Abstract
Plastids are organelles unique to plant cells and are responsible for photosynthesis and other metabolic functions. Despite their important cellular roles, relatively little is known about the mechanism of plastidial DNA replication and repair. Recently, we identified a novel DNA polymerase in Oryza Sativa L. (OsPOLP1, formerly termed OsPolI-like) that is homologous to prokaryotic DNA polymerase Is (PolIs), and suggested that this polymerase might be involved in plastidial DNA replication and repair. Here, we propose to rename the plant PolI homologs as DNA polymerase π (POLP), and investigate the biochemical properties of full-length OsPOLP1. The purified OsPOLP1 elongated both DNA and RNA primer hybridized to a DNA template, and possessed a 3′ exonuclease activity. Moreover, OsPOLP1 displayed high processivity and fidelity, indicating that this polymerase has the biochemical characteristics appropriate for DNA replication. We found that POLPs have two extra sequences in the polymerase domain that are absent in prokaryotic PolIs. Deletion of either insert from OsPOLP1 caused a decrease in DNA synthetic activity, processivity, and DNA binding activity. In addition, OsPOLP1 efficiently catalyzed strand displacement on nicked DNA with a 5′-deoxyribose phosphate, suggesting that this enzyme might be involved in a repair pathway similar to long-patch base excision repair. These results provide insights into the possible role of POLPs in plastidial DNA replication and repair.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plastids, one of the major types of plant cell organelle, are essential to the viability of plants and participate in photosynthesis and other metabolic functions. It is believed that plastids arose from cyanobacteria through endosymbiosis. During development, proplastids can differentiate into several plastid types such as chloroplasts, amyloplasts, and chromoplasts. Plastids usually contain multiple copies of their DNA, which resemble cyanobacterial genomes in size and shape (Bendich 2004; Sato 2001). DNA synthesis in plastids is independent of nuclear DNA replication. Additionally, chloroplasts are known to generate a large amount of reactive oxygen species during photosynthesis, suggesting that oxidative DNA damage to their genomes might often occur (Kimura and Sakaguchi 2006). On the basis of the differences between plastidial and nuclear genomes, it seems likely that plastidial DNA replication and repair might involve different metabolic pathways from those required for nuclear DNA replication and repair. Currently, however, relatively little is known of the machinery of plastidial DNA replication and repair.
DNA polymerases (Pols) are indispensable enzymes in DNA replication, repair and recombination. To date, at least 14 species of DNA polymerase (Polα, β, γ, δ, ε, ζ, η, θ, ι, λ, κ, μ, ν, and PolI-like) have been identified in eukaryotic cells, and these polymerases are classified into 4 families (A, B, X, and Y) based on their amino acid sequences (Burgers et al. 2001; Kimura et al. 2002; Marini et al. 2003). With the exception of Polζ, B-family polymerases (Pol α, δ, and ε) are involved in nuclear DNA replication; Polζ and Y-family polymerases (Pol η, ι, and κ) participate in translesion synthesis (Burgers et al. 2001; Hübscher et al. 2002). Polβ, λ, and μ belong to the X family, and have been suggested to be involved in base excision repair and non-homologous end joining (Burgers et al. 2001; Dianov et al. 2003; Lee et al. 2004). A-family polymerases are homologs of Escherichia coli (E. coli) PolI. Three A-family genes are present in the human genome: Polγ is essential for mitochondrial DNA replication and repair; and Polθ and ν (also termed POLQ and POLN, respectively) appear to function in bypassing some types of DNA damage (Graziewicz et al. 2006; Seki et al. 2004; Takata et al. 2006).
In contrast, two types of A-family polymerase (Pol θ and PolI-like) are encoded in the rice genome, which lacks Polγ and Polν genes (Burgers et al. 2001; Kimura and Sakaguchi 2006). Recently, we characterized a plant-specific polymerase in Oryza sativa L. that is homologous to prokaryotic PolIs (OsPOLP1, formerly termed OsPolI-like) and also identified two orthologs of this polymerase in Arabidopsis thaliana (renamed AtPOLP1 and AtPOLP2 in this study) (Kimura et al. 2002; Mori et al. 2005). These polymerases contain two conserved regions, the 3′ exonuclease and the polymerase regions. We found that OsPOLP1 was included in the plastidial fraction in subcellular fractionation experiments and that the green fluorescent proteins containing the N-terminal polypeptides of AtPOLPs localized in plastids. In addition, OsPOLP1 mRNA was highly expressed in meristematic cells containing undifferentiated proplastids, and a similar result was obtained from an in situ hybidization analysis using RNA probes complementary to AtPOLP mRNAs. Chloroplast DNA replication is largely or entirely limited to meristematic cells, and the number of chloroplast DNA copies is reduced in non-meristematic cells (Bendich 2004). Our observations therefore suggest that the plant-specific polymerases might be involved in plastidial DNA replication and repair. At that time, we also tried to investigate the biochemical properties of the full-length polymerases, but failed to achieve expression of the proteins. In the present report, we propose to rename the plant PolI homologs as DNA polymerase π (POLP), and investigate the biochemical characteristics of a GST fusion protein containing full-length OsPOLP1. Based on the results of these analyses, the possible role of POLPs in plastids is discussed.
Materials and methods
Expression and purification of GST fusion protein containing OsPOLP1
Full-length OsPOLP1 cDNA was amplified by PCR using the full-length cDNA clone, AK103236 (Kikuchi et al. 2003) with the two primers, 5′-TAGAATTCATGGCGGTGGCGCCGCCGCT-3′ and 5′-TACGCTCGAGTCACTTATCGTCGTCATCCTTGTAATCCTTGGCAGCATACCAGCTCTTCGCG-3′. The PCR fragment tagged with a 3′-terminal FLAG was cloned into the pGEX 6P-1 expression vector (GE Healthcare Bioscience), and transformed into E. coli BL21(DE3) (Novagen). One colony was incubated in 10 ml of terrific broth (TB) medium containing 1% (w/v) glucose and 50 μg/ml ampicillin. The culture was grown overnight at 30°C and transferred into 1 liter of fresh TB medium containing 1% (w/v) glucose and 50 μg/ml ampicillin. The cells were grown at 16°C until the A600 value reached 0.3–0.4, at which point isopropyl-β-thio-galactopyranoside was added to 50 μM, and the culture continued at the same temperature for 12 h. The cells were harvested by centrifugation at 4°C, and were washed with cold Phosphate Bufferd Saline (PBS). They were then resuspended in 5 ml of HS buffer (50 mM Hepes-NaOH (pH 7.7) and 10% (w/v) sucrose), and were frozen in liquid nitrogen. The frozen cells were thawed in an ice bath, and were mixed with 90 ml of TEDG buffer (50 mM Tris–HCl (pH 7.5), 1 mM EDTA, 1 mM dithiothreitol (DTT), 10% (v/v) glycerol, 0.01% (v/v) Nonidet P-40) containing 0.5 M NaCl, 1 μg/ml each of pepstatin A and leupeptin, 1 mM phenylmethylsulfonyl fluoride, and 1 mg/ml lysozyme. The suspension was incubated on ice for 30 min, warmed at 37°C for 90 s, and then placed on ice for 30 min. The cell lysate was centrifuged for 30 min at 30,000 × g at 4°C. After filtration through a 0.45 μm PVDF membrane, the supernatant was loaded onto 0.5 ml of a glutathione sepharose 4B column equilibrated with TEDG buffer containing 0.4 M NaCl, and extensively washed with TEDG buffer containing 2 M NaCl, and TEDG buffer supplemented with 0.4 M NaCl, 2 mM adenosine triphosphate and 5 mM MgCl2. GST fusion protein containing OsPOLP1 (GST-OsPOLP1) was eluted from the column with TEDG buffer (adjusted to pH 8.5) containing 100 mM NaCl and 25 mM glutathione (reduced form). The eluted material was loaded onto a HiPrep 16/60 sephacryl S-300 column equilibrated with S-300 buffer (50 mM Tris–HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, 1 mM DTT and 10% (v/v) glycerol). The fractions containing GST-OsPOLP1 were collected, frozen in liquid nitrogen, and stored at −80°C until use.
The polymerase- or 3′ exonuclease-deficient mutants (D780A or D362A, respectively) and the deletion mutants lacking amino acid residues 635-674 or 827-852 (Δinsert I or Δinsert II, respectively) were overexpressed and purified as for the wild type protein.
Other proteins
Klenow fragment was purchased from New England Biolabs Inc. Recombinant DNA polymerase β from Rattus norvegicus (Polβ) were purified as described previously (Takeuchi et al. 2004).
Oligonucleotides
The oligonucleotide sequences used in this study are shown in Supplementary Table 1. T4 polynucleotide kinase (TaKaRa Bio. Inc., Japan) and [γ-32P]-ATP were used for 5′ labeling.
DNA polymerase assay
To determine the optimal reaction coditions, oligo(dT)12-18 hybridized to poly(dA) at a ratio of 1:10 (w/w) (poly(dA)/oligo(dT)10:1) was used as the DNA template. The standard reaction mixture (20 μl) contained 50 mM Tris–HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 10% (v/v) glycerol, 0.5 μM dTTP, 0.1 μM [3H]-dTTP, 50 μg/ml poly(dA)/oligo(dT)10:1, and 100 fmol of GST-OsPOLP1. After incubation for 1 h at 37°C, the reaction mixture was spotted onto a Whatmann DE81 ion-exchange filter (2 × 2 cm2), air-dried, and washed 3 times with 5% (w/v) Na2HPO4 (5 ml/filter), 3 times with distilled water, and twice with ethanol. The radioactivity of each dried filter was measured in a toluene-based scintillator. The processivity assay was carried out with the 36G/67C template (Supplementary Table 1) or 5′ labeled oligo(dT)20 annealed to poly(dA) at a ratio of 1:20 (5′ labeled poly(dA)/oligo(dT)20:1). The reaction mixture (20 μl) contained 50 mM Tris–HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 10% (v/v) glycerol, 25 μM each of dNTPs (for 36G/67C) or dTTP (for 5′ labeled poly(dA)/oligo(dT)20:1), 100 fmol of the radiolabeled DNA template, and the indicated amounts of GST-OsPOLP1 or 10 fmol of Klenow fragment. The strand displacement assay was carried out under the same reaction condition as the processivity assay using the 16/30G, dRP2 pretreated by E. coli uracil-N-glycosylase (TREVIGEN) at 37°C for 30 min, or the Nick DNA templates (Supplementary Table 1). After incubation at 37°C, reactions were terminated by adding 20 μl of the loading buffer (10 mM Tris–HCl (pH 7.5), 1 mM EDTA, 95% (v/v) formamide, 0.1% (w/v) bromophenol blue, and 0.1% (w/v) xylenecyanol), and the reaction products were separated on an acrylamide gel containing 8 M urea. After drying, the gel was exposed to BioMax MS-1 (Kodak) and quantified using a BAS-3000 imaging analyzer (Fuji Film, Japan).
Elongation of an RNA primer
The reaction mixture (20 μl) contained 50 mM Tris–HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 10% (v/v) glycerol, 25 μM dNTPs, 100 fmol of the radiolabeled 16D/30 or 16R/30 template (Supplementary Table 1), and 200 fmol of GST-OsPOLP1. After incubation at 37°C, the reaction products were separated on a 20% acrylamide gel containing 8 M urea. After drying, the gel was exposed to BioMax MS-1.
3′ exonuclease assay
The reaction mixture (20 μl) containing 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 10% (v/v) glycerol, 100 fmol of the 36G/67T substrate (Supplementary Table 1), and 300 fmol of GST-OsPOLP1 was incubated at 37 °C. The reaction was terminated by addition of 20 μl of the loading buffer, and the reaction products were separated on a 20% acrylamide gel containing 8 M urea. After drying, the gels were exposed to BioMax MS-1.
Electrophoretic mobility shift assay
The assay was carried out based on the method described previously (Masuda et al. 2001). Briefly, the reaction mixture (20 μl) containing 20 mM Tris–HCl (pH 7.5), 30 mM NaCl, 1 mM DTT, 2% (v/v) glycerol, 0.2 mg/ml bovine serum albumin, 10 fmol of the 16/30G substrate (Supplementary Table 1), and the indicated amounts of GST-OsPOLP1 was incubated on ice for 10 min, and separated on a prerunning 4% acrylamide gel containing 5 mM Tris–HCl (pH 7.4), 6 mM sodium acetate, and 0.1 mM EDTA at 4°C. After drying, the gel was exposed to BioMax MS-1 (Kodak) and quantified using a BAS-3000 imaging analyzer (Fuji Film, Japan).
Fidelity assay
The nucleotide selectivity was examined using the 16/30G, 16/30A, 16/30T, and 16/30C templates (Supplementary Table 1) under running-start conditions. The reaction mixture (20 μl) containing 50 mM Tris–HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 10% (v/v) glycerol, 25 μM dGTP (for 16/30G, 16/30A, and 16/30T) or dTTP (for 16/30C), 0.1 μM of the correct nucleotides or 250 μM of the incorrect nucleotides for the target site, 100 fmol of the radiolabeled template, and 200 fmol of GST-OsPOLP1 was incubated at 30°C for 5 min. The base substitution fidelity for the 16/30G and 16/30A DNA templates was measured in single completed hit conditions using the gel kinetic methodology (Bloom et al. 1997; Chen et al. 2000; Fygenson and Goodman 1997). The reactions were run at 30°C for 5 min to incorporate the right nucleotides, and for 10 min to incorporate wrong nucleotides. The reaction mixture (20 μl) contained 50 mM Tris–HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 5 mM MgCl2, 10% (v/v) glycerol, 25 μM dGTP, different concentrations of one of dNTPs, 100 fmol of the radiolabeled template, and 200 fmol of GST-OsPOLP1. The reaction was terminated by adding 20 μl of the loading buffer, and the reaction products were separated on a 20% acrylamide gel containing 8 M urea. After drying, the gel was exposed to BioMax MS-1 and quantified using a BAS-3000 imaging analyzer. The nucleotide incorporation frequency opposite the target site was calculated by I T Σ/I T−1 , where I T Σ is the integrated band intensity of the primers extended to the target site and beyond (positions T, T + 1, T + 2, etc.) and I T−1 is the integrated band intensity of primers extended to the one base before the target site (position T−1). A plot of the relative incorporation rate, I T Σ/I T−1 as a function of the dNTP concentration, resulted in a rectangular hyperbola. Apparent K m and relative V max values were obtained using at least squares fit to a rectangular hyperbola. The relative V max value is equal to the maximum value of I T Σ/I T−1. The misincorporation frequencies (f inc ), which is the inverse of fidelity, is given by the ratio shown in Eq 1, where the subscripts W and R refer to incorporation of the wrong or right nucleotide, respectively.
5′-deoxyribose phosphate (dRP) lyase assay
The dRP1 substrate (Supplementary Table 1) was pretreated with E. coli uracil-N-glycosylase at 37°C for 30 min to remove deoxyuracil residues. The reaction mixture (20 μl) containing 50 mM Tris–HCl (pH 7.5), 50 mM NaCl, 1 mM DTT, 10% (v/v) glycerol, 100 fmol of the pretreated dRP1 substrate and 500 fmol of GST-OsPOLP1 or 18 fmol of Polβ was incubated at 37°C. The mixture was mixed with 10 μl of 1 M sodium borohydride and incubated on ice for 30 min. The reaction products were subjected to a 20% acrylamide gel containing 8 M urea. After drying, the gels were exposed to BioMax MS-1.
Results
Two types of DNA polymerase π (POLP) are encoded in the rice genome.
We previously identified a novel polymerase in Oryza sativa L. that was homologous to prokaryotic PolIs (OsPOLP1). We subsequently reported that two orthologs were present in Arabidopsis thaliana (Kimura et al. 2002; Mori et al. 2005). The presence of two orthologs in Arabidopsis suggested that rice cells might also contain a second POLP. To investigate this possibility, we searched the rice genome database and found a second OsPOLP gene (OsPOLP2), located on chromosome 4 (GenBank accession number AL606657). The predicted product of OsPOLP2 was 1032 amino acids, and showed homology to OsPOLP1 (74% identity and 83% similarity) (Supplementary Fig. 1), suggesting that the biochemical properties of the two OsPOLP proteins might be similar. We therefore focused on the enzymatic activities of OsPOLP1.
DNA synthetic activity of GST fusion protein containing OsPOLP1
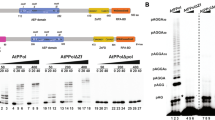
To obtain a recombinant OsPOLP1 protein, we induced expression of a GST fusion protein containing OsPOLP1 with a 3′ terminal FLAG epitope-tag (GST-OsPOLP1) in E. coli. Soluble GST-OsPOLP1 was purified on glutathione sepharose and sephacryl S-300 columns (Fig. 1A), and the purified enzyme displayed substantial DNA synthetic activity (Fig. 1B). We attempted to eliminate the N-terminal GST tag from the purified protein using rhinovirus protease (PreScission Protease), but this resulted in the unexpected fragmentation of OsPOLP1 (data not shown). OsPOLP1 seems to include some non-specific sites susceptible to protease cleavage. Therefore, we investigated the biochemical properties of the polymerase using the GST fusion protein; we could not rule out the possibility that the GST tag influenced enzymatic activities. To confirm the absence of contaminating E. coli polymerases, we constructed a polymerase-deficient mutant (D780A) in which Ala is substituted for a highly conserved Asp in Motif 3 of the polymerase domain. This substitution has been reported to disrupt the DNA synthetic activity of prokaryotic PolI in vivo (Patel and Loeb 2000). The D780A mutant was expressed and purified by the same procedures as the wild type protein (left panel in Fig. 1A). The purified D780A mutant showed no DNA synthetic capability, but did display 3′ exonuclease activity (Fig. 1B), probably because separate structural domains of GST-OsPOLP1 are responsible for the two different enzymatic activities as observed in a Klenow fragment-DNA co-crystal structure (Beese et al. 1993). Next, we determined the optimal reaction conditions for GST-OsPOLP1 using poly(dA)/oligo(dT)10:1 as the template. The optimal reaction conditions were similar to those for the truncated OsPOLP1 protein containing only the polymerase domain (Kimura et al. 2002) (Supplementary Table 2). We also tested the ability of this polymerase to elongate an RNA primer with dNTPs. GST-OsPOLP1 elongated both 16-nt DNA and RNA primers hybridized to a 30-nt DNA template with the same sequence context (Fig. 1C).
Purification and enzymatic activities of GST-OsPOLP1. (A) Left, SDS-PAGE analysis of the wild type (WT) and the polymerase-deficient mutant GST-OsPOLP1 (D780A). Right, SDS-PAGE analysis of WT and the 3′ exonuclease-deficient mutant GST-OsPOLP1 (D362A). Each protein (200 ng) was separated on a 10% acrylamide gel and visualized by silver staining. (B) DNA synthetic activity. Each protein (500 fmol) was incubated with 100 fmol of the 36G/67C template at 37°C. (C) Elongation of an RNA primer hybridized to a DNA template. GST-OsPOLP1 (200 fmol) was incubated with 100 fmol of the 16D/30 (DNA primer) or 16R/30 (RNA primer) template at 37°C. (D) 3′ exonuclease activity. Each protein (300 fmol) was incubated with 100 fmol of the 36G/67T substrate at 37°C
3′ exonuclease activity of GST-OsPOLP1
POLPs have a 3′ exonuclease domain that contains three highly conserved segments, ExoI, ExoII, and ExoIII (Bernad et al. 1989) (Supplementary Fig. 1). To confirm that GST-OsPOLP1 possesses 3′ exonuclease activity, we prepared a mutant protein, D362A, that has a replacement of Asp by Ala in the ExoII domain (right panel in Fig. 1A). Mutation at the homologous position of Klenow fragment (D424) created a mutant protein that abolished 3′ exonuclease activity (Derbyshire et al. 1991). In similar fashion to Klenow fragment, the D362A mutant of GST-OsPOLP1 showed the similar DNA synthetic activity to that of the wild type protein (data not shown). However, the mutant displayed no nuclease activity, whereas the wild type protein excised the 3′ terminal bases of the substrates (Fig. 1D).
Processivity of GST-OsPOLP1
To measure the processivity of GST-OsPOLP1, we first used a 5′-labeled 36-nt primer annealed to a 67-nt oligonucleotide as the DNA template. We confirmed that more than 98% of the primers hybridized to the template (data not shown). Under conditions in which few primers were extended, most of the reaction products were elongated to the end of the 67-nt template by GST-OsPOLP1 (Fig. 2A). In contrast, Klenow fragment, which is a middle processive enzyme, mainly gave reaction products of less than 51-nt (Bambara et al. 1978). We next measured the processivity using 5′ labeled poly(dA)/oligo(dT)20:1. GST-OsPOLP1 extended the 5′ labeled 20-nt primer to more than 600-nt (Fig. 2B). Moreover, reaction products of approximately 600-nt were also generated when an excess amount of unlabeled poly(dA)/oligo(dT)10:1 as a trapping reagent was added to prevent the polymerase from binding to the radiolabeled template on multiple occasions after the radiolabeled template was mixed with the polymerase (lanes 3–5 in Fig. 2C). In contrast, when the protein was added after the radiolabeled template was mixed with the trapping reagent, no reaction product was detectable (lanes 6–8 in Fig. 2C). These results indicate that GST-OsPOLP1 is a highly processive polymerase.
Processivity of GST-OsPOLP1. (A) One hundred fmol of WT and D780A and 10 fmol of Klenow fragment (KF) were incubated with 100 fmol of the 36G/67C template at 37°C. (B) The wild type GST-OsPOLP1 (lane 2, 200 fmol; lane 3, 100 fmol) was incubated with 100 fmol of 5′ labeled poly(dA)/oligo(dT)20:1 at 37°C for 5 min. Control fraction with no protein is shown in lane 1. (C) One hundred fmol of 5′ labeled poly(dA)/oligo(dT) 20:1 and GST-OsPOLP1 were incubated with different amounts of unlabeled poly(dA)/oligo(dT)10:1 as trapping reagent (0.5, 1.0, 2.0 pmol in lanes 3–5 and 6–8, respectively). Control fraction with no protein is shown in lane 1. In lanes 6–8, the trapping reagent was added first before the protein was mixed, in order to confirm the efficacy of the trap
POLPs have two extra sequences in the polymerase domain
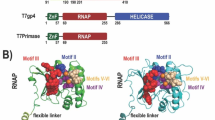
POLPs are A-family polymerases and their amino acid sequences more closely resemble those of prokaryotic PolIs than of eukaryotic Polγ or Polθ (Kimura et al. 2002). Sequence alignment analysis showed that POLPs have two extra sequences in the polymerase domain that are absent in prokaryotic PolIs (Fig. 3A); the 3′ exonuclease domain did not contain any additional sequences (data not shown). One of the additional inserts (amino acid residues 635-674 of OsPOLP1, insert I) consisted of 40-54 amino acids situated between Motifs 1 and 2, within the tip of the thumb subdomain. According to previous reports (Beese et al. 1993; Patel et al. 2001), this region of Klenow fragment forms two helical loops, and interacts with the minor groove of duplex DNA. Deletion of this region causes reductions in DNA binding affinity, processivity, and frameshift fidelity (Minnick et al. 1996). In addition, human Polθ has three sequence insertions in the polymerase domain, and one of these sequences is inserted into the tip of the thumb subdomain, suggesting that this extra sequence causes the unusual activities of Polθ (Seki et al. 2004). The second insert of POLPs (amino acid residues 827-852 of OsPOLP1, insert II) consisted of 26 amino acids lying at the N-terminal side of Motif 4 (Fig. 3A). Several amino acid residues in this motif are known to bind an incoming nucleotide (Beese et al. 1993; Li et al. 1998; Patel et al. 2001).
POLPs have two extra sequences in the polymerase domain that are absent in prokaryotic PolIs. (A) Sequence alignment of the polymerase domain of POLPs and prokaryotic A-family polymerases. The alignment was carried out using the ClustalW program. The bars indicate the inserts and the conserved motifs 1–6. OsPOLP1 and OsPOLP2, Oryza sativa DNA polymerase π1 and π2; AtPOLP1 and AtPOLP2, Arabidopsis thaliana DNA polymerase π1 and π2; TaqPolI, Thermus aquaticus PolI; BstPolI, Bacillus stearothermophilus PolI; EcPolI, E. coli PolI. (B) SDS-PAGE analysis of the wild type (WT) and the deletion mutants of GST-OsPOLP1 lacking either insert I or insert II (Δinsert I or Δinsert II). Each protein (100 ng) was separated on a 10% acrylamide gel and visualized by silver staining. (C) DNA synthetic activity. One hundred fmol of 5′ labeled poly(dA)/oligo(dT) 20:1 was incubated with Δinsert I, Δinsert II (lanes 2 and 4, 400 fmol; lanes 3 and 5, 200 fmol), and WT (lane 6, 200 fmol; lane 7, 100 fmol) at 37°C for 5 min. (D) DNA binding activity. The reaction mixtures containing 10 fmol of the 16/30G template and the indicated enzymes (100, 200, and 400 fmol in lanes 2–4, 5–7, and 8–10, respectively) were loaded on a 4% acrylamide gel. Control fraction with no enzyme is shown in lane 1. ND, not detected; F, free DNA; C, protein–DNA complex
To investigate the biological roles of these inserts, we created deletion mutants lacking either insert I or insert II (Δinsert I or Δinsert II, respectively) (Fig. 3B). The purified Δinsert I showed a much lower DNA polymerase activity than the wild type protein, and elongated a primer by less than 10-nt on 5′-labeled poly(dA)/oligo(dT)20:1 (lanes 2 and 3 in Fig. 3C). This result indicates that deletion of insert I decreased DNA synthetic activity and processivity. In contrast, we did not observe DNA synthesis by Δinsert II, but did find small amounts of the 3’ exonucleolytic products (lanes 4 and 5 in Fig. 3C). From this result, we hypothesized that the deletion of the inserts might influence binding affinity to the template. To investigate this possibility, we carried out an electrophoretic mobility shift assay. We found that both Δinsert I and Δinsert II exhibited lower DNA binding activities than the wild type protein (Fig. 3D).
Fidelity of GST-OsPOLP1
We examined the base substitution fidelity of GST-OsPOLP1 using the gel kinetic methodology, in which the kinetic parameters of the polymerase activity are measured at specific sites on a DNA template in the presence of the 3′ exonuclease activity (Bloom et al. 1997; Chen et al. 2000; Fygenson and Goodman 1997). We first tested nucleotide selectivity for each of the four template bases. In the presence of only dNTP complementary to the first two template bases (i.e. dGTP or dATP), GST-OsPOLP1 elongated the primer to a position one base short of the target site (T−1) (lanes 2, 8, 14, and 20 in Fig. 4). When other dNTPs were added to the reaction mixture (lanes 3–6, 9–12, 15–18, and 21–24 in Fig. 4), the polymerase incorporated the nucleotide opposite the target template base (T). Comparison of the ratios between gel band intensities at or beyond the T site and those at T−1 showed that the correct nucleotides were efficiently inserted at the T site even at a low concentration of nucleotides (0.1 μM), and that misincorporation occurred with a moderate frequency in the presence of an excess of a wrong nucleotide (250 μM). Where misincorporation occurred, replacement of one purine (A or G) by the other, or of one pyrimidine (T and C) by the other was more frequent than replacement of a purine by a pyrimidine, or vice versa.
Nucleotide selectivity of GST-OsPOLP1. GST-OsPOLP1 (200 fmol) was incubated with 100 fmol of the 16/30G, 16/30A, 16/30T, or 16/30C template in the presence of 25 μM dGTP (lane 2–6, 8–12, 14–18) or dCTP (lane 20–24) and the indicated dNTP (0.1 μM correct nucleotides or 250 μM incorrect nucleotides) at 30°C for 5 min. Control fraction with no enzyme is shown in lanes 1, 7, 13, and 19
We next measured base substitution fidelity opposite the template G and A bases using gel kinetic analysis (see Materials and methods). GST-OsPOLP1 misincorporated nucleotides opposite a target G base with a frequency of approximately 10−4–10−5, and most frequently incorporated dTMP (Table 1). The values for the misincorporation frequency (f inc ) for a target A site were also similar to those for a target G site (Table 1). These data are consistent with those shown in Fig. 4, indicating that the base substitution fidelity of GST-OsPOLP1 displayed relatively high fidelity (approximately 10-1000 fold higher than low fidelity polymerases) (Kunkel 2004), and that transition mutations occurred more frequently than transversion mutations in a similar fashion to a general polymerase (Boosalis et al. 1987; 1989; Chen et al. 2000). To examine the influence of the 3′ exonuclease activity of GST-OsPOLP1 on fidelity, we measured base substitution fidelity of the 3′ exonuclease-deficient mutant D362A for a template G base. The f inc values of the D362A mutant were slightly higher (1.4–4.9 folds) than those of the wild type protein (Table 1). The behavior of this mutant is similar to that of Klenow fragment (Bebenek et al. 1990).
GST-OsPOLP1 catalyzes strand displacement on nicked DNA with a 5’-deoxyribose phosphate
Chloroplasts are one of the principal locations in plant cells for generation of reactive oxygen species. As such, it is likely that oxidative DNA damage occurs frequently in the chloroplast genome (Kimura and Sakaguchi 2006). The lesions in the nuclear genome of mammalian cells are mainly eliminated by base excision repair (BER), which contains two subpathways, short-patch BER and long-patch BER (Dianov et al. 2003). Therefore, it is possible that a pathway similar to BER repairs oxidative DNA damage in the plastidial genome. In the short-patch BER pathway, a 5′-deoxyribose phosphate (dRP), created by a class II apurinic/apyrimidinic endonuclease, is usually removed by dRP lyase activity. Several polymerases show this activity, for example, prokaryotic PolIs, Polβ, and Polγ (Matsumoto and Kim 1995; Pinz and Bogenhagen 2000). We therefore tested GST-OsPOLP1 for dRP lyase activity. For this assay, nicked DNA with a 5′ labeled dRP was created by pretreatment with E. coli uracil-N-glycosylase, which eliminated more than 95% of deoxyuracil residues from the substrate (data not shown). Under our experimental conditions, GST-OsPOLP1 had no detectable dRP lyase activity, whereas the positive control, Polβ, displayed clear activity (Fig. 5A). This finding led us to hypothesize that OsPOLP1 might repair a nick with a dRP by catalyzing the strand displacement that occurs during long-patch BER pathway (Dianov et al. 2003). To investigate this hypothesis, we examined the strand displacement activity of GST-OsPOLP1 on nicked DNA with a 5′-phosphate and a dRP. The polymerase yielded reaction products for all three types of template (upper panels in Fig. 5B). However, it was possible that GST-OsPOLP1 elongated the primer of the DNA template following the degradation of downstream oligonucleotides by its exonuclease activity. To check this possibility, GST-OsPOLP1 was incubated under the same conditions above with nicked DNA with a 5′ labeled dRP, as used in Fig. 5A. We found that there was almost no hydrolysis of the 5′ labeled downstream oligonucleotide (lower panels in Fig. 5B). Overall, these results demonstrate that GST-OsPOLP1 efficiently catalyzed strand displacement on nicked DNA with a dRP.
GST-OsPOLP1 displays the strand displacement activity for nicked DNA with a dRP. Asterisks indicate the position of the 32P label. (A) dRP lyase activtity. One hundred fmol of the dRP1 substrate was incubated with 500 fmol of GST-OsPOLP1 or 18 fmol of Polβ at 37°C. (B) Strand displacement activity. GST-OsPOLP1 (200 fmol) was incubated with 100 fmol of the nicked template with a 5′-phosphate (left panel, circles), or with a dRP (middle panel, squares), or the recessed template without a downstream primer (right panel, triangles) at 37°C. The strand displacement activity for the nicked template with a dRP was calculated from the band intensities except for the 17-nt template
Discussion
Analysis of the rice genome database indicated that two POLPs are encoded in the rice genome. The 3′ exonuclease and polymerase domains are highly conserved between OsPOLP1 and the predicted OsPOLP2 gene product, whereas the N-terminal 230-amino acid region, which probably has no enzymatic activity, is relatively unique. Therefore, these two polymerases might possess similar enzymatic characteristics, but might interact with different proteins via their N-terminal regions.
The polymerase domain of POLPs contains two extra sequences that are absent in prokaryotic PolIs. Deletion of insert I from GST-OsPOLP1 caused a decrease in DNA synthetic activity, processivity and DNA binding activities, suggesting that this sequence is essential for efficient binding affinity to the template. A similar effect has been reported in the complex between T7 DNA polymerase and thioredoxin (Tabor et al. 1987). Thioredoxin binds to an extended loop situated in the thumb subdomain of T7 DNA polymerase, and is thought to function like a sliding clamp to stabilize the polymerase-DNA complex (Doublie et al. 1998; Tabor et al. 1987). From these observations, we suggest that insert I of POLPs has a similar role to thioredoxin. Previous studies demonstrated that the processivity of polymerases generally correlates with their frameshift fidelity (Kunkel 1985a,b; Minnick et al. 1996). For instance, Polγ displays high processivity and high frameshift fidelity, whereas Polβ is distributive and error-prone causing frameshift mutations (Kunkel 1985a, b). Therefore, insert I of POLPs might also have an important role in preventing frameshift errors by elevating their processivity. In contrast, Δinsert II showed no DNA synthetic activity but, like Δinsert I, showed low DNA binding activity to a template. This indicates that insert II functions in both nucleotide incorporation into a template and binding affinity to a template. One other possibility that we have yet to exclude is that deletion of these inserts simply causes structural deformation of the polymerase domain.
GST-OsPOLP1 showed relatively high fidelity for base substitutions (approximately 10−4–10−5). However, it is likely that the 3′ exonuclease of GST-OsPOLP1 has little function for proofreading mispairs on DNA, and its fidelity is 10–100 fold lower than those of replicative polymerases such as bacteria PolIII and eukaryotic Polγ (Bloom et al. 1997; Graziewicz et al. 2006). General replicative polymerases are known to form complexes with accessory subunits and DNA binding proteins, and their enzymatic activities are altered by these protein-protein interactions (Bloom et al. 1997; Chen et al. 2000; Hübscher et al. 2002; Longley et al. 2001). Therefore, POLPs might require interactions with other proteins in order to more accurately accomplish plastidial DNA replication.
The machinery of plastidial DNA replication is largely unknown. A previous study showed that a primase activity is present in chloroplasts as well as nuclei (Nielsen et al. 1991). In addition, according to the rice genome database, rice cells contain the homolog of phage T7 gene 4 protein, which has primase and helicase domains (GenBank accession number AP005610). The human homolog of this protein (Twinkle), which lacks the primase domain, is thought to be involved in the maintenance of the mitochondrial genome (Spelbrink et al. 2001). Moreover, phage T7 gene 4 protein interacts with T7 DNA polymerase, which is involved in the T7 DNA replication complex (Nakai and Richardson 1986). From these observations, we speculate that RNA primers are produced by the phage T7 gene 4-like protein at the first step of plastidial DNA replication, and that POLPs catalyze DNA synthesis from the RNA primers. Contrary to this suggestion, however, it was recently proposed that chloroplast DNA replication is initiated by DNA recombination. In this model, RNA synthesis is not essential and the initial stage is D-loop formation between the ends of a DNA monomer and another DNA molecule (Bendich 2004; Sato 2001). Further molecular biological and genetic experiments are needed to elucidate the mechanism of plastidial DNA replication.
We also explored the possible role of POLPs in plastidial DNA repair. Unlike prokaryotic PolIs and eukaryotic Polγ (Pinz and Bogenhagen 2000), we could not detect a dRP lyase activity of GST-OsPOLP1. However, since the dRP lyase reaction by A-family polymerases proceeds much more slowly than that by Polβ in vitro (Pinz and Bogenhagen 2000), we cannot rule out the possibility that GST-OsPOLP1 has the ability to eliminate a dRP. In contrast, GST-OsPOLP1 efficiently catalyzed strand displacement on nicked DNA with a dRP. Human Y-family polymerases with low fidelity inefficiently catalyze DNA synthesis on such a template, probably because DNA synthesis by these polymerases increases the incidence of mutagenesis in BER (Haracska et al. 2003). Our observation therefore suggests that dRP residues in the plastidial genome might be primarily repaired by a pathway involving POLPs that is similar to long-patch BER.
In the present study, we investigated the biochemical properties of GST-OsPOLP1. Results from a series of experiments suggested that POLPs have the necessary characteristics to participate in DNA replication and repair. Moreover, since the mitochondrial polymerase, Polγ, is absent in plant cells (Burger et al. 2001; Kimura and Sakaguchi 2006), we speculate that POLPs are involved in maintenance of the mitochondrial genome as well as that of plastids. Further analyses of protein networks and of transgenic plants should provide insights into the biological roles of POLPs. Information on the functions of the polymerases would help us to understand DNA replication and repair in plant organelles.
Abbreviations
- Pol:
-
DNA polymerase
- POLP:
-
DNA polymerase π
- GST:
-
Glutathione S-transferase
- E. coli :
-
Escherichia coli
- dNTP:
-
Deoxynucleoside-5-triphosphate
- BER:
-
Base excision repair
- dRP:
-
5′-deoxyribose phosphate
References
Bambara RA, Uyemura D, Choi T (1978) On the processive mechanism of Escherichia coli DNA polymerase I. Quantitative assessment of processivity. J Biol Chem 253:413–423
Bebenek K, Joyce CM, Fitzgerald MP, Kunkel TA (1990) The fidelity of DNA synthesis catalyzed by derivatives of Escherichia coli DNA polymerase I. J Biol Chem 265:13878–13887
Beese LS, Derbyshire V, Steitz TA (1993) Structure of DNA polymerase I Klenow fragment bound to duplex DNA. Science 260:352–355
Bendich AJ (2004) Circular chloroplast chromosomes: the grand illusion. Plant Cell 16:1661–1666
Bernad A, Blanco L, Lazaro JM, Martin G, Salas M (1989) A conserved 3'-5' exonuclease active site in prokaryotic and eukaryotic DNA polymerases. Cell 59:219–228
Bloom LB, Chen X, Fygenson DK, Turner J, O'Donnell M, Goodman MF (1997) Fidelity of Escherichia coli DNA polymerase III holoenzyme. The effects of β, γ complex processivity proteins and ε proofreading exonuclease on nucleotide misincorporation efficiencies. J Biol Chem 272:27919–27930
Boosalis MS, Mosbaugh DW, Hamatake R, Sugino A, Kunkel TA, Goodman MF (1989) Kinetic analysis of base substitution mutagenesis by transient misalignment of DNA and by miscoding. J Biol Chem 264:11360–11366
Boosalis MS, Petruska J, Goodman MF (1987) DNA polymerase insertion fidelity. Gel assay for site-specific kinetics. J Biol Chem 262:14689–14696
Burgers PM, Koonin EV, Bruford E, Blanco L, Burtis KC, Christman MF, Copeland WC, Friedberg EC, Hanaoka F, Hinkle DC, Lawrence CW, Nakanishi M, Ohmori H, Prakash L, Prakash S, Reynaud CA, Sugino A, Todo T, Wang Z, Weill JC, Woodgate R (2001) Eukaryotic DNA polymerases: proposal for a revised nomenclature. J Biol Chem 276:43487–43490
Chen X, Zuo S, Kelman Z, O'Donnell M, Hurwitz J, Goodman MF (2000) Fidelity of eucaryotic DNA polymerase δ holoenzyme from Schizosaccharomyces pombe. J Biol Chem 275:17677–17682
Derbyshire V, Grindley ND, Joyce CM (1991) The 3'-5' exonuclease of DNA polymerase I of Escherichia coli: contribution of each amino acid at the active site to the reaction. Embo J 10:17–24
Dianov GL, Sleeth KM, Dianova II, Allinson SL (2003) Repair of abasic sites in DNA. Mutat Res 531:157–163
Doublie S, Tabor S, Long AM, Richardson CC, Ellenberger T (1998) Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 391:251–258
Fygenson DK, Goodman MF (1997) Appendix. Gel kinetic analysis of polymerase fidelity in the presence of multiple enzyme DNA encounters. J Biol Chem 272:27931–27935
Graziewicz MA, Longley MJ, Copeland WC (2006) DNA polymerase γ in mitochondrial DNA replication and repair. Chem Rev 106:383–405
Haracska L, Prakash L, Prakash S (2003) A mechanism for the exclusion of low-fidelity human Y-family DNA polymerases from base excision repair. Genes Dev 17:2777–2785
Hübscher U, Maga G, Spadari S (2002) Eukaryotic DNA polymerases. Annu Rev Biochem 71:133–163
Kikuchi S, Satoh K, Nagata T, Kawagashira N, Doi K, Kishimoto N, Yazaki J, Ishikawa M, Yamada H, Ooka H, Hotta I, Kojima K, Namiki T, Ohneda E, Yahagi W, Suzuki K, Li CJ, Ohtsuki K, Shishiki T, Otomo Y, Murakami K, Iida Y, Sugano S, Fujimura T, Suzuki Y, Tsunoda Y, Kurosaki T, Kodama T, Masuda H, Kobayashi M, Xie Q, Lu M, Narikawa R, Sugiyama A, Mizuno K, Yokomizo S, Niikura J, Ikeda R, Ishibiki J, Kawamata M, Yoshimura A, Miura J, Kusumegi T, Oka M, Ryu R, Ueda M, Matsubara K, Kawai J, Carninci P, Adachi J, Aizawa K, Arakawa T, Fukuda S, Hara A, Hashizume W, Hayatsu N, Imotani K, Ishii Y, Itoh M, Kagawa I, Kondo S, Konno H, Miyazaki A, Osato N, Ota Y, Saito R, Sasaki D, Sato K, Shibata K, Shinagawa A, Shiraki T, Yoshino M, Hayashizaki Y, Yasunishi A (2003) Collection, mapping, and annotation of over 28,000 cDNA clones from japonica rice. Science 301:376–379
Kimura S, Sakaguchi K (2006) DNA repair in plants. Chem Rev 106:753–766
Kimura S, Uchiyama Y, Kasai N, Namekawa S, Saotome A, Ueda T, Ando T, Ishibashi T, Oshige M, Furukawa T, Yamamoto T, Hashimoto J, Sakaguchi K (2002) A novel DNA polymerase homologous to Escherichia coli DNA polymerase I from a higher plant, rice (Oryza sativa L.). Nucl Acids Res 30:1585–1592
Kunkel TA (2004) DNA replication fidelity. J Biol Chem 279:16895–16898
Kunkel TA (1985a) The mutational specificity of DNA polymerase-β during in vitro DNA synthesis. Production of frameshift, base substitution, and deletion mutations. J Biol Chem 260:5787–5796
Kunkel TA (1985b) The mutational specificity of DNA polymerases-α and -γ during in vitro DNA synthesis. J Biol Chem 260:12866–12874
Lee JW, Blanco L, Zhou T, Garcia-Diaz M, Bebenek K, Kunkel TA, Wang Z, Povirk LF (2004) Implication of DNA polymerase λ in alignment-based gap filling for nonhomologous DNA end joining in human nuclear extracts. J Biol Chem 279:805–811
Li Y, Korolev S, Waksman G (1998) Crystal structures of open and closed forms of binary and ternary complexes of the large fragment of Thermus aquaticus DNA polymerase I: structural basis for nucleotide incorporation. Embo J 17:7514–7525
Longley MJ, Nguyen D, Kunkel TA, Copeland WC (2001) The fidelity of human DNA polymerase γ with and without exonucleolytic proofreading and the p55 accessory subunit. J Biol Chem 276:38555–38562
Marini F, Kim N, Schuffert A, Wood RD (2003) POLN, a nuclear PolA family DNA polymerase homologous to the DNA cross-link sensitivity protein Mus308. J Biol Chem 278:32014–32019
Masuda Y, Takahashi M, Tsunekuni N, Minami T, Sumii M, Miyagawa K, Kamiya K (2001) Deoxycytidyl transferase activity of the human REV1 protein is closely associated with the conserved polymerase domain. J Biol Chem 276:15051–15058
Matsumoto Y, Kim K (1995) Excision of deoxyribose phosphate residues by DNA polymerase β during DNA repair. Science 269:699–702
Minnick DT, Astatke M, Joyce CM, Kunkel TA (1996) A thumb subdomain mutant of the large fragment of Escherichia coli DNA polymerase I with reduced DNA binding affinity, processivity, and frameshift fidelity. J Biol Chem 271:24954–24961
Mori Y, Kimura S, Saotome A, Kasai N, Sakaguchi N, Uchiyama Y, Ishibashi T, Yamamoto T, Chiku H, Sakaguchi K (2005) Plastid DNA polymerases from higher plants, Arabidopsis thaliana. Biochem Biophys Res Commun 334:43–50
Nakai H, Richardson CC (1986) Interactions of the DNA polymerase and gene 4 protein of bacteriophage T7. Protein–protein and protein–DNA interactions involved in RNA-primed DNA synthesis. J Biol Chem 261:15208–15216
Nielsen BL, Rajasekhar VK, Tewari KK (1991) Pea chloroplast DNA primase: characterization and role in initiation of replication. Plant Mol Biol 16:1019–1034
Patel PH, Loeb LA (2000) DNA polymerase active site is highly mutable: evolutionary consequences. Proc Natl Acad Sci USA 97:5095–5100
Patel PH, Suzuki M, Adman E, Shinkai A, Loeb LA (2001) Prokaryotic DNA polymerase I: evolution, structure, and “base flipping” mechanism for nucleotide selection. J Mol Biol 308:823–837
Pinz KG, Bogenhagen DF (2000) Characterization of a catalytically slow AP lyase activity in DNA polymerase γ and other family A DNA polymerases. J Biol Chem 275:12509–12514
Sato N (2001) Was the evolution of plastid genetic machinery discontinuous? Trends Plant Sci 6:151–155
Seki M, Masutani C, Yang LW, Schuffert A, Iwai S, Bahar I, Wood RD (2004) High-efficiency bypass of DNA damage by human DNA polymerase Q. Embo J 23:4484–4494
Spelbrink JN, Li FY, Tiranti V, Nikali K, Yuan QP, Tariq M, Wanrooij S, Garrido N, Comi G, Morandi L, Santoro L, Toscano A, Fabrizi GM, Somer H, Croxen R, Beeson D, Poulton J, Suomalainen A, Jacobs HT, Zeviani M, Larsson C (2001) Human mitochondrial DNA deletions associated with mutations in the gene encoding Twinkle, a phage T7 gene 4-like protein localized in mitochondria. Nat Genet 28:223–231
Tabor S, Huber HE, Richardson CC (1987) Escherichia coli thioredoxin confers processivity on the DNA polymerase activity of the gene 5 protein of bacteriophage T7. J Biol Chem 262:16212–16223
Takata K, Shimizu T, Iwai S, Wood RD (2006) Human DNA polymerase N (POLN) is a low fidelity enzyme capable of error-free bypass of 5S-thymine glycol. J Biol Chem 281:23445–23455
Takeuchi R, Oshige M, Uchida M, Ishikawa G, Takata K, Shimanouchi K, Kanai Y, Ruike T, Morioka H, Sakaguchi K (2004) Purification of Drosophila DNA polymerase ζ by REV1 protein-affinity chromatography. Biochem J 382:535–543
Acknowledgements
We thank Dr. Kei-ichi Takata (University of Pittsburgh Cancer Institute, Hillman Cancer Center, Research Pavilion, Pittsburgh, PA) for the helpful discussion of this work. R.T. and S.K. were supported by Research Fellowships of the Japan Society for the Promotion of Science.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Takeuchi, R., Kimura, S., Saotome, A. et al. Biochemical properties of a plastidial DNA polymerase of rice. Plant Mol Biol 64, 601–611 (2007). https://doi.org/10.1007/s11103-007-9179-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-007-9179-2