Abstract
We have previously reported the graft transmission of target specificity for RNA silencing using transgenic Nicotiana benthamiana plants expressing the coat protein gene (CP, including the 3′ non-translated region) of Sweet potato feathery mottle virus. Transgenic plants carrying the 5′ 200 and 400 bp regions of CP were newly produced. From these plants, two silenced and two non-silenced lines were selected to investigate the manifestation of transitive RNA silencing by graft experiments. Non-silenced scions carrying the entire transgene were grafted onto either 5′ or 3′ silencing inducer rootstocks. When non-silenced scions were grafted onto 5′ silencing inducer rootstocks, RNA silencing was induced in the non-silenced scions and spread toward the 3′ region of the transgene mRNA. Similarly, when non-silenced scions were grafted onto 3′ silencing inducer rootstocks, RNA silencing was induced in the non-silenced scions, but was restricted to the 3′ region of the transgene and did not spread to the 5′ region. In addition, results from crossing experiments, involving non-silenced and 3′ silencing inducer plants, confirmed the above finding. This indicates that RNA silencing spreads in the 5′–3′ direction, not in the 3′–5′ direction, along the transgene mRNA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gene silencing, a sequence specific RNA degradation process in eukaryotic cells, is believed to play roles in adaptive protection against viruses (Covey et al. 1997; Al-Kaff et al. 1998) and mobile DNA elements (Ketting et al. 1999), as well as in developmental regulation of gene expression (Grishok et al. 2001). Transgene induced post-transcriptional gene silencing (PTGS) has been reported in plants (co-suppression) (Napoli et al. 1990; van der Krol et al. 1990; Smith et al. 1990) and fungi (quelling) (Romano and Macino 1992). In animals, PTGS can be induced by double-stranded RNA (dsRNA) in a process known as RNA interference (RNAi) (Fire et al. 1998). In plants, PTGS can also be induced by viruses expressing host genes in a process termed virus-induced gene silencing (VIGS) (Baulcombe 1996). Since PTGS, quelling and RNAi all function at the RNA level, these processes are collectively referred to as RNA silencing (Baulcombe 1999).
Double-stranded RNA is a key trigger for RNA silencing. dsRNAs can be generated in different ways: as a replicating intermediate of viruses, by which they can induce RNA silencing and become targets for RNA degradation (Lindbo et al. 1993), transcription of inverted repeats (IR) (Chuang and Meyerowitz 2000; Smith et al. 2000) or RNA-dependent RNA polymerase (RDR also known as RdRP)-mediated synthesis that uses aberrant RNA (abRNA) as a template (Baulcombe 2004). Dicer, a ribonuclease (RNase) III family enzyme, initiates silencing by cleaving the dsRNA to 21–25 nucleotides (nt) duplexes containing a 3′ two nucleotide overhang and a 5′ phosphate group, called short interfering RNAs (siRNAs) (Bernstein et al. 2001; Elbashir et al. 2001). The next step of RNA silencing process is the incorporation of siRNAs into Argonaute (Ago) containing multicomponent RNA-induced silencing complex (RISC). After an ATP-dependent unwinding of RISC-bound siRNA, the Ago acts as a sequence specific ribonuclease (also called Slicer) that cleaves single stranded RNAs at discrete positions (Elbashir et al. 2001; Martinez et al. 2002; Baumberger and Baulcombe 2005).
The gene families encoding common components of RNA silencing and their effector complex machineries are conserved across kingdoms. However, the inside mechanisms have become diversified and specialized between and within kingdoms. These diversifications and specializations are most evident in plants, such as Arabidopsis thaliana, in which the genes corresponding to RNA silencing machineries are extensively studied. Arabidopsis contains at least three active RdRP genes (Baulcombe 2004), termed RDR1, RDR2 and RDR6 (also known as SDE1/SGS2) (Dalmay et al. 2000; Mourrain et al. 2000). Mutation analyses have revealed that different RDRs may involve dsRNA generation from different templates (Xie et al. 2001, 2004; Yu et al. 2003; Herr 2004). Four Dicer-like genes (DCL1 to DCL4) have also been reported in Arabidopsis with distinct or overlapping activities necessary to process these different dsRNAs to two distinct classes of siRNAs (21–22 nt short siRNA and 24–26 long siRNA) (Hamilton et al. 2002) and micro RNAs (miRNAs). DCL1 is a nuclear protein involved in miRNA processing (Finnegan et al. 2003; Papp et al. 2003; Xie et al. 2004; Mlotshwa et al. 2005). DCL2 is likely to be associated with viral dsRNA processing, producing siRNAs of short class, while DCL3 generates endogenous siRNAs that belong primarily to a longer size class and are required for chromatin modification (Xie et al. 2004). Recently, DCL4 is found to be associated with developmental regulation through biogenesis of trans-acting siRNAS (ta-siRNAs) (Xie et al. 2005; Dunoyer et al. 2005).
Although RNA silencing is a sequence specific process, spreading of silencing has been reported in several studies. Cellular RdRPs can synthesize complementary RNA (cRNA) using the abRNAs as templates (unprimed synthesis) or siRNAs can be used as primers to synthesize cRNA using ssRNA as template (Sijen et al. 2001; Tang et al. 2003; Baulcombe 2004). The subsequent processing of the resultant dsRNAs by DCL enzymes can lead to suppression of adjacent sequences along the template, a phenomenon referred to as transitive RNA silencing (Sijen et al. 2001).
Transmission of RNA silencing from silenced rootstocks to non-silenced scions is a well known phenomenon (Palauqui et al. 1997; Voinnet and Baulcombe 1997; Sonoda and Nishiguchi 2000; Crete et al. 2001; Garcia-Perez et al. 2004). The signaling molecules underlying RNA silencing can move to neighboring cells through plasmodesmata or systemically to the entire plant through the vascular system (Fagard and Vaucheret 2000; Mlotshwa et al. 2002; Himber et al 2003). Molecular analysis revealed that short distance movement of RNA silencing is mediated by the 21–22 nt siRNAs and long distance or systemic silencing involves an amplification of the same silencing signal utilizing cellular RdRP (Himber et al. 2003; Schwach et al. 2005).
We have previously reported the graft transmission of target specificity for RNA silencing in transgenic N. benthamiana expressing the coat protein gene (CP) of Sweet potato feathery mottle virus (SPFMV) (Sonoda and Nishiguchi 2000). We found that when non-silenced scions were grafted onto silenced plants, in which the silencing was targeted solely at the 3′ region of the transgene (i.e. 3′ silencing inducer plant), RNA silencing was induced, but was restricted to the 3′ one-third region of the transgene. These results led us to conduct additional experiments. What would happen if non-silenced scions were grafted onto silenced rootstocks with target specificity for the 5′ region of the transgene? To address this question, we produced transgenic silenced plants carrying the 5′ region of the transgene (i.e. 5′ silencing inducer plants) and analyzed the induction of transitive RNA silencing in non-silenced plants by graft experiments. Here we present results showing that RNA silencing was induced in the non-silenced scions when grafted onto the 5′ silencing inducer rootstocks, and the induced RNA silencing spread to the 3′ region along the transgene transcript. RNA silencing was induced but did not spread to the 5′ region along the transgene mRNA when grafted onto the 3′ silencing inducer rootstocks. Thus, in this system, spreading of RNA silencing occurred in the 5′–3′ direction, not the 3′–5′ direction.
Materials and methods
Plant materials
Transgenic Nicotiana benthamiana, lines 4.07 and 4.09, carrying CP of SPFMV were previously described (Sonoda et al. 1999). Line 4.07 is a non-silenced over-expressor and 4.09 is a silenced line, although silencing was targeted only to the 3′ region of the transgene.
Plasmid construction
DNA corresponding to the 5′ 200 or 400 bp regions of CP of SPFMV were generated by polymerase chain reaction (PCR) from a cDNA clone, pVC1 (Mori et al. 1995), using the specific forward primer, SPFMV.CP-F, 5′-GGCCGGATCCAACAACAATGtctagtgaacgtactgaattcaaagatgcggga-3′ and the reverse primers, SPFMV.CP-R200, 5′-GCAGATCTGAGCTCgattccttctccttgctagtgtc-3′ and SPFMV.CP-R400, 5′-GCAGATCTGAGCTCgagtcgaccgggtgtttgcaacc-3′. Letters in lower case represent CP sequence of SPFMV (accession number D 86371; 9652–9684 nt, 9828–9851 nt, 10028–10051 nt for SPFMVCP-F, SPFMV.CP-R200 and SPFMV.CP-R400, respectively). They contained the recognition sites (bold letters) for the restriction enzymes BamHI for SPFMV.CP-F and SacI for SPFMV.CP-R200 and SPFMV.CP-R400. PCR was performed using Pfu polymerase (Stratagene) under conditions of an initial denaturation step for 3 min at 94°C and 25–35 cycles of 30 s at 94°C denaturation step, 45 s at 51°C annealing step and 1 min at 75°C extension step. The CP fragments were then digested with BamHI and SacI and ligated to the GUS deleted vector pBI121 treated with the same restriction enzymes. The resultant plasmids, pSPFMV-CP 5′ 400 and pSPFMV-CP 5′ 200 (Fig. 1A, B), were used for transformation of N. benthamiana.
Southern blot analysis of transgenic plants. (A, B) Schematic representation of DNA constructs used for plant transformation. 5′ 200 and 400 bp of CP were inserted into GUS deleted vector pBI121. RB, right border; pNOS, nopaline synthase promoter; NPTII (kan+), kanamycin resistance gene; NOS, nopaline synthase terminator; p35S, CaMV35S promoter; LB, left border. (C) Genomic DNA (20 μg) was digested with BamHI, size fractionated on 1.0% agarose gel, transferred to a nylon membrane and hybridized with a 32P labeled 5′ 400 bp of CP probe. Plants transformed with GUS deleted vector, pBI121, were used as a control (ΔGUS)
Plant transformation
Nicotiana benthamiana was transformed by the leaf disc method using Agrobacterium tumefaciens LBA4404, containing pSPFMV-CP 5′ 400 or pSPFMV-CP 5′ 200 as previously described (Horsch et al. 1985). The shoots were rooted in the presence of kanamycin (100 μg/ml) in a growth chamber at 25°C under a 16 h-light and 8 h-dark regimen. Rooted plants were transferred to soil and were grown in the greenhouse. From these plants, two silenced (lines 200.4 and 400.15) and two non-silenced lines (lines 200.1 and 400.89) were selected for this study (see Results for detail).
PVX constructs and virus infection
Polymerase chain reaction amplified 5′ 200 or 400 bp of CP fragments were inserted into the EcoRV site of the PVX vector pP2C2S, a derivative of pGC3 (Chapman et al. 1992) which can be used to express foreign genes in plants (kindly provided by D. C. Baulcombe, The Sainsbury Laboratory, Norwich, UK). Resulting PVX constructs, PVX-5′CP 200 (harboring 5′ 200 bp of CP) and PVX-5′CP 400 (harboring 5′ 400 bp of CP), were linearized with SpeI and used for in vitro transcription reactions to produce infectious PVX, as described in Sonoda et al. (1999). Virus-infected leaf homogenates, diluted (1:10) in 10 mM phosphate buffer (pH 7.0), were used for virus inoculation. Inoculated plants were monitored daily for the appearance of symptoms, and the leaves were collected 7–8 days after inoculation for RNA extraction to check virus accumulation.
Grafting procedure
Approximately 8 week old plants were used for the cleft grafting, as described by Palaqui et al. (1997). Rootstocks were prepared by removing the shoots above at least two basal leaves and then creating a vertical cut of 1–2 cm at the center of the stem. Scions (3–5 cm long) were prepared by removing leaves and trimming the base of the scion to a wedge. The scion/rootstock junction was wrapped with Parafilm and a clip. Plants were covered with plastic bags to avoid dehydration for 1 week or until the graft had taken.
Generation of hybrid plants
Crosses were made between lines 4.09 and 4.07 as female and male parents, respectively, and vice versa, using standard procedure; immature flowers of the female parents were emasculated and the stigmas were brushed with the pollen taken from the male parent for 3–4 consecutive days. Pollinated flowers were covered with paper bags until harvesting. The progeny seeds were grown on MS media in presence of kanamycin (100 μg/ml) and, finally, grown in a greenhouse, as previously described.
Southern blot analysis
Genomic DNA was extracted from leaf tissues, as described by Sonoda et al. (1999). Total DNA (20 μg) was digested with BamHI and size fractionated in a 1% agarose gel, transferred to a nylon membrane (Hybond N+, Amersham Biosciences) and hybridized with a randomly primed 32P labeled CP specific probe.
RNA gel blot analysis
Total RNA was isolated from the leaf tissues with Tri reagent, according to the manufacturer’s instructions (Molecular Research Center), followed by two purification steps using equal volumes of chloroform/isoamylalcohol. Total RNA (10 μg) was size-fractionated on a 0.8% agarose gel containing 0.66 M formaldehyde and was transferred to a nylon membrane (Hybond N+) through capillary transfer. Blots were prehybridized with 6 × SSC, 5 × Denhardt’s solution and 0.1% SDS at 65°C for 2–3 h and hybridized at 65°C for about 16 h with [α-32P]dCTP-labeled DNA probes. The membranes were washed once with 5 × SSC at room temperature for 5 min and twice with 2 × SSC and 0.5% SDS at 65°C for 30 min. The membranes were exposed to Imaging Plate for 12–16 h (BAS III, Fuji Photofilm) and were analyzed with the Bio Image Analyzer (BAS-2000, Fuji Photofilm).
siRNA detection
Total RNA was resuspended in water and larger RNAs were removed by selective precipitation with 5% polyethylene glycol (PEG, mol. wt. 8000) in 0.5 M NaCl. Small RNAs were precipitated from the supernatant by adding 3 volumes of absolute ethanol and 0.1 volume of sodium acetate pH 5, incubated overnight at −20°C and, finally, dissolved in formamide.
The enriched small RNAs (50 μg) were separated on a 15% (v/w) polyacrylamide gel (19:1 acrylamide:bisacrylamide) containing 7 M urea in 0.5 × TBE buffer, electrotransferred to a nylon membrane (Hybond N+ membrane) at 160 mA for 1 h in 0.5 × TBE by a semidry electroblotter (Nihon Eido), and cross-linked by ultraviolet irradiation (1.2 × 105 μJ/cm2). Blots were prehybridized with 5 × SSC, 5 × Denhardt’s solution and 0.5% SDS at 50°C for 2–3 h and hybridized with 32P labeled probe DNA at 50°C for about 16 h. The membranes were washed once with 2 × SSC at 50°C for 5 min and three times with 2 × SSC and 1% SDS at 50°C for 30 min. They were then analyzed as mentioned above.
Probe DNA preparation
CP, including the 3′ non-translated region of SPFMV, was divided into three regions which were used as probes for this study: 5′ region (5′CP), middle region (MCP) and 3′ region (3′CP) (Fig. 5A). PCR amplified 5′ 400 bp of CP (9652–10051 nt of SPFMV) was used as the 5′CP probe. The cDNA clone, pVC1, was digested with either the combination of SalI and EcoT221 or EcoT221 and EcoRI. The gel purified nearly 350 bp (10045–10389 nt of SPFMV) and 430 bp DNA fragments (10390–10820 nt of SPFMV) were used as MCP and 3′CP probes, respectively. PVX accumulation was checked using CP of PVX as probe, which was previously amplified from plasmid pP2C2S using the PCR primers: 5′-ATGTCAGCACCAGCTAGC-3′ and 5′-GCGTCGGTTATGTAGACG-3′. [α-32P]dCTP-labeled probe DNAs were prepared using the Megaprime DNA Labeling System (Amersham Biosciences), according to the manufacturer’s instructions.
Results
Generation and characterization of transgenic plants carrying 5′ 200 or 400 bp of CP
Nicotiana benthamiana was transformed with the plasmids carrying either 5′ 200 or 5′ 400 bp of CP (see Materials and Methods for detail). Ten transgenic lines with pSPFMV-CP 5′ 200 (Fig. 1A) and sixteen lines with pSPFMV-CP 5′ 400 (Fig. 1B) were regenerated in the presence of kanamycin. All transgenic lines appeared normal in morphology and development. Each line was selfed, and, from the resulting collection of T1 progeny, two lines each from the ten and sixteen lines were selected for further analyses. Lines 200.1 and 200.4 carried the 5′ 200 bp, and lines 400.15 and 400.89 carried the 5′ 400 bp of CP.
Total RNA from these four transgenic plant lines was used for Northern blot analyses. CP mRNA was almost undetectable in lines 200.4 and 400.15, in contrast to high levels of mRNA accumulation in lines 200.1 and 400.89 (Fig. 2A). Southern blot analysis demonstrated that lines 200.4 and 400.15 carried two copies of the transgene, and lines 200.1 and 400.89 carried one copy of the transgene (Fig. 1C). Transgene-related siRNAs, the hallmark of RNA silencing (Hamilton and Baulcombe 1999), were detected in lines 200.4 and 400.15 with a radiolabeled DNA probe corresponding to the 5′ 400 bp of CP (5′CP), whereas no such siRNAs could be detected in lines 200.1 and 400.89 (Fig. 2B). From the total RNA and siRNA analyses, lines 200.4 and 400.15 were selected as silenced, and lines 200.1 and 400.89 as non-silenced lines.
Northern blot analysis of transgenic plants. (A) Total RNA (10 μg) was extracted from leaves of approximately 4 weeks old plants, subjected to denaturing gel electrophoresis, transferred to a nylon membrane and hybridized with 32P labeled 5′ 400 bp of CP specific probe. An ethidium bromide stained RNA gel photographed before transfer is also shown as control for equal loading. (B) Small RNA fractions (50 μg) were isolated from leaf tissue, separated on 15% polyacrylamide gels, blotted onto a nylon membrane and hybridized with 32P labeled 5′ 400 bp of CP specific DNA probe. DNA oligomers (25 bp) were used for size control (indicated at the left side of the figure). ΔGUS Plants were used as controls. As a control for equal loading, ethidium bromide stained tRNA fractions are shown below
Virus resistance of silenced and non-silenced lines to recombinant PVX, engineered to contain the transgene sequence
It has been reported that silenced lines of transgenic plants show resistance to the virus carrying the transgene sequence (English et al. 1996). For further clarification of the silencing state in these four lines, they were inoculated with the recombinant PVX, PVX-5′CP 200 and PVX-5′CP 400, which have been engineered to contain the 5′ 200 bp and 5′ 400 bp of CP, respectively. Lines 200.4 and 400.15 were resistant against both the recombinant viruses as no symptoms were found at 7–8 days after virus inoculation (dpi). On the other hand, lines 200.1 and 400.89 were susceptible to recombinant PVX. Viral RNA accumulated at low levels in 200.4 and 400.15, but at high levels in 200.1 and 400.89 lines at 7–8 dpi (Fig. 3). This observation confirmed that lines 200.4 and 400.15 were post-transcriptionally silenced.
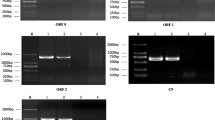
Analysis of virus resistance of transgenic plants against recombinant PVX. Transgenic N. benthamiana plants were inoculated with recombinant PVX carrying either 5′ 200 bp (PVX-5′CP 200) or 400 bp (PVX-5′CP 400) of CP, and total RNA (10 μg) isolated from leaf tissues 7–8 days after virus inoculation was analyzed by Northern hybridization with a 32P labeled PVX-CP specific DNA probe. (A) Accumulation of viral RNA in transgenic plants inoculated with PVX-5′ CP 200. (B) Accumulation of viral RNA in transgenic plants inoculated with PVX-5′CP 400. ΔGUS plants inoculated with recombinant PVX were used as a control. Instead of viral RNA, water was inoculated in ΔGUS plants as a mock inoculation. An ethidium bromide stained RNA gel photographed before transfer is also shown as control for equal loading
Graft transmission of RNA silencing using transgenic lines carrying 5′ 200 or 400 bp of CP
To investigate whether the newly generated silenced lines could induce RNA silencing in non-silenced scions by grafting, non-silenced scions (lines 200.1 and 400.89) were grafted onto silenced rootstocks (lines 200.4 and 400.15). Total RNA and small RNAs were extracted from one or two leaves closest to the graft junction of the scions 6 weeks after grafting. Transgene mRNA levels were significantly reduced in the non-silenced scions (Fig. 4A). siRNAs were also detected in these scions (Fig. 4B). Shoots of the decapitated lower part of non-silenced plants were kept as controls and tested at the same time; they accumulated transgene mRNA at high levels and had no siRNAs. This result indicates that RNA silencing was induced in non-silenced scions by silenced rootstocks, as previously reported (Palauqui et al. 1997; Palauqui and Vaucheret 1998; Voinnet et al. 1998; Crete et al. 2001; Sonoda and Nishiguchi 2000; Garcia-Perez et al. 2004). These results were reproduced in three individual experiments using eight grafted plants.
Induction of RNA silencing in non-silenced scions by silenced rootstocks. (A) Total RNA (10 μg), extracted from the leaves of non-silenced scions approximately 6 weeks after grafting onto silenced rootstocks (200.1/200.4 and 400.89/400.15), were analyzed by Northern hybridization with 32P labeled probes specific to 5′ 400 bp of CP. RNA extracted from beheaded non-silenced plants was used as a control (200.1 and 400.89). An ethidium bromide stained RNA gel photographed before transfer is also shown as a control for equal loading. (B) Small RNA fractions (50 μg), isolated from leaves of non-silenced scions approximately 6 weeks after grafting, were analyzed by Northern hybridization with 32P labeled 5′ 400 bp of CP-specific DNA probe. DNA oligomers were used for size control (size indicated in nucleotides). As a control for equal loading, ethidium bromide stained tRNA fractions are shown below
Grafting using silenced rootstocks carrying 5′ 200 or 400 bp of CP
To explore whether the graft-induced RNA silencing could spread along the transgene mRNA in the scions, we devised a set of graft experiments. At first, non-silenced scions carrying the entire transgene (line 4.07) (Sonoda et al. 1999) were grafted onto the silenced rootstocks of 200.4 and 400.15. Total and small RNA fractions extracted from the grafted scions 6 weeks after grafting were analyzed by Northern blot analysis using radiolabeled probe corresponding to the 5′ 400 bp of CP (5′CP) (Fig. 5A). Transgene mRNA levels were reduced in the grafted scions (Fig. 5B). Transgene derived siRNAs were also detected in these grafted scions (Fig. 5E). After removing the probe, the RNA membranes were re-hybridized with probes specific to either the middle region of CP (MCP) or the 3′ region of CP (3′CP) (see Materials and Methods for detail). Using either probe, reduced levels of transgene mRNA accumulation were observed (Fig. 5C, D) and their corresponding siRNAs were detected (Fig. 5F, G). Shoots of the beheaded lower part of the 4.07 plants were kept as controls and were tested at the same time. High accumulation of transgene mRNAs and no siRNAs were observed in the control using all three probes (Fig. 5, lane 1 in B–G). From these results, it was inferred that RNA silencing was induced in the grafted scions by the silenced rootstocks, and that the graft-induced RNA silencing could spread from the 5′ to 3′ direction along the transgene mRNA.
Analysis of transitive RNA silencing in non-silenced scions after grafting onto silenced rootstocks. (A) Schematic representation of the CP (including the non-translated region, NTR) including position of the different regions of the transgene used as probes. (A–C) Total RNA (10 μg), extracted from non-silenced scions 6 weeks after grafting onto silenced rootstocks (4.07/200.4 and 40.7/400.15), were analyzed by Northern hybridization with 32P labeled probes specific to 5′CP (B), MCP (C) and 3′CP (D). RNA, extracted from beheaded non-silenced plants, was used as a control (4.07). An ethidium bromide stained RNA gel photographed before transfer is shown as a control for equal loading. (E–G) Small RNA fractions (50 μg), isolated from similar leaf samples (A, B and C), were analyzed by Northern hybridization with 32P labeled probes specific to 5′CP (E), MCP (F) and 3′CP (F). DNA oligomers were used for size control (size indicated in nucleotides). As a control for equal loading, ethidium bromide stained tRNA fractions are shown below
To further analyze the spreading of RNA silencing, silenced scions of 4.09 were grafted onto the silenced rootstocks of 200.4 and 400.15. Line 4.09 carries the entire CP but the target specificity for RNA silencing is located only at 3′ region of the transgene (Sonoda et al. 1999). Total and small RNA fractions extracted from the grafted scions 6 weeks after grafting were analyzed by Northern blot analysis using radiolabeled probes corresponding to the different regions of CP (Fig. 5A). Reduced levels of transgene mRNA accumulation were observed in the grafted scions (Fig. 6A, B) and their corresponding siRNAs were also detected with either 5′CP or MCP probe (Fig. 6C, D). Shoots of the beheaded lower part of the 4.09 plants were kept or grafted on the non-silenced rootstocks (200.1 and 400.89) as a control, and were tested at the same time as the above experiment. Using either probe, they accumulated the transgene mRNAs at high levels (Fig. 6, lanes 5–7 in A, B) and had no siRNAs (Fig. 6, lanes 5–7 in C, D). This observation showed that RNA silencing could spread from the 5′ to 3′ direction along the transgene mRNA.
Analysis of graft-induced transitive RNA silencing in 4.09 scions. (A, B) Total RNA (10 μg), extracted from scions 6 weeks after grafting onto silenced rootstocks (4.09/200.4 and 4.09/400.15), were analyzed by Northern hybridization with 32P labeled probes specific to 5′CP (A) and MCP (B). RNA extracted from beheaded 4.09 plants or 4.09 scions grafted onto non-silenced rootstocks (4.09/200.1 and 4.09/400.89) were used as a control. An ethidium bromide stained RNA gel photographed before transfer is shown as a control for equal loading. (C, D) Small RNA fractions (50 μg), isolated from similar leaf samples, were analyzed by Northern hybridization with 32P labeled probes specific to 5′CP (C) and MCP (D). DNA oligomers were used for size control (size indicated in nucleotides). As a control for equal loading, ethidium bromide stained tRNA fractions are shown below
Grafting of silenced rootstocks with target specificity for the 3′ CP gene
Silenced rootstocks of line 4.09 were used as silencing inducers to investigate whether the RNA silencing could spread along the upstream sequences of the silencing inducer. Non-silenced scions carrying the entire transgene (line 4.07) were grafted onto silenced rootstocks (line 4.09). Total RNA and small RNA fractions of the grafted scions were analyzed with radiolabeled probes corresponding to different regions of CP (Fig. 5A). With the 3′CP probe, lower levels of transgene mRNA accumulation were observed in the scions 6 weeks after grafting (Fig. 7C). siRNAs corresponding to the 3′ region of the transgene were also detected in grafted scions (Fig. 7F). When the RNA membranes were re-hybridized with the MCP or 5′CP probes, transgene mRNA accumulation was observed in the grafted scions (Fig. 7A, B), and no siRNAs corresponding to these regions could be detected (Fig 7D, E). Shoots of beheaded 4.07 plants were kept as a control. They accumulated the full length transgene transcript at high levels and no siRNA could be detected using all three probes. In order to better understand these results, non grafted 4.09 plants were tested at the same time. The 4.09 plants accumulated the truncated transgene transcript (as compared to the non-silenced line 4.07) and generated siRNAs corresponding to the 3′ region of CP. This finding reinforced our previous observation that RNA silencing was targeted only to the 3′ region of CP in this line (Sonoda et al. 1999). These results show that RNA silencing was induced in non-silenced scions after grafting onto silenced rootstocks (Fig. 7C, F), and that the target specificity for RNA silencing was the same as that of rootstocks. This suggested that RNA silencing could not spread from the 3′ to 5′ direction along the transgene transcript.
Analysis of graft-induced transitive RNA silencing in the upstream region of the silencing inducer. (A–C) Total RNA (10 μg), extracted from 4.07 scions 6 weeks after grafting on the 4.09 rootstocks (4.07/4.09) were analyzed by Northern hybridization with 32P labeled probes specific to 5′CP (A), MCP (B) and 3′CP (C). RNA samples extracted from beheaded plants of 4.07 and age-matched 4.09 plants were used as controls. An ethidium bromide stained RNA gel photographed before transfer is shown as control for equal loading. (D–F) Small RNA fractions (50 μg), isolated from similar leaf samples, were analyzed by Northern hybridization with 32P labeled probes specific to 5′CP (D), MCP (E) and 3′CP (F). As a control for equal loading, Ethidium bromide stained tRNA fractions are shown below. (G) Total RNA (10 μg), extracted from non-silenced scions grafted onto 4.09 rootstocks (200.1/4.09, 400.89/4.09), were analyzed by Northern hybridization with radiolabeled 5′CP probe. RNA extracted from beheaded plants of 200.1 and 400.89 were used as a control. An ethidium bromide stained RNA gel photographed before transfer is shown as a control for equal loading. (H) Small RNA fractions (50 μg), isolated from non-silenced scions 6 weeks after grafting onto 4.09 rootstocks (200.1/4.09, 400.89/4.09), were analyzed by Northern hybridization with radiolabeled 5′CP probe. As a control for equal loading, ethidium bromide stained tRNA fractions are shown below
To clarify whether the silencing signals from the 3′ portion of the transgene transcript had any effects on the 5′ portion of transgene expression in the scions in the absence of any common sequences, non-silenced scions carrying the 5′ portion of the transgene (lines 200.1 and 400.89) were grafted onto silenced rootstocks (line 4.09). RNA samples from scions 6 weeks after grafting were analyzed with a radiolabeled 5′CP probe. No induction of RNA silencing was found in grafted scions. No reduction of transgene mRNA accumulation was observed in grafted scions, as compared with the beheaded plant controls (Fig. 7G). In addition, no siRNAs were detected in the grafted scions (Fig. 7H). This indicated that both the systemic silencing signals and the transgene mRNA in scions need to share any transgene sequences for the induction of RNA silencing in scions.
Analysis of transitive RNA silencing in the hybrid plants
To further confirm the finding that RNA silencing could not spread from the 3′–5′ direction in the 4.07 scions after being grafted onto the 4.09 rootstocks, we analyzed RNA silencing in hybrid plants obtained by crossing lines 4.09 and 4.07. Total RNA and small RNA fractions of the hybrid plants were analyzed with probes corresponding to different regions of CP. Lower levels of transgene mRNA accumulation were observed with a 3′CP probe (Fig. 8A) and their corresponding siRNAs were detected (Fig. 8D). However, higher accumulation of the truncated transgene transcript was found using the MCP and 5′CP probes (Fig. 8B, C), and no siRNAs corresponding to these regions of transgene could be detected (Fig. 8E, F). This showed that RNA silencing was induced in non-silenced lines by the silencing inducer locus of line 4.09, but the induced-RNA silencing did not spread along the transgene in the 5′ direction. This data indicated that RNA silencing could not spread from the 3′–5′ direction along the transgene transcript in these hybrid plants.
Analysis of transitive RNA silencing in hybrid plants. (A–C) Total RNA (10 μg), extracted from the leaves of approximately four week old plants, was analyzed by Northern hybridization with 32P labeled probes specific to 3′CP (A), MCP (B) and 5′CP (C). RNA samples, extracted from age-matched 4.09 and 4.07 plants, were used as controls. An ethidium bromide stained RNA gel photograph is shown as a control for equal loading. (D–F) Small RNA fractions (50 μg), isolated from similar leaf samples, were analyzed by Northern hybridization with 3′CP (D), MCP (E) and 5′CP (F) specific probes. As a control for equal loading, ethidium bromide stained tRNA fractions are shown below
Discussion
In this paper, we show RNA silencing can spread to downstream sequences of the silencing target region, but not to upstream sequences along the transgene transcript. The newly produced transgenic plants carrying the 5′ portion of CP showed a correlation between the copy number of transgene and their silencing state.
Based on analysis of total RNA, siRNA and virus resistance against the recombinant PVX, engineered to contain the transgene sequence, it was confirmed that the transgene in the transgenic plant lines 200.4 and 400.15 was post-transcriptionally silenced. Southern blot analysis revealed that silenced and non-silenced plants (lines 200.1 and 400.89) carried two copies and one copy of the transgene, respectively (Fig. 1). These results are in alignment with the report that plants with multiple copies of transgene and/or high levels of transgene expression are more likely to exhibit RNA silencing than plants with a single copy transgene or a low level of its transcription (Atkinson et al. 1998).
In our study, RNA silencing was found to spread from the 5′–3′ direction along the transgene mRNA. When non-silenced scions harboring the entire CP were grafted onto the 5′ silencing inducer rootstocks (silenced rootstocks carrying the 5′ portion of CP are mentioned as 5′ silencing inducer rootstocks), reduced levels of transgene mRNA and accumulation of siRNAs were observed with three different probes (Fig. 5). This indicates that the silencing signals derived from 5′ silencing inducer rootstocks were transmitted to non-silenced scions, and that they may have been amplified by the cellular RDR (RDR6) to induce transitive RNA silencing along the entire transgene mRNA (Van Houdt et al. 2003; Garcia-Perez et al. 2004; Schwach et al. 2005). Such amplification of the silencing signals might require the conversion of single-stranded CP transcript to dsRNA by the putative RDR. A primer-dependent process was employed to explain the spreading of RNA silencing in Caenorhabditis elegans and Drosophila melanogastar, in which spreading was observed in the 3′–5′ direction along the transgene resulting in the hypothesis that the siRNAs derived from the initial trigger would serve as primers for the synthesis of dsRNA by cellular RDR (Lipardi et al. 2001; Sijen et al. 2001). In order to apply this mechanism to the observed spreading of RNA silencing from the 5′ to 3′ direction, an antisense RNA would be needed as a template for RDR. Such antisense transcript has already been found in Arabidopsis (Yamada et al. 2003) and humans (Yelin et al. 2003; Rosok and Sioud 2004). An unprimed synthesis of cRNA from the initial trigger can also be speculated to explain the 5′–3′ spreading of RNA silencing in our study. It is notable that the RDR of tomato and wheat germ extract do not require any primer to synthesize cRNA in vitro (Schiebal et al. 1993a, b; Tang et al. 2003).
When non-silenced scions (line 4.07) were grafted onto the 3′ silencing inducer rootstocks (silenced rootstocks in which the silencing was targeted to the 3′ region of CP are mentioned as 3′ silencing inducer rootstocks), reduction of mRNA levels and accumulation of siRNAs were restricted to the 3′ region of the transgene in the scions (Fig. 7C). No spreading of RNA silencing was observed to the 5′ region or even to the middle region of the transgene transcript (Fig. 7A, B). We checked the hybrid plants obtained by a cross between the 3′ silencing inducer plants (line 4.09) and the non-silenced plants (line 4.07) to assess whether the RNA silencing could spread in the 5′ direction along the transgene mRNA when the silencing inducer and silencing target loci were both present in the same cell. RNA silencing was induced only at the 3′ region of the transgene and no transitivity towards the 5′ direction was found in the hybrid plants (Fig. 8). This experiment rules out the possibility of an inefficient transport of the silencing signal through the vascular system in the graft junction, which could be considered a reason for the failure of transitive silencing in the 3′–5′ direction. Although this finding conflicts with several observations in which RNA silencing has been reported to spread in both the 5′–3′ and 3′–5′ directions in transgenic plants (Voinnet et al. 1998; Klahre et al. 2002; Vaistij et al. 2002; Miki et al. 2005), it agrees with our previous observation (Sonoda and Nishiguchi 2000). A non-silenced scion (line 4.07) was susceptible to a recombinant PVX (engineered to carry 5′ 700 bp of transgene) when grafted onto the 3′ inducer rootstocks (4.09), suggesting that no spreading of RNA silencing occurred towards the 5′ sequences of the silencing inducer. It can be considered that plants have some mechanism to inhibit the amplification of silencing signals towards the upstream sequences of the silencing inducer, explaining why RNA silencing could not spread to the entire transgene transcript. Although, at present there are no reports to explain how such inhibitory mechanism may operate and under what circumstances they are activated, there are findings that suggest their existence. Only the 3′ region of the transgene mRNA was found to be targeted for RNA silencing and no spreading of silencing toward the upstream sequences was observed in transgenic plants (Sijen et al. 1996; English et al. 1996; Lee et al. 1997; Han and Grierson 2002). siRNAs could be detected predominantly from downstream sequences rather than upstream sequences of the silencing inducers in transgenic N. benthamiana, in which RNA silencing was induced by VIGS (Braunstein et al. 2002 and Petersen and Albrechtsen 2005). It is speculated that a similar inhibitory mechanism impedes the 3′–5′ spreading of RNA silencing in these experiments. Alternatively, an unprimed synthesis of cRNA from the downstream sequences of the silencing inducer may simply be favored over primed or unprimed synthesis of cRNA from the upstream sequences of the silencing inducer. Such cRNA synthesis was observed in a biochemical study of in vitro tomato RDR, in which the unprimed cRNA synthesis initiates preferentially at the 3′ terminus of the RNA template (Schiebal et al. 1993a, b).
However, several observations in plants have suggested that RNA silencing can spread from the 3′ to 5′ direction as well as from the 5′ to 3′ direction along transgene mRNA (Voinnet et al. 1998; Klahre et al. 2002; Vaistij et al. 2002; Miki et al. 2005). RNA silencing was found to spread along the entire region of GFP after induction with either the 5′ or the 3′ region of the transgene in transgenic N. benthamiana, and was accompanied by methylation of the corresponding transgene sequence (Vaistij et al. 2002). Biolistic delivery of siRNAs corresponding to the central region of GFP induced siRNA generation from both 5′ and 3′ regions of the transgene (Klahre et al. 2002).
Generation of transgenic plants carrying the middle and 3′ regions of the transgene and to use them as silencing inducers in graft experiments will be valuable to understand the mechanism of graft-induced transitive RNA silencing in more detail. The detailed transgene analysis of the 3′ silencing inducer plants (line 4.09) will aid to understand the assumed inhibitory mechanism that hindered the spreading of RNA silencing from the 3′ to 5′ direction.
Abbreviations
- abRNA:
-
Aberrant RNA
- Ago:
-
Argonaute
- CP :
-
Coat protein gene
- dsRNA:
-
Double-stranded RNA
- IR:
-
Inverted repeat
- PTGS:
-
Post-transcriptional gene silencing
- PVX:
-
Potato virus X
- RdRP:
-
RNA-dependent RNA polymerase
- RISC:
-
RNA-induced silencing complex
- siRNA:
-
Short interfering RNA
- SPFMV:
-
Sweet potato feathery mottle virus
References
Al-Kaff NS, Covey SN, Kreike MM, Page AM, Pinder R, Dale PJ (1998) Transcriptional and posttranscriptional plant gene silencing in response to a pathogen. Science 279:2113–2115
Atkinson RG, Bieleski LRF, Gleave AP, Janssen BJ, Morris BAM (1998) Post-transcriptional silencing of chalcone synthase in petunia using a geminivirus-based episomal vector. Plant J 15:593–604
Baulcombe DC (1996) RNA as a target and an initiator of post-transcriptional gene silencing in transgenic plants. Plant Mol Biol 32:79–88
Baulcombe DC (1999) Gene silencing: RNA makes RNA makes no protein. Curr Biol 9(16):R599–601
Baulcombe D (2004) RNA silencing in plants. Nature 431:356–363
Baumberger N, Baulcombe DC (2005) Arabidopsis Argonaute1 is an RNA slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci USA 102:11928–11933
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409:363–366
Braunstein TH, Moury B, Johannessen M, Albrechtsen M (2002) Specific degradation of 3′ regions of GUS mRNA in posttranscriptionally silenced tobacco lines may be related to 5′-3′ spreading of silencing. RNA 8:1034–1044
Chapman SN, Kavanagh TA, Baulcombe DC (1992) Potato virus X as a vector for gene expression in plants. Plant J 2:549–557
Chuang CH, Meyerowitz EM (2000) Specific and heritable genetic interference by double-stranded RNA in Arabidopsis thaliana. Proc Natl Acad Sci USA 97:4985–4990
Covey SN, Al-Kaff NS, Langara A, Turner DS (1997) Plants combat infection by gene silencing. Nature 385:781–782
Crete P, Leuenberger S, Iglesias VA, Suarez V, Schob H, Holtorf H, van Eeden S, Meins Jr F (2001) Graft transmission of induced and spontaneous post-transcriptional silencing of chitinase genes. Plant J 28(5):493–501
Dalmay T, Hamilton A, Rudd S, Angell S, Baulcombe DC (2000) An RNA-dependent RNA polymerase gene in Arabidopsis is required for posttranscriptional gene silencing mediated by a transgene but not by a virus. Cell 101:543–553
Dunoyer P, Himber C, Voinnet O (2005) DICER-LIKE 4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal. Nat Genet 37(12):1356–1360
Elbashir SM, Lendeckel W, Tuschl T (2001) RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev 15:188–200
English JJ, Mueller E, Baulcombe DC (1996) Suppression of virus accumulation in transgenic plants exhibiting silencing of nuclear genes. Plant Cell 8:179–188
Fagard M, Vaucheret H (2000) Systemic silencing signal(s). Plant Mol Biol 43:285–293
Finnegan EJ, Margis R, Waterhouse PM (2003) Posttranscriptional gene silencing is not compromised in the Arabidopsis CARPEL FACTORY (DICER-LIKE1) mutant, a homolog of Dicer-1 from Drosophila. Curr Biol 13:236-240
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Garcia-Perez RD, Van Houdt H, Depicker A (2004) Spreading of post-transcriptional gene silencing along the target gene promotes systemic silencing. Plant J 38:594–602
Grishok A, Pasquinelli AE, Conte D, Li N, Parrish S, Ha I, Baillie DL, Fire A, Ruvkun G, Mello CC (2001) Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106:23–34
Hamilton AJ, Baulcombe DC (1999) A novel species of small antisense RNA in post-transcriptional gene silencing. Science 286:950–952
Hamilton A, Voinnet O, Chappell L, Baulcombe DC (2002) Two classes of short interfering RNA in RNA silencing. EMBO J 21:4671–4679
Han Y, Grierson D (2002) Relationship between small antisense RNAs and abRNAs associated with sense transgene mediated gene silencing in tomato. Plant J 29:509–519
Herr AJ (2004) Silence is green. Biochem Soc Trans 32:946–951
Himber C, Dunoyer P, Moissiard G, Ritzenthaler C, Voinnet O (2003) Transitivity-dependent and -independent cell-to-cell movement of RNA silencing. EMBO J 22:4523–4533
Horsch RB, Fry JE, Hoffman NL, Eicholtz D, Rogers S, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231
Ketting R, Haverkamp T, van Luenen H, Plasterk R (1999) mut-7 of C. elegans, required for transposon silencing and RNA interference, is a homolog of Werner syndrome helicase and RNaseD. Cell 99:133–141
Klahre U, Crete P, Leuenberger SA, Iglesias VA, Meins FJ (2002) High molecular weight RNAs and small interfering RNAs induce systemic posttranscriptional gene silencing in plants. Proc Natl Acad Sci USA 99:11981–11986
Lee KY, Baden C, Howie WJ, Bedbrook J, Dunsmuir J (1997) Post-transcriptional gene silencing of ACC synthase in tomato results from cytoplasmic RNA degradation. Plant J 12:1127–1137
Lindbo JA, Silva-Rosales L, Proebsting WM, Dougherty WG (1993) Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 5:1749–1759
Lipardi C, Wei Q, Paterson BM (2001) RNAi as random degradative PCR: siRNA primers convert mRNA into dsRNAs that are degraded to generate new siRNAs. Cell 107:297–307
Martinez J, Patkaniowska A, Urlaub H, Lührmann R, Tuschl T (2002) Single-stranded antisense siRNAs guide target RNA cleavage in RNAi. Cell 110:563–574
Miki D, Itoh R, Shimamoto K (2005) RNA silencing of single and multiple members in a gene family of rice. Plant Physiol 138:1903–1913
Mlotshwa S, Schauer SE, Smith TH, Mallory AC, Herr JM Jr, Roth B, Merchant DS, Ray A, Bowman LH, Vance VB (2005) Ectopic DICER-LIKE1 expression in P1/HC-Pro Arabidopsis rescues phenotypic anomalies but not defects in microRNA and silencing pathways. Plant Cell 17(11):2873–2885
Mlotshwa S, Voinnet O, Mette MF, Matzke M, Vaucheret H, Ding SW, Pruss G, Vance VB (2002) RNA silencing and the mobile silencing signal. Plant Cell 14:S289–S301
Mori M, Sakai J, Kimura T, Usugi T, Hayashi T, Hanada K, Nishiguchi M (1995) Nucleotide sequence analysis of two nuclear inclusion body and coat protein genes of a sweet potato feathery mottle virus severe strain (SPFMV-S) genomic RNA. Arch Virol 140:1473–1482
Mourrain P, Beclin C, Elmayan T, Feuerbach F, Godon C, Morel JB, Jouette D, Lacombe AM, Nikic S, Picault N, Remoue K, Sanial M, Vo T, Vaucheret H (2000) Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 101:533–542
Napoli C, Lemieux C, Jorgensen R (1990) Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2:279–289
Palauqui JC, Elmayan T, Pollien JM, Vaucheret H (1997) Systemic acquired silencing: transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions. EMBO J 16:4738–4745
Palauqui JC, Vaucheret H (1998) Transgenes are dispensable for the RNA degradation step of cosuppression. Proc Natl Acad Sci USA 95:9675–9680
Papp I, Mette MF, Aufsatz W, Daxinger L, Schauer SE, Ray A, Winden J, Matzke M, Matzke AJ (2003) Evidence for nuclear processing of plant micro RNA and short interfering RNA precursors. Plant Physiol 132:1382–1390
Petersen BO, Albrechtsen M (2005) Evidence implying only unprimed RdRP activity during transitive gene silencing in plants. Plant Mol Biol 58:575–583
Romano N, Macino G (1992) Quelling: transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol Microbiol 6:3343–3353
Rosok O, Sioud M (2004) Systemic identification of sense-antisense transcripts in mammalian cells. Nat Biotechnol 22:104–108
Schiebel W, Haas B, Marinkovic S, Klanner A, Sanger HL (1993a) RNA-directed RNA polymerase from tomato leaves. I. Purification and physical properties. J Biol Chem 268:11851–11857
Schiebel W, Haas B, Marinkovic S, Klanner A, Sanger HL (1993b) RNA-directed RNA polymerase from tomato leaves. II. Catalytic in vitro properties. J Biol Chem 268:11858–11867
Schwach F, Vaistij FE, Jones L, Baulcombe DC (2005) An RNA-dependent RNA polymerase prevents meristem invasion by Potato virus X and is required for the activity but not the production of a systemic silencing signal. Plant Physiol 138:1842–1852
Sijen T, Fleenor J, Simmer F, Thijssen KL, Parrish S, Timmons L, Plasterk RHA, Fire A (2001) On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107:465–476
Sijen T, Wellink J, Hiriart JB, van Kammen A (1996) RNA-Mediated virus resistance: role of repeated transgenes and delineation of targeted regions. Plant Cell 8:2277–2294
Smith CJ, Watson CF, Bird CR, Ray J, Schuch W, Grierson D (1990) Expression of a truncated tomato polygalacturonase gene inhibits expression of the endogenous gene in transgenic plants. Mol Gen Genet 224(3):477–81
Smith NA, Singh SP, Wang MB, Stoutjesdijk PA, Green AG, Waterhouse PM (2000) Total silencing by intron-spliced hairpin RNAs. Nature 407:319–320
Sonoda S, Mori M, Nishiguchi M (1999) Homology-dependent virus resistance in transgenic plants with the coat protein gene of sweet potato feathery mottle potyvirus: Target specificity and transgene methylation. Phytopathology 89:385–391
Sonoda S, Nishiguchi M (2000) Graft transmission of post-transcriptional gene silencing: target specificity for RNA degradation is transmissible between silenced and non-silenced plants, but not between silenced plants. Plant J 21:1–8
Tang G, Brenda JR, David PB, Zamore PD (2003) A biochemical framework for RNA silencing in plants. Genes Dev 17:49–63
Vaistij FE, Jones L, Baulcombe DC (2002) Spreading of RNA targeting and DNA methylation in RNA silencing requires transcription of the target gene and a putative RNA-dependent RNA polymerase. Plant Cell 14:857–867
van der Krol AR, Mur LA, Beld M, Mol JN, Stuitje AR (1990) Flavonoid genes in petunia: addition of a limited number of gene copies may lead to a suppression of gene expression. Plant Cell 2:291–299
Van Houdt H, Bleys A, Depicker A (2003) RNA target sequences promote spreading of RNA silencing. Plant Physiol 131:245–253
Voinnet O, Baulcombe DC (1997) Systemic signalling in gene silencing. Nature 389:553
Voinnet O, Vain P, Angell S, Baulcombe D (1998) Systemic spread of sequence-specific transgene RNA degradation in plants is initiated by localized introduction of ectopic promoterless DNA. Cell 95:177–187
Xie Z, Allen E, Wilken A, Carrington JC (2005) DICER-LIKE 4 functions in trans-acting small interfering RNA biogenesis and vegetative phase change in Arabidopsis thaliana. Proc Natl Acad Sci USA 102(36):12984–12989
Xie Z, Fan B, Chen C, Chen Z (2001) An important role of an inducible RNA-dependent RNA polymerase in plant antiviral defense. Proc Natl Acad Sci USA 98:6516-6521
Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, Zilberman D, Jacobsen SE, Carrington JC (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2:E104
Yamada K et al (2003) Empirical analysis of transcriptional activity in the Arabidopsis genome. Science 302:842–846
Yelin R, Dahary D, Sorek R, Levanon EY, Goldstein O, Shoshan A, Diber A, Biton S, Tamir Y, Khosravi R, Nemser S, Pinner E, Walach S, Bernstein J, Savitsky K, Rotman G (2003) Widespread occurrence of antisense transcription in the human genome. Nat Biotechnol 21:379–386
Yu D, Fan BSA, MacFarlane, Chen Z (2003) Analysis of the involvement of an inducible Arabidopsis RNA-dependent RNA polymerase in antiviral defense. Mol Plant Microbe Interact 16:206–216
Acknowledgements
The authors would like to thank David Baulcombe (The Sainsbury Laboratory, Norwich, UK) and his other lab members for kindly providing the PVX vector and specific comments on the crossing of N. benthamiana. They are also grateful to Sk. Md. Fazle Akbar (School of Medicine, Ehime University, Japan) and Thangavelu U. Arumugam (Faculty of Agriculture, Ehime University) for critical reading of the manuscript, and Charles Lecellier (Institut de Biologie Moleculaire des Plantes - CNRS UPR2357, France) for suggestions concerning the siRNA detection experiment. This work was financially supported in part by Ministry of Education, Culture, Sports, Science and Technology of Japan and by Ministry of Agriculture, Forestry and Fisheries of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Haque, A.K.M.N., Tanaka, Y., Sonoda, S. et al. Analysis of transitive RNA silencing after grafting in transgenic plants with the coat protein gene of Sweet potato feathery mottle virus . Plant Mol Biol 63, 35–47 (2007). https://doi.org/10.1007/s11103-006-9070-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-006-9070-6