, McCourt P (2004) Hormone signaling from a developmental context. J Exp Bot 55:247–251. doi:10.1093/jxb/erh032
Coruzzi GM, Zhou L (2001) Carbon and nitrogen sensing and signaling in plants: emerging ‘matrix effects’. Curr Opin Plant Biol 4:247–253. doi:10.1016/S1369-5266(00)00168-0
Dekkers BJ, Schuurmans JA, Smeekens SC (2004) Glucose delays seed germination in Arabidopsis thaliana. Planta 218:579–588. doi:10.1007/s00425-003-1154-9
Ephritikhine G, Fellner M, Vannini C, Lapous D, Barbier-Brygoo H (1999) The sax1 dwarf mutant of Arabidopsis thaliana shows altered sensitivity of growth responses to abscisic acid, auxin, gibberellins and ethylene and is partially rescued by exogenous brassinosteroid. Plant J 18:303–314. doi:10.1046/j.1365-313X.1999.00454.x
Finkelestein RR, Gampala SS, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14:S15–S45
Finkelstein RR, Gibson SI (2002) ABA and sugar interaction regulating development. Cross-talk or voices in a crowd? Curr Opin Plant Biol 5:26–32. doi:10.1016/S1369-5266(01)00225-4
Finkelstein RR, Lynch TJ (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12:599–609
Finkelstein RR, Somerville CR (1990) Three classes of abscisic acid (ABA)-insensitive mutations of Arabidopsis define genes that control overlapping subsets of ABA responses. Plant Physiol 94:1172–1179. doi:10.1104/pp.94.3.1172
Finkelstein RR, Wang ML, Lynch TJ, Rao S, Goodman HM (1998) The Arabidopsis abscisic acid response locus ABI4 encodes an APETALA 2 domain protein. Plant Cell 10:1043–1054
Fujii H, Verslues PE, Zhu JK (2007) Identification of two protein kinases required for abscisic acid regulation of seed germination, root growth, and gene expression in Arabidopsis. Plant Cell 19:485–494. doi:10.1105/tpc.106.048538
Gazzarrini S, McCourt P (2001) Genetic interactions between ABA, ethylene and sugar signaling pathways. Curr Opin Plant Biol 4:387–391. doi:10.1016/S1369-5266(00)00190-4
Gibson SI (2005) Control of plant development and gene expression by sugar signaling. Curr Opin Plant Biol 8:93–102. doi:10.1016/j.pbi.2004.11.003
Giraudat J, Hauge BM, Valon C, Smalle J, Parcy F, Goodman HM (1992) Isolation of the Arabidopsis ABI3 gene by positional cloning. Plant Cell 4:1251–1261
Hendricks SB, Taylorson RB (1975) Breaking of seed dormancy by catalase inhibition. Proc Natl Acad Sci USA 72:306–309. doi:10.1073/pnas.72.1.306
Himmelbach A, Yang Y, Grill E (2003) Relay and control of abscisic acid signaling. Curr Opin Plant Biol 6:470–479. doi:10.1016/S1369-5266(03)00090-6
Hirt H (1997) Multiple roles of MAP kinase in plant signal transduction. Trends Plant Sci 2:11–15. doi:10.1016/S1360-1385(96)10048-0
Hirt H, Asard H (2000) An international conference with a high ambition: meeting report. Trends Plant Sci 5:3–4. doi:10.1016/S1360-1385(99)01508-3
Huang YD, Li CY, Biddle KD, Gibson SI (2008) Identification, cloning and characterization of sis7 and sis10 sugar-insensitive mutants of Arabidopsis. BMC Plant Biol 8:1–24. doi:10.1186/1471-2229-8-104
Huijser C, Kortstee A, Pego J, Weisbeek P, Wisman E, Smeekens S (2000) The Arabidopsis SUCROSE UNCOUPLED-6 gene is identical to ABSCISIC ACID INSENSITIVE-4: involvement of abscisic acid in sugar responses. Plant J 23:577–585. doi:10.1046/j.1365-313x.2000.00822.x
Koornneef M, Reuling G, Karssen CM (1984) The isolation and characterization of abscisic acid-insensitive mutants of Arabidopsis thaliana. Physiol Plant 61:377–383. doi:10.1111/j.1399-3054.1984.tb06343.x
Koornneef M, Bentsink L, Hilhorst H (2002) Seed dormancy and germination. Curr Opin Plant Biol 5:33–36. doi:10.1016/S1369-5266(01)00219-9
Kuhn JM, Schroeder JI (2003) Impacts of altered RNA metabolism on abscisic acid signaling. Curr Opin Plant Biol 6:463–469. doi:10.1016/S1369-5266(03)00084-0
Kwak JM, Mori IC, Pei ZM, Leonhardt N, Torres MA, Dangl JL, Bloom RE, Bodde S, Jones JDG, Schroeder JI (2003) NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J 22:2623–2633. doi:10.1093/emboj/cdg277
Laby RJ, Kincaid MS, Kim D, Gibson SI (2000) The Arabidopsis sugar-insensitive mutants sis4 and sis 5 are defective in abscisic acid synthesis and response. Plant J 23:587–596. doi:10.1046/j.1365-313x.2000.00833.x
Laloi C, Apel K, Danon A (2004) Reactive oxygen signaling: the latest news. Curr Opin Plant Biol 7:323–328. doi:10.1016/j.pbi.2004.03.005
Lefebvre V, North H, Frey A, Sotta B, Seo M, Okamoto M, Nambara E, Marion-Poll A (2006) Functional analysis of Arabidopsis NCED6 and NCED9 genes indicates that ABA synthesized in the endosperm is involved in the induction of seed dormancy. Plant J 45:309–319. doi:10.1111/j.1365-313X.2005.02622.x
Leon P, Sheen J (2003) Sugar and hormone connections. Trends Plant Sci 8:110–116. doi:10.1016/S1360-1385(03)00011-6
Ligterink W, Kroj T, zur Nieden U, Hirt H, Scheel D (1997) Receptor-mediated activation of a MAP kinase in pathogen defense of plants. Science 276:2054–2057. doi:10.1126/science.276.5321.2054
Lopez-Molina L, Mongrand S, Chua NH (2001) A post germination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc Natl Acad Sci USA 98:4782–4787. doi:10.1073/pnas.081594298
Lu C, Han MH, Guevara-Garcia A, Fedoroff NV (2002) Mitogen-activated protein kinase signaling in postgermination arrest of development by abscisic acid. Proc Natl Acad Sci USA 99:15812–15817. doi:10.1073/pnas.242607499
Mizoguchi T, Ichimura K, Shinozaki K (1997) Environmental stress response in plants: the role of mitogen-activated protein kinases. Trends Biotechnol 15:15–19. doi:10.1016/S0167-7799(96)10074-3
Nakagami H, Pitzschke A, Hirt H (2005) Emerging MAP kinase pathways in plant stress signaling. Trends Plant Sci 10:339–346. doi:10.1016/j.tplants.2005.05.009
Nambara E, Marion-Poll A (2003) ABA action and interactions in seeds. Trends Plant Sci 8:213–217. doi:10.1016/S1360-1385(03)00060-8
Nishimura N, Yoshida T, Kitahata N, Asami T, Shinozaki K, Hirayama T (2007) ABA-Hypersensitive Germination1 encodes a protein phosphatase 2C, an essential component of abscisic acid signaling in Arabidopsis seed. Plant J 50:935–949. doi:10.1111/j.1365-313X.2007.03107.x
Ooms J, Leon-Kloosterziel KM, Bartels D, Koornneef M, Karssen CM (1993) Acquisition of desiccation tolerance and longevity in seeds of Arabidopsis thaliana. A comparative study using abscisic acid-insensitive abi3 mutants. Plant Physiol 102:1185–1191
Parcy F, Valon C, Raynal M, Gaubier-Comella P, Delseny M, Giraudat J (1994) Regulation of gene expression programes during Arabidopsis seed development. Roles of the ABI3 locus and of endogenous abscisic acid. Plant Cell 6:1567–1582
Price J, Li TC, Kang SG, Na JK, Jang JC (2003) Mechanisms of glucose signaling during germination of Arabidopsis. Plant Physiol 132:1424–1438. doi:10.1104/pp.103.020347
Qin X, Zeevaart JAD (2002) Overexpression of a 9-cis-epoxycarotenoid dioxygenase gene in Nicotiana lumbaginifolia increases absisic acid and phaseic acid levels and enhances drought tolerance. Plant Physiol 128:544–551. doi:10.1104/pp.010663
Quarrie SA, Whitford PN, Appleford NEJ, Wang TL, Cook SK, Henson IE, Loveys BR (1988) A monoclonal antibody to (S)-abscisic acid: its characterization and use in a radioimmunoassay for measuring abscisic acid in crude extracts of cereal and lupin leaves. Planta 173:330–339. doi:10.1007/BF00401020
Rolland F, Winderickx J, Thevelein JM (2001) Glucose-sensing mechanisms in eukaryotic cells. Trends Biochem Sci 26:310–317. doi:10.1016/S0968-0004(01)01805-9
Rolland F, Moore B, Sheen J (2002) Sugar sensing and signaling in plants. Plant Cell 14:S185–S205
Rolland F, Baena-Gonzalez E, Sheen J (2006) Sugar sensing and signaling in plants: conserved and novel mechanisms. Annu Rev Plant Biol 57:675–709. doi:10.1146/annurev.arplant.57.032905.105441
Schroeder JI, Kwak JM, Allen GJ (2001) Guard cell abscisic acid signalling and engineering drought hardiness in plants. Nature 410:327–330. doi:10.1038/35066500
Seo M, Hanada A, Kuwahara A, Endo A, Okamoto M, Yamauchi Y, North H, Marion-Poll A, Sun TP, Koshiba T et al (2006) Regulation of hormone metabolism in Arabidopsis seeds: phytochrome regulation of abscisic acid metabolism and abscisic acid regulation of gibberellin metabolism. Plant J 48:354–366. doi:10.1111/j.1365-313X.2006.02881.x
Sheen J, Zhou L, Jan JC (1999) Sugars as signaling molecules. Curr Opin Plant Biol 2:410–418. doi:10.1016/S1369-5266(99)00014-X
Smeekens S (2000) Sugar-induced signal transduction in plants. Annu Rev Plant Physiol Plant Mol Biol 51:49–81. doi:10.1146/annurev.arplant.51.1.49
Thompson AJ, Jackson AC, Symonds RC, Mulholland BJ, Dadswell AR, Blake PS, Burbidge A, Taylor IB (2000) Ectopic expression of a tomato 9-cis-epoxycarotenoid dioxygenase gene causes over-production of abscisic acid. Plant J 23:363–374. doi:10.1046/j.1365-313x.2000.00789.x
Toh S, Imamura A, Watanabe A, Nakabayashi K, Okamoto M, Jikumaru Y, Hanada A, Aso Y, Ishiyama K, Tamura N et al (2008) High temperature-induced abscisic acid biosynthesis and its role in the inhibition of gibberellin action in Arabidopsis seeds. Plant Physiol 146:1368–1385. doi:10.1104/pp.107.113738
Ullah H, Chen JG, Wang S, Jones AM (2002) Role of a heterotrimeric G protein in regulation of Arabidopsis seed germination. Plant Physiol 129:897–907. doi:10.1104/pp.005017
Xing Y, Jia WS, Zhang JH (2007) AtMEK1 mediates stress-induced gene expression of CAT1 catalase by triggering H2O2 production in Arabidopsis. J Exp Bot 58:2969–2981. doi:10.1093/jxb/erm144
Xing Y, Jia WS, Zhang JH (2008) AtMKK1 mediates ABA-induced CAT1 expression and H2O2 production via AtMPK6-coupled signaling in Arabidopsis. Plant J 54:440–451. doi:10.1111/j.1365-313X.2008.03433.x
Xiong L, Zhu JK (2003) Regulation of abscisic acid biosynthesis. Plant Physiol 133:29–36. doi:10.1104/pp.103.025395
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. doi:10.1146/annurev.arplant.57.032905.105444
Yoshida T, Nishimura N, Kitahata N, Kuromori T, Ito T, Asami T, Shinozaki K, Hirayama T (2006) ABA-Hypersensitive Germination3 Encodes a Protein Phosphatase 2C (AtPP2CA) That Strongly Regulates Abscisic Acid Signaling during Germination among Arabidopsis Protein Phosphatase 2Cs. Plant Physiol 140:115–126. doi:10.1104/pp.105.070128
Zhou L, Jang JC, Jones TL, Sheen J (1998) Glucose and ethylene signal transduction crosstalk revealed by an Arabidopsis glucose-insensitive mutant. Proc Natl Acad Sci USA 95:10294–10299. doi:10.1073/pnas.95.17.10294
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273. doi:10.1146/annurev.arplant.53.091401.143329
Acknowledgments
We are grateful to grants support from Hong Kong University Grants Committee (AoE/B-07/99), Hong Kong Research Grants Council (HKBU262708) and Hong Kong Baptist University Matching Research Fund. Wensuo Jia is grateful to grants support from the Ph.D. Programs Foundation of Ministry of Education of China (200800190019) and the High-Tech Research and Development (863) Program of China (2006AA100202).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xing, Y., Jia, W. & Zhang, J. AtMKK1 and AtMPK6 are involved in abscisic acid and sugar signaling in Arabidopsis seed germination. Plant Mol Biol 70, 725–736 (2009). https://doi.org/10.1007/s11103-009-9503-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-009-9503-0