Abstract
Leafy (LFY) and LFY-like genes control the initiation of floral meristems and regulate MADS-box genes in higher plants. The Cucumber-FLO-LFY (CFL) gene, a LFY homolog in Cucumis sativus L. is expressed in the primordia, floral primordia, and each whirl of floral organs during the early stage of flower development. In this study, functions of CFL in flower development were investigated by overexpressing the CFL gene in gloxinia (Sinningia speciosa). Our results show that constitutive CFL overexpression significantly promote early flowering without gibberellin (GA3) supplement, suggesting that CFL can serve functionally as a LFY homolog in gloxinia. Moreover, GA3 and abscisic acid (ABA) treatments could modulate the expression of MADS-box genes in opposite directions. GA3 resembles the overexpression of CFL in the expression of MADS-box genes and the regeneration of floral buds, but ABA inhibits the expression of MADS-box genes and flower development. These results suggest that CFL and downstream MADS-box genes involved in flower development are regulated by GA3 and ABA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There are more than 80 genes involved in the control of flowering time. Several models have been proposed to explain how these genes govern the establishment and maintenance of floral meristem identity (Chou and Yang 1998, 1999; Levy and Dean 1998; Parcy et al. 1998; Pineiro and Coupland 1998). During flower development in Arabidopsis, Leafy (LFY) has been found to play a pivotal role, interacting with and coordinating other flowering-related genes (Weigel and Meyerowitz 1993; Wagner et al. 1999, 2004; Sablowski 2007). LFY can directly target APETALA1 (AP1), AGAMOUS (AG), and APETALA3 (AP3) (Parcy et al. 1998; Busch et al. 1999; Wagner et al. 1999; Lamb et al. 2002). All these LFY-targeted genes belong to the MADS-box gene family of transcription factors, which play distinct roles in flower development (Wagner et al. 2004). LFY homologs have also been identified in other plant species such as apple, orange, morning glory, rice, tobacco, tomato and cucumber (Liu et al. 1999). Our previous studies showed that, Cucumber-FLO-LFY (CFL), a LFY-like gene from Cucumis sativus L., was strongly expressed in the primordia, floral organ primordia, and each whirl of floral organs at the early stage of floral formation, whereas weak expression was observed in floral organs at the late stage of flower development. In addition, the CFL gene was also expressed in the meristem of vegetative bud, leaf primordium and young leaves. However, it was not detectable in mature vegetative tissues (Wang et al. 2004). Thus, the CFL gene may play an important role in the differentiation and formation of both floral and vegetative primordia. This hypothesis, however, needs to be verified.
Flower development can be modulated by various plant hormones, such as gibberellin (GA3) and abscisic acid (ABA) (Martinez-Zapater et al. 1994; Blazquez and Weigel 2000). Recent reports show that GA3 promotes normal development of floral organs owing partly to up-regulation of the expression of floral homeotic genes AP3, PI and AG (Yu et al. 2004). Moreover, GA3 acts as a functional inhibitor of two DELLA proteins, RGA and RGL2 (Cheng et al. 2004; Yu et al. 2004). Therefore, GA3 signaling in flower development is likely to be involved in a regulatory network comprising floral homeotic genes and DELLA proteins (Yu et al. 2004). The effect of GA3 on LFY is mediated through a GA3-responsive element of LFY promoter with a binding motif similar to that of GA-myb. Mutation of this motif can result in the failure of up-regulating a minimal LFY promoter in short days other than long days (Blazquez and Weigel 2000). Mutants with reduced ABA biosynthesis can promote early flowering under non-inducing conditions in Arabidopsis, suggesting that ABA inhibits flowering (Martinez-Zapater et al. 1994). Auxin is present in flower primordia, and can promote outgrowth of flower buds (Benkova et al. 2003; Reinhardt et al. 2003). However, little is known about effect of ABA and auxin on LFY as well as its targeted genes.
Gloxinia is originated from tropical South American, and is widely grown as an ornamental flower plant in China. Gloxinia belongs to Gesneriaceae, a member of the Lamiales (Cubas 2004), and has several floral homeotic variants. This makes it popular as a houseplant (Pang et al. 2003). Gloxinia has a relatively long vegetative phase. Thus, early flowering is necessary for the flower-plant market. However, little is currently known about the molecular mechanism of flower morphogenesis in gloxinia. In the present study, we overexpress CFL in gloxinia to establish the function of CFL and investigate its potential to alter flowering time of gloxinia.
Materials and methods
Plant materials and growth conditions
Gloxinia plants were grown in pearl rock in a glasshouse at 24 ± 1°C, 16 h light/8 h darkness and the ambient humidity (≥85%). Transformation was performed as described by Koncz et al. (1989) and Bechtold et al. (1993). Flower samples were divided into two parts: one part was used immediately for tissue culture and the other was frozen in liquid nitrogen for RNA isolation.
Plasmid constructs
The coding sequence of the CFL gene was amplified and with forward primer (5′-CCGAGCTCTTGACAAGAGAGACTGAAAT-3′) and reverse primer (5′-GACCCGGGATGGATCCAGAAACCCTCTCC-3′), using plasmid pBS-CFL (courtesy of Wang Li-Lin) as a template. The PCR product of 1.2 kb was purified from gel and subcloned into the restriction sites SalI and SacI in the pCAMBIA13011. All cloning was performed using standard methods.
Agrobacterium-mediated transformation
Agrobacterium strain EHA105 containing pCAM13011/CFL was used in the transformation. Leaf pieces from in vitro-grown plants were co-cultivated with agrobacterium for 2 days, and then transferred to the hygromycin (20 mg l−1) selection medium. After two cycles of culture in the selection medium, hygromycin-resistant, green shoots were obtained for production of plantlets.
Southern blot
Genomic DNA was isolated from young leaves using the method described by Murray (Murray and Thompson 1980; Bousquet et al. 1990), digested with HindIII, fractionated on a 0.8% agarose gel, and transferred to a nylon membrane. The DNA was fixed to the membrane with UV light. The membrane was prehybridized for 2 h at 42°C in standard buffer with 50% formamide, and was then hybridized overnight with 25 ng probe ml−1. Probes were labeled and detected using DIG Random Labeling Mix and Detection Kit I according to the manufacturer’s instruction (Roche, Germany). After hybridization, the membrane was washed for 3 × 5 min with 2 × SSC + 0.1% SDS at room temperature, and then for 3 × 15 min with 0.5 × SSC + 0.1% SDS at 68°C. Finally, the hybridized DNA was prepared for color detection with DAB.
RNA isolation, reverse transcription and cloning of partial cDNAs
Total RNA was extracted with TRIZOL Reagent (Promega, USA) according to the manufacturer’s instructions. The total RNA was treated with DNase I (Promega, USA) at 37°C for 30 min before the first-strand synthesis. DNase was removed by TRIZOL extraction. RNA integrity was checked by electrophoresis, and the concentration and quality of RNA were also determined by OD260/230 and OD260/280. Then, one microgram of DNase-digested RNA was used as a template for the first-strand cDNA synthesis according to the manufacturer’s instructions (Promega, USA). The degenerated primers (see follow) which were designed according to sequences of Arabidopsis, tobacco and other homologous genes were used to amplify partial cDNA from gloxinia. The amplified fragments were isolated and cloned into T-easy™ (Promega, USA). At least two clones were selected for sequencing. The degenerate PCR primers used in the experiments are as follows: 5′-GCTCATGAGATCTCTGTT(G)CTC(T)TGTG-3′ and 5′-CTGC(G)TGCTCC(A)AGATTCTGAAG-3′ for SsAP1-1; 5′-CACAACGAATCGTCAAGTCAC-3′ and 5′-GCTCATTCTTC(T)TTGGATCG(T)G-3′ for SsAG-1; 5′-GAGATAAAAAGAATAGAGAACTCAAGC-3′ and 5′-CTCCTGAAGATTAGGCAGCATTGGC-3′ for SsAP3-1; 5′ CAATAAATTGCGTGTTGCTCCTGAG 3′ and 5′-TGTTTCCGTACCGATCCTTTCTGATA-3′ for Ssactin-1. The size of amplified cDNA fragments was 273 bp for SsAP1-1 (Genbank accession number: EF428184), 394 bp for SsAG-1 cDNA (Genbank accession number: EF428183, 599 bp for SsAP3-1 cDNA (Genbank accession number: EF428185), 571 bp for SsActin-1 (Genbank accession number: EF428182).
Flower buds and sepals culture in vitro
Flower buds of 5–6 mm in diameter which contained sepals but remained close were sterilized for 30 s in 70% ethanol and 5 min in 10% NaClO at room temperature. Then the flower buds were rinsed 4 times with sterile water and air dried in a laminar flow hood. Sepals were cut out from flower buds and cultured in flower-induction medium (FI) supplemented with 0.1 mg l−1 6-benzylaminopurine (6-BA). FI medium is a modified MS medium which contained different concentrations of Ca and other macronutrients (15 mM KNO3, 8 mM NH4NO3, 2 mM CaCl2, 1.5 mM KH2PO4, 1 mM MgSO4). Different concentrations of GA3 (0, 0.5, 1.0, 1.5 mg l−1, Sigma, USA), ABA (0, 10, 20 and 30 μM, Sigma, USA) and IAA (0, 0.2, 0.5 and 1.0 mg l−1, Sigma, USA) were added to FI medium separately to determine their effects on development of flower buds, floral regeneration from sepals and analysis of gene expression. All experiments were repeated at least 3 times.
Semi-quantitative RT-PCR analyses
PCR cycles for semi-quantitative RT-PCR were optimized for each of five specific primer pairs by amplifying a set of standards over a range of cycles. Standards were produced from dilutions of amplified cDNA. The yield of cDNA was measured according to the signal density of the SsACT1 PCR product with 0.1 μl of cDNA solution after 18–32 cycles of amplification on a Biometra PCR machine (MJ Research Inc., MA, USA). The concentration of each cDNA pool was adjusted to achieve the same exponential phase in the signal density of PCR products for SsACT1 after 25 cycles. The PCR products were electrophoresed on 1.5% agarose gels, stained with ethidium bromide and photographed. All PCR analyses were repeated at least 3 times to produce independent replication, and the amplification of a set of standards was also included in each PCR run.
Results
Generation of transgenic gloxinia lines overexpressing CFL
Twelve independent gloxinia lines constitutively expressing CFL were generated using agrobacterium-mediated transformation. The integration of CFL into the gloxinia nuclear genome was confirmed by Southern blot analysis (Fig. 1a). Analysis showed that the expression of CFL was detectable in all organs of the transgenic lines tested with reverse transcriptase polymerase chain reaction (RT-PCR) (Fig. 1b), but not in wild-type lines (data not shown).
Transgene analyses and comparison of flowering time between wild-type and transgenic CFL gloxinia plants. (a) Southern blot analysis of CFL in different transgenic lines. Total genomic DNA (30 μg) was digested with HindIII and hybridized with the CFL probe. Lane 1, wild type; lanes 2–5, four different transgenic lines. (b) RT-PCR analysis of CFL in different organs of transgenic gloxinia. Ib, inflorescence branches; Yl, young leaves; Se, sepals; Fb, flower buds (5 mm in size); Pe, petals and St, stamens. (c) Comparison of flowering time between wild-type and transgenic plants. Seedlings were grown to maturity under 16/8 h (day/night). The date of appearance of the first flower bud was recorded for total 96 transgenic plants (white columns) and 80 wild-type plants (black columns)
Morphological and developmental alterations in transgenic gloxinia
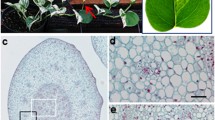
Transgenic seedlings were grown under long-day condition until maturity to determine flowering time and morphological characters. Although transgenic gloxinia plants were smaller in size, had fewer branches and were bushier in growth habit (Fig. 2a–d), most of transgenic gloxinia plants (71%) flowered 26–32 days earlier than wild-type plants (Fig. 1c). Up to 90% of the transgenic plants had terminal flowers emerging directly from shoot apex with no inflorescence branches while approximately 40% of wild-type plants generated more than one inflorescence branches (Fig. 2e). Importantly, approximately 30% of 35S::CFL gloxinia plants showed flower buds opening earlier than wild type (Fig. 3). Based on morphological characters, the development of flower buds was divided into four stages, 15 days after flower buds emerged as stage I (Fig. 3a, b), 15–25 days as stage II (Fig. 3c, d), 25–35 days as stage III (Fig. 3e, f) and 35–45 days as stage IV (Fig. 3g, h). As flower bud development progresses, grooves became wider, allowing sepal blades to spread out and flower buds to open completely (Fig. 3c–h).
Morphological and developmental alterations of transgenic gloxinia. (a) No flower buds appeared in wild-type plants after 90 days. (b) Terminal flower buds (white arrow) appeared in 35S::CFL transgenic gloxinia plants after 90 days. (c) Flower buds (white arrow) in the wild type appeared after 115 days. (d) Flowers of transgenic gloxinia opened in 115 days. (e) Comparison of numbers of inflorescence branches between wild-type and transgenic plants. Total 96 transgenic and 80 wild-type plants were grown to maturity under 16/8 h (day/night) for 125 days and inflorescence branches were counted for each plant
Comparison of flower bud development at four growth stages between wild-type and 35S::CFL gloxinia. The flower buds of wild-type were tightly close 15 days after the appearance of flower buds (a), but the flower buds of 35S::CFL gloxinia opened slightly (b). The flower buds of wild-type remained green in color and opened slightly 25 days after the appearance of flower buds (c), whereas the flower buds of 35S::CFL gloxinia plants were green but widely opened (d). The flower buds of wild-type opened slightly and the petals became pinkish-red 35 days after the appearance of flower buds (e), while the petals of 35S::CFL gloxinia flowers spread out with red color and pistils emerged from the centre of the flower buds (f). Some of the petals of wild-type spread 45 days after the appearance of flower buds (g), but all petals of 35S::CFL gloxinia spread out (h)
Effect of GA3 supplement on development of flower buds and regeneration of floral buds from sepals
Different GA3 concentrations were applied to the culture medium to investigate GA3 effects on the development of excised flower buds and the regeneration of floral buds from sepals in wild-type and 35S::CFL gloxinia lines. Results showed that the expansion of flower buds of 35S::CFL gloxinia occurred approximately 2 days earlier than wild type regardless of whether GA3 (1 mg l−1) was added (Fig. 4e, f) or not (Fig. 4a, b). The flower buds of both wild-type and transgenic plants became red in 8 days, but the expansion of the flower buds was observed only in 35S::CFL gloxinia (Fig. 4a, b).
Effects of three hormones on development of flower buds and regeneration of floral buds from sepals. The flower buds of wild-type expanded and became red after 8-day culture in the flower induction (FI) medium without either of three hormones, GA3, ABA or IAA (a), whereas the flower buds of 35S::CFL gloxinia enlarged and became pinkish-red (b). The sepals of wild-type turned brown and senesced after 17-day culture in FI medium (c), but floral buds were regenerated, and expanded from the sepals of 35S::CFL gloxinia (d). When 1 mg l−1 GA3 was added to FI medium, flower buds expanded and turned red in both wild-type (e) and 35S::CFL gloxinia (f) after 12-day culture. Floral buds were regenerated from the sepals in both wild-type (g) and 35S::CFL gloxinia (h) in the FI medium supplemented with 1 mg l−1 GA3 after 17 day culture. When 20 μM ABA was added to FI medium, the flower buds of wild-type exhibited senescence symptoms (i), but the flower buds of 35S::CFL gloxinia developed normally with red and bright color (j) in 12-day culture. Floral buds could not be regenerated from the sepals of either wild-type (k) or 35S::CFL gloxinia (l) in the FI medium supplemented with 20 μM ABA, and showed brown in colour in 17-day culture. When 0.5 mg l−1 IAA was added in FI medium, the flower buds of wild-type remained tightly closed in 8-day culture in FI medium (m), whereas the flower buds of 35S::CFL gloxinia enlarged and opened partly without appearance of red color (n). The sepals of wild-type showed brown in colour and senescenced in the FI medium supplemented with 0.5 mg l−1 IAA despite of the bottom half of the sepal remaining green after 20-day culture (o), whereas the floral buds were regenerated from the sepals of 35S::CFL gloxinia, and developed with red color in 20-day culture (p)
The effect of GA3 on the regeneration of floral buds from sepals in wild-type and 35S::CFL gloxinia lines was also studied. The term “floral buds” was used for flower buds regenerated from sepals in vitro to differentiate them from those developed in gloxinia plants. 35S::CFL gloxinia lines could regenerate floral buds directly from the surface of sepals after 8 days without GA3 supplement. The floral buds from the sepals expanded in 17-day culture with healthy appearance (Fig. 4d). By contrast, the sepals of wild-type could not regenerate floral buds without GA3 supplement and senesced in 17-day culture (Fig. 4c). GA3 supplement in the culture medium was essential for the regeneration of floral buds from the sepals of wild-type. The minimum concentration of GA3 was 1.0 mg l−1 or higher for a flower frequency equivalent to that of transgenic CFL (Figs. 5a, 4g, h).
Effect of ABA supplement on development of flower buds and regeneration of floral buds from sepals
Different ABA concentrations were applied to the culture medium to investigate ABA effects on the development of excised flower buds and the regeneration of floral buds from sepals in wild-type and 35S::CFL gloxinia lines. The ABA supplement in the culture medium led to the inhibition of both development of flower buds and regeneration of floral buds. Floral tissues in wild-type flowers lost turgor and became translucent and browning (Fig. 4i). By contrast, the flowers in 35S::CFL gloxinia lines turned red and bright although they could not open completely (Fig. 4j). The flower buds from both wild-type and 35S::CFL gloxinia lines eventually wilted. When sepals from either 35S::CFL or wild-type were cultured in the medium supplemented with 20 μM ABA, no floral buds could be regenerated (Fig. 4k, l).
Effect of IAA supplement on development of flower buds and regeneration of floral buds from sepals
IAA supplement was used to treat flower buds and sepals. The flower buds excised from both wild-type and transgenic gloxinia plants expanded and opened partly in the culture medium supplemented with IAA, but no red color appeared in the flower buds from both wild-type and 3S::CFL gloxinia (Fig. 4m, n). The regeneration of floral buds from the sepals was observed only in 35S::CFL gloxinia lines (Fig. 4o, p). The frequency of the regeneration of floral buds from the sepals in transgenic gloxinia was not affected by IAA (Fig. 5b).
Effect of overexpression of CFL on gloxinia MADS-box genes
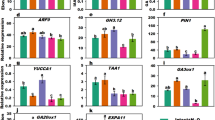
To reveal CFL functions in gloxinia, we isolated partial cDNA fragments of three CFL-targeted genes from gloxinia, using primers designed based on AP1, AP3, AG of Arabidopsis, respectively. These gloxinia cDNAs referred as to SsAP1-1, SsAP3-1 and SsAG-1, respectively shared a high similarity with their homologs in tobacco and Arabidopsis (supplemental Fig. 1). Semi-quantitative RT-PCR showed that these three genes were expressed in the reproductive organ of wild-type gloxinia with different spatial and temporal patterns (Fig. 6a). The expression of SsAP1-1, SsAP3-1 and SsAG-1 was determined in different stages of flower development ranging from 1 to 55 days after flower buds emerged. In wild-type, SsAP1-1 was expressed in young flower buds (day 1–15), but no expression was observed during later flower development day 35–55 (Fig. 6a). SsAP3-1 expression was basically restricted to day 5–15, which was related to the development of petals and stamens. SsAG-1 was expressed between day 5 and 45, which suggested that it was related to the development of stamens and carpels (day 25–35).
Transcript levels of SsAP1-1, SsAP3-1, SsAG−1 in flower buds and sepals treated with different hormones (a) Expression of SsAP1-1, SsAP3-1, SsAG-1 and SsActin-1 (control gene) in the flower buds of wild-type collected at different days. The flower buds and sepals of wild-type (W) and 35S::CFL gloxinia (T) were cultured in the FI medium supplemented with either 1.0 mg l−1 GA3 (c), 20 μM ABA (d) or 0.5 mg l−1 IAA (e). Total RNA was isolated from different flower tissues, and semi-quantitative RT-PCR were performed and repeated for three times. Representative results are presented
The expression of three gloxinia MADS-box genes as well as CFL was also determined in the flower buds and sepals of both wild-type and 35S::CFL gloxinia, which were treated with either GA3, ABA or IAA. CFL was expressed constitutively at a high level in transgenic flower buds and sepals as expected (Fig. 6b–e). When there was no hormone supplement in the culture media, the MADS-box genes (SsAP1-1, SsAP3-1 and SsAG-1) were highly expressed in the 35S::CFL gloxinia line compared to the wild type (Fig. 6b). The low transcript levels of three gloxinia MADS-box genes were observed in the wild-type, which may be due to the low expression of the endogenous gloxinia LFY gene (Fig. 6b–e). The treatment of GA3 (1 mg l−1) supplement gradually enhanced the expression of SsAP1-1 in the flower buds and sepals of wild-type, with its peak expression at day 8 and then declining from day 12–17 (Fig. 6c). However, the level of SsAP1-1 in the flower buds and sepals of 35S::CFL gloxinia was sustained regardless of whether GA3 was added or not (Fig. 6c). When ABA was added to the culture medium, SsAP1-1 expression in the wild type fell gradually to undetectable levels in both flower buds and sepals (Fig. 6d). SsAP1-1 expression was inhibited in the flower buds and sepals of 35S::CFL gloxinia, and the inhibition was more in wild-type (Fig. 6d). The effect of GA3 and ABA on the expression of SsAP3-1 and SsAG-1 in the flower buds and sepals was similar to SsAP1-1 (Fig. 6c, d). IAA had little impact on the expression of three gloxinia MADS-box genes in the flower buds and sepals of both wild-type and 35S::CFL gloxinia (Fig. 6e).
Discussion
Overexpression of CFL promotes early flowering in gloxinia
CFL was previously suggested to strongly express in primordia and floral primordia, implying its potential role in control of flower development (Wang et al. 2004). Here, we show in gloxinia plants that transgenic 35S::CFL can result in bushy growth habits as well as fewer branches, and promote early flowering (approximately 30 days earlier than that of wild type). This observation is quite similar to the constitutive expression of LFY in Arabidopsis (Weigel and Nilsson 1995). LFY mutants delay inflorescence development and prevent flower production in Arabidopsis (Schultz and Haughn 1991; Huala and Sussex 1992; Weigel et al. 1992). Constitutive expression of LFY driven by the CaMV35S promoter attenuates development of both vegetative and inflorescence phases, leading to the production of a terminal flower in the primary shoot of Arabidopsis (Weigel and Nilsson 1995). Our results imply that CFL acts as a functional homolog of LFY in gloxinia despite being derived from a species which is distant from gloxinia in the phylogenetic tree (Soltis et al. 1999). Cucumber belongs to the order of Cucurbitales, but gloxinia is in the order of Lamiales (Soltis et al. 1999). Our results also provide evidence for a broad functional conservation of LFY-like genes and the regulatory MADS-box genes downstream of LFY in dicotyledonous plants (Fig. 6).
Similar effects of both overexpression of CFL and GA3 supplement on flower development in gloxinia
Floral buds were regenerated directly from the sepals of 35S::CFL gloxinia (Fig. 4d), but not the wild type (Fig. 4c), indicating that the overexpression of CFL is sufficient for the formation of floral buds for the sepals. Interestingly, GA3 supplement can also lead to the regeneration of floral buds from the sepals of the wild type (Fig. 4g), indicating that GA3 treatment and CFL overexpression has a similar effect on the regeneration of floral organs from sepals. In Arabidopsis, LFY expression with a constitutive promoter induces flowering in a ga1-3 mutant background (Blazquez et al. 1998), implying that LFY acts downstream of GA3 signaling pathway. LFY plays a role as the central depot in flower morphogenesis, which positively or negatively regulates levels or activities of several flowering-related genes (Parcy et al. 1998; Busch et al. 1999; Wagner et al. 1999; Lamb et al. 2002). Morphological alterations observed in flower development are not solely due to the CFL gene, but also other genes such as MADS-box genes. Our data imply that GA3 is by and large as effective as CFL overexpression in the induction of MADS-box genes such as SsAP1-1 in the wild type (Fig. 6b, c). The overexpression of CFL in gloxinia up-regulates SsAP1-1, SsAP3-1 and SsAG-1 (Fig. 6b), and therefore leads to the formation of floral organs and early flowering, suggesting that CFL promotes flowering by the initiation of floral organs via the interactions of LFY and MADS-box genes.
ABA inhibits the expression of MADS-box genes
The interplay in flower development between GA and the expression of LFY as well as MADS-box genes has been described in Arabidopsis (Cheng et al. 2004; Yu et al. 2004), but the effects of other hormones such as ABA and IAA and relevant gene expression on flower development are still unclear. We investigated the effects of ABA and IAA on flower development, and the expression of SsAP1-1, SsAP3-1 and SsAG-1 during flower development and regeneration of floral buds from sepals. ABA inhibited the expression of three gloxinia MADS-box genes in the flower buds of both wild-type and 35S::CFL, but had a little effect on the expression of CFL (Fig. 6d). Consequently, ABA induced senescence of flower buds, while it inhibited the expansion of flowers and the regeneration of floral buds from sepals (Fig. 4i–l). The results suggest that the inhibition of ABA on flower development may be partly due to its down-regulation of three gloxinia MADS-box genes. IAA has been shown to be active in flower primordia at the early stage of flower buds and promote outgrowth of flower buds (Benkova et al. 2003; Reinhardt et al. 2003). However, our results showed that IAA had a little effect on the expression CFL and three MADS-box genes (Fig. 6e) and the regeneration of floral buds from the sepals of wild-type (Figs. 4o, 5b), but the inhibition of formation of red color in the flower buds (Fig. 4m, n). These results suggest that IAA is unlikely to be directly involved in the regulation of LFY homologs and their downstream genes in flower initiation.
Abbreviations
- cucumber:
-
Cucumis sativus L.
- LFY :
-
Leafy
- FI:
-
Flower-induced
- CFL :
-
Cucumber-FLO-LFY
- gloxinia:
-
Sinningia speciosa
References
Bechtold N, Ellis J, Pelletier G (1993) In planta agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C R Acad Sci Paris Life Sci 316:1194–1199
Benkova E, Michniewicz M, Sauer M, Teichmann T, Seifertova D, Jurgens G, Friml J (2003) Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115:591–602
Blazquez MA, Weigel D (2000) Integration of floral inductive signals in Arabidopsis. Nature 404:889–892
Blazquez MA, Green R, Nilsson O, Sussman MR, Weigel D (1998) Gibberellins promote flowering of Arabidopsis by activating the LEAFY promoter. Plant Cell 10:791–800
Bousquet J, Simmon L, Lalonde M (1990) DNA amplification from vegetative and sexual tissues of trees using polymerase chain reaction. Can J For Res 20:254–257
Busch MA, Bomblies K, Weigel D (1999) Activation of a floral homeotic gene in Arabidopsis. Science 285:585–587
Cheng H, Qin L, Lee S, Fu X, Richards DE, Cao D, Luo D, Harberd NP, Peng J (2004) Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 131:1055–1064
Chou ML, Yang CH (1998) FLD interacts with genes that affect different developmental phase transitions to regulate Arabidopsis shoot development. Plant J 15:231–242
Chou ML, Yang CH (1999) Late-flowering genes interact with early-flowering genes to regulate flowering time in Arabidopsis thaliana. Plant Cell Physiol 40:702–708
Cubas P (2004) Floral zygomorphy, the recurring evolution of a successful trait. Bioessays 26:1175–1184
Huala E, Sussex IM (1992) LEAFY interacts with floral homeotic genes to regulate Arabidopsis floral development. Plant Cell 4:901–913
Koncz C, Martini N, Mayerhofer R, Koncz-Kalman Z (1989) High frequency T-DNA-mediated gene tagging in plants. Proc Natl Acad Sci USA 86:8467–8471
Lamb RS, Hill TA, Tan QK, Irish VF (2002) Regulation of APETALA3 floral homeotic gene expression by meristem identity genes. Development 129:2079–2086
Levy YY, Dean C (1998) The transition to flowering. Plant Cell 10:1973–1989
Liu FQ, Zhu GL, Luo D, Wu XY, Xu ZH (1999) Cloning and analysis of CFL: a LFY-like gene from cucumber. Acta Bot Sin 41:813–819
Martinez-Zapater JM, Coupland G, Dean C, Koornneef M (1994) The transition to flowering. In: Meyerowitz EM, Somerville CR (eds) Arabidopsis. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp 403–433
Murray GC, Thompson WF (1980) Rapid isolation of high molecular weight DNA. Nucleic Acids Res 8:4321–4325
Pang JL, Wang LL, Hu JQ, Liang HM, Li WA (2003) Floral homeotic variants of in vitro seedlings in Sinningia speciosa Hiern. J Mol Cell Biol 1:76–80
Parcy F, Nilsson O, Busch MA, Weigel D (1998) A genetic framework for floral patterning. Nature 395:561–566
Reinhardt D, Pesce ER, Stieger P, Mandel T, Baltensperger K, Bennett M, Traas J, Friml J, Kuhlemeier C (2003) Regulation of phyllotaxis by polar auxin transport. Nature 426:255–260
Pineiro M, Coupland G (1998) The control of flowering time and floral identity in Arabidopsis. Plant Physiol 117:1–8
Sablowski R (2007) Flowering and determinacy in Arabidopsis. J Exp Bot 58:899–907
Schultz EA, Haughn GW (1991) LEAFY, a homeotic gene that regulates inflorescence development in Arabidopsis. Plant Cell 3:771–781
Soltis PS, Soltis DE, Chase MW (1999) Angiosperm phylogeny inferred from multiple genes as a tool for comparative biology. Nature 402:402–404
Wagner D, Sablowski RW, Meyerowitz EM (1999) Transcriptional activation of APETALA1 by LEAFY. Science 285:582–584
Wagner D, Wellmer F, Dillks K, William D, Smith MR, Kumar PP, Riechmann JL, Greenland AJ, Meyerowitz EM (2004) Floral induction in tissue culture: a system for the analysis of LEAFY-dependent gene regulation. Plant J 39:273–282
Wang LL, Pang JL, Liang HM, Zhu MY (2004) Expression of CFL gene during differentiation of floral and vegetative buds in cucumber cotyledonary nodes cultured in vitro. J Plant Physiol Mol Biol 30:644–650
Weigel D, Meyerowitz EM (1993) Activation of floral homeotic genes in Arabidopsis. Science 261:1723–1726
Weigel D, Nilsson O (1995) A development switch for flower initiation in diverse plants. Nature 377:495–500
Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM (1992) LEAFY controls floral meristem identity in Arabidopsis. Cell 69:843–859
Yu H, Ito T, Zhao Y, Peng J, Kumar P, Meyerowitz EM (2004) Floral homeotic genes are targets of gibberellin signaling in flower development. Proc Natl Acad Sci USA 101:7827–7832
Acknowledgments
We sincerely thank CY Huang for critical reading of this manuscript, and CR Leach for English corrections. This research was supported by National Science Foundation of China (No. 30771347, 30300017, 39770420, 30671059), National Science Foundation of Zhejiang Province (300030, Y307116, Y304128) and Program for Science and Technology from Zhejiang Province (No. 2005C22G2010075, 2004C32015).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, MZ., Ye, D., Wang, LL. et al. Overexpression of the cucumber LEAFY homolog CFL and hormone treatments alter flower development in gloxinia (Sinningia speciosa). Plant Mol Biol 67, 419–427 (2008). https://doi.org/10.1007/s11103-008-9330-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9330-8