Abstract
Indole-3-butyric acid (IBA) and 2,4-dichlorophenoxybutyric acid (2,4-DB) are metabolised by peroxisomal β-oxidation to active auxins that inhibit root growth. We screened Arabidopsis mutants for resistance to IBA and 2,4-DB and identified two new 2,4-DB resistant mutants. The mutant genes encode a putative oxidoreductase (SDRa) and a putative acyl-activating enzyme (AAE18). Both proteins are localised to peroxisomes. SDRa is coexpressed with core β-oxidation genes, but germination, seedling growth and the fatty acid profile of sdra seedlings are indistinguishable from wild type. The sdra mutant is also resistant to IBA, but aae18 is not. AAE18 is the first example of a gene required for response to 2,4-DB but not IBA. The closest relative of AAE18 is AAE17. AAE17 is predicted to be peroxisomal, but an aae17 aae18 double mutant responded similarly to aae18 for all assays. We propose that AAE18 is capable of activating 2,4-DB but IBA activating enzymes remain to be discovered. We present an updated model for peroxisomal pro-auxin metabolism in Arabidopsis that includes SDRa and AAE18.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant peroxisomes play a central role in energy metabolism. They house β-oxidation reactions that metabolise fatty acids to produce energy and work in concert with mitochondria and chloroplasts in photorespiration. Nevertheless, peroxisomal function is relatively poorly characterised. Proteome studies (Reumann et al. 2007) in combination with in silico predictions (Kamada et al. 2003; Reumann et al. 2004) indicate that plant peroxisomes may contain up to 300 proteins. The subcellular localisation of many of these remains unconfirmed. Of those that have been localised to peroxisomes and functionally characterised, the majority are involved in β-oxidation (Baker et al. 2006; Goepfert and Poirier 2007). Other proteins contribute to protein import, photorespiration, or oxidation of specific substrates in reactions that evolve oxygen radicals (Baker and Sparkes 2005; Reumann et al. 2004; Reumann and Weber 2006).
In Arabidopsis, up to 70 proteins are predicted to have similarity to core or auxiliary β-oxidation enzymes (Reumann et al. 2004). The core β-oxidation pathway metabolises saturated long chain fatty acids to supply energy during seed germination and seedling establishment. The pathway includes five steps (Baker et al. 2006): (1) CoA activation by acyl-activating enzymes/long chain acyl-CoA synthetases (LACS6 and LACS7); (2) oxidation by acyl-CoA oxidases (ACX1-4), (3) hydration by enoyl-CoA hydratase/isomerase; (4) oxidation by a dehydrogenase (steps 3 and 4 mediated by multifunctional proteins AIM1 and MFP2); (5) thiolytic cleavage of acetyl-CoA and release of an acyl-CoA shortened by 2C (catalysed by a 3-ketoacyl-CoA thiolase (KAT1, 2 or 5). Citrate synthase then utilizes acetyl-CoA and oxaloacetate in synthesis of citrate (Baker et al. 2006). Other substrates, such as unsaturated fatty acids and hormone precursors may be metabolised by a variety of auxiliary enzymes before being fed into the pathway at various points. Alternatively, depending on the substrates, different isoforms may substitute for core enzymes’ activities (Baker et al. 2006; Goepfert and Poirier 2007). For example the jasmonic acid (JA) precursor OPC:8 is activated by 4-Cinnamoyl CoA ligase (4-CL)-like enzymes (Kienow et al. 2008; Schneider et al. 2005) before entering β-oxidation and indole-3-butyric acid (IBA) is initially oxidized preferentially by IBR3 and ACX3 (Zolman et al. 2007).
Classic indicators of defects in β-oxidation include dependence on exogenously supplied carbon sources such as sucrose for seedling establishment and resistance to pro-auxins such as 2,4-dichlorophenoxybutyric acid (2,4-DB) and IBA. In wild type plants, biologically inactive 2,4-DB is metabolized by one cycle of β-oxidation to the active auxin 2,4-dichlorophenoxyacetic acid (2,4-D). Similarly, IBA is metabolised by a cycle of β-oxidation to indole-3-acetic acid (IAA). Primary root elongation is inhibited by both 2,4-D and IAA. β-Oxidation mutants display root elongation on 2,4-DB and IBA (but not on 2,4-D and IAA) (Baker et al. 2006). In this study we have taken a reverse genetic approach to elucidating roles for auxiliary enzymes of β-oxidation. We targeted uncharacterised, putatively peroxisomal acyl-activating enzymes, hydratases, dehydrogenases and thiolases and screened them for sucrose dependence and 2,4-DB resistance. Using these assays we identified two new proteins with roles in β-oxidation, including the first acyl-activating enzyme required for auxin response.
Methods
Identification of mutants and plant growth
SALK (Alonso et al. 2003), FLAG (Samson et al. 2004), GABI-Kat (Rosso et al. 2003), and CSHL (Sundaresan et al. 1995) T-DNA lines were obtained from stock centres and screened to homozygosity. When available, multiple alleles per gene were ordered. Primer design utilised the T-DNA Express primer design server (http://signal.salk.edu/tdnaprimers.2.html). RT-PCR primers were designed to bound the insertion sites and used to screen each line for the absence of full-length transcript as an indicator that they were transcript knockouts (Table 1). For seed germination and growth assays, half strength MS media (Sigma) was buffered with 0.07% MES, solidified with 0.8% agar and supplemented with 1% sucrose and hormones as indicated. Seed were imbibed on plates at 4°C for 48 h prior to assays. All in vitro assays were conducted at 25°C under continuous white light (approx 120 μE). All knockout lines were initially screened for root elongation on 1 μM 2,4-DB (as an indicator of resistance). They were also screened for defects in etiolation in the absence of sucrose (as described below).
Mutant characterisation
Knockout lines that displayed resistance to 2,4-DB were further characterised by dose responses to 2,4-DB and IBA. They were grown for 6 days on media supplemented with 1% sucrose and 0, 0.5, 1, 2 or 4 μM 2,4-DB or with 0, 5, 10 or 20 μM IBA. In addition, response to active auxins was assayed using media containing 100 nM NAA, 50 nM 2,4-D or 100 nM IAA. IAA and IBA containing plates as well as controls for those experiments were incubated under yellow filters (Lee filter 101) to reduce the photochemical degradation of indole containing compounds (Stasinopoulos and Hangarter 1990). Lateral root assays were conducted by growing seedlings for 3 days on media containing 0.5 × MS and 1% sucrose. Seedlings were then transferred to similar media supplemented either with no hormone, 5 μM IBA or 80 nM NAA and the number of lateral roots was counted 2 and 3 days after transfer. Etiolation was assessed by sowing seeds to 0.5 × MS media containing either 1% or 0% sucrose, exposing them to light for 6 h, then transferring to the dark. Hypocotyl lengths were measured after 4 days dark growth. Digital photographs were taken of assay plates and Image J (Research Services Branch, NIH) was used to measure root and hypocotyl lengths. All root and hypocotyl measurement experiments were conducted at least twice and means were calculated on least 15 seedlings. Single experiments are depicted in the results.
GFP localisation
GFP fusions of SDRa and AAE18 were constructed according to a modification of the fluorescent tagging of full-length proteins (FTFLP) method of Tian et al. (2004). FTFLP introduces GFP into genomic clones (including native promoters and terminators) in a position corresponding to approximately 10 amino acids from the C-terminus and enables analysis of spatial and temporal localisation of proteins during plant development. The position close to the C terminus largely avoids structural and functional protein domains and preserves both N- and C- terminal targeting sequences and membrane anchoring signals in full-length proteins (Tian et al. 2004). We adapted the method for use in transient expression assays. Thus, GFP was inserted within a stretch of hydrophilic residues close to the C-terminus but this was done using coding sequence alone and driven by 35S promoter.
Enhanced GFP (eGFP) was amplified including adapters as described by Tian et al. (2004). The N-terminal portions of SDR and AAE18 were amplified from total cDNA using appropriate 25-mer primers (primers P1 and P2). Oligos P3 and P4, corresponding to the C-terminal 10 amino acids (AAE18) and 13 amino acids (SDRa) plus the stop codon of the protein (i.e. 33–42 nucleotides), were synthesised to comprise a template for direct inclusion in a second round of PCR. Primers P1 and P4 included adapters to enable amplification by universal primers. Primers P2 and P3 had adapters that overlapped with those bounding the GFP fragment. Three-template PCR (TT-PCR) thus included three overlapping templates (the P1–P2 bound fragment, GFP and P3–P4 oligo) and used universal primers with attB cloning sequences. Universal primers and adapters are as described in Tian et al. (2004). TT-PCR products were cloned into Gateway vector pDONR207 and sequenced to check PCR accuracy. The GFP constructs were then introduced into a pGREEN (Hellens et al. 2000) vector modified for Gateway. We used pGREEN0179 containing CAMV 2x35S promoter and CAMV terminator with the Gateway A cassette inserted between them in the SmaI site of the pGREEN multiple cloning site. For colocalisation studies pGREEN0049-RFP-SRL (Pracharoenwattana et al. 2005) was used as a peroxisomal marker.
Plasmids were precipitated onto 1 μm gold particles and biolistically transformed into onion epidermal peel as described in Thirkettle-Watts et al. (2003). Fluorescence images were obtained using Olympus BX61 epifluorescence microscope with GFP (U-MGFPHQ) and RFP filters (U-MRFPHQ) and manipulated with CellR software.
Fatty acid analysis
Plant tissue (approx. 100 mg) was snap-frozen in liquid nitrogen, ground into a fine powder and extracted with 400 μl isopropyl alcohol (containing 0.01% BHT (2,6-di-tert-butyl-4-methylphenol) as an antioxidant) at 75°C for 15 min. Fifty micrograms of C19:0 fatty acid was added as an internal standard to each sample before the extraction. After cooling down to room temperature 600 μl of n-hexane was added to the tube and the mixture sonicated for 5 min. The isopropyl alcohol-hexane mixture was separated from the cell debris and 500 μl was dried under vacuum. The fatty acids in the extract were converted to their methyl esters by heating at 80°C for 2 h in 300 μl of methanol with 2.0% H2SO4 (v/v). After cooling to room temperature and mixing with 300 μl of 0.9% NaCl, fatty acid methyl esters were extracted with 300 μl of n-hexane for GC/MS analysis.
GC–MS analysis utilised an Agilent 6890GC coupled with 5795 mass selective detector. The temperature program was: initial temperature 1 min at 70°C; temperature was then ramped to 76°C at 1°C/min, then ramped to 325°C at 6°C/min and held at 325°C for 10 min. The GC capillary column used for analysis was Varian factor 4 (VF-5 ms, 30 M × 0.25 mm ID and 0.25 μm film). Fatty acid peak areas were normalized to that of the internal standard and the weight of starting tissue to yield mg/g fresh weight for each fatty acid.
Results
Isolation of 2,4-DB resistant mutants
We selected genes encoding predicted peroxisomal proteins with similarity to known β-oxidation proteins for reverse genetic analysis. These included uncharacterised acyl-activating enzymes, hydratases, dehydrogenases/oxidoreductases and thiolases (Table 1). Homozygous mutants were isolated from one EMS (kat2-2) and 37 T-DNA lines representing 20 genes. Of the T-DNA lines, RT-PCR confirmed 23 lines from 17 genes to be lacking full-length transcripts (Table 1; Supplementary Figure 1). The transcript knockout lines so obtained are listed in Table 1 by their allelic designations. kat2-2 is a new Col-0 EMS mutant for this gene originally identified in a screen for new sucrose dependent mutants (Eastmond 2006). We backcrossed kat2-2 twice and sequencing revealed the lesion to be S140F (TCT → TTT), a change that presumably affects protein topology near the catalytic Cys138 residue (see Sundaramoorthy et al. 2006). kat2-1 is in Ws background (Germain et al. 2001).
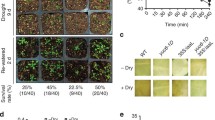
We screened mutants listed in Table 1 for resistance to pro-auxins 2,4-DB and IBA. Screening of homozygous lines on 1 μM 2,4-DB yielded two new 2,4-DB resistant mutants, sdra and aae18. (Fig. 1). SDRa is a short chain dehydrogenase/reductase family protein and AAE18 an acyl-activating enzyme (acyl-CoA synthetase) family protein. When screened on 10 μM IBA, the two sdra alleles were resistant to IBA, but all three aae18 alleles were sensitive (Fig. 1). No further new IBA resistant mutants were revealed amongst the lines listed in Table 1. We also characterized chy1-4, a new mutant allele of CHY1. It is similar to other characterised strong alleles of CHY1 which are highly resistant to pro-auxins (Lange et al. 2004; Zolman et al. 2001a, b). Similarly, we obtained ibr3-9 as a characterised, partially resistant control (see Zolman et al. 2007). Both of the sdra alleles and two of the aae18 alleles were subsequently characterised in detail, for auxin response and other characteristics.
sdra and aae18 are resistant only to pro-auxins
The new 2,4-DB resistant lines were characterised for dose responses to 2,4-DB. On increasing concentrations of 2,4-DB, aae18 and sdra appeared to be as resistant for root elongation as chy1 up to 1 μM 2,4-DB, but were less resistant above this concentration (Fig. 2a). aae18-1, aae18-2 and sdra-1 were more resistant to 2,4-DB than ibr3-9. The sdra-2 mutant is in Ler background and shows similar trends but both the Ler wild-type and sdra-2 are apparently more sensitive to 2,4-DB than Col-0 and sdra-1 and are thus presented separately (Fig. 2b). Both kat2-1 and kat2-2 appear to be as resistant as chy1-4 to 2,4-DB (Fig. 2c). The kat2 alleles were more resistant to higher than lower concentrations of IBA and 2,4-DB (Fig. 2c, f). This was previously noted by Lange et al. (2004) for kat2 grown on 2,4-DB, but there is no obvious explanation for this phenomenon. Neither of the kat2 alleles germinated on 4 μM 2,4-DB. Lethality of kat2-1 at high concentrations of 2,4-DB was not previously reported (Lange et al. 2004).
Response of mutants to varying concentrations of 2,4-dichlorobutyric acid (2,4-DB) and indole butyric acid (IBA). a–c Response to 2,4-DB. d–f Response to IBA. a, d sdra-1 and aae18 mutants. b, e Comparison of sdra-1 (Col-0) and sdra-2 (Ler) and their respective wild types. c, f kat2-1 and kat2-2 mutants and their respective wild types (Ws and Col-0). In (a), (c), (d) and (f), chy1-4 and ibr3-9 are included for comparative purposes. Plants were grown for 6 days in continuous light on half strength MS media supplemented with 1% sucrose and the indicated concentration of 2,4-DB or IBA. Each value represents the mean root length with standard error indicated (n ≥ 15)
When grown on IBA, chy1-4, ibr3-9, kat2-1 and kat2-2 all displayed resistance consistent with that reported in previous studies (Fig. 1). Compared to growth on media lacking hormone, root elongation of all lines (mutant and wild type) was markedly inhibited by 5 μM IBA (Fig. 2d–f). This response is at variance with previous reports for ibr3 and chy1 where root growth inhibition was less severe in response to 5 μM IBA (Zolman et al. 2007). Similarly, the greater inhibition of root elongation at 5 μM compared to 10 μM IBA for the stronger mutants (chy1-4 and kat2-2) was not reported earlier (c.f. Zolman et al. 2001a, 2007). These two discrepancies are most likely due to different media and growth conditions of the studies. In accordance with Zolman et al. (2007), ibr3-9 was less severe in its response to IBA than chy1-4 (Fig. 2d). kat2 mutants were at least as resistant as chy1-4 (Fig. 2f). The sdra-1 mutant appeared to be almost as resistant to IBA as chy1-4 at all concentrations tested and was more resistant than ibr3-9 to IBA (Fig. 2d). sdra-2 displayed a similar trend of high resistance with its difference to sdra-1 most likely attributable to the ecotype differences of the alleles (Fig. 2e). Interestingly, all three aae18 alleles behaved like wild type in their IBA response (Figs. 1, 2d; aae18-3 not shown).
sdra, aae18, kat2, chy1 and ibr3 respond normally to the active auxins IAA, 2,4-D and NAA (Figs. 1, 3) indicating there is no general block to auxin signaling. When grown on 100 nM IAA, 50 nM 2,4-D or 100 nM NAA, all genotypes showed inhibition of root length comparable to their respective wild types. Ws (and mutants in Ws background) had a low level of natural resistance to 2,4-D and 2,4-DB compared to Col-0 and Ler (Figs. 1, 3). The resistance of Ws to low concentration of 2,4-DB has been previously reported (Lange et al. 2004). Interestingly though, both kat2-1 (Ws) and kat2-2 (Col-0) appeared to be somewhat resistant to 2,4-D (Fig. 3).
To further characterise their auxin response, all 2,4-DB-resistant mutants were assessed for their capacity to produce lateral roots. Seeds were germinated and grown for 3 days on medium containing 1% (w/v) sucrose, then plants were transferred to medium containing hormones for 2 days. With the exception of kat2, all mutants produced numbers of lateral roots comparable to their wild types when grown on media without added hormone. Both kat2 mutants produced less lateral roots under these conditions than their wild types (Fig. 4). This was also the case for kat2 grown for differing durations on sucrose up to 8 days in total (data not shown).
Lateral root formation in sdra and aae18 mutants. Plants were grown for 3 days in continuous light on half strength MS medium supplemented with 1% sucrose, then transferred to similar base media supplemented with the indicated hormones. The number of lateral roots was counted 2 days after transfer. Each value represents the mean number of lateral roots with standard error indicated (n ≥ 15)
IBA stimulated lateral root formation considerably in wild type controls as well as in aae18. In contrast, sdra-1, sdra-2, kat2-1, kat2-2, chy1-4 and ibr3-9 all produced fewer lateral roots on IBA than their respective wild types (Fig. 4). Marked increases in the number of lateral roots were observed in all cases when the seedlings were transferred to media supplemented with 80 nM NAA, indicating no general block to lateral rooting capacity.
SDRa and AAE18 localise to the peroxisome
GFP fusions with SDRa and AAE18 were constructed by inserting GFP in frame into the coding sequence close to the C-terminus. By this method SDRa-GFP and AAE18-GFP both localised to peroxisomes when compared to a peroxisomal RFP-SRL marker (Fig. 5). SDRa possesses a C-terminal-SRL and AAE18 a C-terminal-SRI. These are both canonical plant PTS1 signal peptides (Reumann et al. 2004), consistent with the observation of peroxisomal GFP. SDRa has moreover been identified in the proteome of leaf peroxisomes (Reumann et al. 2007).
In vivo targeting of SDRa and AAE18. GFP was cloned in frame 10–13 amino acid residues upstream from the stop codon in the coding sequence of AAE18 and SDRa. These constructs were co-transformed with the peroxisomal marker RFP-SRL into onion epidermal cells by biolistic bombardment. Localisation of fluorescent proteins was assessed one day after transformation by fluorescence microscopy. The length of the scale bar is 20 μm
sdra and aae18 are not dependent on exogenous sucrose for seedling establishment
To examine whether any of the lines were also affected in triacylglycerol (TAG) metabolism, we grew seedlings in the dark on media with or without added sucrose. sdra and aae18 alleles were able to etiolate normally in the absence of sucrose, indicating that TAG metabolism was not affected sufficiently to disrupt seedling germination and growth (Fig. 6a). As expected (Germain et al. 2001), neither of the kat2 alleles emerged from seed coats during this experiment, and chy1-4 grew poorly without sucrose, as reported previously for CHY1 mutants (Zolman et al. 2001a, b). Seedling establishment and early development of sdra and aae18 alleles in the light without exogenously supplied sucrose appeared normal (data not shown).
TAG metabolism is normal in sdra and aae18. a Etiolation of sdra and aae18 is normal in the absence of sucrose. Plants were germinated and grown for 4 days in the dark on half strength MS medium with or without supplement of 1% sucrose. b Fatty acid analysis of wild-type, kat2-1 and sdra-1 mutants. Fatty acids were extracted from seedlings grown for 2 days on half strength MS medium supplemented with 1% sucrose, and then identified and quantified using GC–MS
Fatty acid metabolism in sdra-1
Although it is expressed during early seedling establishment (Supplementary Figure 1), further information about expression of the AAE18 gene is lacking because it is not represented on the Affymetrix genechip ATH1. However, much information about expression of SDRa is available. SDRa expression was examined using publicly available microarray data (Schmid et al. 2005). Using Genevestigator V3 (Zimmermann et al. 2004), SDRa is highly expressed throughout the plant life cycle, but is more highly expressed in germinated seed. During development, similar expression patterns are observed in SDRa, ECH1a, ECH1b, MFP2, ACX2, IBR3, ACX3, ACX4, KAT2 and NQR (not shown). In line with the apparent co-regulation with β-oxidation genes, SDRa also clusters with NQR and β-oxidation genes when analysed using ATTED (Obayashi et al. 2007). Many β-oxidation genes are in the top 300 genes coexpressed with SDRa including NQR (position 1), MFP2 (5), IBR3 (79) and AIM1 (105). Analysis of transcript abundance during the diurnal cycle (Smith et al. 2004) also indicates that in mature leaf tissue, SDRa is co-regulated with β-oxidation genes (Supplemental Figure 2).
Despite the apparent link between SDRa and β-oxidation genes, no evidence was obtained to show that mobilisation of TAG for seedling growth was impaired in sdra-1 or sdra-2 (Fig. 6a). Nevertheless, the possibility that β-oxidation of specific fatty acids could be impaired was investigated. Total fatty acids were isolated for GC-MS analysis from two day old seedlings, a stage when TAG breakdown is taking place most rapidly (Germain et al. 2001). The amounts of numerous medium and long chain fatty acids were about the same in Col-0 and sdra-1, including the major TAG fatty acids linoleic acid (C18:2) and eicosenoic acid (C20:1) (Fig. 6b). In contrast kat2-1 contained elevated levels of these fatty acids due to impaired TAG breakdown. These features indicate that fatty acid metabolism in sdra-1 is indistinguishable from wild type, consistent with the hypothesis that SDRa participates preferentially in auxin synthesis.
aae17 aae18 double knockout remains sensitive to IBA
AAE18 is a member of a large family of acyl-activating enzymes and redundancy of peroxisomal isoform functions has already been demonstrated in the family, with the closely related LACS6 and LACS7 involved in activating TAG-derived fatty acids, but not auxins (Fulda et al. 2004). For this reason we generated a double mutant of AAE18 and AAE17 to examine auxin responses. AAE17 also has a PTS1 (C-terminal SKL). It is the closest relative of AAE18 in the Arabidopsis genome and a good candidate for double mutant analysis. The aae17-1 aae18-1 double mutant was no more resistant to 2,4-DB than aae18-1 (Fig. 7a), normally sensitive to IBA (Fig. 7b) and mature auxins (Fig. 7c), produced normal numbers of lateral roots when stimulated by IBA and NAA (Fig 7d) and was not dependent on sucrose for etiolation (Fig. 7e).
aae17 does not contribute to metabolism of pro-auxins. a–c Root length of aae17, aae18 and aae17 aae18 double mutants grown on different auxin containing media. Root lengths of aae17-1 and aae17-1 aae18-1 double mutants were not significantly different to wild-type and aae18-1, respectively when grown on different concentrations of 2,4-DB (a) and IBA (b), but responded normally to IAA, 2,4-D and NAA (c). d Lateral root initiation in response to IBA and NAA. (e) aae17-1, aae18-1 and aae17-1 aae18-1 are not dependent on sucrose for hypocotyl elongation
Discussion
A model for IBA and 2,4-DB metabolism
The metabolism in peroxisomes of pro-auxins IBA and 2,4-DB to the biologically active forms IAA and 2,4-DB has been used extensively in screening for peroxisomal mutants with reduced capacity for β-oxidation (Baker et al. 2006; Hayashi et al. 1998; Woodward and Bartel 2005). However, some of the enzymes involved remain to be elucidated. In this work we employed a systematic screen of available T-DNA lines for peroxisomal genes with predicted roles in β-oxidation. We identified two PTS1-containing, peroxisome-targeted proteins that we propose to act as, firstly, an acyl-CoA synthetase (acyl-activating enzyme 18, AAE18) that activates 2,4-DB, and, secondly, an oxidoreductase (SDRa) that oxidises 3-hydroxyacyl-CoA derivatives of both 2,4-DB and IBA. Knockouts for SDRa and AAE18 were shown to be compromised in their ability to respond to pro-auxins but were not dependent on sucrose for seedling establishment. They are thus potentially members of the class of β-oxidation genes that have as their primary function hormone metabolism rather than energy provision (Zolman et al. 2000).
Based on the mutant phenotypes reported here and in other studies, we were able to elaborate pathways for β-oxidation of 2,4-DB and IBA (Fig. 8). By this model 2,4-DB is activated by AAE18. It is subsequently oxidized by IBR3 (Zolman et al. 2007), possibly in conjunction with ACX3 and ACX4, mutants of which are also partially resistant to 2,4-DB (Rylott et al. 2003; Zolman et al. 2000). AIM1 likely contributes the hydratrase/isomerase activity after which SDRa oxidises the 3-hydroxyacyl-CoA derivative. The thiolase activity is provided primarily by KAT2. Ultimately, the products of β-oxidation must be released from CoA, a step that could be catalysed by one of the several thioesterases predicted to be targeted to peroxisomes (Reumann et al. 2004). We propose that the oxidation of IBA proceeds via a similar pathway (Fig. 8), except that enzymes that might activate IBA remain unknown. Also, in addition to IBR3 and ACX3, the first oxidative step in the β-oxidation of IBA may be catalyzed by ACX1 and ACX2, mutants of which are resistant to IBA (Adham et al. 2005). Alternatively, acx1 and acx2 mutants may cause sequestration of the peroxisomal CoA supply and indirectly yield IBA-resistance (Adham et al. 2005).
Model for β-oxidation of 2,4-DB and IBA. This model is compiled based on mutant phenotypes and other data and includes the two new genes presented in this work (SDRa and AAE18). Fatty acid β-oxidation is presented for comparison and to highlight differences between it and the auxin pathways. (Abbreviations: IBE-CoA indole butenoyl-CoA; I-3HB-CoA indole-3-hydroxybutenyl-CoA; I-3 KB-CoA indole-3-keto-butenyl-CoA; DBE-CoA 2,4-dichlorobutenoyl-CoA; D-3HB-CoA 2,4-dichloro-3-hydroxybutenyl-CoA; D-3 KB-CoA 2,4-dichloro-keto-butenyl-CoA)
It is likely that AIM1 is the major hydratase/isomerase in 2,4-DB and IBA oxidation because the aim1 mutant is the only hydratase/isomerase containing protein identified to date that is 2,4-DB and IBA resistant (see Richmond and Bleecker 1999; Zolman et al. 2000). In contrast, mfp2 is sensitive to 2,4-DB (Rylott et al. 2006) and IBA (data not shown), as are the ECHI (enoyl-CoA hydratase/isomerase) knockouts we examined. Nevertheless, AIM1 may still act with other hydratase/isomerase proteins and it would be instructive to make double mutants with the various ECHI genes that were individually sensitive to these hormone precursors. The double knockout aim1 mfp2 is embryo lethal and thus not testable in this manner (Rylott et al. 2006).
Although AIM1 is at least a trifunctional protein containing epimerase/hydratase and dehydrogenase domains, we propose that that SDRa contributes the bulk of dehydrogenase activity in the oxidation of pro-auxins (rather than AIM1). For example, sdra-1 is almost as resistant to IBA as kat2 and is markedly more resistant than ibr3, consistent with evidence that IBR3 and ACX3 are partially redundant during the initial oxidative step (Zolman et al. 2007). Similarly, sdra-1 has a stronger 2,4-DB phenotype than ibr3. These observations do not rule out contribution of another dehydrogenase activity at this step. Future studies might examine this by partially complementing aim1 with truncated protein variants containing either the hydratase/isomerase or the dehydrogenase domains.
KAT2 is likely to be the major thiolase in the pathway as other thiolase mutants (kat1 and kat5) are as sensitive as wildtype to IBA and 2,4-DB. The kat2 mutant was as resistant to IBA and 2,4-DB as chy1 (Figs. 1 and 2) in which thiolase activity generally is inhibited by accumulation of the toxic intermediate methacrylyl-CoA (Lange et al. 2004; Zolman et al. 2001a). We observed that kat2 had a slightly shorter root (Fig. 3) and grew fewer lateral roots (Fig. 4) than wild-type on sucrose medium (without hormones). One possible explanation for this is that kat2 was delayed in development due to the severe β-oxidation lesion. Alternatively, a block in the conversion of endogenous IBA to IAA may inhibit lateral root initiation. This has previously been suggested to explain the similar phenotype of peroxisomal ABC transporter (pxa1) mutants (Zolman et al. 2001b).
We have argued that SDRa fills the role as an oxidoreductase during IBA and 2,4-DB metabolism. This is based on the phenotypes of IBA and 2,4-DB mutants. A possible alternative function of SDRa is that it might constitute the as yet unidentified 3-hydroxy-isobutyrate dehydrogenase (HIBDH) that immediately follows the 3-hydroxy-isobutyryl CoA-hydrolase (HIB-CoAH) encoded by CHY1 (Taylor et al. 2004). The auxin-phenotypes observed in sdra could potentially phenocopy those of chy1 if there was feedback inhibition of CHY1 due to buildup of the 3-hydroxy-isobutyrate in sdra. We view this as unlikely, however, because TAG breakdown in sdra is unaffected, indicating that the lesion is specific to auxin metabolism and not fatty acid β-oxidation.
Endogenous role of AAE18
Metabolic mutants published to date that are 2,4-DB resistant have for the most part also been resistant to IBA (Woodward and Bartel 2005; Zolman et al. 2007). Exceptions are the acx2 mutant and acx1 acx2 double knockout that are resistant only to IBA (Adham et al. 2005). AAE18 is the first mutant found to be resistant only to 2,4-DB. This indicates either that IBA is not a substrate of AAE18, or that there is genetic redundancy for the activation of IBA. A double knockout of aae18 with aae17, the closest relative of AAE18 (Shockey et al. 2003) was made and shown also to be sensitive to IBA (Fig. 7). Thus, it is unlikely that either AAE18 or AAE17 have a role in β-oxidation of IBA. This raises the questions of if, how and where IBA is activated?
There are 63 members in the AAE superfamily (Shockey et al. 2003), 17 of which have a PTS (Reumann et al. 2004). The peroxisome-targeted AAE proteins do not present as good candidates to be an IBA activating enzyme. None of the AAE mutants we tested were IBA resistant, the lacs6 lacs7 double mutant is sensitive to pro-auxins (Fulda et al. 2004), AAE7 can activate acetate and butyrate (Shockey et al. 2003; Turner et al. 2005), AAE11 can activate medium chain length fatty acids (C6–C8) (Shockey et al. 2003), and BZO1 is a benzoyl-CoA ligase in the pathway of benzoyloxyglucosinolate synthesis (Kliebenstein et al. 2007). Additionally, a systematic in vitro substrate screen of the peroxisomal 4CL-like proteins indicated that they could not utilise IBA as a substrate (Kienow et al. 2008). An alternative possibility is that IBA-CoA may be synthesised in the cytosol and imported to the peroxisome by the ABC transporter CTS/PXA1/PED1. CTS imports acyl-CoAs (Footitt et al. 2002; Fulda et al. 2004; Zolman et al. 2001b) and also potentially imports some unesterified substrates (e.g. OPDA during JA biosynthesis (Kienow et al. 2008; Theodoulou et al. 2005)). Potential extra-peroxisomal activation enzymes include the 19 member JAR1/GH3 sub-family of AAE enzymes, some of which are able to conjugate JA (JAR1), SA and IAA (one gene), or IAA (six genes) to amino acids or other compounds (Shockey et al. 2003; Staswick et al. 2002, 2005).
What is the natural substrate for AAE18? No clues can be garnered through microarray data as AAE18 is not represented on the ATH1 chip, and it does not respond significantly in the CATMA arrays (http://www.catma.org/). We can look to compounds that are structurally similar to 2,4-DB. For example, trans-cinnamic acid is potentially metabolised in peroxisomes to benzoic acid during salicylic acid synthesis (Wildermuth 2006). However, this may be difficult to dissect genetically because it has been previously shown that direct synthesis of SA via isochorismate synthase accounts for the vast majority of defense related SA synthesis, and a variety of alternative routes probably contribute to basal SA levels only (Wildermuth 2006; Wildermuth et al. 2001).
Concluding remarks
Recent in silico and proteome studies (Reumann et al. 2004; 2007) have highlighted the dominance of fatty acid β-oxidation in peroxisome function, with perhaps 30% of the peroxisome proteins involved in such metabolism. Here we have screened mutants in 13 uncharacterised putatively peroxisomal genes, but found none that affect seedling growth (and by implication, TAG metabolism) or other visual aspects of plant growth and development (data not shown). We have identified two new proteins that potentially contribute to auxin metabolism, including the first example of a mutant resistant to 2,4-DB but normally responsive to IBA. This indicates that it may be possible to genetically engineer resistance to pro-auxin herbicides without compromising the normal functions of endogenous auxins. Understanding the roles of other peroxisomal proteins represents a major challenge that will require concerted and multidisciplinary approaches.
References
Adham AR, Zolman BK, Millius A, Bartel B (2005) Mutations in Arabidopsis acyl-CoA oxidase genes reveal distinct and overlapping roles in beta-oxidation. Plant J 41:859–874
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Baker A, Sparkes IA (2005) Peroxisome protein import: some answers, more questions. Curr Opin Plant Biol 8:640–647
Baker A, Graham IA, Holdsworth M, Smith SM, Theodoulou FL (2006) Chewing the fat: beta-oxidation in signalling and development. Trends Plant Sci 11:124–132
Eastmond PJ (2006) SUGAR-DEPENDENT1 encodes a patatin domain triacylglycerol lipase that initiates storage oil breakdown in germinating Arabidopsis seeds. Plant Cell 18:665–675
Footitt S, Slocombe SP, Larner V, Kurup S, Wu Y, Larson T, Graham I, Baker A, Holdsworth M (2002) Control of germination and lipid mobilization by COMATOSE, the Arabidopsis homologue of human ALDP. EMBO J 21:2912–2922
Fulda M, Schnurr J, Abbadi A, Heinz E, Browse J (2004) Peroxisomal Acyl-CoA synthetase activity is essential for seedling development in Arabidopsis thaliana. Plant Cell 16:394–405
Germain V, Rylott EL, Larson TR, Sherson SM, Bechtold N, Carde JP, Bryce JH, Graham IA, Smith SM (2001) Requirement for 3-ketoacyl-CoA thiolase-2 in peroxisome development, fatty acid beta-oxidation and breakdown of triacylglycerol in lipid bodies of Arabidopsis seedlings. Plant J 28:1–12
Goepfert S, Poirier Y (2007) Beta-oxidation in fatty acid degradation and beyond. Curr Opin Plant Biol 10:245–251
Hayashi M, Toriyama K, Kondo M, Nishimura M (1998) 2, 4-Dichlorophenoxybutyric acid-resistant mutants of Arabidopsis have defects in glyoxysomal fatty acid beta-oxidation. Plant Cell 10:183–195
Hellens RP, Edwards EA, Leyland NR, Bean S, Mullineaux PM (2000) pGreen: a versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol Biol 42:819–832
Kamada T, Nito K, Hayashi H, Mano S, Hayashi M, Nishimura M (2003) Functional differentiation of peroxisomes revealed by expression profiles of peroxisomal genes in Arabidopsis thaliana. Plant Cell Physiol 44:1275–1289
Kienow L, Schneider K, Bartsch M, Stuible HP, Weng H, Miersch O, Wasternack C, Kombrink E (2008) Jasmonates meet fatty acids: functional analysis of a new acyl-coenzyme A synthetase family from Arabidopsis thaliana. J Exp Bot 59:403–419
Kim HU, van Oostende C, Basset GJ, Browse J (2008) The AAE14 gene encodes the Arabidopsis o-succinylbenzoyl-CoA ligase that is essential for phylloquinone synthesis and photosystem-I function. Plant J 54:272–283
Kliebenstein DJ, D’Auria JC, Behere AS, Kim JH, Gunderson KL, Breen JN, Lee G, Gershenzon J, Last RL, Jander G (2007) Characterization of seed-specific benzoyloxyglucosinolate mutations in Arabidopsis thaliana. Plant J 51:1062–1076
Lange PR, Eastmond PJ, Madagan K, Graham IA (2004) An Arabidopsis mutant disrupted in valine catabolism is also compromised in peroxisomal fatty acid beta-oxidation. FEBS Lett 571:147–153
Obayashi T, Kinoshita K, Nakai K, Shibaoka M, Hayashi S, Saeki M, Shibata D, Saito K, Ohta H (2007) ATTED-II: a database of co-expressed genes and cis elements for identifying co-regulated gene groups in Arabidopsis. Nucleic Acids Res 35:D863–D869
Pracharoenwattana I, Cornah JE, Smith SM (2005) Arabidopsis peroxisomal citrate synthase is required for fatty acid respiration and seed germination. Plant Cell 17:2037–2048
Reumann S, Weber APM (2006) Plant peroxisomes respire in the light: Some gaps of the photorespiratory C2 cycle have become filled—others remain. Biochim Biophys Acta 1763:1496–1510
Reumann S, Ma C, Lemke S, Babujee L (2004) AraPerox. A database of putative Arabidopsis proteins from plant peroxisomes. Plant Physiol 136:2587–2608
Reumann S, Babujee L, Ma C, Wienkoop S, Siemsen T, Antonicelli GE, Rasche N, Luder F, Weckwerth W, Jahn O (2007) Proteome analysis of Arabidopsis leaf peroxisomes reveals novel targeting peptides, metabolic pathways, and defense mechanisms. Plant Cell 19:3170–3193
Richmond TA, Bleecker AB (1999) A defect in beta-oxidation causes abnormal inflorescence development in Arabidopsis. Plant Cell 11:1911–1924
Rosso MG, Li Y, Strizhov N, Reiss B, Dekker K, Weisshaar B (2003) An Arabidopsis thaliana T-DNA mutagenized population (GABI-Kat) for flanking sequence tag-based reverse genetics. Plant Mol Biol 53:247–259
Rylott EL, Rogers CA, Gilday AD, Edgell T, Larson TR, Graham IA (2003) Arabidopsis mutants in short- and medium-chain acyl-CoA oxidase activities accumulate acyl-CoAs and reveal that fatty acid beta-oxidation is essential for embryo development. J Biol Chem 278:21370–21377
Rylott EL, Eastmond PJ, Gilday AD, Slocombe SP, Larson TR, Baker A, Graham IA (2006) The Arabidopsis thaliana multifunctional protein gene (MFP2) of peroxisomal beta-oxidation is essential for seedling establishment. Plant J 45:930–941
Samson F, Brunaud V, Duchene S, De Oliveira Y, Caboche M, Lecharny A, Aubourg S (2004) FLAGdb ++: a database for the functional analysis of the Arabidopsis genome. Nucleic Acids Res 32:D347–D350 database issue
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Schneider K, Kienow L, Schmelzer E, Colby T, Bartsch M, Miersch O, Wasternack C, Kombrink E, Stuible HP (2005) A new type of peroxisomal acyl-coenzyme A synthetase from Arabidopsis thaliana has the catalytic capacity to activate biosynthetic precursors of jasmonic acid. J Biol Chem 280:13962–13972
Shockey JM, Fulda MS, Browse J (2003) Arabidopsis contains a large superfamily of acyl-activating enzymes. Phylogenetic and biochemical analysis reveals a new class of acyl-coenzyme a synthetases. Plant Physiol 132:1065–1076
Smith SM, Fulton DC, Chia T, Thorneycroft D, Chapple A, Dunstan H, Hylton C, Zeeman SC, Smith AM (2004) Diurnal changes in the transcriptome encoding enzymes of starch metabolism provide evidence for both transcriptional and posttranscriptional regulation of starch metabolism in Arabidopsis leaves. Plant Physiol 136:2687–2699
Stasinopoulos TC, Hangarter RP (1990) Preventing photochemistry in culture media by long-pass light filters alters growth of cultured tissues. Plant Physiol 93:1365–1369
Staswick PE, Tiryaki I, Rowe ML (2002) Jasmonate response locus JAR1 and several related Arabidopsis genes encode enzymes of the firefly luciferase superfamily that show activity on jasmonic, salicylic, and indole-3-acetic acids in an assay for adenylation. Plant Cell 14:1405–1415
Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, Maldonado MC, Suza W (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17:616–627
Sundaramoorthy R, Micossi E, Alphey MS, Germain V, Bryce JH, Smith SM, Leonard GA, Hunter WN (2006) The crystal structure of a plant 3-ketoacyl-CoA thiolase reveals the potential for redox control of peroxisomal fatty acid [beta]-oxidation. J Mol Biol 359:347–357
Sundaresan V, Springer P, Volpe T, Haward S, Jones JD, Dean C, Ma H, Martienssen R (1995) Patterns of gene action in plant development revealed by enhancer trap and gene trap transposable elements. Genes Dev 9:1797–1810
Taylor NL, Heazlewood JL, Day DA, Millar AH (2004) Lipoic acid-dependent oxidative catabolism of alpha-keto acids in mitochondria provides evidence for branched-chain amino acid catabolism in Arabidopsis. Plant Physiol 134:838–848
Theodoulou FL, Job K, Slocombe SP, Footitt S, Holdsworth M, Baker A, Larson TR, Graham IA (2005) Jasmonic acid levels are reduced in COMATOSE ATP-binding cassette transporter mutants. Implications for transport of jasmonate precursors into peroxisomes. Plant Physiol 137:835–840
Thirkettle-Watts D, McCabe TC, Clifton R, Moore C, Finnegan PM, Day DA, Whelan J (2003) Analysis of the alternative oxidase promoters from soybean. Plant Physiol 133:1158–1169
Tian GW, Mohanty A, Chary SN, Li S, Paap B, Drakakaki G, Kopec CD, Li J, Ehrhardt D, Jackson D, Rhee SY, Raikhel NV, Citovsky V (2004) High-throughput fluorescent tagging of full-length Arabidopsis gene products in planta. Plant Physiol 135:25–38
Turner JE, Greville K, Murphy EC, Hooks MA (2005) Characterization of Arabidopsis fluoroacetate-resistant mutants reveals the principal mechanism of acetate activation for entry into the glyoxylate cycle. J Biol Chem 280:2780–2787
Wildermuth MC (2006) Variations on a theme: synthesis and modification of plant benzoic acids. Curr Opin Plant Biol 9:288–296
Wildermuth MC, Dewdney J, Wu G, Ausubel FM (2001) Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414:562–565
Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot (Lond) 95:707–735
Zimmermann P, Hirsch-Hoffmann M, Hennig L, Gruissem W (2004) GENEVESTIGATOR. Arabidopsis microarray database and analysis toolbox. Plant Physiol 136:2621–2632
Zolman BK, Yoder A, Bartel B (2000) Genetic analysis of indole-3-butyric acid responses in Arabidopsis thaliana reveals four mutant classes. Genetics 156:1323–1337
Zolman BK, Monroe-Augustus M, Thompson B, Hawes JW, Krukenberg KA, Matsuda SP, Bartel B (2001a) chy1, an Arabidopsis mutant with impaired beta-oxidation, is defective in a peroxisomal beta-hydroxyisobutyryl-CoA hydrolase. J Biol Chem 276:31037–31046
Zolman BK, Silva ID, Bartel B (2001b) The Arabidopsis pxa1 mutant is defective in an ATP-binding cassette transporter-like protein required for peroxisomal fatty acid beta-oxidation. Plant Physiol 127:1266–1278
Zolman BK, Nyberg M, Bartel B (2007) IBR3, a novel peroxisomal acyl-CoA dehydrogenase-like protein required for indole-3-butyric acid response. Plant Mol Biol 64:59–72
Zolman BK, Martinez N, Millius A, Adham AR, Bartel B (2008) Identification and characterization of Arabidopsis indole-3-butyric acid response mutants defective in novel peroxisomal enzymes. Genetics 180:237–251
Acknowledgement
This project was funded by the Australian Research Council (Grants FF0457721 and CE0561495), the Western Australian Government’s Centre of Excellence Program and an Australian Postgraduate Award to AAGW. We thank Peter Eastmond for kind donation of sucrose dependent lines from which we obtained kat2-2. We thank also ABRC, SALK, INRA Versailles, NASC, GABI and CSHL for T-DNA seed lines.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1
RT-PCRs of new T-DNA lines reported in this study. Seedlings were grown for 5–7 days on half-strength MS media supplemented with 1% sucrose. RNA was isolated using Biorad Aurum RNA isolation kit. RT utilised Biorad iScript. Where insertions interrupted putative transcripts, primers were designed to anneal to transcribed sequences bounding the insertion sites. In cases of insertions in UTRs, 300–400 bp, amplicons were designed using primer3. RT-PCR of actin2 was used as a control. Lines are assumed to be knockouts if transcript was absent; knockouts are indicated by allelic designation if transcript was absent and by their name if transcript remained in the line. AGIs, and a full list of alleles and T-DNA names can be found in Table 1. (740 kb)
Supplementary Figure 2
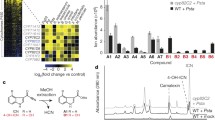
SDRa expression correlates with β-oxidation genes during the diurnal cycle. Many β-oxidation genes are co-ordinately down-regulated during the light period. Core β-oxidation genes (LACS6, LACS7, ACX2, MFP2, AIM1, KAT2, PMDH1) and transcripts of some putative auxiliary genes (e.g. CSY3, ECH1a, ECH2, SCP-2) experience a maximum during the dark period and reduced expression in the light. SDRa follows a similar pattern. There are rapid changes in transcript levels upon transition from light to dark and dark to light. For many of these genes, there is also an increase in expression during the light period similar to that seen for starch-degrading enzymes (Smith et al. 2004) that may be anticipatory of a dark phase where net energy reserves are consumed. Putative PTS1- and PTS2-encoding transcripts (identified using ARAPEROX; Reumann et al. 2004) and from other peroxisomal, non-PTS containing genes were identified in the transcriptome data from Smith et al. (2004). These 254 transcripts were further filtered to remove those for which the maximum normalised signal across the 11 time points was less than 50. The signal data for the remaining 158 transcripts were log2 transformed, then normalised against their time 0 values to obtain log2 fold changes through the diurnal cycle. This matrix was analysed with the CAST algorithm of TMEV (http://www.tm4.org/mev.html) using Pearson correlation coefficient and a threshold of 0.8. The cluster containing SDRa is depicted. ACX2 did not cluster in this group, but is superimposed for comparison. (1418 kb)
Rights and permissions
About this article
Cite this article
Wiszniewski, A.A.G., Zhou, W., Smith, S.M. et al. Identification of two Arabidopsis genes encoding a peroxisomal oxidoreductase-like protein and an acyl-CoA synthetase-like protein that are required for responses to pro-auxins. Plant Mol Biol 69, 503–515 (2009). https://doi.org/10.1007/s11103-008-9431-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9431-4