Abstract
Maize PBF (prolamin-box binding factor) belongs to the Dof class of plant specific transcription factors containing one highly conserved zinc finger DNA-binding domain, called Dof (DNA binding with one finger) domain. Maize PBF trans-activates the γ-zein gene (γZ) promoter in developing maize seeds as shown by transient expression in maize endosperms. Co-transfection of a γZ:GUS construct with 35S:PBF resulted in a sevenfold increase in GUS expression, however, PBF mutation in Cys residues within the Dof domain abolishes both, binding to DNA and the capacity to activate γZ promoter. We present two pieces of evidence that PBF transactivates γZ promoter by binding to the Pb3 motif (TGTAAAG). First, recombinant Dof domain of PBF (bdPBF) specifically recognized Pb3 site as shown by gel mobility shift assays and second, co-expression of PBF with γZ promoter mutated in Pb3 motif suppressed PBF trans-activation capacity. Immunocytochemical analysis on developing endosperm sections shows that PBF is localized in the nuclei of the peripheral layer cells of starchy endosperm, the tissue in which the initial accumulation of γ-zein protein occurs. By contrast, PBF is detected in the cytosol of the starchy endosperm cells newly differentiated from aleurone daughter cells, where γ-zein was absent. Taken together these data indicate that maize PBF plays an essential role in the regulation of the temporal and spatial expression of γZ gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prolamins, the most abundant cereal seed storage proteins, accumulate in a spatial and developmental stage-specific pattern in the highly specialized endosperm tissue. The expression of prolamins, then, appears to be highly regulated and several cis-motifs present in their promoters as well trans-acting transcription factors that interact with them have been described. The best characterized cis-consensus motifs are the bifactorial endosperm box (Kreis et al. 1985) and the 5′ AACA 3′ motif. The 5′ AACA 3′ motif is the binding site of OsMYB5 transcription factor in rice (Wu et al. 1998) and HvGAMYB in barley (Díaz et al. 2002; Wu et al. 1998). The bifactorial endosperm box is largely conserved along cereal prolamin genes such as those for zeins in maize (Marzábal et al. 1998; Schmidt et al. 1992), hordeins in barley (Entwistle et al. 1991; Forde et al. 1985), secalins in rye (Hull et al. 1991) or gliadins (Sumner-Smith et al. 1985) and glutenins (Hammond-Kosack et al. 1993) in wheat. The endosperm box is composed of two motifs separated by few nucleotides: the 7 bp prolamin box (P-box: 5′ TG (T/C/A)AAAG 3′) motif (Boronat et al. 1986; Ottoboni et al. 1993; Thompson and Larkins 1989) and the GCN4-like motif (5′ G(A)TGA(G)GTCAT 3′), which shares homology with yeast GCN4 (Hill et al. 1986) and mammalian AP1 (Piette et al. 1988) binding sites. The GCN4-like sequence is bound by basic leucine zipper (bZIP) class of transcription factors, the best characterized on regulation of prolamin expression belonging to the Opaque2 (O2) subfamily (Albani et al. 1997; Oñate et al. 1999; Schmidt et al. 1990; Wu et al. 1998), and the P-box motif is recognized by the Dof transcription factors called PBFs (Mena et al. 1998; Vicente-Carbajosa et al. 1997; Yanagisawa 2004).
Dof proteins are plant-specific transcription factors containing a highly conserved DNA-binding domain, the Dof domain [reviews in Yanagisawa (2002, 2004)], which includes one single C2–C2 zinc finger (Umemura et al. 2004), hence the name Dof (DNA-binding with one finger). In previous years, it was reported that numerous Dof proteins were involved in diverse plant processes such as seed germination (Isabel-LaMoneda et al. 2003; Mena et al. 2002; Papi et al. 2000; Washio 2001), photosynthesis (Yanagisawa 2000; Yanagisawa and Sheen 1998), secondary metabolism (Skirycz et al. 2006), phytohormone expression (Baumann et al. 1999) and seed storage protein expression (Díaz et al. 2005; Mena et al. 1998) in both monocots and dicots. The assumption that Dof proteins are plant specific was verified in a comparative study of Arabidopsis, Drosophila, yeast and Caenorhabditis elegans genomes (Riechmann et al. 2000). A recent phylogenetic analysis of Dof sequences allowed the identification of six major clusters of orthologs and seven subfamilies from green algae to plants (Moreno-Risueño et al. 2007). These proteins usually contain two principal domains: the N-terminal conserved DNA-binding Dof domain (Yanagisawa 1995, 2002) and the variable C-terminal domain, which can include activation domains (Díaz et al. 2002; Kang and Singh 2000; Yanagisawa 2002). Like other zinc fingers (Mackay and Crossley 1998), Dof proteins are also involved in protein-protein interactions. OBP1 and OBP2 are members of the Arabidopsis Dof transcription factor family (Kang and Singh 2000; Zhang et al. 1995) that interact with OBF bZIP transcription factors, and are able to significantly stimulate the binding of OBF4 protein to octopine synthase (ocs) elements characteristic of pathogen-responsive promoters.
In cereals, the maize prolamin-box binding factor, PBF, is able to interact in vitro with a prolamin-box motif present in the 22-kDA α-zein gene promoter and the maize bZIP protein O2 (Vicente-Carbajosa et al. 1997). In vivo maize PBF activity was demonstrated by transient expression assays in an heterologous system, developing rice endosperms (Hwang et al. 2004). PBF is a 36 kD protein that contains a single copy of the Dof domain in its N-terminal region linked by a stretch of serines to a C-terminal domain. Maize PBF orthologs have been isolated from wheat (WPBF) and barley (BPBF) and have been shown to be transcriptional activators of endosperm gene expression in wheat, barley and coix (de Souza Filho et al. 1999; Mena et al. 1998; Oñate et al. 1999). However, it seems that WPBF transcriptional activity is not specific of storage protein genes since it was not only expressed in seeds but also in other wheat tissues (Dong et al. 2007). Recently, a rice PBF ortholog (RPBF) has been reported to activate endosperm rice genes in cooperation with the transcription factor RISBZ1, which recognizes the GCN4-motif observed in many storage protein genes (Yamamoto et al. 2006).
It has also been described that Dof proteins interact with transcriptional factors of the GAMYB family. For instance, BPBF interacts with HvGAMYB through its C-terminal domain to activate barley endosperm specific genes during seed development (Díaz et al. 2002). SAD, another Dof protein from barley, also interacts in vivo through its C-terminal domain with HvGAMYB. This interaction enhances the transcriptional activation of a gibberellin induced protease promoter in aleurone cells (Díaz et al. 2005; Isabel-LaMoneda et al. 2003). Thus, Dof proteins not only bind to DNA cis-elements but also mediate protein-protein interactions through its Dof domain or C-terminal domain. This dual activity of Dof proteins could be essential to understand how each Dof protein can regulate the correct target in vivo. It is likely that this dual function participates in the correct choice as well as in selective activation or repression of target genes in vivo (Yanagisawa 2002).
Maize endosperm cells express a large family of zein genes. γ-zein gene (γZ), highly regulated during the endosperm development, is present in only one or two copies in the maize genome (Boronat et al. 1986). The γZ gene product, (γ-zein) is a 27 kD protein that represents as much as 15% of the total endosperm protein content. Several cis-elements have been identified on the proximal γZ gene promoter (Marzábal et al. 1998) that include four possible prolamin-boxes (Pb1, Pb2, Pb3 and Pb4), one GCN4-like motif (GZM) and a 5′ AACA 3′ motif. The functional role of the prolamin-box Pb1 was established by Ueda et al. (1994) by using transient expression in protoplasts isolated from cultured endosperm cells. The role of the bifactorial endosperm box containing the prolamin-box Pb3 and the GCN4-like motifs in γZ gene regulation was analyzed by using transient expression in the homologous maize endosperm tissue (Marzábal et al. 1998). Both prolamin-box and GCN4-like motif were shown to be necessary and act co-ordinately for γZ gene regulation. Protein regulators of γZ gene transcription have not been isolated so far. While α-zeins are regulated by O2 (Schmidt et al. 1990, 1992) it seems that this bZIP factor is not involved in γ-zein gene expression, since it is not affected in opaque2 maize mutants (Geetha et al. 1991; Motto et al. 1989).
In this report, in an effort to elucidate functional transcriptional factors involved in γZ gene expression, we explore the role of maize Dof protein PBF by using transient expression in maize developing endosperms. PBF is shown to be a transcriptional activator of γZ gene. Mutation analysis of prolamin-box domains showed that gene activation is the result of Dof domain binding to γZ prolamin-box sequences and that the C-terminal domain is also necessary for this activation. Moreover, we confirm that PBF up-regulates γZ gene during endosperm development and localizes in nuclei of transcriptionally active sub-aleurone cells.
Materials and methods
Plasmid constructs
Vector pET28a (Novagen) was used to express recombinant PBF derived proteins fused to His-tag in E. coli. The maize PBF cDNA and its Cys65 (Cys → Ala) mutated version (hereafter mPBF) were cloned in the pGEX-2TK vector (Pharmacia). The coding regions of PBF and mPBF from pGEX derived constructs were cloned as BamHI–NcoI fragments into pET28a vector and the new vectors were named pET28A-PBF and pET28A-mPBF. The N-terminal domains of PBF and mPBF (residues 1 to 105) were obtained through PCR amplification by using 5′GTAATACGACTCACTATAGGG3′ and 5′GCTCAATTGCTAGAGCTAGC3′ as primers (annealing regions are underlined) and were also cloned into pET28a vector by BamHI–SacI to obtain the new constructs pET-bdPBF and pET-mbdPBF. These N-terminal fragments are hereafter referred as bdPBF (from PBF binding domain) and mbdPBF. pGEX-CtPBF was obtained by the amplification of the C-terminal domain of maize PBF by PCR using 5′GGATCCATGAGCATCAACAAACATATG3′ and 5′CTCGAGTCATTATTGTCCCTTGTTGTT3′ as primers (annealing regions are underlined) over the construct pET28A-PBF and cloning into pGEX-4T3 vector restricted with BamHI–XhoI.
For endosperm transient expression experiments we used 444γZ and 527γZ constructs containing the proximal region (from −444 or −527 bp, respectively, to +61 bp of the transcriptional start) of γZ promoter fused to the GUS reporter gene and the nos 3′ region (Tnos) of the pBI 101.1 vector (Jefferson et al. 1987). These constructs as well as the corresponding mutated versions in Pb and GZM boxes were obtained as previously described (Marzábal et al. 1998). Constructs containing maize PBF and barley PBF DNA sequences were obtained by cloning PBF sequences into the expression vector PMF6 under the control of the CaMV 35S constitutive promoter, the first intron of maize AdhI gene and the nos 3′ regions as previously described (Mena et al. 1998). PMF6-PBF, PMF6-mPBF and PMF6-O2 constructs, containing respectively the cDNA sequences of maize PBF, its Cys65 (Cys → Ala) mutated version or the maize opaque 2 (O2) sequence. PMF6-bdPBF contained the N-terminal domain of maize PBF and was obtained by cloning the BamHI–SacI fragment from pET-bdPBF into PMF6 vector restricted by the same enzymes. PMF6-CtPBF was obtained by BamHI-XhoI restriction over pGEX-Ct-PBF and was introduced into PMF6 vector restricted with the same enzymes. PMF6-bdofPBF contained the Dof domain of barley PBF and was obtained after PCR amplification using 5′CCCGGGATGGAGGAAGTGTTTTCGTC3′ and 5′GGATCCAGGGCGTTTGGGCTTGCG3′ as primers (annealing regions are underlined) over the construct pGBT9-DofBPBF (kindly provided by Dr. Vicente-Carbajosa) and cloning into PMF6 vector restricted with BamHI. PMF6-ChPBF contained the Dof domain of barley PBF fused to the C-terminal domain of maize PBF and was obtained by cloning the BamHI restriction fragment from pET28A-PBF, containing the PBF C-terminal domain, into the PMF6-bdofPBF plasmid. Lastly, the plasmid pCaMV35SLUC, used as an internal control in transient expression experiments, was obtained as previously described (Torrent et al. 1997).
Microprojectile bombardment and enzyme assays
Developing maize seeds (from 8 to 30 days after pollination (DAP)) of the inbred line W64A were sterilized. The embryo, the pericarp and the aleurone cell layer were removed from seeds under a low-powered microscope using special forceps (Dumoxel 4, Dumont). Tangential sections of endosperms were cut in order to expose a large surface of subaleurone endosperm cells. After dissection, six to nine endosperms were placed in sterile Petri dishes on filter papers soaked in Murashige and Skoog medium (Duchefa). Endosperms were transformed by particle bombardment as previously described (Torrent et al. 1997) using 2 μg of 444γZ:GUS or 527γZ:GUS derived constructs, and variable amounts of effector constructs (from 0.5 to 2 μg). The endosperms were co-transformed with 1 μg of the pCaMV35SLUC construct used as internal standard (Torrent et al. 1997). All of the samples were bombarded twice and subsequently incubated for 24 h at 26°C in the dark before freezing in liquid nitrogen.
For quantitative enzyme assays, samples were thawed and homogenized on ice in a buffer containing 25 mM Tris, pH 7.8, 2 mM CDTA, 2 mM DTT, 10% glycerol and 1% Triton X-100. After centrifugation at 12,000g for 5 min at 4°C, the supernatants were decanted and the total soluble protein in the extracts was quantified using the Bradford Assay (BioRad). Around 60 to 100 μg of protein extract were used in luciferase (LUC) and GUS assays. LUC activity was determined by luminometry, using the Luciferase Assay System Kit (Promega) and GUS activity was determined using the chemiluminescent assay GUS-LightTM Kit (Tropix), according to the manufacturer’s instructions. For cytological localization of 444γZ:GUS expression, transversal sections of 15 DAP maize kernels were bombarded with 2 μg of 444γZ:GUS as described (Torrent et al. 1997) and GUS activity was detected by hystochemistry using X-Gluc (Duchefa) as substrate essentially as described (Jefferson et al. 1987).
Eletrophoretic mobility shift assays
DNA oligonucleotides used in electrophoretic mobility assays (EMSAs) are described in Fig. 2a. The full-length oligonucleotides were purified as previously described (Busk et al. 1997), and complementary oligonucleotides were annealed. The double-stranded oligonucleotides were labeled with α-32P-dCTP (3000 Ci mmol−1, Amersham) by filling in with the Klenow fragment of DNA polymerase, and purified on NAP5 columns (Pharmacia). The radioactive probe (25,000 cpm) was incubated with different recombinant proteins in binding buffer (25 mM HEPES, pH 7.8, 75 mM KCl, 10% glycerol, 500 μM EDTA, 5 mM MgCl2, 450 μM DTT, 800 ng Poly(dI-dC) (Sigma) and 25 μg BSA (Sigma) for 20 min at room temperature as previously described by Marzábal et al. (1998). The recombinant proteins (PBF, bdPBF, mbdPBF and Ct-PBF) were produced either by in vitro transcription and translation using the kit from Promega (TnT) as indicated by the manufacturers, or by overexpression of transformed BL21 E. coli cells with the constructs described above. Proteins produced in E. coli were induced with 1 mM IPTG. After 2 h of induction, the recombinant proteins were purified by Ni-affinity columns as described by the manufacturers (Amersham). The DNA-protein complexes were resolved on 4% polyacrylamide native gels in 0.5× Tris-borate-EDTA (TBE). Electrophoresis was performed at 10 V cm−1 for 3 h at 4°C. In supershift assays, the protein tested was pre-incubated in binding buffer for 10 min at room temperature with either the anti T7-tag antibody (Novagen) or the PBF antiserum. Non-immune serum was used as a control.
Antibodies and immunocytochemistry
Polyclonal antibodies against PBF were raised in rabbits injected with purified recombinant His-PBF. The antiserum was purified against recombinant bdPBF by affinity chromatography, using Hi-Trap NHS-activated columns (Amersham Biosciences) and following the manufacturer’s instructions. Anti-γ-zein antibodies (anti-G2) were raised in rabbits as described (Ludevid et al. 1985). For immunolocalization, 15 DAP maize seeds (Zea mays) of pure inbred line W64A were fixed for 1 hour at room temperature with 2% paraformaldehyde and 1% glutaraldehyde in PBS pH 7.4. After the dehydration process (Langdale 1994), samples were embedded in paraffin. Sections (8 μm thick) of paraffin-embedded samples were blocked with 0,5% BSA and 5% Normal Goat Serum (NGS, Gibco) in PBS pH 7.4 and incubated either with purified anti-PBF or anti-G2 (Ludevid et al. 1985) antibodies (dilutions 1:10 and 1:300, respectively). For immunodetection the samples were incubated with Biotin conjugated goat-antirabbit antibody (Jackson ImmunoResearch, 1:500 dilution) and stained with either Streptavidin-X-Rodamin or Streptavidin-X-Oregon-green 488 (Jackson ImmunoResearch, 1:100 dilution). Negative controls were performed by using non-immune sera. When indicated, the samples were stained with the fluorescent dye DAPI (Sigma-Aldrich), and mounted on Mowiol (Calbiochem). Samples were observed in a Zeiss AxioPhot microscope and in a Leica TCS SP confocal laser scanning microscope (Heidelberg, Germany).
Total and nuclear protein extracts and immunoblot
Protein extracts were obtained from around 50 maize seeds at different stages of development (8 to 25 DAP) dissected in order to remove pericarp, embryo and the central endosperm tissue. After grinding the endosperms in frozen nitrogen using a mortar, the fine powder was extracted with 70% ethanol in order to remove α-zeins. Nuclear extracts from 8 to 25 DAP endosperms were carried out as described (Marzabal et al. 1998). Samples were processed and analyzed by SDS-PAGE and immunoblot as previously described (Torrent et al. 1997). For immunoblot, nitrocellulose sheets were incubated o/n at 4°C with anti-PBF (1:200 dilution) purified antibody or 1 h at room temperature with anti-G2 (1:10.000 dilution) antiserum. Immunoreactive bands were detected by chemiluminiscence (ECL, Amersham).
Results
PBF activates the γ-zein promoter through Pb1 and Pb3 motifs
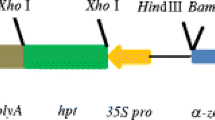
In order to study in vivo the activity of maize PBF (mPBF) over the γZ promoter in the homologous tissue, transient transformation assays were performed on 15 DAP maize endosperms by biolistic methods. Maize endosperms were co-transformed with the proximal γZ promoter (444γZ) fused to the GUS reporter gene (Marzábal et al. 1998) and the effector 35S:PBF (Fig. 1a) at 1:3 molar ratio. As it can be seen in Fig. 1b, as much as sevenfolds of activation were observed in co-transformation experiments when compared to the basal GUS activity determined after co-transformation using a control vector lacking the PBF transcription factor cDNA (Fig. 1b, lane c). In the 444γZ proximal promoter used in these transient assays, two putative functional Pbs have been described: Pb1 and Pb3 (Marzábal et al. 1998). To determine whether PBF activation could be the result of its binding to these cis-elements, GUS reporter constructs were generated containing site directed mutations at the 444γZ promoter either in Pb3 (Fig. 1a, M1), Pb1 (Fig. 1a, M2) or in both elements (Fig. 1a, M3). The results obtained after transformation (Fig. 1c) showed that the promoters bearing mutations in Pb1 and Pb3 (lanes M2 and M1) exhibit a significant lower GUS activity as compared with the wild type construct (wt), indicating that the transactivation capacity of PBF is accomplished by these two cis-elements. Nevertheless, it should be noted that only the mutation in Pb3 was able to completely abolish the PBF transcriptional activation of the 444γZ promoter (Fig. 1c, lane M1). As expected, when both Pb1 and Pb3 were mutated in the same construct (Fig 1a, M3) no any effect of the effector 35S:PBF was observed (Fig. 1c, M3).
PBF effect on the expression of the proximal γZ promoter on transiently transformed 15 DAP maize endosperms. (a) Schematic representation of wild type (WT) and mutated (M1, M2, M3 and M4) 444γZ:GUS constructs used as reporter genes and 35S:PBF and 35S:O2 constructs used as effectors. The mutated cis-elements Pb1, GZM and Pb3 are represented by the empty boxes indicated by arrowheads. (b) Relative activation of the expression of 444γZ:GUS (wt) in maize endosperms co-transformed with the effectors 35S:PBF, 35S:O2 or the corresponding empty vector (c) used as control. (c) Relative transcriptional activity of wild type (WT) and mutated (M1, M2, M3 and M4) 444γZ:GUS constructs after co-transformation with the 35S:PBF vector. In all of the cases the mean value of at least three independent experiments is included and the error bars indicate the standard deviation from the mean
These results show that Pb3 plays an essential role in the trans-activation of the 444γZ promoter by PBF. The GZM motif is located at 7 bp downstream of Pb3 of γZ promoter. In order to test the likely existing functional interaction between these two cis-elements, a new mutant affecting the GZM box was produced (Fig. 1a, M4). The co-expression of M4 and 35S:PBF resulted in a reduced PBF trans-activation capacity (Fig. 1c, M4) suggesting that the presence of a functional GZM element in the γZ promoter is important for PBF activity. Lastly, since the GZM motif is a bZIP binding cis-element, we analyzed whether O2 could have some functional relevance in γZ expression. As it can be seen in Fig. 1b, the 35S:O2 effector promotes more than threefolds of activation of the 444γZ proximal promoter.
The Dof domain of PBF interacts specifically with the Pb3 motif
γZ activity assays indicate that maize PBF is able to activate the γZ promoter through the Pb3. To study whether PBF is able to interact directly and specifically with this cis-element, the PBF protein synthesized by in vitro transcription and translation method was used in EMSA experiments using probes containing both elements, Pb3 and GZM (Fig. 2a). Additional EMSA experiments were performed with the N-terminal half of PBF containing the Dof domain, hereafter referred to as PBF binding domain (bdPBF), as well as the C-terminal half of PBF (Ct-PBF). The incubation of the Ct-PBF with the bifactorial box probe (Fig. 2a, oBB) did not produce a shift (Fig. 2b, lane 3), but the bdPBF was able to interact with the oBB probe and a clear shift was observed (Fig. 2b, lane 5, arrow). In addition, a mutated bdPBF protein (m-bdPBF) was produced where one Cys of the zinc finger was replaced by Ala. The disruption of the Zn finger abolished the interaction capacity of bdPBF with the oBB probe (Fig. 2b, lane 4), indicating that the Zinc finger was essential for the protein-DNA interaction.
The PBF binding domain (bdPBF) interacts in vitro with the prolamin box Pb3 motif. (a) Oligonucleotides oBB, oBBΔPb3 and oBBΔGZM used in the electrophoretic mobility shift assays (EMSA) showing the bifactorial box motifs Pb3 and GZM in boxes. The nucleotide substitutions in the oligonucleotides oBBΔPb3 and oBBΔGZM, showing respectively the Pb3 and GZM mutated cis-elements, are in lower case and indicated by arrows. (b) EMSA assays with the oBB probe using PBF domains produced by in vitro transcription and translation. The specific interaction of oBB probe with bdPBF (lane 5) is indicated by an arrow, and the unspecific binding to some protein present in reticulocyte extracts is indicated by an arrowhead (lanes 2–5). (c) EMSA assays with bdPBF produced in E. coli. The specific interactions of bdPBF and the intact Pb3 (lanes 2, 4 and 7) are indicated by a black arrow. The supershift observed after pre-incubation of bdPBF with an anti-T7 antibody is indicated by an empty arrow (lane 6). The arrowhead shows the unspecific interaction of some protein present in the rabbit serum with the oBB probe (lanes 7, 8 and 9). Negative controls: probe incubated with the binding buffer (lanes 1 in b and c) or with the transcription and translation reaction performed with an empty vector (lane 2 in b). Ct-PBF, C-terminal domain of PBF; bdPBF, PBF N-terminal half including the Dof binding domain; m-bdPBF, PBF binding domain mutated at Cys65 residue. PreI: Rabbit pre-immune serum; Im: anti-PBF antibody
In order to further analyze the specificity of the interaction of bdPBF with its site in the bifactorial box, bdPBF was produced in E. coli. The incubation of recombinant bdPBF with the oBB probe produced a clear shift (Fig. 2c, lane 2). Nevertheless, when recombinant bdPBF was incubated with a mutated oBB probe within the Pb3 motif (Fig. 2a, oBBΔPb3), the interaction was completely abolished (Fig. 2c, lane 3). In contrast, mutations within the GZM motif did not disrupt the interaction of bdPBF with Pb3 site (Fig. 2c, lane 4). As a control, band shift experiments using m-bdPBF, which contained a non functional Dof domain, were also carried out, and no shift was detected (Fig. 2c, lane 5).
To demonstrate that bdPBF promoted the shift observed, bdPBF was preincubated with a T7 monoclonal antibody. A supershift corresponding to the antibody-PBF-probe complex was observed (Fig. 2c, lane 6, white arrow). When an antiserum against the bdPBF was used, the interaction was also abolished (Fig. 2c, lane 8), however we did not observe a supershift. The lack of supershift could be explained by the polyclonal origin of the antibody, which recognized some epitopes of bdPBF and disturbed its DNA binding. A non-specific complex was detected in immuno-EMSA experiments (Fig. 2c, lanes 7, 8, 9; arrowhead). This upper band was present either in absence of bdPBF (lane 9) or in control experiments using non-immune serum (lane 7).
The C-terminal domain of PBF is involved in γZ gene activation
PBF as a transcriptional activator of γZ gene might be included in a transcriptional complex that determines the spatial and temporal expression of this storage protein. As it occurs with the Dof proteins described so far, Dof domain of PBF could be a bifunctional domain for DNA-binding on AAAG motif and protein–protein interactions (Yanagisawa 2004). In addition to Dof domain, the non-conserved C-terminal domain of PBF could also play a relevant role in the up-regulation of γZ gene through protein-protein interactions. To explore activation domains within PBF protein, a first set of transient co-transformation assays over 15 DAP endosperms were carried out using the proximal γZ promoter (444γZ) and it was found that PBF activated the 444γZ promoter in a dose-dependent manner (Fig. 3b, PBF). In addition, it was observed that the PBF activity was dependent on the Dof domain, because no transactivation of the 444γZ promoter was observed when the PBF mutated at the Zinc finger domain (Fig. 3a, m-PBF) was used as effector in the co-transformation experiments (Fig. 3b, m-PBF). These results evidenced that a functional Dof binding domain was essential for PBF to activate 444γZ promoter. However, we cannot discard that the loss of transactivation was due to an overall change in PBF conformation created by Cys mutation. Nevertheless, the Dof domain of PBF (bdPBF) is insufficient by itself to induce the transcription of the 444γZ promoter. As it can be seen in Fig. 3b, when overexpressing bdPBF a slight but consistent decrease of γZ promoter activity was observed. It is likely that this decrease is due to some competition of bdPBF with endogenous PBF for Pb3 site, as it has been described for the DNA binding domains of maize Dof1 transcription factors over the C4pepc promoter (Yanagisawa 2000).
The C-terminal half of PBF is necessary for γZ promoter activation. (a) On the left, schematic representation of the reporter constructs showing the principal cis-elements identified in the 444γZ and 527γZ promoters. On the right, the effector constructs coding for PBF or PBF domains are shown. Maize PBF sequences are indicated in light grey, the Dof domain in intense grey. The chPBF construct codes for a chimeric PBF containing the barley N-terminal domain of BPBF (including the corresponding Dof domain bDof) fused in frame to the maize PBF C-terminal domain. Ct-PBF and bdPBF are as in Fig. 2 and m-PBF corresponds to PBF with the Dof domain mutated at Cys65 residue. (b) Transient expression of 444γZ:GUS in maize endosperms co-transformed with increasing amounts of PBF, m-PBF or bdPBF effectors. The results are expressed as folds of activation with regard to the basal expression of 444γZ:GUS co-transformed with equivalent increasing amounts of an empty vector. (c) Transient expression of 527γZ:GUS in maize endosperms co-transformed with increasing amounts of PBF, Ct-PBF or chPBF effectors. The results are expressed as folds of activation with regard to the basal expression of 527γZ:GUS co-transformed with equivalent amounts of an empty vector. In all of the cases the mean value of at least three independent experiments is included and the error bars indicate the standard deviation from the mean
These results suggested that the C-terminal region of PBF was necessary to transactivate the γZ promoter. This hypothesis was supported by the result of a set of transient transformation assays using the proximal 527γZ promoter fused to GUS as a reporter (Fig. 3a). We decided to use 527γZ promoter, because it contained an AACA box. The presence of this motif in prolamin genes from barley has been demonstrated to be relevant for quantitative expression (Díaz et al. 2002). Thus, we expected some differences by adding additional 5′ regulatory sequences. However, it was not the case, we found similar overall activation results by using both proximal promoters, 444γZ and 527γZ. The co-transformation with the 35S:Ct-PBF effector construct (Fig. 3a) had no effect on the basal 527γZ promoter activity (Fig. 3c, Ct-PBF), indicating that if any protein–protein interaction takes place in this region it is insufficient for activation and depends on the DNA binding capacity of the Dof domain. Furthermore, a chimeric construct was designed containing the Dof domain of the PBF from barley (BPBF; Mena et al. 1998) fused in frame to the Ct-PBF from maize (Fig. 3a, chPBF). Co-transformation of the 527γZ:GUS construct used as reporter and the 35S:chPBF as effector showed restored levels in the transcriptional activation capacity of maize PBF (Fig. 3c, ch-PBF). Interestingly, the chimeric chPBF promotes a slightly higher activation of the 527γZ:GUS reporter than the wild type PBF construct (Fig. 3c compare PBF and chPBF), maybe due to a better anchorage to its target sequence.
The presence of PBF protein in the developing endosperm is temporally correlated with γZ gene transcription
We further examined the correlation between the 444γZ transcriptional activity and the presence of PBF protein through the maize endosperm development. In order to detect PBF protein we used a polyclonal antibody affinity-purified against bdPBF. To determine γZ transcriptional activity, endosperms at different DAP (from 8 to 30 DAP) were transiently transformed with the 444γZ:GUS reporter gene, using the LUC vector as a control of the transformation efficiency. As it can be seen in Fig. 4a, the GUS/LUC relative activity indicates that the γZ promoter is active from 10 to 25 DAP, the main transcriptional activity being restricted to a narrow window during the endosperm development (from 10 to 20 DAP) with a maximum at around 12 DAP. It is worth noting that, except at 30 DAP, the luciferase values were high in all the stages analyzed (data not shown), indicating that the tissue was functional and the transformation was efficient at these developmental stages. The lack of LUC expression at 30 DAP agrees with the cessation of transcription activity linked to the endosperm desiccation developmental stage. When endosperm extracts were analysed by western blot using the anti-PBF antibody, one single band was detected (Fig. 4b, PBF). We observed a correlation between the presence of PBF transcription factor and the activity of the 444γZ promoter (see Fig. 4a) being the larger amount of PBF detected at 12 and 15 DAP (Fig. 4b) when γZ gene reached the maximal transcriptional activity. In this context it is worth noting that PBF was also detected by western blot in nuclear extracts of 12 and 15 DAP maize endosperms. The correlation between the presence of PBF and the initial process of γ-zein accumulation in the maize endosperm cells was also evidenced. As it can be seen in Fig. 4b, γ-zein was immuno-detected from 10 DAP endosperms and accumulated progressively throughout the endosperm development. Note that although PBF was mainly present in 12 and 15 DAP nuclear extracts, a small amount of PBF was also detected at earlier DAP in over-exposed films (data not shown).
Temporal correlation of PBF accumulation and activity with γZ gene expression during endosperm development. (a) Expression of the 444γZ:GUS reporter construct in maize endosperms transformed at different DAP. The results are expressed as GUS activity relative to the efficiency of transformation determined by co-transformation with LUC reporter constructs (see methods). (b) Western blot showing the presence of PBF in total protein extracts and in nuclear extracts of endosperms at different DAP. Total protein extracts were also immuno-labeled for γ-zein by using a γ-zein antibody. PBF was detected by using a polyclonal antibody against bdPBF. Molecular weights are indicated on the right. (c) Transient expression of 444γZ:GUS in maize endosperms at different DAP co-transformed with PBF. The results are expressed as folds of activation with regard to the basal expression of 444γZ:GUS co-transformed with and equivalent amount of an empty vector. In graphs (a) and (c) the mean value of at least three independent experiments is included and the error bars indicate the standard deviation from the mean
PBF gene was previously found to be expressed specifically in maize endosperms and its mRNA was first detected at 10 DAP reaching a peak at 15 DAP (Vicente-Carbajosa et al. 1997). As shown in Fig. 4a, γZ was not expressed at the early stages of seed development in our transient experiments. GUS activity was detected after 10 DAP reaching a peak at 12 DAP and decreasing afterwards to undetectable values at 30 DAP. To test whether PBF determines the starting point of γZ expression, we explored the transactivation capacity of PBF over the 444γZ promoter along the endosperm development by using the effector 35S:PBF and the reporter 444γZ:GUS. The results in Fig. 4c show that the over-expression of PBF increased the expression of GUS only at the developmental stages where the activity of 444γZ detected in our transient expression assays was high (compare Fig. 4a and c). Effectively, at 8 and 10 DAP no significant γZ activity increase was observed by PBF over-expression but almost seven and fourfolds of γZ promoter activation were determined respectively when 12 and 15 DAP endosperms were used. There is a clear correlation between the basal expression detected after transformation with the 444γZ promoter and the trans-activation capacity of PBF in the co-transformation experiments (r = 0.97), thus demonstrating that PBF upregulates γZ expression along the endosperm development. Nevertheless, high levels of PBF by itself are insufficient to trigger the mechanisms responsible for the induction of the γZ expression; indicating that other protein factors bound to PBF should also play an essential role.
PBF localizes in the nucleus of peripheral cells of 15 DAP endosperms
In developing maize endosperm mitotic phase occurs after 4 DAP and completes at around 12 DAP. During this period of time, cells in peripheral position differentiate in aleurone layer cells (Fig. 5a, (a)), over the main vascular tissue in transfer cells and central cells in starchy endosperm (Olsen 2001). After mitotic phase, cell divisions only occur in the subaleurone layer and the central cells initiate the accumulation of starch and storage proteins (Lopes and Larkins 1993). As described, γ-zein protein accumulates both in peripheral and central cells of developing endosperms (Fig. 5c). However, the transcription of γZ was restricted to the peripheral cells of the endosperm (Fig. 5e). To investigate the cellular localization of PBF and whether it correlates with those cells which are more active in γZ expression, we analysed the endosperm cortical region by immunocytochemistry over paraffin-embedded sections of maize seeds. Transversal sections of 15 DAP seeds (Fig. 5a) show the classical morphology at this stage of development. Pericarp (P), nucella (N), aleurone layer (a) and endosperm (En) were clearly recognisable with the aid of the autofluorescence associated with the cell walls. Starchy endosperm is composed of large cells that diminished in size from central endosperm to peripheral cells. This cell size gradient was more evident when nuclei were labelled with DAPI (Fig. 5f), the smallest cells, most of them still dividing, being detected in the subaleurone region (black arrowheads). To localize PBF, the grain sections were incubated with an anti-PBF antibody and labelled with FITC (Fig. 5d). Immunolabelling of PBF indicated that this protein localized exclusively in the endosperm cells since no specific label was detected in the external peripheral layers. No significant background was observed in the control sections with the exception of some autofluorescence detected in cell walls of aleurone layer (Fig. 5b).
PBF localizes in the nuclei of 15 DAP maize endosperm cells. (c) Transversal sections of 15 DAP maize grains incubated with anti-γ-zein antibody showing the specific γ-zein immunolabeling (in red) in the endosperm tissue. Arrows in (c) indicate the absence of label in the aleurone layer (a). In (a) the autofluorescence associated with the cell walls is shown. (b) and (d): high magnification of grain sections incubated with (d) or without (b) purified anti-PBF antibodies showing the PBF pattern by immunofluorescence (d, in green). The DAPI staining (f) of the section in d shows the presence and position of the nuclei in the sub-aleurone endosperm cells (black arrowheads), in the rest of the cortical endosperm cells (white arrowheads) and in the aleurone (a) and the nucelle (N) tissues. (e) GUS activity detected in the maize endosperm peripheral region after bombardment of transversal kernel halves with the 444γZ:GUS construct. En, Endosperm; a, aleurone; N, nucelle and P, pericarp. White arrows and black arrowheads indicate the first subaleurone cell layers
It can be observed that PBF label was detected in nuclei and also broadly distributed along the cytoplasm in dividing cells close to the aleurone layer (compare Fig. 5, d and f, black arrowheads). However, we observed that PBF localizes preferentially in the nucleus in the cell layers extending from the periphery to the central region (Fig. 5. d and f, white arrowheads). Note that in those cells where PBF label was distributed both, in nuclei and cytoplasm (black arrowheads in d), the γ-zein label was not detected (white arrows in c). This indicates that, as expected, PBF biosynthesis temporally precedes γ-zein accumulation.
Discussion
Since the original cloning of the maize Dof protein PBF (Vicente-Carbajosa et al. 1997) there have been no reports demonstrating PBF function on maize storage protein genes. Circumstantial evidence, however, suggests that PBF is a regulator of the 22 kD α-zein, since it specifically binds to the prolamin box in the gene promoter and interacts in vitro with the O2 protein (Vicente-Carbajosa et al. 1997), which is known to regulate 22 kD α-zein expression (Schmidt et al. 1992). Nevertheless, previous attempts to verify PBF transactivation of the α-zein promoter have failed, both in transient transformation studies performed on suspension cultures of maize endosperm cells (Vicente-Carbajosa, personal communication), as well as in whole maize endosperms (supplemental data). A recent report has confirmed that maize PBF and O2 proteins act as effective stimulators of rice glutelin gene expression in rice endosperms (Hwang et al. 2004).
PBF is a critical regulator of the γZ gene promoter
In this study, we provide experimental evidences that maize PBF is a γ-zein gene regulator in maize endosperms. PBF is an efficient activator of γ-zein gene, most likely by binding to AAAG motifs (prolamin boxes) present in the γZ regulatory sequence. This binding activity is consistent with those reported previously for Dof proteins in other cereals (Onodera et al. 2001; Vicente-Carbajosa et al. 1997; Yanagisawa and Schmidt 1999). Through transient expression assays we found that maize PBF is able to activate by up to sevenfold the transcription of the proximal region of γZ promoter by binding to Pb3 and Pb1 sequences. Although previous research showed Pb boxes to have merely a quantitative role (Wu et al. 1998), our data indicate that Pb3 plays a central role in the control of γZ expression. The transactivation capacity of PBF was completely abolished when γZ promoter is mutated in the Pb3 site, in contrast, PBF was able to slightly activate γZ expression when Pb1 was mutated. The cis-element Pb2, previously described in γZ promoter (Marzábal et al. 1998) has not been considered in this study since it lacks the core sequence 5′ (A/T)AAAG3′ necessary for prolamin box functionality (Ueda et al. 1994).
γZ promoter contains a unique binding site (GZM) that matches the GCN4-type consensus (TGACATG). γZ promoter mutations in the GZM site also reduced significantly the PBF capacity to transactivate the γZ promoter, confirming the functional interaction between Pb3 and GZM that we reported in a previous study (Marzábal et al. 1998). It is well known that during seed development several trans-acting factors including bZIP and Dof proteins act together in regulating the expression of storage protein genes (Isabel-LaMoneda et al. 2003; Mena et al. 1998, 2002). The GCN4 motif is the recognition site of bZIP proteins such as maize O2 (Schmidt et al. 1992), barley BLZ2 (Oñate et al. 1999), wheat SPA1 (Albani et al. 1997) and rice RISBZ1 (Onodera et al. 2001). O2 has been previously described as an important activator of α-zein multigenic family (Cord Neto et al. 1995; Schmidt et al. 1992) as well as of other endosperm specific proteins in maize (Hunter et al. 2002; Lohmer et al. 1991). Maize O2 was able to activate 444γZ:GUS expression in functional assays and to bind specifically to the GZM box in EMSA experiments (supplemental data). However, it is well known that γZ expression is not affected by the opaque2 mutation in maize endosperms (Geetha et al. 1991; Marzábal et al. 1998; Motto et al. 1989). These data suggest that in maize endosperm there should be a component of the bZIP family other than O2, which up-regulates γZ promoter in a redundant manner.
Functional domains of PBF
Maize PBF, like other Dof proteins, is a bi-functional transcription factor. The N-terminal PBF region (first 145 amino acid residues, bdPBF) contains the Cys2/Cys2 Zn2+ DNA-binding domain (52 residues) that specifically binds to Pb3 site within target promoter. When Cys65 was replaced by Ala, the protein lost its capacity to bind Pb3 and also transactivate γZ. This indicates that the Cys2/Cys2 Zn2+ structure is necessary to maintain the proper conformation for DNA binding, as it has been previously described for other Dof transcription factors (Mena et al. 1998; Shimofurutani et al. 1998; Yanagisawa 1995). In addition to Cys residues, it appears that aromatic residues (Y and W) in the C-terminal part of Dof domain contribute to stabilize the Dof structure (Umemura et al. 2004). BdPBF binds to Pb3 site but is unable by itself to transactivate γZ when overexpressed in endosperm cells. The C-terminal region (last 183 amino acid residues), which does not bind Pb3, also contributes to γZ transactivation, although, as expected, over-expression of Ct-PBF could not induce transactivation on its own (Fig. 3c, Ct-PBF). In fact, both N-terminal (bdPBF) and C-terminal (Ct-PBF) caused a slight inhibition of the basal γZ promoter activity. It is noteworthy that when the Ct-PBF was fused to a heterologous Dof domain (Dof-BPBF), the chimeric protein (chPBF) was able to transactivate γZ promoter, even to a higher level than the native maize PBF protein (Fig. 3c, chPBF). Therefore, it appears that the PBF C-terminal region needs an anchorage to DNA to show functional activation. Recently it has been reported that the Dof domain was involved in trans-activation of target genes by protein-protein interactions (Dong et al. 2007), yet the role of C-terminal domain of cereal PBFs being still unknown. In maize PBF, the striking presence of an asparagine-rich strain located at the end of the C-terminal half of maize PBF suggests that it may be involved in the activation of the transcriptional machinery via protein–protein interactions. Recently, asparagine rich domains have been reported to be overrepresented in yeast transcriptional activators (Titz et al. 2006) and, interestingly, Asn content correlates with activation capacity.
PBF temporal and spatial expression is consistent with its role in γZ regulation
During maize endosperm development, we observed that the presence of PBF protein correlates temporally with the accumulation of γ-zein protein in maize endosperms. Antibodies against PBF allowed us to detect PBF in both, total protein and nuclear extracts of endosperms of 12 and 15 DAP. However, at later stages of development (20–25 DAP), where γ-zein is highly accumulated, PBF is not detectable either in nuclei or in total protein extracts. These data are consistent with a temporal delay between RNA expression/protein accumulation and the long-life of γ-zein mRNA (Plotnikov and Bakaldina 1996; Woo et al. 2001). However, we cannot exclude the possibility that it might reflect a restricted spatial distribution of PBF within the endosperm tissue. The transcriptional activity of the 444γZ:GUS during the endosperm development correlates with the presence of PBF and is in agreement with previous data on the expression profile of PBF gene along endosperm development (Vicente-Carbajosa et al. 1997). Transiently-expressed PBF failed to transactivate γZ promoter outside the temporal window where endogenous PBF is expressed (from 10 to 20 DAP with a peak at 12 DAP) indicating that, besides PBF, other protein factors are involved in γZ gene regulation. Several protein factors interacting with Dof proteins have been described as co-regulators of target genes. As it occurs with PBFs in other cereals, maize PBF could participate as a partner in the γZ transcriptional complex interacting with other protein factors such as HMG-type proteins, MYB proteins and bZIP O2-like proteins.
444γZ:GUS promoter expression in sections of 15 DAP endosperms is mainly localized in peripheral cells, whereas the central part of the endosperm is γZ transcriptionally inactive and cells are entering the early programmed cell death. Interestingly, PBF is localized in the nucleus of peripheral cell layers and in the cytoplasm of cells that are in contact with aleurone layer. In maize, it seems that aleurone and starchy endosperm cell fate specification occurs through an intrinsic developmental program that discriminates between surface and internal cell positions (Gruis et al. 2006). To expand the aleurone layer and to increase the endosperm volume, endosperms retain a low activity of mitosis in the peripheral cell layers as late as 42 DAP (Mangelsdorf and Jones 1926). Inner daughter cells of dividing aleurone cells become starchy endosperm upon internalization (Costa et al. 2003). The absence of PBF in aleurone, but its immuno-detection in the first layer of peripheral cells might reflect the positional conversion of aleurone cells into starchy endosperm. In inner cells, the nuclear localization of PBF could determine the activation of γZ promoter and the subsequent start of storage protein synthesis. This pattern corresponds to the γ-zein accumulation that is absent in the first two-subaleurone layers but present in the rest of endosperm. The absence of PBF in the central region of the starchy endosperm indicates the loss of transcriptional activity in these cells. In fact, DAPI staining of endosperm sections revealed the presence of few nuclei with irregular shape, suggesting that cells enter programmed cell death (Young and Gallie 2000).
Maize storage protein mutants should be a tool to understand γZ gene expression and to search for γZ regulatory genes. At present, several maize mutants have been described to influence a large number of genes including zein genes in endosperm (Hunter et al. 2002). However, with the exception of opaque2 (Schmidt et al. 1992) and fl2 (Coleman et al. 1997), little is known about the molecular basis of these mutants. Neither PBF nor other γZ regulatory gene mutants have been identified. Among the o2 mutants, γ-zein biosynthesis is altered in the opaque-15 but not in other opaque mutants (Dannenhoffer et al. 1995). There is evidence, however, that the opaque-15 mutant is the result of a mutation of an opaque-2 modified gene not related to a putative γZ gene transcription factor. These genetic approaches together with post-transcriptional gene silencing approaches, recently developed in plants, would help to elucidate mechanisms underlying seed storage gene regulatory pathways (Houmard et al. 2007; Segal et al. 2003).
References
Albani D, Hammond-Kosack MC, Smith C, Conlan S, Colot V, Holdsworth M, Bevan MW (1997) The wheat transcriptional activator SPA: a seed-specific bZIP protein that recognizes the GCN4-like motif in the bifactorial endosperm box of prolamin genes. Plant Cell 9:171–184
Baumann K, De Paolis A, Costantino P, Gualberti G (1999) The DNA binding site of the Dof protein NtBBF1 is essential for tissue-specific and auxin-regulated expression of the rolB oncogene in plants. Plant Cell 11:323–334
Boronat A, Martínez MC, Reina M, Puigdomènech P, Palau J (1986) Isolation and sequencing of a 28-kDa glutelin-2 gene from maize. Common elements in the 5′ flanking regions among zein and glutelin genes. Plant Sci 47:95–102
Busk PK, Jensen AB, Pages M (1997) Regulatory elements in vivo in the promoter of the abscissic acid responsive gene rab17 from maize. Plant J 11:1285–1295
Coleman CE, Clore AM, Ranch JP, Higgins R, Lopes MA, Larkins BA (1997) Expression of a mutant alpha-zein creates the floury2 phenotype in transgenic maize. Proc Natl Acad Sci USA 94:7094–7097
Cord Neto G, Yunes JA, da Silva MJ, Vettore AL, Arruda P, Leite A (1995) The involvement of Opaque 2 on beta-prolamin gene regulation in maize and Coix suggests a more general role for this transcriptional activator. Plant Mol Biol 27:1015–1029
Costa LM, Gutierrez-Marcos JF, Brutnell TP, Greenland AJ, Dickinson HG (2003) The globby1–1 (glo1–1) mutation disrupts nuclear and cell division in the developing maize seed causing alterations in endosperm cell fate and tissue differentiation. Development 130:5009–5017
Dannenhoffer JM, Bostwick DE, Or E, Larkins BA (1995) Opaque-15, a maize mutation with properties of a defective opaque-2 modifier. Proc Natl Acad Sci USA 92:1931–1935
de Souza Filho GA, da Silva MJ, Vettore AL, Yunes JA, Leite A, Arruda P, Ottoboni LM (1999) Identification of a DNA-binding factor that recognizes an alpha-coixin promoter and interacts with a Coix Opaque-2 like protein. Plant Mol Biol 39:95–104
Díaz I, Vicente-Carbajosa J, Abraham Z, Martínez M, Isabel-La Moneda I, Carbonero P (2002) The GAMYB protein from barley interacts with the DOF transcription factor BPBF and activates endosperm-specific genes during seed development. Plant J 29:453–464
Díaz I, Martínez M, Isabel-LaMoneda I, Rubio-Somoza I, Carbonero P (2005) The DOF protein, SAD, interacts with GAMYB in plant nuclei and activates transcription of endosperm-specific genes during barley seed development. Plant J 42:652–662
Dong G, Ni Z, Yao Y, Nie X, Sun Q (2007) Wheat Dof transcription factor WPBF interacts with TaQM and activates transcription of an alpha-gliadin gene during wheat seed development. Plant Mol Biol 63:73–84
Entwistle J, Knudsen S, Muller M, Cameron-Mills V (1991) Amber codon suppression: the in vivo and in vitro analysis of two C-hordein genes from barley. Plant Mol Biol 17:1217–1231
Forde BG, Heyworth A, Pywell J, Kreis M (1985) Nucleotide sequence of a B1 hordein gene and the identification of possible upstream regulatory elements in endosperm storage protein genes from barley, wheat and maize. Nucleic Acids Res 13:7327–7339
Geetha KB, Lending CR, Lopes MA, Wallace JC, Larkins BA (1991) opaque-2 modifiers increase gamma-zein synthesis and alter its spatial distribution in maize endosperm. Plant Cell 3:1207–1219
Gruis DF, Guo H, Selinger D, Tian Q, Olsen OA (2006) Surface position, not signaling from surrounding maternal tissues, specifies aleurone epidermal cell fate in maize. Plant Physiol 141:898–909
Hammond-Kosack MC, Holdsworth MJ, Bevan MW (1993) In vivo footprinting of a low molecular weight glutenin gene (LMWG-1D1) in wheat endosperm. EMBO J 12:545–554
Hill DE, Hope IA, Macke JP, Struhl K (1986) Saturation mutagenesis of the yeast his3 regulatory site: requirements for transcriptional induction and for binding by GCN4 activator protein. Science 234:451–457
Houmard N, Mainville J, Bonin C, Huang S, Luethy M, Malvar T (2007) High-lysine corn generated by endosperm-specific suppression of lysine catabolism using RNAi. Plant Biotechnol J 5:605–614
Hull GA, Halford NG, Kreis M, Shewry PR (1991) Isolation and characterisation of genes encoding rye prolamins containing a highly repetitive sequence motif. Plant Mol Biol 17:1111–1115
Hunter BG, Beatty MK, Singletary GW, Hamaker BR, Dilkes BP, Larkins BA, Jung R (2002) Maize opaque endosperm mutations create extensive changes in patterns of gene expression. Plant Cell 14:2591–2612
Hwang YS, Ciceri P, Parsons RL, Moose SP, Schmidt RJ, Huang N (2004) The maize O2 and PBF proteins act additively to promote transcription from storage protein gene promoters in rice endosperm cells. Plant Cell Physiol 45:1509–1518
Isabel-LaMoneda I, Díaz I, Martínez M, Mena M, Carbonero P (2003) SAD: a new DOF protein from barley that activates transcription of a cathepsin B-like thiol protease gene in the aleurone of germinating seeds. Plant J 33:329–340
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Kang HG, Singh KB (2000) Characterization of salicylic acid-responsive, arabidopsis Dof domain proteins: overexpression of OBP3 leads to growth defects. Plant J 21:329–339
Kreis M, Forde BG, Rahman S, Miflin BJ, Shewry PR (1985) Molecular evolution of the seed storage proteins of barley, rye and wheat. J Mol Biol 183:499–502
Langdale JA (1994) In situ hybridization. In: Freeling MWV (ed) The maize handbook. Springer-Verlag, New York, pp 165–179
Lohmer S, Maddaloni M, Motto M, Di Fonzo N, Hartings H, Salamini F, Thompson RD (1991) The maize regulatory locus Opaque-2 encodes a DNA-binding protein which activates the transcription of the b-32 gene. EMBO J 10:617–624
Lopes MA, Larkins BA (1993) Endosperm origin, development, and function. Plant Cell 5:1383–1399
Ludevid MD, Martínez-Izquierdo JA, Armengol M, Torrent M, Puigdomenech P, Palau J (1985) Immunological relations between glutelin-2 and low molecular weight Zein-2 proteins from maize (Zea mays L.) endosperm. Plant Sci 41:41–48
Mackay JP, Crossley M (1998) Zinc fingers are sticking together. TIBS 23:1–4
Mangelsdorf PC, Jones DF (1926) The expression of Mendelian factors in the gametophyte of maize. Genetics 11:423–455
Marzábal P, Busk PK, Ludevid MD, Torrent M (1998) The bifactorial endosperm box of gamma-zein gene: characterisation and function of the Pb3 and GZM cis-acting elements. Plant J 16:41–52
Mena M, Vicente-Carbajosa J, Schmidt RJ, Carbonero P (1998) An endosperm-specific DOF protein from barley, highly conserved in wheat, binds to and activates transcription from the prolamin-box of a native B-hordein promoter in barley endosperm. Plant J 16:53–62
Mena M, Cejudo FJ, Isabel-Lamoneda I, Carbonero P (2002) A role for the DOF transcription factor BPBF in the regulation of gibberellin-responsive genes in barley aleurone. Plant Physiol 130:111–119
Moreno-Risueño MA, Martínez M, Vicente-Carbajosa J, Carbonero P (2007) The family of DOF transcription factors: from green unicellular algae to vascular plants. Mol Genet Genomics 277:379–390
Motto M, Di Fonzo N, Hartings H, Maddaloni M, Salamini F, Soave C, Thompson RD (1989) Regulatory genes affecting maize storage protein synthesis. Onf Surv Plant Mol Cell Biol 6:87–114
Olsen OA (2001) Endosperm development: cellularization and cell fate specification. Annu Rev Plant Physiol Plant Mol Biol 52:233–267
Onodera Y, Suzuki A, Wu CY, Washida H, Takaiwa F (2001) A rice functional transcriptional activator, RISBZ1, responsible for endosperm-specific expression of storage protein genes through GCN4 motif. J Biol Chem 276:14139–14152
Oñate L, Vicente-Carbajosa J, Lara P, Díaz I, Carbonero P (1999) Barley BLZ2, a seed-specific bZIP protein that interacts with BLZ1 in vivo and activates transcription from the GCN4-like motif of B-hordein promoters in barley endosperm. J Biol Chem 274:9175–9182
Ottoboni LM, Leite A, Yunes JA, Targon ML, de Souza Filho GA, Arruda P (1993) Sequence analysis of 22 kDa-like alpha-coixin genes and their comparison with homologous zein and kafirin genes reveals highly conserved protein structure and regulatory elements. Plant Mol Biol 21:765–778
Papi M, Sabatini S, Bouchez D, Camilleri C, Costantino P, Vittorioso P (2000) Identification and disruption of an Arabidopsis zinc finger gene controlling seed germination. Genes Dev 14:28–33
Piette J, Hirai S, Yaniv M (1988) Constitutive synthesis of activator protein 1 transcription factor after viral transformation of mouse fibroblasts. Proc Natl Acad Sci USA 85:3401–3405
Plotnikov VK, Bakaldina NB (1996) Differential stability of zein mRNA in developing corn kernel. Plant Mol Biol 31:507–515
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Schmidt RJ, Burr FA, Aukerman MJ, Burr B (1990) Maize regulatory gene opaque-2 encodes a protein with a “leucine-zipper” motif that binds to zein DNA. Proc Natl Acad Sci USA 87:46–50
Schmidt RJ, Ketudat M, Aukerman MJ, Hoschek G (1992) Opaque-2 is a transcriptional activator that recognizes a specific target site in 22-kD zein genes. Plant Cell 4:689–700
Segal G, Song R, Messing J (2003) A new opaque variant of maize by a single dominant RNA-interference-inducing transgene. Genetics 1:387–397
Shimofurutani N, Kisu Y, Suzuki M, Esaka M (1998) Functional analyses of the Dof domain, a zinc finger DNA-binding domain, in a pumpkin DNA-binding protein AOBP. FEBS Lett 430:251–256
Skirycz A, Reichelt M, Burow M, Birkemeyer C, Rolcik J, Kopka J, Zanor MI, Gershenzon J, Strnad M, Szopa J, Mueller-Roeber B, Witt I (2006) DOF transcription factor AtDof1.1 (OBP2) is part of a regulatory network controlling glucosinolate biosynthesis in Arabidopsis. Plant J 47:10–24
Sumner-Smith M, Rafalski JA, Sugiyama T, Stoll M, Soll D (1985) Conservation and variability of wheat alpha/beta-gliadin genes. Nucleic Acids Res 13:3905–3916
Thompson GA, Larkins BA (1989) Structural elements regulating zein gene expression. Bioessays 10:108–113
Titz B, Thomas S, Rajagopala SV, Chiba T, Ito T, Uetz P (2006) Transcriptional activators in yeast. Nucleic Acids Res 34:955–967
Torrent M, Alvarez I, Geli MI, Dalcol I, Ludevid D (1997) Lysine-rich modified gamma-zeins accumulate in protein bodies of transiently transformed maize endosperms. Plant Mol Biol 34:139–149
Ueda T, Wang Z, Pham N, Messing J (1994) Identification of a transcriptional activator-binding element in the 27-kilodalton zein promoter, the −300 element. Mol Cell Biol 14:4350–4359
Umemura Y, Ishiduka T, Yamamoto R, Esaka M (2004) The Dof domain, a zinc finger DNA-binding domain conserved only in higher plants, truly functions as a Cys2/Cys2 Zn finger domain. Plant J 37:741–749
Vicente-Carbajosa J, Moose SP, Parsons RL, Schmidt RJ (1997) A maize zinc-finger protein binds the prolamin box in zein gene promoters and interacts with the basic leucine zipper transcriptional activator Opaque2. Proc Natl Acad Sci USA 94:7685–7690
Washio K (2001) Identification of Dof proteins with implication in the gibberellin-regulated expression of a peptidase gene following the germination of rice grains. Biochim Biophys Acta 1520:54–62
Woo YM, Hu DW, Larkins BA, Jung R (2001) Genomics analysis of genes expressed in maize endosperm identifies novel seed proteins and clarifies patterns of zein gene expression. Plant Cell 13:2297–2317
Wu CY, Suzuki A, Washida H, Takaiwa F (1998) The GCN4 motif in a rice glutelin gene is essential for endosperm-specific gene expression and is activated by Opaque-2 in transgenic rice plants. Plant J 14:673–683
Yamamoto MP, Onodera Y, Touno SM, Takaiwa F (2006) Synergism between RPBF Dof and RISBZ1 bZIP activators in the regulation of rice seed expression genes. Plant Physiol 141:1694–1707
Yanagisawa S (1995) A novel DNA-binding domain that may form a single zinc finger motif. Nucleic Acids Res 23:3403–3410
Yanagisawa S (2000) Dof1 and Dof2 transcription factors are associated with expression of multiple genes involved in carbon metabolism in maize. Plant J 21:281–288
Yanagisawa S (2002) The Dof family of plant transcription factors. Trends Plant Sci 7:555–560
Yanagisawa S (2004) Dof domain proteins: plant-specific transcription factors associated with diverse phenomena unique to plants. Plant Cell Physiol 45:386–391
Yanagisawa S, Schmidt RJ (1999) Diversity and similarity among recognition sequences of Dof transcription factors. Plant J 17:209–214
Yanagisawa S, Sheen J (1998) Involvement of maize Dof zinc finger proteins in tissue-specific and light-regulated gene expression. Plant Cell 10:75–89
Young TE, Gallie DR (2000) Programmed cell death during endosperm development. Plant Mol Biol 44:283–301
Zhang B, Chen W, Foley RC, Buttner M, Singh KB (1995) Interactions between distinct types of DNA binding proteins enhance binding to ocs element promoter sequences. Plant Cell 7:2241–2252
Acknowledgements
This work was supported by the MCYT-FEDER (BIO 2004-03202) and Generalitat de Calalunya (CeRBa and 2005 SGR00182). P.M. was recipient of a fellowship from Spanish Ministerio de Educación y Ciencia. E.G was recipient of a fellowship from AGAUR (Generalitat de Catalunya). We thank Robert Schmidt (UCSD, USA) for kindly providing PBF cDNA. We are grateful to Maite Galiñánez for the technical assistance in the antibody production and to Monica Pons for her assistance in immuno microscopy.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. A
The opaque 2 transcription factor (O2) interacts in vitro with the GZM motif. (A1) Oligonucleotides oBB, oBBDPb3 and oBBDGZM used in EMSA assays showing the bifactorial box motifs Pb3 and GZM as well as the nucleotide substitutions introduced in these cis-elements (arrows over lower case). (A2) EMSA assays with the O2 protein produced by in vitro transcription and translation, using as probes the oligonucleotides detailed in A. As it can be seen in the figure, the incubation of O2 with the wild type probe (oBB; lane 2) produced several band shifts. Only the upper band (arrow) corresponds to O2 binding because the rest of the shifts are also observed in the negative control (lane 7). The interaction of the O2 to the oligonucleotide oBB takes place specifically at the GZM motif. The specific band shift observed in lane 2 (arrow) is abolished when GZM motif was mutated (oBBDGZM; lane 4), but not when Pb3 motif was mutated (oBBDPb3; lane 6). Lanes 1, 3 and 5 correspond to the probe incubated in the absence of O2 (TIF 949 kb)
Fig. B
The over-expression of PBF does not affect the transcriptional rate of the αZ promoter of the 22 kD α-zein gene on transiently transformed 15 DAP maize endosperms. (B1) Schematic representation of the reporter (22αZ:LUC) and the effector genes used in functional studies. The O2 (Z1, Z2 and Z3) and Pb (prolamin-box) ciselements present in the reporter gene promoter are shown. (B2) Representation of the folds of activation of the 22αZ promoter when co-expressed with PBF and O2 used as effectors. The values are expressed as folds of activation in relation to the basal 22αZ:LUC activity (lane C) obtained by using an empty vector as effector. The overexpression of O2 increased significantly the expression of the 22αZ promoter (more than 3-folds, lane O2). No effect on the expression of this promoter was observed when PBF was over-expressed (lane PBF). Even when both, O2 and PBF were overexpressed (lane PBF + O2) the slight increase observed on 22αZ:LUC activity as compared to that observed in lane O2 was not significant (TIF 299 kb)
Rights and permissions
About this article
Cite this article
Marzábal, P., Gas, E., Fontanet, P. et al. The maize Dof protein PBF activates transcription of γ-zein during maize seed development. Plant Mol Biol 67, 441–454 (2008). https://doi.org/10.1007/s11103-008-9325-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9325-5