Abstract
For the first time, numerous sequences of Copia and Gypsy retrotransposons from the Cucumis melo genome have been obtained and analyzed. Phylogenetic analyses of sequences of both types of long terminal repeat (LTR) retrotransposons were carried out. The melon genome contains approximately 20,000 Gypsy and 6,800 Copia elements, comprising about 26% of its total size. Starting from a retrotransposon fragment, we have cloned and characterized an entire melon retrotransposon, named Reme1, which is 5,149 bp long. Reme1 belongs to the Superfamily Copia retrotransposons by its protein domain order and sequence similarity to other Copia elements of dicotyledons. The haploid genome of melon (var. “Piel de Sapo”) contains about 120 copies of Reme1. Several copies of Reme1 are transcriptionally active, although at low levels, in melon leaves as analyzed by reverse-transcription PCR (RT-PCR) and sequencing. However, the transcript pool is considerably increased when melon leaves are treated with UV light, as has been seen for various retroelements in many organisms. The cDNAs of Reme1 transcripts showed less diversity than do Reme1 genomic sequences, suggesting that a subfamily of these elements is differentially responsive to UV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transposable elements are ubiquitous in the plant kingdom and are present in high copy number in most plants, comprise more than 50% of many large genomes (reviewed in Kumar and Bennetzen 1999), generally distributed as interspersed repetitive sequences throughout most of the length of host chromosomes. They are divided into two classes. Class I transposable elements or retrotransposons replicate via an RNA intermediate. Class II transposons move as a DNA segment by a “cut-and-paste” mechanism (Wicker et al. 2007). The long terminal repeat retrotransposons, which contain long terminal repeats at their ends, resemble retroviruses in their organization and coding capacity. They are further divided into two major superfamilies, Copia and Gypsy, differing in the relative order of the integrase (int) and reverse transcriptase (rt) domains (Wicker et al. 2007). Their transcripts, expressed by a promoter in the LTR, are reverse-transcribed by their RT into cDNA. The cDNAs are ultimately integrated into the chromosome by integrase (INT), which is also encoded by the retrotransposon itself (Bingham and Zachar 1989; Boeke and Corces 1989). This replicative mode of transposition can readily increase the element copy number and thereby greatly increase the plant genome size (Kumar 1996; SanMiguel and Bennetzen 1998; Vitte and Panaud 2005).
The replicative transposition of retrotransposons, combined with the error-prone nature of transcription and reverse transcription, generates families or “quasispecies” (Casacuberta et al. 1995) of related sequences. The resulting heterogeneity of genomic retrotransposon copies has been well established in many plant species (Flavell et al. 1992a, b; Pearce et al. 1996a, b; Suoniemi et al. 1998; Friesen et al. 2001). Phylogenetic analyses of this variability have made it possible to see the expansion and diversification of retrotransposon populations before the divergence of the host species. This has been studied both within genera including Nicotiana (Vernhettes et al. 1998) and Zea (García-Martínez and Martínez-Izquierdo 2003); and between genera such as Vicia and Pisum, which share related retrotransposon sequences from a heterogeneous group of Copia retrotransposons (Pearce et al. 2000). Sequence variability is found not only in the protein-coding domains, but also in the LTR promoters, leading to the possibility of differential expression of retrotransposons between tissues and conditions (Beguiristain et al. 2001). This can have major evolutionary consequences for the population structure of retrotransposons.
Nevertheless, a few years ago, most retrotransposons were thought to be transcriptionally inactive (Kumar and Bennetzen 1999) or silent in somatic tissues but active during certain stages of plant development or under stress (Grandbastien 1998). Among the stress agents, UV light was reported to activate animal short interspersed nuclear elements (SINEs) (Rudin and Thompson 2001) and LTR retrotransposons in both Xenopus (Shim et al. 2000) and plants (e.g., the Copia element OARE1 of oat; Kimura et al. 2001). McClintock (1984) viewed the UV light -induced activation of maize transposons (Wessler 1996; Walbot 1999) as “genomic stress.” Furthermore, the increasing use of RT-PCR to follow transcription (Neumann et al. 2003) offers the possibility to detect the rare transcripts of retrotransposons, having less dramatic transcriptional activation, which were previously considered inactive using less sensitive filter hybridization methods.
Little is known about the transposable elements or their role in the genome organization of the melons (Cucumis melo L.) and Cucurbitaceae generally, even though this family includes the most economically important horticultural crops following those of the Solanaceae. One Gypsy and two Copia elements were previously reported in work focusing on the sequencing of a region containing a resistance gene (van Leeuwen et al. 2003, 2005). Melon is a diploid species (2n = 2x = 24), with a relatively small genome of 450 Mb (Arumuganathan and Earle 1991), similar in size to that of rice. The question of the presence of active retrotransposons in a small genome is of interest, because of the role of retrotransposons in increasing genome size. We therefore decided to investigate the characteristics and contribution of retrotransposons in the melon genome. We designed degenerate primer pairs corresponding to a consensus sequence from a conserved rt domain of Superfamily Copia elements (Flavell et al. 1992a) and to conserved rt and int domains of plant Gypsy elements (Suoniemi et al. 1998). These were used to amplify products from two different, divergent melon lines. Derived from this research, an entire Copia element, Reme1, was cloned and characterized, including its transcriptional activity, which is induced upon UV light treatment.
Materials and methods
Plant material and DNA extraction
Two melon (Cucumis melo L.) cultivars, “Piel de Sapo” T-111 (PS), an important Spanish commercial line, and the exotic Korean accession “Shongwan Charmi” PI 161375 (Co), a non-commercial cultivar from Korea, were used throughout this work. These were provided by Semillas Fitó S.A. (Barcelona, Spain). Additional material was used from the following plants belonging to the Cucurbitaceae: cucumber (Cucumis sativus, SATH, PI215589 Hardwickii cultivar), Cucumis metuliferus (PI482462), Cucumis africanus (PI203974 line), Cucumis pustulatus (PI343699), Cucumis prophetarum (PI193967), Cucumis ficifolius (FIC, PI196844), watermelon (Citrullus lanatus, Fitó line, Klondike), zucchini (Cucurbita pepo, Fitó line, JK601) and squash (Cucurbita maxima, NSL 20182 line). Plant genomic DNA was extracted from young leaves using the “Dneasy Plant Mini Kit (50)” (Qiagen). The extraction yields ranged from 20 to 30 μg of pure DNA per 100 mg of fresh tissue.
Retrotransposon DNA amplification, cloning and sequencing
Fragments of both Copia- and Gypsy retrotransposons were amplified from melon genomic DNA by the polymerase chain reaction (PCR), using degenerate primers from conserved motifs of Copia RTs as described by Flavell et al. (1992a) and from conserved motifs of Gypsy RTs and INTs as described by Suoniemi et al. (1998).
The inverse PCR (I-PCR) technique was used for isolating an entire retrotransposon using one of the Copia reverse transcriptase clones (Cps21) as the basis. For this purpose, a melon bacterial artificial chromosome (BAC) clone that hybridized with Cps21 was digested with EcoRI. The DNA was diluted to 10 μg/ml in ligase buffer; ligation was carried out with 1 U of T4 ligase at 16°C. After circularization by ligation, the DNA was diluted again and amplified with outward-facing primers designed from the selected Copia clone Cps21. The I-PCR conditions were: 95°C for 3 min; 10 cycles of 94°C for 30 s, 59°C for 1 min, a ramp of 1°C per 3 s and 68°C for 7 min; 20 cycles of 94°C for 30 s, 61°C for 1 min, a ramp of 1°C per 3 s, and 68°C for 7 min; a final cycle of 68°C for 7 min. The oligos used for the I-PCR were: Cps21d’, 5′-GAGACTTGTTTAACCTGCAAACC-3′ and Cps21r’, 5′-GCTAAATGCAGACCCTTGTGC-3′. The priming sites flank nt 3433–3552 in Reme1. The amplified fragments were cloned using the pGEM-T kit (Promega) and sequenced with the PRISM kit (Applied Biosystems) on an automatic sequencer (ABI PRISM 377, Applied Biosystems). Reme1 retrotransposons were amplified from species of the Cucurbitaceae family with primers for the int region, 5′-GATTTCTCCAGAAGGTGTTG-3′ (forward) and 5′-ATTGCAGTTGATGGAGAACG-3′ (reverse), matching nt 2193–2531 in Reme1, and with 5′-GTGCTACATTTGACTTACACC-3′ (forward) and 5′-GTCTCAGTGAATTTGCATTCC-3′ (reverse), matching the rt region at nt 3309–3733 bp.
Sequence analysis
Sequences were aligned using ClustalW (Thompson et al. 1994) and Bioedit (Hall 1999) software. Phylogenetic trees were constructed by the Neighbor-Joining method (Saitou and Nei 1987) with the “complete deletion” option. Poisson genetic distance for amino acid sequences and Kimura-2-parameters for genetic distance between nucleotide sequences were used. Phylogenetic trees were displayed with the MEGA 2.1 program (Kumar et al. 2001).
Copy number determination by slot blot hybridization
Retrotransposon copy number in melon was determined as described previously (Aledo et al. 1995; García-Martínez and Martínez-Izquierdo 2003). Briefly, DNA was blotted onto nylon filters by vacuum filtration through the slots of a manifold device (Hoefer PR600, Amersham Pharmacia Biotech). Filters were hybridized in 0.25 M Na2HPO4, 7% SDS, 1 mM EDTA, pH 7.2, overnight at 65°C. Hybridized filters were consecutively washed with 2× SSC (1× SSC is 0.15 M NaCl and 0.015 sodium citrate) and 0.1% SDS (10 min, at room temperature), twice in 2× SSC and 0.1% SDS (15 min, 65°C), twice in 0.5× SSC and 0.1% SDS (15 min, 65°C), and twice in 0.1× SSC and 0.1% SDS (15 min, 65°C). The 32P present in the probes (described below) was quantified by exposure of filters to an imaging plate followed by scanning in a PhosphoImager (Personal Molecular Imager FX System, BIORAD).
Probes for detection of genomic Copia and Gypsy populations were radioactively labeled by PCR using degenerate primers (Flavell et al. 1992a; Suoniemi et al. 1998). The following primers were used to amplify probes for the various Reme1 regions PCR using radioactive dNTPs: gag region (nucleotide positions 735–1146, in Reme1), 5′-TGGAAGTTGAAGATGAAAGC-3′ (forward) and 5′-ATCATACGAATCAGGAAGAC-3′ (reverse); int region, 5′-GATTTCTCCAGAAGGTGTTG-3′ (forward) and 5′-ATTGCAGTTGATGGAGAACG-3′ (reverse); LTRs (nucleotide positions 55–432 in Reme1), 5′-CCATTGGATCTCACACTTTC-3′ (forward) and 5′-GAGAGGATAGAACACAAGG-3′ (reverse).
Genomic copy number was calculated on the basis of the hybridization signal of the genomic DNA compared with the control DNA on the slot blot as follows: copies ng−1 = genomic PSL ng−1 × fragment copies × fragment PSL−1 (PSL stands for photo-stimulated luminescence units, the output unit for exposure of the phosphoimager screens). Retrotransposon copies per DNA quantity in ng were converted to copy number per genome using the melon genome size. A replicate for each amount of genomic DNA as well as of the amplified Copia Cps21 and the Gypsy Gps46 clones were used to determine average values of retrotransposon copy number. The Copia and Gypsy clones were labeled by random priming.
Southern blot hybridization analysis
Agarose gel electrophoresis of restricted genome DNA from melon and Southern blotting to membranes were performed as described by Oliver et al. (2001). The membranes (kindly provided by IRTA) were hybridized with radioactively labeled retrotransposon probes and washed as described above for slot blot hybridization. Membranes were exposed to films at −80°C for the appropriate times.
RNA extraction
Total RNA was extracted from leaves of PS, either treated or not with UV light, by the guanidine hydrochloride method (Logemann et al. 1987). Plant material was frozen in liquid nitrogen and ground to a fine paste. Three ml of Z6 Buffer (8 M guanidine hydrochloride, 20 mM MES, 20 mM EDTA, pH 8.0, 50 mM β-Mercaptoethanol) and 3 ml phenol:chloroform:isoamyl alcohol (24:24:1) per g of material were then added and this mixture centrifuged 30 min at 4°C, at 13,000 rpm. Following centrifugation, 0.7 volumes ethanol and 0.2 volumes 1 M acetic acid were added to the removed upper phase, which was then incubated on ice for 30 min. This mixture was centrifuged 15 min at 13,000 rpm, the RNA pellet washed twice with pre-cooled 3 M sodium acetate (pH 5.2) and the pellet re-centrifuged 15 min at 13,000 rpm at 4°C. The pellet was washed with cold 70% ethanol, dissolved in DEP water and stored at −70°C.
RT-PCR amplification
After DNAseI digestion (Ambion), a PCR reaction with a reverse transcription step (RT-PCR) reaction was performed with the OneStep RT-PCR kit (Qiagen), under the following conditions: 30 min at 50°C, 16 min at 95°C, 36 cycles of 30 s at 94°C, 30 s at 52°C, 2 min at 72°C, 10 min at 72°C. The primers used to perform RT-PCR from the Reme1 sequence were as follows: 5′UTR (Reme1 nt positions 286–536), 5′-CTTCTTAGAGAGCTTTGTATCC-3′ (forward) and -5′-CTTGGCTTTGATACCACTTG-3′ (reverse); gag, 5′-TGGAAGTTGAAGATGAAAGC-3′ (forward) and 5′-ATCATACGAATCAGGAAGAC-3′ (reverse); int region, 5′-GATTTCTCCAGAAGGTGTTG-3′ (forward) and 5′-ATTGCAGTTGATGGAGAACG-3′ (reverse); rt region, 5′-GTGCTACATTTGACTTACACC-3′ (forward) and 5′-GTCTCAGTGAATTTGCATTCC-3′ (reverse). Two negative controls, in which the reaction was carried out either without RNA and skipping the step of reverse transcription, were included. In the RT-PCR experiment where an internal control gene from melon, as the elongation factor-1α (EF-1α) like gene (Gonzalez-Ibeas et al. 2007; Tremousaygue et al. 1997), was used, the conditions were: 5′UTR, gag and rt, 35 cycles and 54°C annealing, int, 35 cycles and 56°C annealing and EF-1α, 30 cycles and 60°C annealing. In the case of EF-1α gene the following primers were used: 5′-GGACATCGTGACTTTATCAAGAAC-3′ (forward) and 5′-CTTGGAGTATTTGGGAGTGGTG-3′ (reverse).
PCR amplification of Reme1 domains
In order to obtain a set of sequences from various regions of Reme1, PCR was carried out. The primers for this were: LTR region, 5′-CCATTGGATCTCACACTTTC-3′ (forward) and 5′-GAGAGGATAGAACACAAGG-3′ (reverse); 5′UTR region, 5′-CTTCTTAGAGAGCTTTGTATCC-3′ (forward) and -5′-CTTGGCTTTGATACCACTTG-3′ (reverse); gag region, 5′-TGGAAGTTGAAGATGAAAGC-3′ (forward) and 5′-ATCATACGAATCAGGAAGAC-3′ (reverse); int region, 5′-GATTTCTCCAGAAGGTGTTG-3′ (forward) and 5′-ATTGCAGTTGATGGAGAACG-3′ (reverse); rt region, 5′-GTGCTACATTTGACTTACACC-3′ (forward) and 5′-GTCTCAGTGAATTTGCATTCC-3′ (reverse).
Results
Identification and analysis of LTR-retroelements in melon
Degenerate primers designed from conserved motifs of the RT domain of Superfamily Copia retrotransposons (Flavell et al. 1992a) were used to amplify corresponding segments from melon genomic DNA by PCR. Two melon varieties were used: “Pinyonet Piel de Sapo” T-111 (PS) and a Korean variety, “Songwhan Charmi PI161375” (Co). We took the same approach to obtain segments of Gypsy elements in the same two melon varieties, but in this case employed a pair of degenerate primers spanning from rt to int (Suoniemi et al. 1998).
Thirty-two RT clones of about 266 bp, the expected size from Copia elements, were sequenced. For Gypsy elements, we obtained 15 clones of around 1,500 bp, matching the distance from the rt to int priming sites, and sequenced the 5′ and 3′ ends. The conceptual open reading frames (ORFs) of each of the sequences corresponded to the Copia and Gypsy motifs expected from the primer pairs. Phylogenetic analyses of the conceptual translations (Fig. 1) show that both PS and Co melon retrotransposon sequences are distributed throughout the trees without independent clustering by plant variety. These results suggest that the retrotransposons represented by the cloned segments had propagated and diverged prior to the separation of the two melon varieties.
Phylogenetic trees from predicted translations of coding regions of Copia and Gypsy retrotransposon elements from melon. (a) Copia RT region. (b1) Gypsy RT region. (b2) Gypsy INT region. Sequences are labeled according to their source, either “Piel de Sapo” (PS) or Korean Songwhan Charmi (Co) melon varieties, preceded by a “C” for Copia or a “G” for Gypsy. Clones with insertions/deletions are indicated by a rectangle and clones with insertions/delections plus stop codons are shaded and underlined. Trees were constructed using the Neighbor-Joining (Saitou and Nei 1987) method from amino acid sequence alignments corresponding to DNA fragments of Copia RT. Trees were displayed with the MEGA2 program, showing bootstrap values from 1,000 replicates. Horizontal distances are proportional to evolutionary distances according to the scale shown on the bottom
Among the Copia sequences (Fig. 1a), group I members were very similar to one another. All have uninterrupted ORFs for the cloned rt segments. This suggests that they have spread recently and constitute a family of retrotransposons. Group II is more heterogeneous and contains several well-supported sub-clades. The rt segment of three well-studied, Superfamily Copia LTR retrotransposons, Tnt1 from tobacco (Grandbastien et al. 1989), Copia from Drosophila melanogaster (Mount and Rubin 1985) and Ty1 from yeast (Warmington et al. 1985) were included in the analysis of elements from the Copia superfamily of melon. Tnt1 shows affinity to one of the Group II sub-clades; the yeast and Drosophila are distinct from Group I but are on long branches and can be considered outgroups. Analyses of the Gypsy clones from melon (Fig. 1b) showed two strongly supported clades, one major and one minor, for both rt and int regions. Cure1, a Gypsy LTR-retrotransposon from melon (van Leeuwen et al. 2003), is on a separate, robust branch but still within Group II of the Gypsy elements (Fig. 1b).
Copy number and pattern of distribution of LTR retrotransposons in melon
Given the relatively small size of its genome, the abundance of retrotransposons in melon was examined. Copy number was determined by genomic reconstruction in the two melon varieties used in this work, using slot blot hybridization. The total numbers of Copia and Gypsy elements was estimated by probing with the multiple products amplifiable from PS genomic DNA with degenerate primers respectively for the rt of Copia (Flavell et al. 1992a) and the rt-int region of Gypsy (Suoniemi et al. 1998). This approach detected 20,000 Gypsy and 6,800 Copia elements (results not shown) in the PS melon genome. Together, these represent around 26% of the melon genome if we assume that the average retrotransposon size is around 5 kb (e.g., similar in size to Tnt1) for Copia elements and 8 kb for Gypsy elements and if we discount solo LTRs and other fragments as well as highly divergent elements that would not be detected by the probes.
We also estimated the abundance of elements related to the partial Copia clone Cps21 and Gypsy clone Gp46 respectively at 1,000 and 3,800 copies per haploid genome (results not shown). The int and gag domains were used as probes to examine the copy number of the full-length Reme1, isolated with the aid of Cps21 as described below. These yielded an estimate of 116 copies per haploid genome, or 0.11% of the total genome. The average number estimated using the LTR as the probe was around twice that obtained for the internal domain, indicating that Reme1 has relatively few solo LTRs, which can be derived from full-length elements by LTR:LTR recombination (Shirasu et al. 2000), in the haploid genome.
The distribution pattern of LTR retrotransposons in the melon genome was studied by Southern hybridization (Fig. 2). PS and Co melon DNAs were digested with Bst NI, Bam HI, Eco RI and Hind III restriction enzymes and hybridized with Cps21 (Copia retrotransposon) or Gps46 (Gypsy retrotransposon) probes, which matched the rt and rt-int region of the elements respectively. The two probes gave very different hybridization patterns. The Cps21- Copia probe (Fig. 2a) detects discrete bands that are mostly smaller than, or just larger than, the expected full length of an LTR-retrotransposon. Most bands are of similar size in both cultivars, excepting EcoRV and HindIII, which appear to detect internal polymorphisms within the Cps21 family of elements. In contrast, the Gps46 probe displays a high molecular weight smear for all enzymes, suggesting a pattern of dispersion characteristic of highly abundant elements. This is consistent with the copy number estimates for these two families of elements. Likewise, similar hybridization intensities required 30 min for Gps46 but 24 h for Cps21.
Genomic Southern blot analysis of melon LTR retrotransposons. (a) Hybridization to the Cps21 Copia probe. (b) Hybridization to the Gps46 Gypsy probe. Both probes were α-dCTP-32P radiolabeled. Genomic DNA from “Piel de Sapo” (PS) or Korean (Co) varieties of melon were digested with BstNI, BamHI, EcoRI, EcoRV and HindIII, as shown at the bottom of the figure. Both experiments (a) and (b) were done under the same conditions except the film for (a) were exposed 24 h and for (b), half an hour. Numbers on the left are sizes of molecular weight markers (M) in kb
Isolation of a complete Copia element
One of the Copia retrotransposon clones, Cps21, was chosen for further analysis because of its uninterrupted ORF covering the RT segment. The clone was used as a probe to screen a melon BAC library in order to obtain the entire sequence of the element. Subcloning from a positive BAC clone was carried out by inverse PCR and followed by rounds of PCR amplification and sequencing (as described in Materials and methods). This new Copia retrotransposon of melon was named Reme1 (Retrotransposon of melon, accession number AM117493).
Features of Reme1
The cloned element, which has a 37.7% G + C content (A = 1,718, T = 1,491, G = 1,068, C = 872), is 5,149 bp long and flanked by 5 bp target site duplications in the host DNA, which are imperfect due to a dinucleotide inversion (Fig. 3a). The two LTRs (5′, 518 bp; 3′, 515 bp) are 95% identical and contain 5 bp perfect inverted repeats at their ends (5′-TGTTG...CAACA-3′). Reme1 contains all canonical domains of LTR retrotransposons arranged as in Copia elements: sequentially GAG, proteinase, integrase, reverse transcriptase and RNAse H (Fig. 3b).
LTR-retrotransposon Reme1. (a) Structural features of Reme1. The different internal domains are shown: GAG, protease (PR), integrase (INT), reverse transcriptase (RT) and RNase H (RH). The primer binding site (PBS) sequence, complementary to the proposed methionyl-tRNA primer, is indicated. PPT denotes the polypurine tract, the putative purine-rich initiation site, which primes (+)-strand DNA synthesis on the (−)-strand DNA. TS indicates the target site duplication flanking the inserted element. The numbers below the figure are the scale in bp. (b) Amino acid sequence alignments corresponding to the main conserved functional domains of Copia retrotransposons: GAG, PR, INT, RT, RH. Identical amino acids in 60% of sequences are shaded dark grey and similar amino acids in 60% of sequences are shaded gray. Highly conserved amino acids are indicated with a black triangle in GAG, PR and INT; conserved motifs in RT and RH are indicated with a horizontal line above. The accession number for Reme1 is AM117493
Because transcription of retroviruses and LTR retrotransposons is driven by a promoter in the LTR, a search was made for eukaryotic promoter motifs. A putative TATA box was found at position 200–212, 5′-TGGCTATAAATAG-3′, which shows similarity to the consensus TATA box of plants (Joshi 1987a). The central adenine of the putative transcription start site (5′-CCCATGG-3′) was found 19 nt downstream of the initial T of the putative TATA box. The putative CAAT site (5′-CCATT-3′) has a 4–5 nucleotide similarity to the plant consensus. A hypothetical polyadenylation signal (5′-AATAAG-3′) was found at position 311 and it is very similar to the consensus sequence for plants (Joshi 1987b). The putative polyadenylation site (5′-TAGTG-3′) is 37 nt downstream of the polyadenylation signal. Reme1 also contains a potential primer binding site (PBS) for initiation of minus-strand cDNA synthesis. This matches the initiator methionyl tRNA, the one most commonly serving as PBS in LTR retrotransposons. Reme1 contains a polypurine tract (PPT), which serves as the initiation site for plus strand cDNA synthesis, in the canonical position just 5′ to the 3′ LTR. The genomic flanks of the cloned element contain some microsatellites, but no other known sequences.
Phylogenetic analysis of Reme1
Although Cps21, the partial Reme1 clone, contained a complete ORF, the full-length Reme1 from the BAC displays seven frameshifts (due to four stop codons and three small insertions or deletion) and eight stop codons in the predicted reading frame having highest similarity to Copia retrotransposon polypeptides. However, the resulting conceptual translation is highly similar to all the requisite coding domains of an autonomous LTR retrotransposon (Sabot and Schulman 2006; Fig. 3b). The interruptions of the ORF are commensurate with the 5% sequence divergence between the LTRs, together indicating that the BAC insertion is not recent and has accumulated mutations since its integration.
The Reme1 coding domain was compared to that of other LTR retrotransposons on the DNA level (Fig. 4). The phylogenetic analyses strongly support the placement of Reme1 into the Copia clade. Among the Copia elements, Reme1 (from the Order Cucurbitales) clusters most closely with Rtsp1 from Ipomoea batatas of the Order Solanales and with Panzee (Cajanus cajan, Fabales) and secondarily with Tnt1 and Tto1 (both Nicotiana tabacum, Solanales), all species being dicotyledonous. The position of the Reme1 sequence is hence fully consonant with the current close phylogenetic placement of the Solanales and the Cucurbitales (Soltis et al. 2000).
Phylogenetic tree of nucleotide sequences from the Reme1 coding region and other Copia and Gypsy retrotransposons. The tree was constructed with the Neighbor-Joining method and displayed with the MEGA2 program. Bootstrap values from 1,000 replicates are shown. Horizontal distances are proportional to evolutionary distances according to the scale shown on the bottom. Accession numbers: Rtsp1 AB162659 (Ipomoea batatas), Panzee AJ000893 (Cajanus cajan), Tnt1 X13777 (Nicotiana tabacum), Tto1 D83003 (Nicotiana tabacum), Ta12 X53976 (Arabidopsis thaliana), Tst1 X52287 (Solanum tuberosum), PDR1 X66399 (Pisum sativum), BARE1 Z17327 (Hordeum vulgare), Hopscotch U12626 (Zea mays), Opie2 U68408 (Zea mays), RIRE1 D85597 (Oryza australiensis), Cure1 AF499727 (Cucumis melo), Grande1 X97604 (Zea mays), del1 X13886 (Lilium henryi), Reina U69258 (Zea mays), Ty3 S53577 (Saccharomyces cerevisiae), Gypsy P10401 (Drosophila melanogaster)
Reme1 transcription
As described above, the Reme1 element contains promoter motifs in the LTR. We therefore examined the presence of Reme1 transcripts by RT-PCR. Using primers designed to match a 250 bp fragment from the Reme1 5′ UTR, we were able to amplify a product of the expected size on total leaf RNA from untreated PS melon plants, which can be seen as weak band in Fig. 5a (left image, lane N). Because some plant retrotransposons are known to be transcriptionally activated by stresses such as wounding and UV light (Grandbastien 1998; Takeda et al. 1998; Kimura et al. 2001), we decided to investigate the action of various stress agents on the transcriptional levels of Reme1. The agents included UV light, leaf wounding and water stress.
RT-PCR amplification of Reme1 transcripts from total RNA of PS melon leaves. Total RNA was extracted from detached leaves from untreated, control melon plants (C) or from melon plants subjected to UV light treatment (252 nm, in an AV-100 Vertical laminar flow bench) for 6 h (UV). Lanes—RT are RNA samples without the step of reverse transcription. (Panel a) The three images represent PCR amplifications with primers of internal regions from the 5′UTR (left image), gag or int domains (central image), and the rt domain (right image). Lanes M are molecular markers (the 100 bp ladder for 5′UTR and rt RT-PCR amplification analyses and the λ PstI markers for gag and for int ones). Lanes ϕ, reversed transcribed sample without RNA. Lanes N, reversed transcribed RNA sample. The size of DNA products is indicated on the left side of panels. (Panel b) This panel represents the same RT-PCR amplifications than in Panel a, including an internal control elongation factor-1α gene from melon (EF lanes). Lane M is 100 bp ladder molecular marker. DNA bands of 200 and 500 bp long are indicated on the left side of panel
Wounding or water stress treatments of melon plants did not produce any detectable increase in the levels of Reme1 transcripts in melon leaves when compared with non-treated plants (results not shown). Only UV light clearly increases Reme1 transcript pools over that of untreated leaves (Fig. 5a, left image). Similar results were obtained with primers corresponding to the gag, int, and rt domains of Reme1, indicating that full-length elements are transcriptionally induced by UV stress (Fig. 5a, central and right images). In order to examine if the UV induction of Reme1 represented a general cellular response, we tested the induction of a housekeeping gene in a second series of experiments. Primers were designed to match a melon EST (Gonzalez-Ibeas et al. 2007), which has been annotated as an EF-1α-like gene. The homologous genes in Arabidopsis are highly expressed housekeeping genes (Tremousaygue et al. 1997). The results from parallel RT-PCR amplification of melon Reme1 and EF-1α gene transcripts are shown in Fig. 5b. UV light highly increased the Reme1 transcript pools over that of untreated leaves with four different pairs of Reme1 primers (Fig. 5b, lanes UV vs. C). However, no changes in the prevalence of the amplified EF-1α gene product, having an expected size of 214 bp, was seen in UV treated leaves compared with untreated ones (Fig. 5b, lanes EF-UV vs. EF-C).
The RT-PCR products from stress-induced Reme1 transcripts were cloned. In a parallel experiment, the same regions of Reme1 were PCR-amplified from genomic DNA. In addition, a 370 bp segment of the Reme1 LTR (nt 286–536), comprising most of the U3 zone and containing the promoter, was amplified from both RNA and DNA. In contrast to cellular genes, a copy of the promoter is present in retrotransposon transcripts. This is because transcription from the 5′ LTR continues into the 3′ LTR, which also contains the promoter. Independent, randomly selected clones from both RNA and DNA were sequenced and aligned. The sequence heterogeneity found in these was analyzed by calculating genetic variability parameters, as shown in Table 1.
The proportions of polymorphic sites and nucleotide diversity (π) are higher for the genomic sequences than for the RNA transcripts. This, however, is only true for the protein coding regions; the 5′UTR is more polymorphic in the RNA sequences. For the protein-coding regions, rt is the most conserved in both the RNA and DNA sequences, followed by the gag region, whereas int is the most variable (Table 1). The nucleotide diversity values for the Reme1 DNA and RNA sequences are lower than observed in the maize Grande element (García-Martínez and Martínez-Izquierdo 2003; Gomez et al. 2006). Nevertheless, the haplotype diversity (h) values from Reme1 sequences are less than 1 for both RNA and DNA (Table 1). This indicates that some individual sequences in the regions analyzed are identical.
The DNA and RNA sequences in Table 1, excepting genomic LTRs, were aligned and analyzed by neighbor-joining (Fig. 6). Most of the RNA sequences are grouped into one or two clusters at least for the gag, int, and rt trees. Hence, the expressed elements constitute a closely related set, as for Tnt1 in tobacco (Casacuberta et al. 1995). All four trees, but especially those built from protein-coding domains, contain genomic sequences that are topologically close to the clusters of RNA sequences and are therefore similar to the transcriptionally active portions of the Reme1 family. The cloned Reme1 is highly similar to six rt, three gag, two int, and one 5′ UTR sequences in the respective trees.
Neighbor-joining trees of Reme1 retrotransposon DNA and RNA sequences. The trees, from the 5′UTR, gag, int and rt regions, each include both DNA and RNA sequences. DNA sequences are underlined and in bold case and transcript sequence names have RNA as a prefix. Trees were displayed with the MEGA2 program. Horizontal distances are proportional to evolutionary distances according to the scale shown on the bottom
Presence of Cucumis melo Reme1 retrotransposon in other Cucurbitaceae species
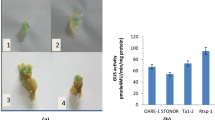
The presence of the Reme1 retrotransposon was surveyed by PCR in nine species of the Cucurbitaceae family. These included watermelon, zucchini, squash, and six species of the Cucumis genus including cucumber (Fig. 7). The presence of int and rt domains related to Reme1 was examined for each (Fig. 7a and b). Reme1-related sequences were found in each of the species. Sequence analysis revealed that the majority of the Reme1 elements in the other Cucurbitaceae samples are highly similar to that of melon (results not shown).
Detection of Reme1 elements in Cucurbitaceae species. PCR products, derived from different accessions of representative plants, were separated by agarose gel electrophoresis. (a) Amplification products from the Reme1 int region (b) Amplification products from the Reme1 rt region. Lane numbers indicate: 1, Cucumis africanus; 2, Cucumis pustulatus; 3, Cucumis prophetarum; 4, Cucumis ficifolius; 5, Cucumis metuliferus; 6, Cucumis sativus (cucumber); 7, Citrullus lanatus (watermelon); 8, Cucurbita pepo (zucchini); 9, Cucurbita maxima (squash); PS, Piel de Sapo melon; Co, Korean melon. M, molecular markers, product sizes are indicated in bp on the left of the figure
Discussion
Abundance and diversity of Copia and Gypsy retrotransposons in the Cucumis melo genome
In this first extensive analysis of retrotransposons in melon, we have found that Gypsy and Copia retrotransposons are a relatively major component. Copia and the three-fold more abundant Gypsy elements together comprise about 26% of the genome (450 Mb). By comparison, 23% of the Citrus sinensis genome (380 Mb) and 17% of the rice genome (430 Mb) are composed of LTR retrotransposons (McCarthy et al. 2002; Rico-Cabanas and Martínez-Izquierdo 2007). Elements of both the Gypsy and Copia groups displayed sequence heterogeneity consonant with that seen elsewhere (Flavell et al. 1992a, b; Pearce et al. 1996a, b; Suoniemi et al. 1998; Friesen et al. 2001).
Reme1, a Copia retrotransposon
Reme1 is, to our knowledge, the first complete Copia retrotransposon cloned, sequenced and characterized in melon. At about 120 copies per haploid genome, Reme1 is not highly abundant, and in the same range as the Copia PDR1 of pea (Ellis et al. 1998) and the Gypsy Cure1 of melon (van Leeuwen et al. 2003). The Reme1 LTR is just twice as abundant as the internal region. This is to be expected if most copies are full-length. Where LTR-LTR recombination actively serves to remove retrotransposons from the genome, thereby counteracting genome expansion, LTRs are much more than twice as abundant as full-length elements (Vicient et al. 1999; Vitte and Panaud 2005). If this mechanism is acting in melon, it appears not to involve Reme1. In this regard, it is noteworthy that the Reme1 LTRs, at 518 bp, are considerably shorter than the LTRs of retrotransposon families that generate many solo LTRs such as BARE1 with LTRs of 1.8 kb (Shirasu et al. 2000). Short LTRs may not efficiently form the loop structures of recombinational intermediates.
Reme1 and host phylogenetics
Analyses of the coding region of Reme1 on the DNA level places it in a clade close to Copia elements of the Solanales and Fabales, consistent with the phylogenetic position of the Violales to which the Cucumis genus of melon belongs. Furthermore, Reme1 itself is present in all of the various Cucurbitaceae species tested, indicating that it was present before their separation. The results suggest that much of the existing diversity among retrotransposons was present before divergence of these orders, because they display divergence by descent in parallel with the rest of the genome.
For the Reme1 sequences analyzed within melon, the proportion of polymorphic sites and nucleotide diversity (π) are higher for the genomic sequences than for those derived from RNA, but only within the protein coding regions. This suggests that the transcribed elements are undergoing purifying selection for protein expression. The nucleotide diversity values for both RNA and DNA are lower than for high-copy elements (García-Martínez and Martínez-Izquierdo 2003; Gomez et al. 2006), consistent with the proposal (Charlesworth 1986) that sequence heterogeneity should be related to copy number.
Reme1 transcriptional activation
The Reme1 LTR contains a putative promoter, which suggested that it could be transcribed. Using RT-PCR, transcription of Reme1 in leaves was observed under normal conditions. This is fairly unusual for a low-copy retrotransposon such as Reme1; transcription in unstressed somatic tissues is typical for the abundant elements of the grasses (Suoniemi et al. 1996; Meyers et al. 2001; Vicient et al. 2001; Echenique et al. 2002; Araujo et al. 2005; Gomez et al. 2006). Transcripts in leaves, however, are unlikely to contribute to genome expansion because leaves do not generally give rise to floral meristems and consequently gametes.
Nevertheless, the stress of UV irradiation induced a sharp increase in Reme1 transcript levels compared to non-treated melon plants, in contrast to a housekeeping EF gene (Tremousaygue et al. 1997) from melon (Gonzalez-Ibeas et al. 2007) where no changes in the level of transcripts upon UV light treatment were observed (Fig. 5b). Various stresses, biotic and abiotic, are known to increase the transcriptional levels of plant retrotransposons (Hirochika 1993; Grandbastien 1998; Takeda et al. 1998), but only UV light was effective for Reme1. This element also differs from others which were shown to be transcriptionally silent in somatic tissues but active only during certain stages of plant development (Pouteau et al. 1991; Pearce et al. 1996b; Turcich et al. 1996). The UV activation of Reme1 was confirmed with RT-PCR from four different Reme1 regions.
Phylogenetic analysis of the RNA transcripts show that they are not only interspersed with genomic sequences on four clades, but also more clustered that the genomic ones. These results suggest that only some elements or subfamilies of Reme1 are transcriptionally active. Altogether, the phylogenetic analyses, diversity measurements, and control experiments support the independent, LTR-driven transcriptional origin of the RNA sequences, rather than the alternatives of either read-through transcription from cellular promoters or contamination of genomic DNA in the RNA preparations.
To date, relative few retroelements have been shown to be transcriptionally activated by UV light. These include SINEs from mammals (Rudin and Thompson 2001), HIV-1 of humans (Valerie et al. 1996) and some LTR retrotransposons from animals (Shim et al. 2000), yeast (Boeke and Corces 1989b; Bradshaw and McEntee 1989) and plants (Kimura et al. 2001). The plant element OARE1 is a Copia retrotransposon from oat similar to BARE1 of barley. However, OARE1 shows a defense-response activation profile, responding to wounding, jasmonic and salicylic acids (Kimura et al. 2001), while Reme1 does not. An increase in the transposition also of DNA transposons by UV light irradiation has been observed for bacteria (Eichenbaum and Livneh 1998) and plants (McClintock 1984; Wessler 1996; Walbot 1999). Whether the increased Reme1 transcription leads to an increase in insertion of new copies, as in yeast (Bradshaw and McEntee 1989), remains an open question. As reported for the tobacco element Tnt1 (Melayah et al. 2001), transcript abundance does not necessarily correlated with success in integration.
Reme1 as an autonomous retrotransposon
In order to be propagated, a genomic copy of a retrotransposon must be transcribed and the transcript packaged into virus-like particles, reverse-transcribed, and integrated back into the nucleus. Although Reme1 is transcribed and encodes all of the proteins needed by retrotransposons for autonomous replication, some of the transcripts contain stop codons. The presence of stop codons on transcripts has been also reported for other plant retrotransposons, such as OARE1 and Grande (Kimura et al. 2001; Gomez-Orte 2002; Gomez et al. 2006). Transcription and reverse transcription are highly error prone, and new retrotransposon copies containing stop codons can be thereby integrated into the genome. Both retroviruses and retrotransposons, however, appear able to cross-complement translationally incompetent RNAs with the proteins either from the same family of elements, by cis-parasitism, or from other groups of elements by trans-parasitism (Escarmis et al. 2006; Kejnovsky et al. 2006; Sabot and Schulman 2006; Tanskanen et al. 2007).
Abbreviations
- BAC:
-
Bacterial artificial chromosome
- bp:
-
Base pairs
- Co:
-
Korean melon variety
- EDTA:
-
Ethylene diamine tetra acetic acid
- EF:
-
Elongation factor
- INT/int :
-
Integrase
- I-PCR:
-
Inverse PCR
- Kb:
-
Kilobases
- LTR:
-
Long terminal repeat
- Mb:
-
Megabases
- MES:
-
2-Morfolinoetansulfonic acid
- ORF:
-
Open reading frame
- PBS:
-
Primer binding site
- PCR:
-
Polymerase chain reaction
- PPT:
-
Polypurine tract
- PS:
-
“Piel de Sapo” melon line
- PSL:
-
Photo-stimulated luminescence
- RT/rt:
-
Reverse transcriptase
- RT-PCR:
-
Reverse-transcription PCR
- SDS:
-
Sodium dodecyl sulphate
- SINE:
-
Short interspersed nuclear elements
- U:
-
Units
- UV:
-
Ultraviolet light
References
Aledo R, Raz R, Monfort A, Vicient CM, Puigdomenech P, Martínez-Izquierdo JA (1995) Chromosome localization and characterization of a family of long interspersed repetitive DNA elements from the genus Zea. Theor Appl Genet 90:1094–1100
Araujo PG, Rossi M, de Jesus EM, Saccaro NL Jr, Kajihara D, Massa R, de Felix JM, Drummond RD, Falco MC, Chabregas SM, Ulian EC, Menossi M, Van Sluys MA (2005) Transcriptionally active transposable elements in recent hybrid sugarcane. Plant J 44:707–717
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9:208–219
Beguiristain T, Grandbastien MA, Puigdomenech P, Casacuberta JM (2001) Three Tnt1 subfamilies show different stress-associated patterns of expression in tobacco. Consequences for retrotransposon control and evolution in plants. Plant Physiol 127:212–221
Bingham P, Zachar Z (1989) Retrotransposons and the FB transposon from Drosophila Melanogaster. In: Berg DE, Howe MM (eds) Mobile DNA. American Society for Microbiology, Washington, pp 485–502
Boeke JD, Corces VG (1989) Transcription and reverse transcription of retrotransposons. Annu Rev Microbiol 43:403–434
Bradshaw VA, McEntee K (1989) DNA damage activates transcription and transposition of yeast Ty retrotransposons. Mol Gen Genet 218:465–474
Casacuberta JM, Vernhettes S, Grandbastien MA (1995) Sequence variability within the tobacco retrotransposon Tnt1 population. EMBO J 14:2670–2678
Charlesworth B (1986) Genetic divergence between transposable elements. Genet Res 48:111–118
Echenique V, Stamova B, Wolters P, Lazo G, Carollo L, Dubcovsky J (2002) Frequencies of Ty1-copia and Ty3-gypsy retroelements within the Triticeae EST databases. Theor Appl Genet 104:840–844
Eichenbaum Z, Livneh Z (1998) UV light induces IS10 transposition in Escherichia coli. Genetics 149:1173–1181
Ellis THN, Poyser SJ, Knox MR, Vershinin AV, Ambrose MJ (1998) Polymorphism of insertion sites of Ty1-copia class retrotransposons and its use for linkage and diversity analysis in pea. Mol Gen Genet 260:9–19
Escarmis C, Lazaro E, Manrubia SC (2006) Population bottlenecks in quasispecies dynamics. Curr Top Microbiol Immunol 299:141–170
Flavell AJ, Dunbar E, Anderson R, Pearce SR, Hartley R, Kumar A (1992a) Ty1-copia group retrotransposons are ubiquitous and heterogeneous in higher plants. Nucleic Acids Res 20:3639–3644
Flavell AJ, Smith JB, Kumar A (1992b) Extreme heterogeneity of Ty1-copia group retrotransposons in plants. Mol Gen Genet 231:232–242
Friesen N, Brandes A, Heslop-Harrison JS (2001) Diversity, origin, and distribution of retrotransposons (gypsy and copia) in conifers. Mol Biol Evol 18:1176–1188
García-Martínez J, Martínez-Izquierdo JA (2003) Study on the evolution of the Grande retrotransposon in the Zea genus. Mol Biol Evol 20:831–841
Gomez-Orte E (2002) Estudio del gen 23 del retrotransposon Grande de maíz. Ph.D. Dissertation, Barcelona University, Spain
Gomez E, Schulman AH, Martínez-Izquierdo JA, Vicient CM (2006) Integrase diversity and transcription of the maize retrotransposon Grande. Genome 49:558–562
Gonzalez-Ibeas D, Blanca J, Roig C, González-To M, Picó B, Truniger V, Gómez P, Deleu W, Caño-Delgado A, Arús P, Nuez F, Garcia-Mas J, Puigdomènech P, Aranda MA (2007) MELOGEN: an EST database for melon functional genomics. BMC Genomics 8:306
Grandbastien MA (1998) Activation of plant retrotransposon under stress conditions. Trends Plant Sci 3:181–187
Grandbastien MA, Spielmann A, Caboche M (1989) Tnt1, a mobile retroviral-like transposable element of tobacco isolated via plant cell genetics. Nature 337:376–380
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hirochika H (1993) Activation of tobacco retrotransposons during tissue culture. EMBO J 12:2521–2528
Joshi CP (1987a) An inspection of the domain between putative TATA box and translation start site in 79 plant genes. Nucleic Acids Res 15:6643–6653
Joshi CP (1987b) Putative polyadenylation signals in nuclear genes of higher plants: a compilation and analysis. Nucleic Acids Res 15:9627–9640
Kejnovsky E, Kubat Z, Macas J, Hobza R, Mracek J, Boris Vyskot B (2006) Retand: a novel family of gypsy-like retrotransposons harboring an amplified tandem repeat. Mol Genet Genomics 276:254–263
Kimura Y, Tosa Y, Shimada S, Sogo R, Kusaba M, Sunaga T, Betsuyaku S, Eto Y, Nakayashiki HI, Mayama S (2001) OARE-1, a Ty1-copia retrotransposon in oat activated by abiotic and biotic stresses. Plant Cell Physiol 42:1345–1354
Kumar A (1996) The adventures of the Ty1-copia group of retrotransposons in plants. Trends Genet 12:41–43
Kumar A, Bennetzen JL (1999) Plant retrotransposons. Annu Rev Genet 33:479–532
Kumar S, Tamura K, Jakobsen IB, Nei M (2001) MEGA2: molecular evolutionary genetics analysis software. Bioinformatics 17:1244–1245
Logemann J, Schell J, Willmitzer L (1987) Improved method for the isolation of RNA from plant tissues. Anal Biochem 163:16–20
McCarthy EM, Liu J, Lizhi G, McDonald JF (2002) Long terminal repeat retrotransposons of Oryza sativa. Genome Biol 3(10):research0053.1–research0053.11
McClintock B (1984) The significance of responses of the genome to challenge. Science 226:792–801
Melayah D, Bonnivard E, Chalhoub B, Audeon C, Grandbastien MA (2001) The mobility of the tobacco Tnt1 retrotransposon correlates with its transcriptional activation by fungal factors. Plant J 28:159–168
Meyers BC, Tingey SV, Morgante M (2001) Abundance, distribution and transcriptional activity of repetitive elements in the maize genome. Genome Res 11:1660–1676
Mount SM, Rubin GM (1985) Complete nucleotide sequence of the Drosophila transposable element copia: homology between copia and retroviral proteins. Mol Cell Biol 5:1630–1638
Neumann P, Pozarkova D, Macas J (2003) Highly abundant pea LTR retrotransposon Ogre is constitutively transcribed and partially spliced. Plant Mol Biol 53:399–410
Oliver M, Garcia-Mas J, Cardús M, Pueyo N, López-Sesé AL, Arroyo M, Gómez-Paniagua H, Arús P, de Vicente MC (2001) Construction of a reference linkage map for melon. Genome 44:836–845
Pearce SR, Harrison G, Li D, Heslop-Harrison P, Kumar A, Flavell AJ, (1996a) The Ty1-copia group retrotransposons in Vicia species: copy number, sequence heterogeneity and chromosomal localisation. Mol Gen Genet 250:305–315
Pearce SR, Kumar A, Flavell AJ (1996b) Activation of the Ty1-copia group retrotransposons of potato (Solanum tuberosum) during protoplast isolation. Plant Cell Rep 15:949–953
Pearce SR, Knox M, Ellis THN, Flavell AJ, Kumar A (2000) Pea Ty1-copia group retrotransposons: transpositional activity and use as markers to study genetic diversity in Pisum. Mol Gen Genet 263:898–907
Pouteau S, Huttner E, Grandbastien MA, Caboche M (1991) Specific expression of the tobacco Tnt1 retrotransposon in protoplasts. EMBO J 10:1911–1918
Rico-Cabanas L, Martinez-Izquierdo JA (2007) CIRE1, a novel transcriptionally active Ty1-copia retrotransposon from Citrus sinensis. Mol Genet Genomics 277:365–377
Rudin CM, Thompson CB (2001) Transcriptional activation of short interspersed elements by DNA-damaging agents. Genes Chromosomes Cancer 30:64–71
Sabot F, Schulman AH (2006) Parasitism and the retrotransposon life cycle in plants: a hitchhiker’s guide to the genome. Heredity 97:381–388
Saitou N, Nei N (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
SanMiguel P, Bennetzen JL (1998) Evidence that a recent increase in maize genome size was caused by the massive amplification of intergene retrotransposons. Ann Bot 81:37–44
Shim S, Lee SK, Han JK (2000) A novel family of retrotransposons in Xenopus with a developmentally regulated expression. Genesis 26:198–207
Shirasu K, Schulman AH, Lahaye T, Schulze-Lefert P (2000) A contiguous 66-kb barley DNA sequence provides evidence for reversible genome expansion. Genome Res 10:908–915
Soltis DE, Soltis PS, Chase MW, Mort ME, Albach DC, Zanis M, Savolainen V, Hahn WH, Hoot SB, Fay MF, Axtell M, Swensen SM, Prince LM, Kress WJ, Nixon KC, Farris JS (2000) Angiosperm phylogeny inferred from 18S rDNA, rbcL, and atpB sequences. Bot J Linn Soc 133:381–461
Suoniemi A, Narvanto A, Schulman AH (1996) The BARE-1 retrotransposon is transcribed in barley from an LTR promoter active in transient assays. Plant Mol Biol 31:295–306
Suoniemi A, Tanskanen J, Schulman AH (1998) Gypsy-like retrotransposons are widespread in the plant kingdom. Plant J 13:699–705
Takeda S, Sugimoto K, Otsuki H, Hirochika H (1998) Transcriptional activation of the tobacco retrotransposon Tto1 by wounding and methyl jasmonate. Plant Mol Biol 36:365–376
Tanskanen JA, Sabot F, Vicient C, Schulman AH (2007) Life without GAG: the BARE-2 retrotransposon as a parasite´s parasite. Gene 390:166–174
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, population-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tremousaygue D, Bardet C, Dabos P, Regad F, Pelese F, Nazer R, Gander E, Lescure B (1997) Genome DNA sequencing around the EF-1 alpha multigene locus of Arabidopsis thaliana indicates a high gene density and a shuffling of noncoding regions. Genome Res 7:198–209
Turcich MP, Bokhari-Riza A, Hamilton DA, He C, Messier W, Stewart CB, Mascarenhas JP (1996) PREM-2, a copia-type retroelement in maize is expressed preferentially in early microspores. Sex Plant Reprod 9:65–74
Valerie K, Laster WS, Cheng L, Kirkham JC, Reavey P, Kuemmere NB (1996) Signal transduction and HIV transcriptional activation after exposure to ultraviolet light and other DNA-damaging agents. Photochem Photobiol 64:280–285
Van Leeuwen H, Monfort A, Zhang HB, Puigdomenech P (2003) Identification and characterisation of a melon genomic region containing a resistance gene cluster from a constructed BAC library. Microlinearity between Cucumis melo and Arabidopsis thaliana. Plant Mol Biol 51:703–718
Van Leeuwen H, Garcia-Mas J, Coca M, Puigdomenech P, Monfort A (2005) Analysis of the melon genome in regions encompassing TIR-NBS-LRR resistance genes. Mol Genet Genomics 273:240–251
Vernhettes S, Grandbastien MA, Casacuberta JM (1998) The evolutionary analysis of the Tnt1 retrotransposon in Nicotiana species reveals the high plasticity of its regulatory sequences. Mol Biol Evol 15:827–836
Vicient CM, Suoniemi A, Anamthawat-Jonsson K, Tanskanen J, Beharav A, Nevo E, Schulman AH (1999) Retrotransposon BARE-1 and its role in genome evolution in the genus Hordeum. Plant Cell 11:1769–1784
Vicient CM, Kalendar R, Schulman AH (2001) Active retrotransposons are a common feature of grass genomes. Plant Physiol 125:1283–1292
Vitte C, Panaud O (2005) LTR retrotransposons and flowering plant genome size: emergence of the increase/decrease model. Cytogenet Genome Res 110:91–107
Walbot V (1999) UV-B damage amplified by transposons in maize. Nature 397:398–399
Warmington JR, Waring RB, Newlon CS, Indge KJ, Oliver SG (1985) Nucleotide sequence characterization of Ty 1-17, a class II transposons from yeast. Nucleic Acids Res 13:6679–6693
Wessler SR (1996) Plant retrotransposons: turned on by stress. Curr Biol 6:959–961
Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell AJ, Leroy P, Morgante M, Panaud O, Paux E, SanMiguel P, Schulman AH (2007) A unified classification system for eukaryotic transposable elements. Nature Rev Genet 12. doi:10.1038/nrg2165
Acknowledgments
The authors are indebted to IRTA and Semillas Fitó for plant material and to IRTA for Southern blot membranes. We are grateful to Dr. Van Leeuwen for providing the melon BAC library, to Dr. A. Caño, A. Mascarell and D. Gonzalez-Ibeas (CSIC) for providing the primers sequences of a melon EF gene and functional information about it and to IBMB (CSIC) technicians for support, especially those from the sequencing service. E.R. was the recipient of a collaborative CSIC-industry Fellowship: CSIC-Semillas Fitó S.A., a contract from the Spanish Biotechnology National Plan (BIO99-1175), a travel grant from the ESF for a stay in the laboratory of A.H.S. where the work was supported by a grant from the Academy of Finland (project 53453), and a grant from Consorci CSIC-IRTA. Some funds for this study were provided by CERBA (Generalitat of Catalonia). This work was mainly supported by the grant BIO99-1175 (CICYT) from the MEC of Spanish Govern to J.A.M.I.
Author information
Authors and Affiliations
Corresponding author
Additional information
Accession Numbers of New Sequences
Cucumis melo Copia retrotransposons, putative reverse transcriptase (partial sequences): From AM182612 to AM182643, clones Cco26, 24, 43, 45, 22, 23, 8, 27, 32, 9, 31, 25, 33; Cps42, 25, 32, 23, 21, 38, 35, 1, 31, 3, 15, 33, 36, 37, 27, 24, 29, 22 and 26, respectively.
Retrotransposon Reme1 (complete sequence): AM1174993.
Cucumis melo Gypsy retrotransposons, reverse transcriptase (partial sequences): From AM182333 to AM182346, clones Gco9rt, 43, 25, 24, 8, 45, 18, 46; Cps1rt, 28, 29, 43, 3 and 46, respectively. Cucumis melo Gypsy retrotransposons, putative integrase (partial sequences): From AM182873 to AM182886, clones Gco9int, 25, 24, 8, 45, 46, 18, 43; Cps1int, 28, 29, 43, 3, and 46, respectively. Reme1retrotransposon elements (partial sequences): From AM228927 to AM228954, clones 5′UTR-1, 2, 4, 6, 9, 10, 11, 13; GAG-13, 5, 11, 1, 2, 10, 9, 14, 3, 6; INT-14, 1, 11, 7; RT-9, 15, 2, 3, 6 and 12, respectively. Reme1 retrotransposon elements, cDNAs (from RNAs) (partial sequences): From AM228981 to AM229008, clones RNA5′UTR-1.6, 2, 28, 29, 30, 31; RNAGAG-17, 18, 34, 1.5, 32, 35, 15, 16; RNA-INT-22, 23, 16, 17, 37, 40, 41, 25; RNA-RT-20, 22, 23, 24, 6 and 7, respectively.
Rights and permissions
About this article
Cite this article
Ramallo, E., Kalendar, R., Schulman, A.H. et al. Reme1, a Copia retrotransposon in melon, is transcriptionally induced by UV light. Plant Mol Biol 66, 137–150 (2008). https://doi.org/10.1007/s11103-007-9258-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-007-9258-4