Abstract
Two putative glycosyltransferases in Arabidopsis thaliana, designated reduced residual arabinose-1 and -2 (RRA1 and RRA2), are characterized at the molecular level. Both genes are classified in CAZy GT-family-77 and are phylogenetically related to putative glycosyltranferases of Chlamydomonas reinhardtii. The expression pattern of the two genes was analyzed by semi-quantitative RT-PCR using mRNA extracted from various organs of bolting Arabidopsis thaliana plants. In addition, promoter::gusA analysis of transgenic Arabidopsis thaliana containing a fusion between either the RRA-1 or -2 promoter fragment and the gusA reporter gene showed that whereas the RRA1 promoter was primarily active in the apical meristem, the expression pattern of the RRA2 promoter was more diverse but also highly active in the meristematic region. In addition, T-DNA mutant insertion lines of both RRA-1 and -2, were identified and characterized at the molecular and biochemical level. Monosaccharide compositional analyses of cell wall material isolated from the meristematic region showed a ca. 20% reduction in the arabinose content in the insoluble/undigested cell wall residue after enzymatic removal of xyloglucan and pectic polysaccharides. These data indicate that both RRA-1 and -2 play a role in the arabinosylation of cell wall component(s).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The cell wall (CW) of higher plants represents a unique type of extra-cellular matrix with numerous structural, protective and growth-regulating functions. Plant CWs need to be strong enough to prevent the protoplast from bursting under turgor and at the same time possess sufficient yielding properties to permit controlled cell expansion (Carpita and Gibeaut 1993).
The chemical composition of plant CW is well characterized, whereas little is known about the biosynthesis of wall polysaccharides and the principles of CW assembly. The earliest detailed CW model was based on studies of Platanus occidentalis suspension culture cells and proposed a CW of covalently interconnected biopolymer networks (Keegstra et al. 1973). Later models emphasized instead the separability of the different classes of CW polysaccharides observed during wall extraction, and proposed a wall structure that relied heavily on hydrogen bonding and other non-covalent forces (Talbott and Ray 1992; Carpita and Gibeaut 1993; Cosgrove 2001; Somerville et al. 2004). Covalent cross-linking was then mostly viewed as wall maturation processes that followed the completion of cell expansion. Recent observations by Popper and Fry (2005) have re-introduced emphasis on covalent cross-links as crucial to the integrity of the expanding primary wall.
Both types of models maintain that some polymers, notably the tethering heteroxylans and xyloglucans, associate strongly with microfibrillar cellulose, and researchers invariably find galactose- (Gal) and arabinose- (Ara) containing remnants bound to cellulose, even after exhaustive chemical extraction.
The major Ara- and Gal-containing CW polysaccharides are arabinans and type I (arabino)galactans associated with the pectic polysaccharides and the arabinoxylans (Bacic et al. 1988). The CW polysaccharides are, however, not the only carbohydrate-containing components that may bind strongly to cellulose or resist sequential extraction. Some of the less readily extractable polymers may be glycoproteins. The apoplastic space contains a substantial number of glycoproteins, of which a major group are the hydroxyproline-rich glycoproteins (HRGPs). These include the readily soluble arabinogalactan-proteins (AGPs), a class of highly abundant proteoglycans, which contain up to 98% carbohydrate composed mainly of D-Gal and L-Ara (Showalter 1993; Johnson et al. 2003). Two types of glycomodules occur in the AGPs, the complex AGP-glycomodules and the extensin-type glycomodule associated with the Ser-Hyp3 or Ser-Hyp4 motif. Extensins are characterized by the Tyr-X-Tyr or Val-Tyr-Lys crosslinking motif in addition to the extensin glycomodule motifs (for reviews see Showalter 1993; Johnson et al. 2003). The extensin glycomodule consists of short arabinan chains and single galactosyl residues attached directly to the protein. Extensins are deposited in the wall as monomers, and remain extractable for a very short time before they are rendered resistant to extraction (Lamport 1963) probably by peroxidase catalyzed cross-linking (e.g., Propper and Fry, 2005).
Initially, CAZy GT-family-77 (Coutinho and Henrissat 1999; Coutinho et al. 2003; http://afmb.cnrs-mrs.fr/CAZY/index.html) was formed using two retaining Arabidopsis thaliana xylosyltransferases (XylTs), At4g01770 and At4g01750 (Egelund et al. 2004). The two XylTs have been shown to transfer xylose on to fucose forming an α-(1,3)-linkage and have been shown to be involved in the synthesis of pectic rhamnogalacturonan-II (Egelund et al. 2006). Together with the UDP-Gal:fucoside (1,3)-α-d-galactosyltransferase (GalT) from Dictyostelium discoideum, these are presently the only known activities in CAZy GT-family-77.
In the present work, two homologous Arabidopsis thaliana genes, At1g75120 and At1g75110, belonging to CAZy GT-family-77, are characterized by promoter::gusA mediated expression analysis and by analysis of the corresponding T-DNA insertion lines. CWs isolated from the meristematic tissues from T-DNA insertion lines in both At1g75120 and At1g75110, displayed a 20% reduction in the Ara content of the cellulosic residue following enzyme mediated extraction of the wall and were thus designated, reduced residual arabinose 1 and -2 (rra1; At1g75120 and rra2; At1g75110), respectively. Possible functions in the synthesis of less readily extractable CW polymers and glycoproteins are discussed.
Materials and methods
Sequence retrieval and phylogenetic analysis
The identification of the six putative GTs (At4g01220, At1g56550, At4g01770, At4g01750, At1g75120 and At1g75110) has recently been published (Egelund et al. 2004). All other accessions are available through the CAZy database (http://www.afmb.cnrs-mrs.fr/CAZY/index.html). Alignments and phylogenetic trees were generated using Muscle (Edgar 2004) and modified using TreeView version 1.6.6 (Page 1996).
Plant material and growth conditions
Arabidopsis thaliana ecotype Columbia (Col-0; WT) and T-DNA insertional mutants, were grown in soil in a controlled-environment growth chamber (Percival AR-60 I, Boone, IA, USA) at a photosynthetic flux of 100–120 mol photons m−2 s−1 at 8 h light/16 h dark cycle, 20°C and 70% relative humidity.
T-DNA insertional mutants in At1g75120 (Garlic_76_G04 and SAIL_590_G09), and in At1g75110 (Garlic_244_A03 and SAIL_70_D08), were obtained through Syngenta (Garlic lines, http://www.tmri.org/pages/collaborations/garlic_files/GarlicAnalysis.html) and the Salk Institute (SAIL lines, Alonso et al. 2003), respectively.
Transcripts detected by RT-PCR
RNA was extracted from root, rosette leaves, cauline leaves, stem, siliques and flower from 4-week old bolting Arabidopsis thaliana plants. The RNA was extracted using the TRIzol reagent (Invitrogen A/S, Taastrup, Denmark) according to the manufacture’s instructions. RT-PCR using 3 μg of RNA from each tissue type, was performed according to Mikkelsen et al. (2003), with the exception of the cycle parameters in the PCR which were as follows: 3 min at 94°C, (25 cycles for 18S and 45 cycles for At1g75120 and At1g75110) of: 94°C for 30 s, 50°C for 30 s and 68°C for 30 s followed by 12 min at 72°C. The following primer sets were used to amplify each transcript specifically: At1g75120_forward, 5′-tgagaaaggatgaggaaatga-3′; At1g75120_reverse, 5′-atggttccaattttcctcaa-3′; At1g75110_forward, 5′-gaaagaatcaagaactgaaga-3′; At1g75110_reverse, 5′-aggataactgattataacaaca-3′; 18S_forward, 5′-taaggattgacagactgagagct-3′; 18S_reverse, 5′-aatacatcagtgtagcgcgcgt-3′. To ensure reproducibility the experiment was replicated; both experiments produced identical results.
Screening for homozygous T-DNA insertional mutants
Homozygous plants were identified by PCR, using a pair of gene-specific primers designed to anneal outside of the T-DNA insertion, which in case of homozygozity does not produce a band of the predicted size (negative selection): forward primer (5′-atggcggttcgtaaagag-3′ for At1g75120, 5′-atggcgggtcgcagagac-3′ for At1g75110) and reverse primer (5′-ctatgaaccatcacggaac-3′ for At1g75120, 5′-tcaatctgaaccatcggg-3′ for At1g75120). As a control for the negative selection, a set of At4g01750 specific primers (5′-atggcgagaaacaacag-3′ forward and 5′-ttactgcaatttccctaatg-3′), were used. Genomic DNA was isolated using the Nucleon PhytoPure kit (Amersham Biosciences, UK).
Strategy for cloning of the RRA-1 and -2 promoters
The promoter regions 2.0 kb upstream from the start codon of RRA-1 (At1g75120) and -2 (At1g75110) were amplified from genomic DNA using PCR as described in the “Polymerase chain reaction” section and sub-cloned into pCR2.1-TOPO (as described by the manufacturer, Invitrogen Life Technologies) and cloned into pCambia 1301 (NcoI and KpnI).
The final pCambia 1301 vectors containing either the At1g75120 or At1g75110 promoter fragments were sequenced in both orientations, using the services provided by MWG (MWG-Biotech AG, Germany, http://www.mwg-biotech.com/html/all/index.php).
The pCambia 1301 vector (Clunies Ross Street at Dickson Rd, Black Mountain ACT 2601, Australia) contains the 35S::gusA cassette. The catalase intron (GIS) is inserted within the coding sequence of gusA, which ensures that the gene is not expressed in bacteria but only upon transfer to plants.
Arabidopsis thaliana transformation
The binary plasmid pCambia 1301, containing the promoter::gusA fusion construct, was mobilized into Agrobacterium tumefaciens strain pGV3850 by electroporation. Overnight cultures in LB medium supplemented with rifampicin (100 mg/l) and kanamycin (50 μg/ml) were grown to OD600 = 2, centrifuged and the cells re-disolved in infiltration medium (0.22% Murashige and Skoog medium (Sigma-Aldrich, Denmark), 0.32% g Gamborgs B5 medium (Sigma), 5% w/v sucrose, 0.03% v/v Silwet (Lehle Seed)) to give a OD600 = 0.8. Arabidopsis thaliana plants were placed upside-down in the solution for 15 min and placed in the greenhouse. Seeds were harvested after ca. 8 weeks. The T1 transgenic plants were selected on MS plates supplemented with hygromycin (50 μg/ml). Before plating on MSO plates, seeds were sterilized by incubating 2 × 10 min in 5% NaOCl; 0.02% Triton X-100, followed by washes in sterile water, and then germinated on MS agar containing 50 μg/ml hygromycin until positive transformants could be identified and transferred to soil. The transformed plants were subjected to PCR to confirm the T-DNA inserts, and 10 independent T2 lines of the RRA1 and RRA2 promoter::gusA transformed Arabidopsis thaliana plants were analyzed for β-glucuronidase (GUS) activity.
Polymerase Chain Reaction
The Polymerase Chain Reaction (PCR) was used to clone promoter regions from gDNA, and to check transformed Arabidopsis thaliana plants to ensure the presence of the T-DNA construct harboring the promoter::gusA construct in the genome. Taq polymerase (Invitrogen Life Technologies) was used for analytical purposes and PFU-polymerase (Stratagene) was used to generate the various constructs. Reaction mixture, mixed in 0.2 ml MicroAmp@ Reaction Tube With Cap (Perkin Elmer): 3.0 μl Gibco BRL@ 10 × PCR buffer (Invitrogen Life Technologies), 1 μl 50 mM MgCl2, 1.0 μl 5 mM dNTP’s , 2.0 μl 10 pM 5′ primer, 2.0 μl 10 pM 3′ primer, MQ water up to 30 μl, approximately 1–5 ng template DNA, 0.1 μl Gibco BRL@ Taq polymerase (5 U/μl) (Invitrogen Life Technologies) or 0.5 μl Stratagene@ cloned PFU polymerase (2 U/μl) (Stratagene). MgCl2 was not added when using PFU. The PCR reaction was carried out on a Progene thermal cycler (Techne) using the following standard program: 97°C, 3 min, then 30 cycles at: 94°C, 30 s, followed by 50°C 30 s (annealing) and elongation for l min per kilobase (kb) fragment at 72°C, followed by 10 min at 72°C and ∞ at 4°C. Three microliter of the PCR reaction was tested on 1% agarose gels. The following primers were used (all listed from 5′ to 3′): At1g75120_promoter_forward, atatggatccggtaccaaataccaagattggctggta; At1g75120_promoter_reverse, atatgtcgacccatggtgccaattgaagttatccgaa; At1g75110_promoter_forward, atatggatccggtacctatatgcttgttaagcaagata; At1g75110_promoter_reverse, atatgtcgacccatggtgccgaaatgacgcagaagccaa.
Analysis of β-glucuronidase expression
Transformed Arabidopsis thaliana plants carrying the promoter::gusA reporter gene constructs were tested for GUS activity, using the protocol from Jefferson et al. (1987), with a few modifications. Plant material was incubated in GUS staining buffer (50 mM sodium phosphate buffer (pH 7.2), 0.1% Triton X-100, 10 mM EDTA, 5 mM K4Fe(CN)6, 5 mM K3Fe(CN)6, 0.1% X-gluc (w/v)), and incubated at 37°C O/N. Samples were then cleared in 80% ethanol. All experiments were performed in triplicate with 10 independent transgenic lines from each construct. For each construct, all individual lines had a similar GUS staining pattern, with slight differences in intensity.
All pictures were taken using a digital camera (Nikon Coolpix 990) and were later modified using the Adobe Photoshop Version 7.0. © Adobe Systems Incorporated software. Small details were viewed using a microscope (Olympus BH-2, Olympus Instruments, UK) equipped with a normal light source.
Crude cell wall extraction: alcohol insoluble residues (AIR)
Alcohol insoluble residues were isolated as described by Abdulrazzak et al. (2005), with few modifications. Briefly, plant material was harvested and frozen in liquid nitrogen. The tissue was then homogenized using a Retschmill machine (model MM200, Retsch, Haan, Germany) at 25 Hz for 1 min. The ground tissue was then suspended in 70% ethanol, vortexed, and pelleted by centrifugation at 10,000 g for 10 min. The ethanol was decanted, and this procedure was then repeated using chloroform:methanol (1:1,v/v), until all chlorophyll was removed. The pellet was then washed twice in acetone. The remaining pellet was dried under vacuum for 5 min and either processed directly or stored until further use.
Fractionation of the cell wall polymers
Isolation and characterization of xyloglucan (XG) was performed as follows. The isolated CW pellet (100 μg AIR) was incubated with 3 U of xyloglucanase (XEG was provided by Novozymes (Bagsvaerd, Denmark), and purified according to Pauly et al. 1999), in 1 ml of 50 mM ammonium formate buffer, for 36 h at 37°C (shaking). After 24 h of incubation a further 3 U of XEG was added to maximize the removal of XG from the CW pellet (1 U of XEG releases 1 μmol of reducing XG oligosaccharide per hour). Samples were then briefly centrifuged to pellet the insoluble CW components, and the supernatant containing the solubilized XG collected for analysis using matrix assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry (MS) as described by Lerouxel et al. (2002). Xylan was extracted in a similar manner by digestion of CW material using a 50 mM ammoinium formate buffer (pH = 4.5) with 3 U of xylanase M1 (Megazyme, Bray Ireland). After incubation for 24 h at 37°C (shaking) the supernatant was collected by centrifugation (10 min at 20,000g) and stored for analysis.
The remaining CW pellet was washed twice in 50 mM ammonium formate buffer, and used for extraction of the pectic polysaccharides as described by Sørensen et al. (2000), with the following modifications. The CW pellet was incubated with 3 U of pectin methylesterase (PME) and 20 U of endo-polygalacturonanase (EPG), in 1 ml of 50 mM ammonium formate buffer, for 36 h at 37°C (shaking). After 24 h of incubation a further 3 U of PME and 20 U of EPG were added to maximize the removal of the pectic polysaccharides from the CW pellet. Samples were then briefly centrifuged to pellet the large insoluble CW components, and the supernatant containing the solubilized pectic polysaccharides collected and further separated into rhamnogalacturonan I (RG I), rhamnogalacturonan II (RG II) and homogalacturonan (HG), using size exclusion chromatrography on a Superose 12 column (1 × 30 cm, Amersham Pharmacia), as described by Sørensen et al. (2000). The remaining CW pellet containing insoluble and undigested CW material was washed twice in 50 mM ammonium formate buffer, and twice in acetone before further processing. Extraction of water-soluble polysaccharides and proteoglycans was done as described by Schultz et al. (2000). Precipitation and collection of AGPs from the water soluble extract were done by using the β-glucosyl Yariv reagent (β-GlcY) as described by Gane et al. (1995). All CW extracted fractions were freeze-dried and stored for further analysis. The following CW fractions were obtained: total CWs (AIR), RG I, RG II, HG, XG, xylan and the remaining insoluble/undigested CW pellet.
Monosaccharide composition analysis
For neutral sugar analysis the freeze-dried CW fractions were hydrolysed with TFA, reduced and acetylated to form their corresponding alditol acetates and then analysed by gas chromatography mass-spectrometry (GC-MS) as previously described (Albersheim et al. 1967; Sims and Bacic 1995). Measurement of uronic acid was done using the protocol described by Blumenkrantz and Asboe-Hansen (1973).
Results
Organization of CAZy GT-family-77
The CAZy GT-family-77 presently comprises 36 accessions, 18 from Arabidopsis thaliana, 15 from Oryza sativa, one from Linum usitatissimum and the Dictyostelium discoideum (1,3)-α-d-GalT, AX3, (Fig. 1A). At5g40900 appears to be incomplete and as the flanking sequences do not immediately suggest how to reconstruct the full gene. We hypothesize that it is a pseudogene and have thus omitted it from the phylogenetic analysis. The Dictyostelium discoideum (1,3)-α-d-GalT has neither a signal peptide nor a membrane anchor and has been shown to galactosylate Skp1, a cytoplasmic signalling molecule (Ketcham et al. 2004). It is noteworthy that two of the plant sequences are also predicted to be soluble glycosyltransferases (GTs) (At1g28695 and At1g70630) whereas the remainder are predicted to be classical type II transmembrane proteins. All of the sequences are homologous, albeit quite divergent, and the Dictyostelium discoideum (1,3)-α-d-GalT clearly forms an outgroup relative to the higher plant accessions. The phylogenetic tree shown in Fig. 1A cannot be assumed to reliably represent evolutionary relationships due to the low level of identity as well as large differences in length of the coding sequences (Supplemental Figure 1). However, the four clades labeled A to D appear to be robust to different choices of alignment algoritms (e.g., ClustalX v1.81, data not shown) and clade members are joined by 33% or higher identity.
Phylogenetic analysis of the CAZy GT-family-77. (A) A rooted cladogram comprising 35 accessions of CAZy GT-family-77 (17 from Arabidopsis thaliana, 16 from Oryza sativa, one from Linum usitatissimum, the Dictyostelium discoideum (1,3)-α-d-GalT (AX3) and the two additional Chlamydomonas reinhardtii accessions) was generated as described in “Material and methods” section, resulting in four clades (A to D), Clade A contains the two genes RRA1 (At1g75120) and RRA2 (At1g75110), selected for analysis in the present study. Accessions starting with a ‘P’ are from Oryza sativa. Initially CAZy GT-family-77 was seeded using the two Arabidopsis thaliana (1,3)-α-d-XylTs, RGXT1 and RGXT2 (Egelund et al., 2006), which are found in clade B. Bootstrap values were generated from 1.000 re-samplings. (B) RRA-1 and -2 and the two Chlamydomonas reinhardti sequences were aligned over a range of 100 a.a. comprising the D×D-motif (indicated by ‘stars’). The Dictyostelium discoideum (1,3)-α-d-GalT (AX3) was added to this alignment using the profile option in muscle
RRA1 (At1g75120), RRA2 (At1g75110) and a third Arabidopsis thaliana gene, At1g19360 belong to the A-clade, which, we propose, also comprises two Chlamydomonas reinhardtii accessions Chlre3/scaffold_5:813768–817127 (acegs_kg.scaffold_5000061) and Chlre3/scaffold_26:1159362–1161511 (estExt_gwp_1W.C_260040) accessible via the Chlamyomonas reinhardtii database v3.0 (http://genome.jgi-psf.org/Chlre3/Chlre3.home.html). The algal sequences are 33–36% identical to the Arabidopsis thaliana genes and 35–37% identical to the rice sequence. Alignment of the sequences surrounding the catalytic domain (100 a.a. containing the D×D motif, Fig. 1B) between the two Chlamydomonas reinhardtii accessions and the RRA genes display up to 86% identity, indicating a strong evolutionary relationship. The significance of the Chlamydomonas reinhardtii accessions in the A-clade will be elaborated upon in the “Discussion” section. Both RRA1 and RRA2 are located on chromosome 1, separated by only 1.2 kb. In the Arabidopsis thaliana genome, many highly homologous genes are situated in clusters created by recent, local duplication (Coutinho et al. 2003) and it is thus likely that the two genes are paralogs.
The B-clade comprises the two highly identical (1,3)-α-d-XylTs (89% identity), two additional very similar Arabidopsis thaliana genes with unknown function, and the Linum usitatissimum accession. Each of the clades includes one Oryza sativa sequence and clades C and D may represent orthologous pairs of Oryza sativa and Arabidopsis thaliana genes.
RRA1 and RRA2 encode polypeptides of 402 and 428 a.a., respectively. They are predicted to be type II membrane proteins and are 17–23% identical to the accessions in the B-clade. Similarities are in all cases most pronounced in the vicinity of the D×D motif, e.g., as seen in the focused alignment of the out-group accession Dictyostelium discoideum (1,3)-α-d-GalT (AX3), the RRA genes and the two Chlamydomonas reinhardti sequences (Fig. 1B). Corresponding expressed sequence tags (ESTs) have been detected in various monocot and dicot plants, e.g., as seen in Table 1. ESTs representing the RRA genes have also been found, but the two genes are so similar that individual EST-sequences usually cannot be assigned to either gene. ESTs thus provide very little information about RRA1 and RRA2 expression profiles. Homologous sequences from organisms outside the plant kingdom other than the Dictyostelium discoideum (1,3)-α-d-GalT, have not been found.
Expression analysis
The level of RRA-1 and -2 transcripts in various tissues was investigated using RT-PCR (Fig. 2). Whereas a strong 18S rRNA derived band was obtained (18S in Fig. 2), neither RRA-1 nor -2-derived transcripts were detectable after 25 cycles of PCR (data not shown). After 45 cycles of PCR amplification, RRA2 mRNA was detectable (Fig. 2), with strongest expression in roots, flowers and siliques, medium expression in rosette leaves and cauline leaves and low expression in stems. In comparison, RRA1 mRNAs only produced a very faint band in siliques (Fig. 2).
Detection of RRA-1 and -2 transcripts in bolting Arabidopsis thaliana plants, using RT-PCR. RT-PCR was performed with specific primer sets for RRA1, RRA2 and for ribosomal 18S (control) as described in the “Material and methods” section. The RNA used was isolated from tissues of 5-weeks old bolting WT plants. H2O was included as control. Each RT-PCR reaction was performed twice to ensure reproducibility
In addition, the expression pattern of RRA1 and RRA2 was analyzed by promoter::gusA fusions in Arabidopsis thaliana. Expression of the RRA1 promoter was restricted to meristimatic tissue and only seen from the stage in which development of the first true leaves had begun and subsequently throughout the lifespan of the plant (Fig. 3A–D). Tissues where vascular connections are established with the developing leaves also stain intensely (Fig. 3B). In addition, expression was occasionally observed in points of cauline leaf attachments on the primary stem (Fig. 3E). The absence of RRA1 promotor mediated GUS staining, where RRA1 transcripts were detected by RT-PCR, might result from the absense of potential enhancer elements present in, e.g., intron- or the 3′UTR regions of the RRA1 gene. Similar observations have, e.g., been reported from RT-PCR versus GUS analysis of the UDP-d-glucoronate-4-epimerase in Arabidopsis thaliana (Usadel et al. 2004).
Expression of RRA1 promoter::gusA in Arabidopsis thaliana. (A) GUS activity was found in meristematic tissue in seedlings, at the stage in which development of the first true leaves had begun. (B) Close up view of GUS staining of the meristematic region in seedlings, in which the development of the third pair of leaves had begun. (C) GUS staining of the meristematic region in seedlings, in which the development of the third pair of leaves had begun. (D) GUS staining in the meristematic region of adult non-bolting plants. (E) GUS staining observed on the primary stem, adjacent to the point of cauline leaf attachment
Strong GUS activity was found throughout germinating seedlings of RRA2 promoter::gusA transformed Arabidopsis thaliana (Fig. 4A), and throughout seedlings, in which the development of the first true leaves had begun (Fig. 4B). GUS staining activity was rapidly decreasing, though still found throughout all tissues in plants in which the development of fourth pair of leaves had begun (Fig. 4C), and in mature plants GUS activity was restricted to rosette leaf hydathodes and trichome support cells in the adaxial epidermis (Fig. 4D). In bolting Arabidopsis thaliana plants GUS activity was found in cauline leaves (Fig. 4E), in petals (Fig. 4E, G–I), and in both the proximal and distal ends of siliques (Fig. 4F–G). A more detailed analysis revealed that high GUS activity was found in leaves of 2–3-week old plants (Fig. 5A), which rapidly weakened later in development (Fig. 5B–C). The GUS activity was weak in leaves from mature plants and found in rosette leaf hydathodes and trichome support cells in the adaxial epidermis (Fig. 5D–F). GUS activity was found in rosette leaf hydathodes (Fig. 5G), and in trichome support cells (Fig. 5G–H). Strong GUS activity was also found in the apical meristem, in the junction of newly developing leaves (Fig. 5I). GUS activity was found in the vascular tissue in both root tip (Fig. 5J), and in older non-elongating roots (Fig. 5K). These data are consistent with the RT-PCR analysis where RRA1 transcripts could not be detected in roots, rosette leaves, cauline leaves, stems and flowers from 4 week bolting WT plants whereas RRA2 transcripts were detected in all tissues tested (Fig. 2).
Expression of RRA2 promoter::gusA in Arabidopsis thaliana. (A–B) Strong GUS activity was found throughout newly developing seedlings (A), and throughout seedlings, in which the development of the first true leaves had begun (B). (C–D) GUS activity was rapidly decreasing, though still found throughout all tissues, in plants in which the development of forth pair of leaves had begun (C) and in mature plants GUS activity was restricted to rosette leaf hydathodes and trichome support cells in the adaxial epidermis (D). (E–I) In bolting Arabidopsis thaliana plants GUS activity was found in cauline leaves (E), in petals (E, G, H and I), and in both proximal and distal ends of siliques (F and G)
Detailed expression analysis of RRA2 promoter::gusA in Arabidopsis thaliana (A–C) High GUS expression was found in leaves from 2- to 3-weeks old plants (A), and then the GUS activity rapidly weakened in more mature leaves (B and C). (D–F) The GUS activity was weak in leaves from mature plants and only found throughout cell hydathodes and trichome support cells in the ad axial epidermis (D, E and F). (G–H) GUS activity was found in rosette leaf hydathodes (G), and in trichome support cells (G and H). (I) Strong GUS activity was found in the apical meristem, in the junction of newly developing leaves. (J–K) GUS activity was found in root tips (J), and in older non-growing roots (K)
T-DNA insertional mutants for At1g75120 and At1g75110: rra1 and rra2
Two T-DNA insertional mutants for RRA1, Garlic_76_G04 & SAIL_590_G09, with insertions in the first intron and in the last exon, respectively (Fig. 6A) and two T-DNA insertional mutants for RRA2, Garlic_244_A03 & SAIL_70_D08, with insertion in the last exon (Fig. 6B), as determined by sequencing from the T-DNA left border (Alonso et al. 2003) were selected for this investigation. Homozygous plants for all four T-DNA insertional lines were identified using PCR-based screening (Fig. 6C–D).
T-DNA insertion mutants in RRA-1 and -2. Positions of the two allelic T-DNA insertional mutants in both of the RRA1 and -2 gene. Boxes indicate exons and lines indicate introns. (A) The T-DNA is inserted in the first intron (Garlic_76_G04) and in the last exon (SAIL_590_G09). (B) Both T-DNAs are inserted into the second exon (Garlic_244_A03 and SAIL_70_D08). (C) Homozygous lines identified in SAIL_590_G09 and Garlic_76_G04, respectively. (D) Homozygous lines identified in Garlic_244_A03 and SAIL_70_D08, respectively. The PCR analysis was carried out using a set of gene specific primers outside of the T-DNA insertions. ‘DNA control’ refers to the amplification of At4g01750. Triangles indicate homozygous lines. The results were confirmed using three independent analyses, and were also confirmed in the T2 generation (data not shown)
No obvious morphological differences between the WT and the four mutant lines were observed. Histological analysis of root and leaf tissue of 1-week old seedlings, and of 5-week old bolting plants did not reveal any differences between WT and mutant plants. In addition, 1-week old seedlings were analyzed using a panel of antibodies raised against CW specific epitopes both by applying the “whole-mount” immunolabelling technique to the primary roots (Willats et al. 2001) and by ELISA of fractionated CW-material from whole seedlings. The panel of antibodies comprised: LM6 (RG I, (1,5)-α-l-arabinan), LM5 (RG I, (1,4)-β-d-galactan), MAC 207 (plasma membrane AGPs), LM2 (AGPs, β-d-glucuronosyl), JIM5 (low esterified HG) and IGII (RG I, unesterified GalUA). No significant differences between the T-DNA mutants and WT were observed (our unpublished results).
Monosaccharide composition of alcohol insoluble residues, from cell walls of the T-DNA insertional mutants rra-1 and -2
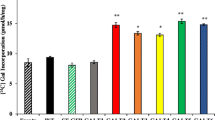
Alcohol insoluble residues (AIR) were prepared from CW material isolated from the following tissues: roots and leaves isolated from seedlings, root, leaves, stems and flowers of 5-week old bolting plants, and the apical meristem of 3-week old plants. AIR of all the tissues were sequentially extracted to yield RG I, RG II, HG and xyloglucan. In addition, water-soluble extracts and AGPs were isolated from rosette leaves and meristematic tissues of 3-week old plants. The fractions were then analyzed by either oligosaccharide mass profiling (xyloglucan) or by monosaccharide composition analysis. No significant differences were observed in these analyses (data not shown). However, a significant reduction of Ara was observed in the CW pellet after enzymatic removal of the pectic polysaccharides and xyloglucan and this reduction was only observed in CWs isolated from meristematic tissue (Table 2). The reduction was observed in both rra1 mutants (Garlic_76_G04: p ≤ 0.00002 & SAIL_590_G09: p ≤ 0.0002), and in both rra2 mutants (Garlic_244_A03: p ≤ 0.0010 & SAIL_70_D08: p ≤ 0.010).
The Garlic_244_A03 line (rra2 mutant) contains elevated levels of Glc (Table 2). No clear trend was seen in the relative levels of other monosaccharides compensating for the Ara reduction. The two alleles appear not to respond similarly, but it is important to appreciate that these differences are not statistically significant. However, the reduction of Ara was seen in both alleles of RRA2.
The method used for determining the monosaccharide compositions of the neutral sugars, does not allow for the simultaneous quantification of acidic sugars such as e.g., GalA, and these were therefore analyzed separately (no differences were observed, data not shown).
The monosaccharide profile of the CW remaining pellet suggests the presence of tightly bound non-cellulosic polysaccharides, notably xyloglucans and (arabino)xylans. While xyloglucan in solanaceaous plants contains Ara, xyloglucan in Arabidopsis thaliana does not (Zablackis et al. 1995), so changes in xyloglucan composition cannot account for the reduced Ara content of the CW-pellet. Glucuronoarabinoxylans do contain Ara (Zablackis et al. 1995; Bacic et al. 1988), and the xylans were therefore digested with a xylanase and the solubilized degradation products analyzed for arabinose content. No differences between WT and mutants were observed (data not shown) suggesting that the Ara is not derived from heteroxylans.
Discussion
CAZy GT-family-77: retaining glycosyltransferases that form (1,3)-linkages
A phylogenetic analysis of CAZy GT-family-77 lead us to propose a sub-division of some members into four clades labeled A to D, and a large remainder of accessions that could not be subdivided. If clades A and B reflect not only evolutionary relatedness, but also correlate with catalytic function, it would appear that Arabidopsis thaliana devotes several GT-genes where Oryza sativa employs one. It is, however, premature to conclude that the Arabidopsis thaliana genes of clade A and B, respectively, are redundant; currently only RGXT1 and RGXT2 of clade B have been functionally characterized, the annotation as a GalT of the Linum sequence, which is also assigned to clade B, is premature (C. Morvan personal communication). The accessions not yet assigned to clades seem to split according to species rather than being organized as sets of orthologous Oryza sativa and Arabidopsis thaliana genes (see bottom of the phylogenetic tree in Fig. 1A). This may hint at functions in the synthesis of carbohydrates that are typical of Graminaceous monocots and other angiosperms, there are several polysaccharides and proteoglycans that differ between Type-I and Type-II CWs, sensu Carpita and Gibeaut (1993). We recommend that definition of a clade of e.g., Poaceae-specific genes should await the functional characterization of some of the Oryza sativa accessions.
The (1,3)-α-d-GalT (AX3) from the slime mold Dictyostelium discoideum has also been classified to CAZy GT-family-77. It is fairly distantly related to the plant sequences (below 10%) and is not included in any of the plant clades. The Dictyostelium discoideum (1,3)-α-d-GalT is a soluble protein, which takes part in the glycosylation of Skp1, a cytoplasmic and nuclear protein involved in cell cycle regulation (Ketcham et al. 2004). The Skp1 pentasaccharide (to which the (1,3)-α-d-GalT transfers the penultimate galactosyl residue) is attached to a hydroxyproline residue and is assembled in the cytoplasm. This cytoplasmic O-glycosylation pathway is known from lower eukaryotes, possibly including Chlamydomonas reinhardtii (West et al. 2004), but has yet to be demonstrated in vascular plants, like Arabidopsis thaliana. The Chlamydomonas reinhardtii accessions, which clearly belong in clade A, might suggest the existence of a similar pathway in higher plants (the CAZy database does not list sequences from genome projects. Bernard Henrissat personal communication). Another possibility is that the Chlamydomonas reinhardtii genes encode GTs involved in CW biosynthesis. Interestingly RG II is not present in the CWs of algae (O’Neill et al. 2004) and the CW of Chlamydomonas reinhardtii is unusual in that it is almost entirely composed of HRGPs, which are related to the extensins of higher plants (Waffenschmidt et al. 1993). It may thus be speculated that the Chlamadymonas reinhardtii genes could encode either GalTs or AraTs involved in glycosylation of extensin-like cell wall proteins.
Organ specific and developmentally determined expression of RRA-1 and -2
Both RRA-1 and -2 were shown by promoter::gusA studies to be active in the meristematic region. Pectic arabinan, but not galactan, has been shown to be enriched in apical meristems of potato stolons (Bush et al. 2001) and has also been shown to be essential to meristematic function in potato (Borkhardt et al. 2005). The observation that arabinans are associated with meristematic regions in carrot (Willats et al. 1999) may be relevant to the observed GUS expression in root tips of the RRA2 promoter::gusA transformants.
The expression pattern in mature plants in specialised tissues and cell types: hydathodes, trichome support cells in the adaxial epidermis, cauline leaves, petals, mature roots and in both the proximal and distal ends of siliques, does not allow us to make inferences with regard to the function(s) of RRA-1 and -2.
Characterization of the rra1 and rra2 mutants
T-DNA insertional lines, rra-1 and -2, for At1g75120 and At1g75110, respectively, are similar to WT-plants with regard to morphology and growth rate. A significantly altered CW monosaccharide profile was only observed in the residual CW pellet after enzymatic removal of the pectic and xyloglucan polysaccharides, and only observed in tissues isolated from meristematic tissue. A significant reduction in Ara was observed in both mutants for RRA-1 and -2. These results suggest that both genes may encode arabinosyltransferases (AraTs), although other more complex explanations such as indirect effects on arabinosylation cannot be ruled out.
Inferring the catalytic function of RRA-1 and -2
A mutant phenotype reduced in Ara in a tightly bound wall fraction would be most easily explained if RRA-1 and -2 encode AraTs. A reduction in Ara can, however, also be the consequence of a knocked out GalT. Extensins contain α-linked galactosyl residues as single substituents directly on serine residues of the protein backbone, and the arabinosylation af adjacent Hyp residues could, for example, depend on correct galactosylation of the Ser-Hyp4 repeat.
Ara is a major component in various plant polysaccharides e.g., the pectic polysaccharides RG I and RG II, in the non-cellulosic arabinoxylans, in extensins as well as in type II AGs/AGPs (reviewed by Ishii et al. 2005a). Ara is predominately found in the arabinofuranose (Araf) form in plants (Carpita and Gibeaut 1993). Arabinopyranose (Arap) has been found as a terminal sugar in galactans from soybean (Huisman et al. 2001), in AGPs from maize (Bacic et al. 1986), in the pectic polysaccharide RG II (O’Niell et al. 2004), and in arabinans isolated from various plant species (reviewed by Ishii et al. 2005a). The biochemistry of Araf incorporation into pectic polysaccharides from UDP-Arap has received much attention recently (Numan and Scheller 2003; Ishii et al. 2005b; Konishi et al. 2006; Harholt et al. 2006), but is unresolved at present. As long as this is the case, it cannot be predicted with certainty whether a GT-family-77 AraT would form α- or β-glycosidic linkages, although we consider the latter more likely, however, α-linked Araf is much more prevalent in the CW than β-linked Ara. The terminal Arap of soybean RG I has been suggested to be β-(1,4)-linked (Huisman et al. 2001). β-(1,3)-Linked Arap has so far only been found in the gymnosperm Larix dahurica AG II (Odonmaig et al. 1994). The internal Ara residues of the extensin oligo-arabinoside side chains are Araf-β-(1,2)-linkages (Akiyama and Kato 1977). Extensins become tightly bound to the CW soon after their deposition, so a change in CW composition of the mutants should be observed in the pellet rather than in any of the CW extracts. It should be noted, however, that Zykwinska et al. (2005) recently showed that pectic arabinan and galactan side chains are able to bind tightly to specific sites on cellulose microfibrils in vitro.
The most persuasive approach to resolving whether the RRA-1 and -2 play roles (and not necessarily the same role) in either arabinosylation of xylan, RG I side chain or extensin, would be to heterologously express the RRA-1 and -2 gene products and demonstrate transfer of Ara to acceptors originating from the three candidate biopolymers.
Abbreviations
- a.a.:
-
Amino acid
- AG:
-
Arabinogalactan
- AG II:
-
Arabinogalactan type II
- AGP:
-
Arabinogalactan-proteins
- CAZy:
-
Carbohydrate-Active enZYmes
- CW:
-
Cell wall
- EST:
-
Expressed sequence tag
- GT:
-
Glycosyltransferase
- HG:
-
Homogalacturonan
- RG I:
-
Rhamnogalacturonan I
- RG II:
-
Rhamnogalacturonan II
- XG:
-
Xylogalacturonan
- XEG:
-
Xyloglucanase
- GUS:
-
β-glucronidase
References
Abdulrazzak N, Pollet B, Ehlting J, Larsen K, Asnaghi C, Ronseau S, Proux C, Erhardt M, Seltzer V, Renou JP, Ullmann P, Pauly M, Lapierre C, Werck-Reichhart D (2005) A coumaroyl-ester-3-hydroxylase insertion mutant reveals the existence of nonredundant meta-hydroxylation pathways and essential roles for phenolic precursors in cell expansion and plant growth. Plant Physiol 140:30–48
Akiyama Y, Kato K (1977) Structure of hydroxyproline-arabinoside from tobacco cells. Agricultural and Biological Chemistry 41(1):79–81
Albersheim P, Nevins DJ, English PD, Karr A (1967) A method for the analysis of sugars in plant cell wall polysaccharides by gas–liquid chromatography. Carbohydr Res 5:340–345
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Bacic A, Harris PJ, Stone BA (1988) Structure and function of plant cell walls. In: Preiss J (ed) The biochemistry of plants: a comprehensive treatise, vol. 14, Carbohydrates, New York, Academic Press, pp 297–371
Bacic A, Moody SF, Clarke AE (1986) Structural analysis of secreted root slime from maize (Zea mays L). Plant Physiol 80:771–777
Blumenkrantz N, Asboe-Hansen G (1973) New method for quantitative-determination of uronic acid. Anal Biochem 54:484–489
Borkhardt B, Skjøt M, Mikkelsen R, Jørgensen B, Ulvskov P (2005) Expression of a fungal endo-α-1,5-l-arabinanase during stolon differentiation in potato inhibits tuber formation and results in accumulation of starch and tuber-specific transcripts in the stem. Plant Sci 169:872–881
Bush MS, Marry M, Huxam IM, Jarvis MC, McCann MC (2001) Developmental regulation of pectic epitopes during potato tuberization. Planta 213:869–880
Carpita NC, Gibeaut DM (1993) Structural models of primary cell walls in flowering plants: consistency of molecular structure with the physical properties of the walls during growth. Plant J 3:1–30
Cosgrove DJ (2001) Wall structure and wall loosening. A look backwards and forwards. Plant Physiol 125:131–134
Coutinho PM, Deleury E, Davies GJ, Henrissat H (2003) An evolving hierarchical family classification for glycosyltransferases. J Mol Biol 328:307–317
Coutinho PM, Henrissat B (1999) Carbohydrate-active enzymes server at url: http://afmb.cnrs-mrs.fr/∼cazy/CAZY/index.html
Edgar RC (2004), MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
Egelund J, Petersen BL, Motawia S, Damager I, Faik A, Clausen H, Olsen CE, Ishii T, Ulvskov P, Geshi N (2006) Biosynthesis of pectic rhamnogalacturonan II: Molecular cloning and characterization of Golgi-localized α-xylosyltransferases encoded by the RGXT1 and RGXT2 genes of Arabidopsis thaliana. Plant Cell (in press)
Egelund J, Skjodt M, Geshi N, Ulvskov P, Petersen BL (2004) A complementary bioinformatic approach to identify potential plant cell wall glycosyltransferase encoding genes. Plant Physiol 136:2609–2620
Gane AMJ, Craik D, Munro SLA, Howlett GJ, Clarke AE, Bacic A (1995) Structural analysis of the carbohydrate moiety of arabinogalactan-proteins fromstigmas and styles of Nicotiana alata. Carbohydr Res 277:67–85
Harholt J, Jensen JK, Sorensen SO, Orfila C, Pauly M, Scheller HV (2006) Arabinan deficient 1 is a putative arabinosyltransferase involved in biosynthesis of pectic arabinan in Arabidopsis. Plant Physiol 140:49–58
Huisman MMH, Brull LP, Thomas-Oates JE, Haverkamp J, Schols HA, Voragen AGJ (2001) The occurrence of internal (1,5) α-linked arabinofuranose and arabinopyranose residues in arabinogalactan side chains from soybean pectic substances. Carbohydr Res 330:103–114
Ishii T, Ono H, Ohnishi-Kameyama M, Maeda I (2005a) Enzymic transfer of α-l-arabinopyranosyl residues to exogenous 1,4-linked β-d-galacto-oligosaccharides using solubilized mung bean (Vigna radiate) hypocotyls microsomes and UDP-β-l-arabinopyranose. Planta 221:953–963
Ishii T, Konishi T, Ito Y, Ono H, Ohnishi-Kameyama M, Maeda I (2005b) A β(1–3)-arabinopyranosyltransferase that transfers a single arabinopoyranose onto arabino-oligosaccherides in mung bean (Vigna radiate) hypocotyls. Phytochem 66:2418–2425
Jefferson RA, Kavanagh TA, Bevan MV (1987) GUS fusions: β-glucuronisidase as a sensitive and versatile gene fusion maker in higher plants. EMBO J 6:3901–3907
Johnson KL, Jones BJ, Bacic A, Schultz CJ (2003) The fasciclin-like arabnogalactan proteins of Arabidopsis. A multigen family of putative cell adhesion molecules. Plant Physiol 133:1911–1925
Keegstra K, Talmadge KW, Bauer WD, Albersheim P (1973) The structure of plant cell walls. III. A model of the walls of suspension-cultured sycamore cells based on the interconnections of the macromolecular components. Plant Physiol 51:188–196
Ketcham C, Wang F, Fisher SZ, Ercan A, van der Wel H, Locke RD, Sirajud-Doulah K, Matta KL, West CM (2004) Specificity of a soluble UDP-galactose:fucoside alpha 1,3-galactosyltransferase that modifies the cytoplasmic glycoprotein Skp1 in Dictyostelium. J Bio Chem 279(28):29050–29059
Konishi T, Ono H, Ohnishi-Kameyama M, Kaneko S, Ishii T (2006) Identification of a mung bean arabinosyltransferase that transfers arabinofuranosyl residues onto (1,5)-linked α-l-arabino-oligosaccharides. Plant Physiol 141(3):1098–1105
Lamport DTA (1963) Ozygen fixation into hydroxyproline of plant cell wall proteins. J Biochem Chem 238:1438–1440
Lerouxel O, Choo TS, Seveno M, Usadel B, Faye L, Lerouge P, Pauly M (2002) Rapid structural phenotyping of plant cell wall mutants by enzymatic oligosaccharide fingerprinting. Plant Physiol 130:1754–1763
Mikkelsen MD, Petersen BL, Glawishnig E, Jensen AB, Andreasson E, Halkier BA (2003) Modulation of CYP79 genes and glucosinolate profiles in Arabidopsis by defense signaling pathways. Plant Physiol 131:298–308
Numan KJ, Scheller HV (2003) Solubilization of an arabinan arabinosyltransferase activity from mung bean hypocotyls. Plant Physiol 132:331–342
O’Neill MA, Ishii T, Albersheim P, Darvil AG (2004) Rhamnogalacturonan II: Structure and function of a borate cross-linked cell wall pectic polysaccharide. Annu Rev Plant Biol 55:109–139
Odonmaig P, Ebringerova A, Machová E, Alföldi J (1994) Structural and molecular properties of arabinogalactan isolated from Mongolian larchwood (Larix dahurica L.). Carbohydr Res 252:317–324
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Computer Applications in the Biosciences 12:357–358
Pauly M, Anderson LN, Kauppinen S, Kofod LV, York WS, Albersheim P, Darvill AG, (1999) A xyloglucan specific endo-beta-1,4-glucanase from Aspergillus aculeatus: expression cloning in yeast, purification and characterization of the recombinant enzyme. Glycobiology 9:93–100
Popper ZA, Fry SC (2005) Widespread occurrence of a covalent linkage between xyloglucan and acidic polysaccharides in suspension-cultured Angiosperm cells. Ann Bot 96:91–99
Schultz CJ, Johanson KL, Currie G, Bacic A (2000) The classical arabinogalactan protein gene family of Arabidopsis. Plant Cell 12:1751–1767
Showalter AM (1993) Structure and function of plant cell wall proteins. Plant Cell 5:9–23
Sims IM, Bacic A (1995) Extracellular polysaccharides from suspension cultures of Nicotiana plumbaginifolia. Phytochemistry 38:1397–1405
Somerville C, Bauer S, Brinistool G, Facette M, Hamann T, Milne J, Osborne E, Paredez A, Persson S, Raab T, Vorwerk S, Youngs H (2004) Toward a systems approach to understanding plant cell walls. Science 306:2206–2211
Sørensen SO, Pauly M, Bush M, Skjøt M, McCann MC, Borkhardt B, Ulvskov P (2000) Pectin engineering: modification of potato pectin by in vivo expression of an endo-1,4-beta-d-galactanase. Proc Natl Acad Sci USA 97:7639–7644
Talbott LD, Ray PM (1992) Molecular size and separability features of Pea cell wall polysaccharides: Implications for models of primary wall structure. Plant Physiol 98:357–368
Usadel B, Schluter U, Mølhøj M, Gipmans M, Verma R, Kossmann J, Reiter WD, Pauly M (2004) Identification and characterization of a UDP-d-glucuronate-4-epimerase in Arabidopsis. FEBS Lett 596:327–331
Waffenschmidt S, Woessner JP, Beer K, Goodenough UW (1993) Isodityrosine cross-linking mediates insolubilization of cell walls in chlamydomonas. Plant Cell 5:809–820
West CM, van der Wel H, Sassi S, Gaucher EA (2004) Cytoplasmic glycosylation of protein-hydroxyproline and its relationship to other glycosylation pathways. Biochim Biophys Acta Gen Sub 1673(1–2):29–44
Willats WGT, McCartney L, Knox JP (2001) In-situ analysis of pectic polysaccharides in seed mucilage and at the root surface of Arabidopsis thaliana. Planta 213(1):37–44
Willats WGT, Steele-King CG, Marcus SE, Knox JP (1999) Side chains of pectic polysaccharides are regulated in relation to cell proliferation and cell differentiation. Plant J 20:619–628
Zablackis E, Huang J, Muller B, Darvill AG, Albersheim P (1995) Structure of plant-cell walls: characterization of the cell-wall polysaccharides of Arabidopsis-thaliana leaves. Plant Physiol 107:1129–1138
Zykwinska AW, Ralet MCJ, Garnier CD, Thihault JFJ (2005) Evidence for in vitro binding of pectin side chains to cellulose. Plant Physiol 139:397–407
Acknowledgements
We would like to thank Dr. Kirk Schnorr, Novozymes, for the gift of XEG. Ms. Vibeke Strange Petersen and Dorthe Christiansen are thanked for their skilful technical assistance. Dr. Marcia Kieliszewski is acknowledged for fruitful discussions while this manuscript was prepared. This work was supported by the Danish National Research Foundation (B.L.P., P.U., N.G.), by an EU-FP6 grant in the training network WallNet (B.L.P. and P.U.) and by The Carlsberg Foundation (J.E.).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Egelund, J., Obel, N., Ulvskov, P. et al. Molecular characterization of two Arabidopsis thaliana glycosyltransferase mutants, rra1 and rra2, which have a reduced residual arabinose content in a polymer tightly associated with the cellulosic wall residue. Plant Mol Biol 64, 439–451 (2007). https://doi.org/10.1007/s11103-007-9162-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-007-9162-y