Abstract
Allelic differences in expression are important genetic factors contributing to quantitative trait variation in various organisms. However, the extent of genome-wide allele-specific expression by different modes of gene regulation has not been well characterized in plants. In this study we developed a new methodology for allele-specific expression analysis by applying Massively Parallel Signature Sequencing (MPSS™), an open ended and sequencing based mRNA profiling technology. This methodology enabled a genome-wide evaluation of cis- and trans-effects on allelic expression in six meristem stages of the maize hybrid. Summarization of data from nearly 400 pairs of MPSS allelic signature tags showed that 60% of the genes in the hybrid meristems exhibited differential allelic expression. Because both alleles are subjected to the same trans-acting factors in the hybrid, the data suggest the abundance of cis-regulatory differences in the genome. Comparing the same allele expressed in the hybrid versus its inbred parents showed that 40% of the genes were differentially expressed, suggesting different trans-acting effects present in different genotypes. Such trans-acting effects may result in gene expression in the hybrid different from allelic additive expression. With this approach we quantified gene expression in the hybrid relative to its inbred parents at the allele-specific level. As compared to measuring total transcript levels, this study provides a new level of understanding of different modes of gene regulation in the hybrid and the molecular basis of heterosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Allelic variation in gene expression is an important contributor to the natural genetic variation of human disease, and quantitative traits in animals and plants (Cong et al. 2002; Yan et al. 2002; Doebley 2004; Knight 2004), and presumably a primary source accounting for species evolution (Wray et al. 2003; Doebley 2004; Knight 2004; Adams and Wendel 2005; Prud’homme et al. 2006; Wray 2007). The genetic basis of the allelic expression variation may be the result of differences in the cis-regulatory regions (such as promoter, enhancer elements, or intron regions) that affect transcription initiation, transcription rate, and transcript stability in an allele-specific manner. Cis-acting factors interact with trans-acting factors, such as transcriptional factors, to regulate transcriptional expression. The prevalence of cis- and trans-acting effects can be determined by mapping expression quantitative trait loci (eQTL) (West et al. 2006) or alternatively, by allele-specific expression analysis. Cis-regulatory variation may be assessed by comparing expression of two alleles in a common genotype such as an F1 hybrid, where a common cellular environment including trans-acting factors is involved. Trans-acting factors can be evaluated by comparing expression of the same allele present in different genetic backgrounds (Cowles et al. 2002; Wittkopp et al. 2004). Recent studies with allele-specific expression analyses show that such natural allelic expression variation is wide-spread in human and animal genomes (Yan et al. 2002; Lo et al. 2003; Knight 2004; Morley et al. 2004). More than 50% of the genes in human (Lo et al. 2003), and nearly 90% of genes in Drosophila interspecific hybrids (Wittkopp et al. 2004) show differential allelic expression. Trans-regulatory differences affect approximately 55% of genes analyzed in interspecific hybrids in Drosophila (Wittkopp et al. 2004).
In plants much less is known regarding the role of cis- and trans-regulatory effects to gene expression. A maize hybrid and its inbred parents are an ideal system for this type of study. The maize genome has been shown to have abundant allelic sequence diversity (Ching et al. 2002; Springer and Stupar 2007a, b). Allelic differences in cis-regulatory factors can be analyzed in the F1 hybrid, and trans-acting effects can be analyzed by comparing the same allele expression in the hybrid with its inbred parents. Allelic expression variation of non-imprinted genes was first reported in the triploid endosperm tissue of maize hybrids (Guo et al. 2003), and in diploid plant tissues including maize immature ears and seedlings (Guo et al. 2004). These studies also show that allelic expression varies in different tissue types and responds to environmental changes. Further data suggest that the allelic diversity may be functional and have an impact on hybrid performance. Despite the potential biological significance of the allele-specific regulation, limited data is available in plants to address related questions.
Furthermore, in the efforts of understanding the molecular basis of heterosis, most of the studies of genome-wide gene expression in hybrids and their inbred parents have been focused on total level of gene expression, regardless of the contribution of the parental alleles in the hybrid (Auger et al. 2005; Bao et al. 2005; Vuylsteke et al. 2005; Huang et al. 2006; Swanson-Wagner et al. 2006). Heterosis results from the combination of two parental alleles (Birchler et al. 2003; Springer and Stupar 2007a, b). However, how the alleles are differently regulated in the hybrid as compared to the inbred parents has not been characterized. Addressing this type of question requires analyzing gene expression in the hybrid and inbred parents at the allele-specific level, and taking consideration of the allelic contribution to gene expression in the hybrid. This approach may provide a different level of understanding of gene regulation in the hybrid and the molecular basis of heterosis, as compared to measuring total transcript amount without allelic differentiation.
Microarrays are a commonly used technology for genome-wide transcript profiling analysis. However, the microarray technology poses potential limitations because of the dependency on hybridization; for example, cross-hybridization among family members or allele variants can mask allele-specific signals. Designing experiments for allele-specific expression hybridization require sequence information for both parental alleles, which is not commonly available in plant genome databases yet. An alternative technology to microarray chips is the Massively Parallel Signature Sequencing (MPSS™), an open-ended and sequencing-based mRNA profiling technology for quantitative expression analysis (Brenner et al. 2000a, b). MPSS involves deep transcript sampling and sequencing, usually 1–2 million transcripts per library, of a cDNA library on microbeads. The process produces a 17-base sequence tag adjacent to the 3′-most DpnII restriction site in the transcript for each expressed gene, and has the potential to capture virtually all genes expressed in a sample. The signature tags are usually gene specific, and can be assigned to a single locus in the genome (Reinartz et al. 2002; Saha et al. 2002). The expression level of a gene is measured by a normalized count of the signature tag to a corresponding gene per library. The technology has been shown to provide comprehensive coverage, and a sensitive measure of gene expression (Jongeneel et al. 2005; Stolovitzky et al. 2005; Hedgecock et al. 2007). The MPSS profiling has been used in studies to address various biological questions, including a whole-genome transcript analysis in Arabidopsis (Meyers et al. 2004a, b) and in human (Jongeneel et al. 2003), and transcript expression profiling of hybrid and inbred parents in oysters (Hedgecock et al. 2007).
Despite its common use in transcript profiling, the application of the MPSS™ technology in allele-specific expression profiling has not yet been reported. In this study, we applied the MPSS™ technology in a genome-wide allele-specific expression analysis. We developed a methodology for evaluating how parental alleles are regulated when combined in a hybrid background, the extent of cis- and trans-effects on gene regulation in the hybrid, and the relevance of different modes of gene regulation to heterosis. The scope of this study, however, is not to establish the causal relationship of genes to heterosis, although providing a solution to such a question remains a highly interesting topic. The methodology, however, can be applied in future experiments to address questions relating specific genes or pathways to expression phenotype of heterosis.
Materials and methods
Plant materials and tissue sampling
Four genotypes, maize inbred lines B73, Mo17 and their reciprocally crossed hybrids B73/Mo17 and Mo17/B73 were sampled for meristem tissues (Fig. 1). The shoot apical meristems (SAMs) were sampled at V1, V2, and V3 stages when one, two, and three leaves were fully opened, respectively. Meristems were dissected and collected with two leaf primordia, and pooled from approximately 100 plants for each stage. Ear inflorescence meristems (EIMs) were collected from the immature ears at three time points corresponding to stages when 25%, 60% and 100% of the spikelets, respectively, were differentiated. Meristems of each stage were pooled from the primary ears of approximately 50 plants. Meristem tissues were dissected under a dissection scope, frozen in liquid nitrogen immediately, and stored in −80°C until RNA preparation. No biological replicated samples were collected due to the large quantity of meristem tissue and a large number of plants needed for each mRNA sample.
Shoot apical meristem (SAM) and ear inflorescent meristem (EIM) of maize inbred B73, Mo17 and their reciprocally crossed F1 hybrids B73/Mo17, Mo17/B73. SAM and EIM tissues were collected from the four genotypes as shown. SAM1, SAM2, and SAM3 tissues were collected from V1, V2, and V3 stages, respectively, before transitioning to the floral state, usually at V4. EIM1, EIM2, and EIM3 are stages of ear development when 25%, 60%, and 100% of the spikelets are differentiated, respectively
RNA isolation
Total RNA was isolated using Eppendorf’s “Protocol for RNA Isolation using TRIzol Reagent with Phase Lock Gel-Heavy” (Order # 0032 005.152, Eppendorf, Westbury, NY, USA). The tissue was homogenized in TRIzol solution and the cell lysate was added to a pre-spun Phase Lock Gel-Heavy tube. Chloroform was added and the tube was centrifuged to allow separation of the aqueous phase. Aqueous phase containing the RNA was transferred to a fresh tube and precipitated by the addition of isopropyl alcohol. Samples were then centrifuged to cause the RNA to pellet. The supernatant was decanted and ethanol was added to the pellet. After the ethanol wash the pellet was allowed to air dry before re-suspension in nuclease-free water. The mRNA was isolated using Amersham Biosciences mRNA Purification Kit (Cat # 27-9258-02, Amersham Biosciences, Piscataway, NJ, USA). The total RNA sample was prepared for loading on the column and allowed to soak in under gravity before centrifugation to bind the poly(A)+RNA to the column. The column with bound polyA was washed several times with high-salt and then low-salt buffers before the poly(A) RNA was eluted with successive aliquots of elution buffer.
Transcription profiling by Massively Parallel Signature Sequencing (MPSS™)
MPSS profiling produces short tags adjacent to the 3′-most DpnII restriction site in the transcript. The process of cloning of cDNA, preparation of microbead libraries, microbead DNA sequencing, and sequence base calling were previously described (Brenner et al. 2000a, b). Briefly, cDNA with polyA/T tail was prepared and digested by DpnII restriction enzyme. An adaptor with MmeI recognition site was ligated to the 5′-end, followed by MmeI digestion that cuts 21–22 bases downstream. This 21–22 base signature from each transcript was subsequently cloned by a unique adaptor and loaded to a microbead. This MPSS profiling process sampled 1–2 million sequenced transcripts per library on microbeads. Sequenced tags were generated by serial cutting and ligation of decoding adaptors. Sequencing runs were done by using two different cleavage steps which are two different 4-nucleotide sequencing frames offset by two bases (2-step) or three bases (3-step) (Brenner et al. 2000a, b). Two or three technical replicates were conducted for individual sequencing runs. For each sequence run, data were first normalized to PPM (part per million) for every tag in individual samples. For a given tag, its counts were summed and averaged by the cleavage step. The cleavage step with higher count average was selected. Replicated runs of the chosen cleavage step were averaged as the final representation for the tag in a sample.
Identification of Allelic Signature Tag Pairs (ASTPs)
We identified Allelic Signature Tag Pairs (ASTPs) between B73 and Mo17 inbred lines by searching for a single nucleotide polymorphism (SNP) in the 17-mer sequence signature. The tag pairs differed by only one base between B73 and Mo17, for example, ‘GATCGTGTGGGAAAGCC’ and ‘GATCATGTGGGAAAGCC’ have one base different (underlined). There were three possible alternative bases for each of the 13 variable nucleotides (excluding the unchangeable GATC at the 5′end), yielding a total of 3 × 13 = 39 possible signature tags. Each of the tags with a SNP was paired with the original tag and evaluated for the likelihood of allelic relationship. A putative ASTP was identified based on the following criteria: A tag corresponding to the B73 allele is present in only B73 samples (at 25 PPM or higher) and absent in Mo17 samples, and vise versa for the Mo17 allele. The 25 PPM background cutoff was empirically determined to reduce the background noise associated with lowly expressed genes. To further validate the allelic relationship of these putative ASTPs, we crosschecked them in other tissue libraries in the MPSS database, which consisted of 27 B73 libraries with 10 tissue types of various developmental stages (ear, endosperm, stem internode, whole kernel, leaf, meristem, pericarp, root, scutellum, and silk), and eight Mo17 libraries which consisted of three tissue types of different development stages (ear, meristem, and seedling). Given the large number of B73 libraries and potential data outliers (e.g. derived from sequencing error), a putative Mo17 allele tag was allowed to be present only in 1 PPM in less than 10% of the B73 samples. A putative B73 allele must be absent in all of the Mo17 samples because of fewer Mo17 tissue samples.
Data analysis
Statistical hypothesis testing
For each tag expressed in a biological sample we first took an average of the PPM values from technical replicates. To analyze allelic expression in the hybrid, we applied the threshold cutoff such that at least one of the two alleles expressed in the hybrid must be 25 PPM or higher, that is, above the background level. (If both alleles were expressed below the background level, the gene was considered as not expressed in the tissue, and no further analysis was pursued). When calculating allelic expression ratio, the 0 PPM values of the denominators were treated as 1 PPM to make it mathematically feasible. Standard error of the sample mean difference was estimated based on the data of the two reciprocally-crossed hybrids of the same tissue and stage by treating them as if they were biological replicates. Since the reciprocal crosses are only approximations for true replicates, the error estimates should be larger than the variation of true replicates from the same samples, therefore, introducing a conservative bias for detecting differential expression. The fitted spline was then used to predict standard errors from PPM values for all of the tests for differential expression as if the predicted values were estimates from two biological replicates (R Development Core Team 2005). Like a previous use of a spline fit (Nadon et al. 2001) and similar procedures (Jain et al. 2003) our use of the spline fit was based on the observation that standard error estimates are much more variable than mean estimates, especially when there are few biological replicates.
For differential allele expression analyses, we used the paired t-test with one degree of freedom for this study. The null hypothesis was that two allelic tags had equal mean values in a hybrid. We selected the P-value thresholds (significance levels) that yielded the lowest estimated false discovery rates (FDRs) (Benjamini and Hochberg 1995). The FDR was estimated by dividing the product of the total number of the allelic pairs tested and the significance level, by the number of allelic pairs with P values below the significance level, setting FDR estimates greater than 1 to 1. Less conservative estimation methods may improve results by taking into account an estimate of the portion of tags that are differentially expressed (Efron et al. 2001; Storey 2002; Bickel 2004). Based on the datasets from all three developmental stages of the SAM and EIM, the mean reciprocals of the FDR estimates are 1, 1.6, 5.0, 4.5, and 3.9 when the significance levels are 0.01, 0.05, 0.10, 0.15, and 0.20, respectively. To maximize the mean reciprocal FDR estimate, the significance level of 0.10 was therefore used in this study.
Trans-acting effect analysis
To evaluate trans-acting effects we developed a protocol to measure the expression difference of the same allele in the hybrid and the inbred parents. This approach considered each allele independently and detected trans-acting effects by directly comparing the expression level of a specific allele in the hybrid with that of its inbred parents. The assumption is that if different trans-acting effects are present, the expression level of a specific allele is expected to be different in the hybrid as compared to its inbred parents.
We define the two alleles of a gene in a diploid species as 1 and 2. The genotypes for two inbred parents would be g11, g22, respectively, and the corresponding hybrid progeny g12 (or g21 if reciprocal cross). We assume the concentration of the trans-acting factor in a haploid is 1 dose for a gene; the concentration in a diploid inbred and hybrid would be 2 doses. We denote eij as expression contribution from allele i (1 or 2, which is always coupled with the same cis-element i) under the control of trans-factor j; Eg,i as the total expression level of allele i in genotype g, which is what was directly measured in the experiments. The expression level in inbred g11 can be formulated as Eq. 1: E11,1 = 4e11 (2 doses of allele 1 multiplied by 2 doses of trans-factor); E11,2 = 0 (0 dose of allele 2 in inbred 1). Likewise in the inbred g22 the expression level can be formulated as Eq. 2: E22,2 = 4e22; E22,1 = 0. In hybrid g12 (or g21) the expression level can be formulated as Eq. 3: E12,1 = e11 + e12, Eq. 4: E12,2 = e21 + e22. Values of e11 and e22 can be directly derived from Eqs. 1 and 2, and values of e12 and e21 can be resolved by Eqs. 3 and 4. Below is an example showing two different scenarios where the values of each component are obtained using these equations.
Case | e11 | e22 | e12 | e21 |
|---|---|---|---|---|
A | 5 | 2.5 | 5 | 2.5 |
B | 5 | 2.5 | 95 | 47.5 |
Trans-acting effects are measured by using the formula: (e12 − e11)/e11 for allele 1, and (e21 − e22)/e22 for allele 2. The null hypothesis is e12 = e11 or e21 = e22. If e12 = e11 and (e12 − e11)/e11 = 0, which is case A, then there is no different trans-acting effect on allelic expression between the hybrid and inbred backgrounds. On the other hand, a significant deviation from zero will be an indication of the expression difference of this allele, and therefore, different trans-acting effect present between the genotypes e.g., (e12 − e11)/e11 = (95–5)/5 = 18, as in case B. Those ASTPs that had significant (P < 0.1) differences between e12 and e11 or e21 and e22 were identified as exhibiting a different trans-acting effect. Allelic comparison was made for both B73 and Mo17 alleles, respectively, with the hybrid (Mo17/B73) for each tissue stage.
RT-PCR confirmation
We used gene-specific primers to obtain the cDNA from each inbred parent by RT-PCR with Optimase polymerase (Transgenomic, Omaha, NE, USA). The PCR products were then sequenced to identify allele-specific sequence polymorphisms between the inbred lines that would allow separation of the two parental alleles on the WAVE dHPLC system (Transgenomic, Omaha, NE, USA). We designed gene-specific primers in conserved regions that flank a sequence polymorphism between alleles, either a single nucleotide polymorphism (SNP) or insertion/deletion (InDel) to minimize amplification preference of either allele and to optimize the amplicon for analysis on the WAVE. Thirty-cycle PCR was performed with cDNA from hybrids. Three PCR replicates were performed for each RNA sample. The RT-PCR products were then separated and quantified by the WAVE dHPLC system. Detailed WAVE dHPLC analysis has been described previously (Guo et al. 2003, 2004). Pair wise correlation analysis was conducted for the MPSS and RT-PCR/WAVE data by using the allelic ratios from Table 3. Zero PPM values of denominators (B73 allele expression level) in MPSS data were treated as 1 PPM to obtain an allelic ratio.
Novel materials described in this publication may be available for non-commercial research purposes upon acceptance and signing of a material transfer agreement. In some cases such materials may contain or be derived from materials obtained from a third party. In such cases, distribution of material will be subject to the requisite permission from any third-party owners, licensors or controllers of all or parts of the material. Obtaining any permission will be the sole responsibility of the requestor.
Results
MPSS™ profiling of shoot apical meristem (SAM) and ear inflorescent meristem (EIM) of F1 hybrids and the parental inbreds
To gain a global view of the regulation of allelic expression in a hybrid and its inbred parents, we developed a methodology for an open-ended, genome-wide allele-specific transcript profiling by applying the MPSS technology. We profiled reciprocally-crossed F1 hybrids B73/Mo17 and Mo17/B73 (where the first parent denotes the female and the second denotes the male parent), and the inbred parents B73 and Mo17. The tissues used were the shoot apical meristem (SAM) and the ear inflorescence meristem (EIM), corresponding to the vegetative and reproductive meristems of plant growth (Fig. 1). Tissue samples were collected from three stages of each SAM and EIM, which span most of the meristem development. Meristems are developmentally important tissues, and consist of very specific cell types. Since allele-specific expression varies among different tissues (Guo et al. 2004), by focusing on tissue with few cell types such as meristems, analysis of allelic expression was simplified, and less confounded than if using complex tissue types. RNA samples extracted from the meristems were subjected to the cloning and sequencing process of the MPSS profiling. Biological samples were not replicated due to the large quantity of tissue (∼100 meristems per sample) required to make enough mRNA for the profiling. The large number of plants in each sample, although technically is not a biological replicate, does serve the purpose of minimizing plant-plant variation. The in-depth sequencing and repetitive sampling of the mRNA pool for more than a million times, with technical replications, make it a highly robust technology. MPSS has been used successfully without biological replications for studies focusing on global trends (Meyers et al. 2004a, b).
Identification and validation of Allelic Signature Tag Pairs (ASTPs)
To measure allele-specific expression in the F1 hybrid, we took advantage of the Single Nucleotide Polymorphisms (SNP) contained in the MPSS signature tags between alleles of B73 and Mo17, and used the SNPs as markers to discriminate allele-specific tags. The maize genome is highly genetically diverse, and SNP frequency is about 1 per 70 bp among different inbred lines (Ching et al. 2002), which makes this approach feasible. We first identified MPSS signature tag pairs between B73 and Mo17 alleles, in which we allowed only one nucleotide polymorphism in the 17-base tag pairs. These tag pairs presumably represented the two alleles of a common gene. The allelic relationship of the tag pairs was further verified by examining their expression patterns in the B73 and Mo17 tissue samples. Verification was based on the assumption that a signature tag corresponding to a B73 allele should not be expressed in the Mo17 tissue samples, and vise versa for the putative Mo17 allele tags. The lack of expression of a putative allele tag (e.g., Mo17 allele) in tissues of the other genotype (e.g., B73) could be due to a true allelic relationship; alternatively, it could be due to the lack of expression of this gene in specific tissues of B73. To reduce the latter possibility, we further verified the allelic relationship by examining expression of the allelic tag across multiple tissue types and selected those that fit the criteria in multiple tissue libraries (Materials and methods). Using these criteria, we identified a total of 364 unique Allelic Signature Tag Pairs (ASTPs) from all six meristem stages: SAM1, SAM2, SAM3, EIM1, EIM2, and EIM3, with some ASTPs represented in more than one meristem stage (Supplementary Table 1).
To test the accuracy that the selected putative ASTPs match the alleles of corresponding genes, we compared sequence of B73 and Mo17 alleles corresponding to the putative allelic tag pairs by using ESTs from the public and proprietary databases. From the 46 genes that had sequence information for both inbred parents in the databases, 42 (91%) were confirmed for the exact nucleotide difference as in the putative ASTPs identified from MPSS. Sequences of these 42 genes from both parental alleles are listed in Supplemental Materials.
Allelic difference in expression and cis-regulation in the F1 hybrid
Transcript regulation is controlled by cis-acting factors, which are usually linked to the locus, and trans-acting factors, which are encoded elsewhere, such as transcriptional factors. Allelic differences in cis-regulation can be determined by comparing different alleles expressed in a common genetic background, such as in the hybrid, where the alleles are under the influence of the same trans-acting factors. In this study, we identify functional allelic cis-regulatory differences by comparing the differences of allele-specific expression in the maize hybrid. For the 364 ASTPs identified, we selected those that had at least one of the alleles expressed at 25 PPM for further allelic expression analysis. This empirical background cut-off (25 PPM) is higher than that (3 PPM) used in other transcriptome analyses, studies of oyster hybrids (Hedgecock et al. 2007), and Arabidopsis (Meyers et al. 2004a, b). Although this approach may exclude some lowly expressed genes; we feel, in this study, obtaining an accurate estimate from fewer identified ASTPs is more important than attempting an exhaustive coverage of all expressed genes. ASTPs that showed allele-specific expression differences between reciprocally-crossed hybrids (B73/Mo17 and Mo17/B73) were excluded from further analyses to avoid the confounding of differential expression due to parental effects. ASTPs that met these criteria were included in further allele-specific expression analyses (Tables 1, 2, and Supplementary Table 1).
Among the total of 364 unique ASTPs identified from all meristem stages, we found 62–79 (average 73) ASTPs in each meristem stage that matched the selection criteria for allele-specific expression analysis in the hybrid. These ASTPs included those that appeared in more than one stage. Allelic expression of each individual ASTP was analyzed to determine whether they are differentially expressed in the Mo17/B73 hybrids by a t-test. ASTPs were considered as differentially expressed if the allelic expression ratio deviated from 1.0 at a P value <0.1 and FDR < 0.2 (Materials and methods). The fold changes of the allelic expression level of the differentially expressed ASTPs identified by these criteria ranged from nearly 2.0 to mono-allelic expression, where transcript of one parental allele could not be detected. Although no fold-change cut-off was used, of those with a P value <0.1, only a few had an allelic difference ratio less than 2.0 fold, suggesting that the proportion of the differentially expressed genes described may be a conservative estimate.
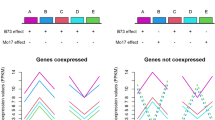
In each meristem stage, approximately 55% of the ASTPs were differentially expressed, that is exhibiting allelic difference in cis-regulation (Table 1), and nearly 15% showed significant expression difference in two or more stages (Supplementary data Table 2). We did not find any ASTPs that had significant allelic expression difference (in the same direction) across all six meristem stages. Even though the difference may not be statistically significant in all six stages, for some genes the allelic expression in the hybrid showed distinct developmental or temporal patterns. Figure 2 shows examples of the differential patterns of allele-specific expression in the developing meristems of F1 hybrid: Mo17 allele dominant expression; B73 allele dominant expression; opposite allele expression between Mo17 and B73, that is, the expression level of one allele increased and the other decreased, as the hybrid meristem developed. The allelic expression patterns during F1 hybrid meristem development provide a view of allelic difference in temporal regulation, in addition to the expression level regulation.
Examples of allelic expression patterns in developing meristems of the F1 hybrid. (a) Mo17 allele dominant expression (polymyositis/scleroderma autoantigen 1, AP004775.3). (b) B73 allele dominant expression (putative fibrillarin, AP004878.3). (c) Opposite allele expression between Mo17 and B73 (Nucleasome assembly factor D protein, AF440223.1). Y-axis: MPSS expression level in PPM, X-axis: developmental stages of shoot apical meristem (SAM), and ear inflorescence meristem (EIM). The error bars are estimated from fitted spline function based on the approximations for true replicates of reciprocal crosses as described in the Materials and methods
Trans-acting effects on allelic expression in the F1 hybrid
Trans-acting factors such as transcriptional factors regulate gene expression via interacting with cis-regulatory elements. While cis-regulatory differences can be tested by subjecting different alleles to one genotype under the influence of a common set of trans-acting factors, trans-acting effects can be assessed by comparing expression of the same allele under different genetic backgrounds. The hypothesis is that if no different trans-acting regulatory effects are involved between the hybrid and inbred backgrounds, the expression of a specific allele is expected to be the same between genotypes; otherwise, an indication of different trans-acting factors present in different genotypes. Therefore, we developed the method to compare expression level of an individual allele in the hybrid with that in the inbred parent, and consider one allele independently from the other. This approach differs from that described in previous studies (Wittkopp et al. 2004; Stupar and Springer 2006; Springer and Stupar 2007a, b), in which trans-acting effect are determined by comparing allelic ratio differences between hybrid and inbred parents.
From the total 364 ASTPs, we selected approximately 100 in each meristem stage based on the background threshold and reciprocal cross criteria, for further analysis between hybrid and inbred parents. By comparing the expression differences of B73 and Mo17 alleles, respectively, with hybrid Mo17/B73, we found 30–45% genes showed significant allelic expression differences between the hybrid and inbred parents, and therefore regulated by different trans-acting effects (Table 2). Allele-specific expression analysis of B73 and Mo17 alleles showed that both inbred parents had similar proportions of genes that were affected by such a trans-acting regulation. However, only a small percentage of genes showed different trans-acting effects in the F1 hybrid from both inbred parents (B73 and Mo17 alleles) (Table 2). This is not surprising knowing that the two parental alleles are often differentially regulated in the F1 hybrid, that is, exhibiting allele-specific expression differences. Among those ASTPs that exhibited either cis- or trans-effect, approximately 50% of them showed both effects, that is, the allelic expression of these genes varied between the hybrid and inbred parents, and also within the hybrid (Supplementary data Table 4). When considering all ASTPs analyzed in each meristem stage, less than 10% had both cis- and trans-effects.
RT-PCR confirmation
We used RT-PCR/WAVE dHPLC method (Guo et al. 2003, 2004) to verify MPSS data of the allele-specific transcript level of three genes expressed in different stages of the SAM and EIM development. RT-PCR validation was not performed in the SAM1 stage due to a lack of the tissue and RNA in this stage (used up in MPSS profiling). MPSS expression of ASTPs was included in the RT-PCR confirmation regardless whether the allelic expression ratio was significantly deviated from 1.0. However, if both alleles had MPSS expression level below 25 PPM, a background cut-off level, the ASTP was not included in the RT-PCR validation. RT-PCR was done in three replicates and the mean is shown in Table 3. In most cases, the allelic expression observed in MPSS data and RT-PCR was biased toward the same direction. A pair-wise correlation analysis between MPSS data and RT-PCR data using the allelic ratio in Table 3 produced a correlation coefficient r = 0.86 (P-value = 0.0004). The results suggest that data from the two platforms are consistent.
Discussion
Although MPSS technology has been used for various genome-wide transcript profiling analyses (Reinartz et al. 2002; Jongeneel et al. 2003, 2005; Meyers et al. 2004a, b; Hedgecock et al. 2007), its application in allele-specific expression profiling has not been reported. In this study, we took advantage of specific features of the MPSS technology and developed a proof-of-concept methodology for genome-wide allele-specific expression analysis in the maize hybrid and parental inbreds. One of the advantages of this methodology over the microarray hybridization platform is that it captures virtually all genes expressed in a given tissue and does not require prior knowledge of sequence data (Brenner et al. 2000a, b). This is particularly important for allele-specific expression analysis in which both parental allele sequences are required and yet, sequence information for both parental alleles is often unavailable in the database. Although genes lacking a DpnII restriction site cannot be detected, the problem can be overcome by digesting the cDNA with an alternative enzyme. Another advantage of this method is that there is no issue of cross-hybridization between alleles as in the case of a hybridization platform. The analysis is dependent on the allele-specific polymorphism between the allelic tag pairs. Because of the highly polymorphic nature of the 3′ un-translated region in the maize genome, we were able to identify nearly 400 allelic tag pairs that potentially represent 400 genes for a genome-wide expression analysis. The accuracy of the ASTPs representing respective alleles of corresponding genes was verified by sequence analysis of 46 genes for both B73 and Mo17 alleles. The ASTPs were confirmed as having exact sequence match in 42 of the 46 genes (91%) (Supplement materials). This is consistent with the high accuracy of matched signature tags reported in MPSS profiling studies of other organisms (Reinartz et al. 2002; Saha et al. 2002; Meyers et al. 2004a, b; Hedgecock et al. 2007). Using allelic tag pairs in this study provides additional verification and would likely increase the accuracy of the tag-gene relationship (Silva et al. 2004). Although identifying genes is not the objective of this study, the result validates that the methodology can be used for this purpose if needed.
Since MPSS profiling sequences a 17-base long tag for each expressed gene, the sequence may be limiting for allele-specific SNP identification and thus, the number of genes for allele-specific expression analysis. By applying additional restriction enzymes in MPSS profiling we can generate more signature tags per gene, more sequence information for allelic SNP identification, and therefore increase the number of genes with SNP markers for allele-specific expression analysis. Alternatively, one could apply the same concept by using other technologies, such as 454 Life Sciences technology (Margulies et al. 2005), which produces a 100–200 base read for each gene, and may significantly increase the allelic SNP discovery. The MPSS application may not differentiate nearly identical paralogs, which occurs in approximately 1% of the genes in the maize genome (Emrich et al. 2007). Nevertheless, this methodology is a unique tool for a genome-wide allele-specific expression profiling in the maize hybrid and inbred parents, and can have a broader application as newer and improved sequencing technologies become available.
We used RT-PCR/WAVE dHPLC to validate the MPSS expression difference of three genes across meristem stages. We selected these genes based on the availability of sequence information of both alleles in the database, the resolvability of the allele sequence polymorphism, and the expression across multiple stages in both SAM and EIM tissues, regardless of expression level. RT-PCR data confirmed the MPSS data in most cases, where the allele expression is biased toward the same direction between the two platforms. Since these genes happened to represent those at the lower end of the distribution of expression level (as a reference, the expression level of the tubulin gene is approximately 2,000 PPM in the MPSS experiment), and considering that lowly expressed genes are generally known as being more sensitive to background or noise, the results from the two platforms were in good agreement. Furthermore, a correlation coefficient r = 0.86 (P-value = 0.0004) between MPSS data and RT-PCR/WAVE data suggests that results from the two platforms were consistent.
Our genome-wide analysis of allele-specific expression in maize hybrid meristems showed that 50–70% of genes exhibited allelic expression variation or allelic difference in cis-regulation. The result is consistent with our previous study of RT-PCR analysis of 15 genes where we found 11 (70%) of them exhibit allelic expression differences in at least one tissue type or one environmental condition (Guo et al. 2004). Other studies report that 70–85% of gene expression differences are attributed to cis-regulatory difference in maize hybrids (Stupar and Springer 2006; Springer and Stupar 2007a, b). A similar percent of cis-regulatory differences have been reported in human (Lo et al. 2003; Pant et al. 2006). These results suggest the abundance of functional cis-regulatory differences between parental alleles in the maize genome. The allelic diversity at the expression level is consistent with the high allelic diversity at the DNA sequence and structural levels (Fu and Dooner 2002; Lai et al. 2005; Morgante et al. 2005; Springer and Stupar 2007a, b).
The wide-spread occurrence of allelic differences in cis-regulation in a heterotic hybrid suggests a prominent role in hybrid gene regulation and potentially heterosis. This notion is consistent with that allelic diversity measured by DNA sequence polymorphisms correlates with heterosis and hybrid performance, assuming the allelic sequence variation contains similar proportions of different modes of regulatory variation among different hybrids. Allelic diversity measured by restriction fragment length polymorphisms or pedigree is positively correlated with hybrid yield and heterosis (Smith et al. 1990; Smith and Smith 1992). Comparing hybrids from crosses of closely related parents (low heterosis) to parents with high genetic diversity (high heterosis), shows that the number of polymorphisms is correlated with heterosis, and a similar proportion of cis- and trans-regulatory variation is involved among different hybrids (Springer and Stupar 2007a, b). Furthermore, our previous study shows that the proportion of genes with differential allele regulation (not an increased level of expression) is positively correlated with yield heterosis based on RNA profiling analysis of a series of 16 hybrids that are produced by crossing inbreds varying in pedigree relationship (Guo et al. 2006). Allelic diversity of inbred parents based on DNA sequence polymorphisms has been increased through historical breeding improvement of commercial hybrids (Feng et al. 2006). The increased genetic diversity may involve, at least in part, the increased allelic differences in cis-regulation. Some of the supporting evidences is that low allelic diversity in expression (mono-allelic expression) is associated with old, low yielding hybrids, whereas bi-allelic expression is associated with a newer and high yielding hybrids (Guo et al. 2004, 2006). As an example, alleles in the hybrid may show differential response to drought stress and plant density stress, one allele may be more favorable than the other in one particular environment (e.g., drought stress) and vise versa in others (e.g., density stress) (Guo et al. 2004). Such allelic diversity may provide the hybrid with biological alternatives or plasticity in responding to environmental stresses, or developmental cues.
Further data suggest that in addition to its role in regulating allelic differences in expression level, cis-regulatory differences may impact allelic expression in other dimensions, such as temporal patterns or differential developmental patterns during meristem development (Fig. 2, Supplementary data Table 2). Changes in temporal and spatial expression as results of cis-regulatory differences have been reported in plants (Wang et al. 1999; Cong et al. 2002) and animals (Wray et al. 2003). Our findings of allelic differences in developmental expression regulation in maize hybrids from this and previous studies (Guo et al. 2003, 2004) suggest the potential roles of cis-regulatory differences in modulating hybrid gene expression beyond the expression level. Hybrid gene expression may exceed its inbred parents in more than one dimension, such as the hybrid may have extended period of expression, broader spatial distribution, or more adapted environments than its inbred parents. The cumulative effects from such an allelic diversity may result in an overall non-allelic additive effect on hybrid performance and heterosis (Birchler et al. 2006).
We also developed a computational protocol to evaluate the trans-acting effects on allele-specific expression in the maize hybrid. Using this method, we found 30–45% of the genes were affected by different trans-acting effect in the hybrid as compared to the inbred parents. A higher percentage of genes exhibiting cis-regulatory differences (45–70%) than trans-acting effects (30–45%) was observed in this study. Similar results have been reported in other studies (Springer and Stupar 2007a, b) where cis-regulatory differences account for 70% of gene expression differences and only 25% of the gene expression differences are regulated by either trans-regulatory difference only, or involve both cis-acting and trans-acting regulatory variation. It is possible that pre-selection of genes with sequence polymorphisms that are required for the allele-specific expression analysis may have favored those genes with cis-regulatory differences (Springer and Stupar 2007a, b). Since cis-acting effects were detected by comparing two alleles expressed in a hybrid, where variations in cellular environments, external environmental cues, developmental or tissue-specific differences are all minimized; such comparison may be more sensitive in detecting cis-acting differences. In trans-acting regulatory analysis, however, different genotypes are involved, and additional variables may reduce the detection power. However, the data may simply indicate that cis-regulatory polymorphisms play a more predominant role in hybrid gene regulation than trans-acting regulation.
Previous studies (Wittkopp et al. 2004; Stupar and Springer 2006; Springer and Stupar 2007a, b) detect trans-acting effects based on allelic ratio differences between a hybrid and inbred parents. The method described in this study detects trans-acting effects by directly comparing allelic expression between hybrid and inbred parents. This approach not only provides a global measurement of trans-acting effects on hybrid gene regulation, but also a quantitative analysis of gene expression in the hybrid with its inbred parents at the allele-specific level. Traditionally, comparison of transcript level between hybrid and inbred parents has been made at the total expression level, without discriminating the parental alleles (Auger et al. 2005; Bao et al. 2005; Vuylsteke et al. 2005; Huang et al. 2006; Swanson-Wagner et al. 2006). In this case, an allelic additive expression level in the hybrid may not be the sum of each parental allele-specific transcript. For example, if a gene is expressed at 100 and 200 PPM in parent A and B, respectively, a mid-parent (additive) expression level predicts 150 PPM in the F1 hybrid. However, the contribution of the two parental alleles may not necessarily be 50 PPM from parent A and 100 PPM from B, respectively. The present allele-specific expression analysis took consideration of the allelic contribution to gene expression in the hybrid and therefore, revealed how the alleles are differently regulated in the hybrid as compared to the inbred parents. Understanding gene regulation in the hybrid at the higher resolution provides new insights to the molecular basis of heterosis.
Non-allelic additive expression in the hybrid indicates different trans-effect in different genotypes (hybrid and inbred parents). What are the roles of trans-acting regulation in heterosis expression? Non-additive expression of total transcript or over-dominant gene expression in the hybrid has been observed in numerous studies in the effort to understand the molecular basis of heterosis (Auger et al. 2005; Bao et al. 2005; Vuylsteke et al. 2005; Huang et al. 2006; Swanson-Wagner et al. 2006). It has been tempting to relate gene expression of this category to heterosis, however there is no evidence to support a correlation of such expression with heterosis of any phenotype (Gibson and Weir 2005). In our previous RNA profiling study of a series of 16 hybrids that vary in their degree of yield heterosis, we do not find any correlation between the number of non-additively expressed genes with hybrid performance and heterosis (Guo et al. 2006). This data does not support the presumed roles of over-dominant gene expression or trans-acting regulation in heterosis. However, since gene regulation is the result of cis- and trans-interaction, the roles of trans-acting effects may be through co-selection with cis-regulatory changes for optimized gene regulation, and therefore contribute to heterosis expression. A connection of any mode of gene regulation to heterosis would require further validation. This study attempted to establish the concept and methodology of applying allele-specific expression analysis to address gene regulation in the hybrid. The data revealed a genome-wide impact of the cis- and trans-regulation and the large number of genes involved, however, a direct connection of particular classes of genes and their modes of regulation to heterosis cannot be made within the scope of this study. Linking specific genes to heterosis can be achieved by applying this tool in future experiments that involve phenotypic expression of heterosis.
References
Adams KL, Wendel JF (2005) Allele-specific, bidirectional silencing of an alcohol dehydrogenase gene in different organs of interspecific diploid cotton hybrids. Genetics 171:2139–2142
Auger DL, Gray AD, Ream TS, Kato A, Coe EH Jr, Birchler JA (2005) Nonadditive gene expression in diploid and triploid hybrids of maize. Genetics 169:389–397
Bao J, Lee S, Chen C, Zhang X, Zhang Y, Liu S, Clark T, Wang J, Cao M, Yang H, Wang SM, Yu J (2005) Serial analysis of gene expression study of a hybrid rice strain (LYP9) and its parental cultivars. Plant Physiol 138:1216–1231
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Bickel DR (2004) Error-rate and decision-theoretic methods of multiple testing: which genes have high objective probabilities of differential expression? Stat Appl Genet Mol Biol 3:8
Birchler JA, Auger DL, Riddle NC (2003) In search of the molecular basis of heterosis. Plant Cell 15:2236–2239
Birchler JA, Yao H, Chudalayandi S (2006) Unraveling the genetic basis of hybrid vigor. Proc Natl Acad Sci USA 103:12957–12958
Brenner S, Johnson M, Bridgham J, Golda G, Lloyd DH, Johnson D, Luo S, McCurdy S, Foy M, Ewan M, Roth R, George D, Eletr S, Albrecht G, Vermaas E, Williams SR, Moon K, Burcham T, Pallas M, DuBridge RB, Kirchner J, Fearon K, Mao J, Corcoran K (2000a) Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat Biotechnol 18:630–634
Brenner S, Williams SR, Vermaas EH, Storck T, Moon K, McCollum C, Mao JI, Luo S, Kirchner JJ, Eletr S, DuBridge RB, Burcham T, Albrecht G (2000b) In vitro cloning of complex mixtures of DNA on microbeads: physical separation of differentially expressed cDNAs. Proc Natl Acad Sci USA 97:1665–1670
Ching A, Caldwell K, Jung M, Dolan M, Smith O, Tingey S, Morgante M, Rafalski A (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19
Cong B, Liu J, Tanksley SD (2002) Natural alleles at a tomato fruit size quantitative trait locus differ by heterochronic regulatory mutations. Proc Natl Acad Sci USA 99:13606–13611
Cowles CR, Hirschhorn JN, Altshuler D, Lander ES (2002) Detection of regulatory variation in mouse genes. Nat Genet 32:432–437
Doebley J (2004) The genetics of maize evolution. Annu Rev Genet 38:37–59
Efron B, Tibshirani R, Storey JD, Tusher V (2001) Empirical Bayes analysis of a microarray experiment. J Am Stat Assoc 96:1151–1160
Emrich SJ, Li L, Wen TJ, Yandeau-Nelson MD, Fu Y, Guo L, Chou HH, Aluru S, Ashlock DA, Schnable PS (2007) Nearly identical paralogs: implications for maize (Zea mays L.) genome evolution. Genetics 175:429–439
Feng L, Sebastian S, Smith S, Cooper M (2006) Temporal trends in SSR allele frequencies associated with long-term selection for yield of maize. Maydica 51:293–300
Fu H, Dooner HK (2002) Intraspecific violation of genetic colinearity and its implications in maize. Proc Natl Acad Sci USA 99:9573–9578
Gibson G, Weir B (2005) The quantitative genetics of transcription. Trends Genet 21:616–623
Guo M, Rupe MA, Danilevskaya ON, Yang X, Hu Z (2003) Genome-wide mRNA profiling reveals heterochronic allelic variation and a new imprinted gene in hybrid maize endosperm. Plant J 36:30–44
Guo M, Rupe MA, Zinselmeier C, Habben J, Bowen BA, Smith OS (2004) Allelic variation of gene expression in maize hybrids. Plant Cell 16:1707–1716
Guo M, Rupe MA, Yang X, Crasta O, Zinselmeier C, Smith OS, Bowen B (2006) Genome-wide transcript analysis of maize hybrids: allelic additive gene expression and yield heterosis. Theor Appl Genet 113:831–845
Hedgecock D, Lin JZ, Decola S, Haudenschild CD, Meyer E, Manahan DT, Bowen B (2007) Transcriptomic analysis of growth heterosis in larval Pacific oysters (Crassostrea gigas). Proc Natl Acad Sci USA 104:2313–2318
Huang Y, Zhang L, Zhang J, Yuan D, Xu C, Li X, Zhou D, Wang S, Zhang Q (2006) Heterosis and polymorphisms of gene expression in an elite rice hybrid as revealed by a microarray analysis of 9198 unique ESTs. Plant Mol Biol 62:579–591
Jain N, Thatte J, Braciale T, Ley K, O’Connell M, Lee JK (2003) Local-pooled-error test for identifying differentially expressed genes with a small number of replicated microarrays. Bioinformatics 19:1945–1951
Jongeneel CV, Iseli C, Stevenson BJ, Riggins GJ, Lal A, Mackay A, Harris RA, O’Hare MJ, Neville AM, Simpson AJ, Strausberg RL (2003) Comprehensive sampling of gene expression in human cell lines with massively parallel signature sequencing. Proc Natl Acad Sci USA 100:4702–4705
Jongeneel CV, Delorenzi M, Iseli C, Zhou D, Haudenschild CD, Khrebtukova I, Kuznetsov D, Stevenson BJ, Strausberg RL, Simpson AJ, Vasicek TJ (2005) An atlas of human gene expression from massively parallel signature sequencing (MPSS). Genome Res 15:1007–1014
Knight JC (2004) Allele-specific gene expression uncovered. Trends Genet 20:113–116
Lai J, Li Y, Messing J, Dooner HK (2005) Gene movement by Helitron transposons contributes to the haplotype variability of maize. Proc Natl Acad Sci USA 102:9068–9073
Lo HS, Wang Z, Hu Y, Yang HH, Gere S, Buetow KH, Lee MP (2003) Allelic variation in gene expression is common in the human genome. Genome Res 13:1855–1862
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Meyers BC, Lee DK, Vu TH, Tej SS, Edberg SB, Matvienko M, Tindell LD (2004a) Arabidopsis MPSS. An online resource for quantitative expression analysis. Plant Physiol 135:801–813
Meyers BC, Vu TH, Tej SS, Ghazal H, Matvienko M, Agrawal V, Ning J, Haudenschild CD (2004b) Analysis of the transcriptional complexity of Arabidopsis thaliana by massively parallel signature sequencing. Nat Biotechnol 22:1006–1011
Morgante M, Brunner S, Pea G, Fengler K, Zuccolo A, Rafalski A (2005) Gene duplication and exon shuffling by helitron-like transposons generate intraspecies diversity in maize. Nat Genet 37:997–1002
Morley M, Molony CM, Weber TM, Devlin JL, Ewens KG, Spielman RS, Cheung VG (2004) Genetic analysis of genome-wide variation in human gene expression. Nature 430:743–747
Nadon R, Shi P, Skandalis A, Woody E, Hubschle H, Susko E, Rghei N, Ramm P (2001) Statistical inference methods for gene expression arrays. In: Bittner ML, Chen Y, Dorsel AN, Dougherty ER (eds) Proceedings of SPIE 2001, Microarrays: Optical Technologies and Informatics, vol 4266. SPIE, San Jose, CA, USA, pp 46–55
Pant PV, Tao H, Beilharz EJ, Ballinger DG, Cox DR, Frazer KA (2006) Analysis of allelic differential expression in human white blood cells. Genome Res 16:331–339
Prud’homme B, Gompel N, Rokas A, Kassner VA, Williams TM, Yeh SD, True JR, Carroll SB (2006) Repeated morphological evolution through cis-regulatory changes in a pleiotropic gene. Nature 440:1050–1053
R Development Core Team (2005) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, URL http://www.R-project.org
Reinartz J, Bruyns E, Lin JZ, Burcham T, Brenner S, Bowen B, Kramer M, Woychik R (2002) Massively parallel signature sequencing (MPSS) as a tool for in-depth quantitative gene expression profiling in all organisms. Brief Funct Genomic Proteomic 1:95–104
Saha S, Sparks AB, Rago C, Akmaev V, Wang CJ, Vogelstein B, Kinzler KW, Velculescu VE (2002) Using the transcriptome to annotate the genome. Nat Biotechnol 20:508–512
Silva AP, De Souza JE, Galante PA, Riggins GJ, De Souza SJ, Camargo AA (2004) The impact of SNPs on the interpretation of SAGE and MPSS experimental data. Nucleic Acids Res 32:6104–6110
Smith OS, Smith JSC (1992) Measurement of genetic diversity among maize hybrids; a comparison of isozymic, RFLP, pedigree, and heterosis data. Maydica 37:53–60
Smith OS, Smith JSC, Bowen SL, Tenborg RA, Wall SJ (1990) Similarities among a group of elite maize inbreds as measured by pedigree, F1 grain yield, heterosis, and RFLPs. Theor Appl Genet 80:833–840
Springer NM, Stupar RM (2007a) Allele-specific expression patterns reveal biases and embryo-specific parent-of-origin effects in hybrid maize. Plant Cell 19:2391–2402
Springer NM, Stupar RM (2007b) Allelic variation and heterosis in maize: how do two halves make more than a whole? Genome Res 17:264–275
Stolovitzky GA, Kundaje A, Held GA, Duggar KH, Haudenschild CD, Zhou D, Vasicek TJ, Smith KD, Aderem A, Roach JC (2005) Statistical analysis of MPSS measurements: application to the study of LPS-activated macrophage gene expression. Proc Natl Acad Sci USA 102:1402–1407
Storey JD (2002) A direct approach to false discovery rates. J R Stat Soc Ser B Stat Methodol 64:479–498
Stupar RM, Springer NM (2006) Cis-transcriptional variation in maize inbred lines B73 and Mo17 leads to additive expression patterns in the F1 hybrid. Genetics 173:2199–2210
Swanson-Wagner RA, Jia Y, DeCook R, Borsuk LA, Nettleton D, Schnable PS (2006) All possible modes of gene action are observed in a global comparison of gene expression in a maize F1 hybrid and its inbred parents. Proc Natl Acad Sci USA 103:6805–6810
Vuylsteke M, van Eeuwijk F, Van Hummelen P, Kuiper M, Zabeau M (2005) Genetic analysis of variation in gene expression in Arabidopsis thaliana. Genetics 171:1267–1275
Wang RL, Stec A, Hey J, Lukens L, Doebley J (1999) The limits of selection during maize domestication. Nature 398:236–239
West MA, Kim K, Kliebenstein DJ, van Leeuwen H, Michelmore RW, Doerge RW, St Clair DA (2006) Global eQTL mapping reveals the complex genetic architecture of transcript level variation in arabidopsis. Genetics 173:1441–1450
Wittkopp PJ, Haerum BK, Clark AG (2004) Evolutionary changes in cis and trans gene regulation. Nature 430:85–88
Wray GA (2007) The evolutionary significance of cis-regulatory mutations. Nat Rev Genet 8:206–216
Wray GA, Hahn MW, Abouheif E, Balhoff JP, Pizer M, Rockman MV, Romano LA (2003) The evolution of transcriptional regulation in eukaryotes. Mol Biol Evol 20:1377–1419
Yan H, Yuan W, Velculescu VE, Vogelstein B, Kinzler KW (2002) Allelic variation in human gene expression. Science 297:1143
Acknowledgements
The Authors thank Jo Ann Dieter and Brian Zeka for assistance with tissue and RNA preparation, Susan Nilles and Shifu Zheng for assistance with growing plants and Li Zeng for statistical analysis, and Lynx Therapeutics, now Solexa/Illumina for the support of MPSS™ technology. The authors also thank the anonymous reviewers that have improved the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Guo, M., Yang, S., Rupe, M. et al. Genome-wide allele-specific expression analysis using Massively Parallel Signature Sequencing (MPSS™) Reveals cis- and trans-effects on gene expression in maize hybrid meristem tissue. Plant Mol Biol 66, 551–563 (2008). https://doi.org/10.1007/s11103-008-9290-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9290-z