Abstract
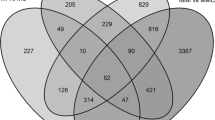
Oidium neolycopersici is a causal agent of tomato powdery mildew. In this paper, gene expression profiles were investigated of susceptible, monogenic- and polygenic resistant tomato genotypes in response to O. neolycopersici infection by using cDNA-AFLP. Around 30,000 TDFs (Transcript Derived Fragments), representing ∼22% of the transcriptome based on in silico estimation, were identified and 887 TDFs were differentially expressed (DE-TDFs) upon inoculation with O. neolycopersici spores. Forty-two percent of the identified DE-TDFs were detected in both the compatible and incompatible interactions, a subset of these were studied for their temporal patterns. All of these common induced DE-TDFs displayed an expression peak at 7 days post incoluation in monogenic resistant response but sustained up-regulation in the susceptible and the polygenic resistant response. While more than half of these common DE-TDFs showed earlier timing in incompatible interactions compared to compatible interaction. Only 2% of the identified DE-TDFs were specific to either the monogenic or the polygenic resistant response. By annotation of the 230 sequenced DE-TDFs we found that 34% of the corresponding transcripts were known to be involved in plant defense, whereas the other transcripts played general roles in signal transduction (11%), regulation (24%), protein synthesis and degradation (11%), energy metabolism (12%) including photosynthesis, photorespiration and respiration.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- DE-TDF:
-
Differentially expressed TDF
- DPI:
-
Days post inoculation
- HPI:
-
Hours post inoculation
- TDF:
-
Transcript derived fragment
- HR:
-
Hypersensitive response
References
Bachem CW, Oomen RJ, Visser RG (1988) Transcript imaging with cDNA-AFLP: a step-by-step protocol. Plant Mol Biol Rep 16:157–173
Bachem CW, van der Hoeven RS, de Bruijn SM, Vreugdenhil D, Zabeau M, Visser RG (1996) Visualization of differential gene expression using a novel method of RNA fingerprinting based on AFLP: analysis of gene expression during potato tuber development. Plant J 9:745–753
Bai Y, Huang CC, van der Hulst R, Meijer-Dekens F, Bonnema G, Lindhout P (2003) QTLs for tomato powdery mildew resistance (Oidium lycopersici) in Lycopersicon parviflorum G1.1601 co-localize with two qualitative powdery mildew resistance genes. Mol Plant Microbe Interact 16:169–176
Bai Y, van der Hulst R, Bonnema G, Marcel TC, Meijer-Dekens F, Niks R, Lindhout P (2005) Tomato defense to Oidium neolycopersici: Dominant Ol genes confer isolate-dependent resistance via a different mechanism than recessive ol-2. Mol Plant Microbe Interact 18:354–362
Chen Q, Vazquez EJ, Moghaddas S, Hoppel CL, Lesnefsky EJ (2003) Production of reactive oxygen species by mitochondria: central role of complex III. J Biol Chem 278:36027–36031
Coaker G, Falick R, Staskawicz B (2005) Activation of a phytopathogenic bacterial effector protein by a Eukaryotic cyclophilin. Nature 308:548–550
Durrant WE, Rowland O, Piedras P, Hammond-Kossak KE, Jones JDG (2000) cDNA-AFLP reveals a striking overlap in the race-specific resistance and wound response expression profiles. Plant Cell 12:963–977
Ditt RF, Nester EW, Comai L (2001) Plant gene expression response to Agrobacterium tumefaciens. PNAS 98:10954–10959
Eulgem T (2005) Regulation of the Arabidopsis defense transcriptome. Trends Plant Sci 10:71–78
Flor HH (1971) Current status of the gene-for-gene concept. Annu Rev Phytopathol 9:275–296
Gjetting T, Carver TL, Skot L, Lyngkjaer MF (2004) Differential gene expression in individual papilla-resistant and powdery mildew-infected barley epidermal cells. Mol Plant Microbe Interact 17:729–738
Gregersen PL, Thordal-Christensen H, Forster H, Collinge DB (1997) Differential gene transcript accumulation in barley leaf epidermis and mesophyll in response to attack by Blumeria graminis f. sp. Hordei (syn. Erysiphe graminis f. sp. hordei). Physiol Mol Plant Pathol 51:85–97
Hammond-Kosack KE, Parker JE (2003) Deciphering plant-pathogen communication: fresh perspectives for molecular resistance breeding. Curr Opin Plant Biol 14:177–193
Fulop K, Pettko-Szandtner A, Magyar Z, Miskolczi P, Kondorosi E, Dudits D, Bako L (2005) The Medicago CDKC; 1-CYCLINT; 1 kinase complex phosphorylates the carboxy-terminal domain of RNA polymerase II and promotes transcription. Plant J 42:810–820
Huang CC, Groot T, Meijer-Dekens F, Niks RE, Lindhout P (1998) The resistance to powdery mildew (Oidium lycopersicum) in Lycopersicon species is mainly associated with hypersensitive response. Eur J Plant Pathol 104:399–407
Huang CC, Cui YY, Weng CR, Zabel P, Lindhout P (2000a) Development of diagnostic markers closely linked to the tomato powdery mildew resistance gene Ol-1 on chromosome 6 of tomato. Theor Appl Genet 101:918–924
Huang CC, van der Putte PM, Haanstra-van der Meer JG, Meijer-Dekens F, Lindhout P (2000b) Characterization and mapping of resistance to Oidium lycopersicum in two Lycopersicon hirsutum accessions: Evidence for close linkage of two Ol-genes on chromosome 6. Heredity 85:511–520
Hückelhoven R, Dechert C, Kogel KH (2003) Overexpression of barley BAX inhibitor 1 induces breakdown of mlo-mediated penetration resistance to Blumeria gramins. PNAS 100(9):5555–5560
Jones H, Whipps JM, Guu SJ (2001) The tomato powdery mildew fungus Oidium neolycopersici, Mol Plant Pathol 2:303–309
Jones H, Whipps JM, Thomas BJ, Carver LW, Guu SJ (2000) Initial events in the colonization of tomatos by Oidium neolycopersici, a distinct powdery mildew fungus of Lycpersicon species. Can J Bot 78:1361–1366
Joosten M, de Wit P (1999) The tomato-Cladosporium fulvum interaction: a versatile experimental system to study plant–pathogen interactions. Annu Rev Phytopathol 37:335–367
Lindhout P, Pet G, van der Beek H (1994a) Screening wild Lycopersicon species for resistance to powdery mildew (Oidium lycopersicum). Euphytica 72:43–49
Lindhout P, van der Beek H, Pet G (1994b) Wild Lycopersicon species as sources for resistance to powdery mildew (Oidium lycopersicum): mapping of resistance gene Ol-1 on chromosome 6 of Lycopersicon hirsutum. Acta Horticult 376:387–394
Maleck K, Levine A, Eulgem T, Morgan A, Schmid J, Lawton KA, Dangl JL, Dietrich RA (2000) The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nat Genet 26:403–420
Mysore KS, Crasta OR, Tuori RP, Folkers O, Swirsky PB, Martin GB (2002) Comprehensive transcript profiling of␣Pto- and Prf-mediated host defense responses to infection by Pseudomonas syringae pv. tomato. Plant J 32:299–215
Panstruga R (2003) Establishing compatibility between plants and obligate biotrophic pathogens. Curr Opin Plant Biol 6:32–326
Reijans M, Lascaris R, Groeneger AO, Wittenberg A, Wesselink E, van Oeveren J, de Wit E, Boorsma A, Voetdijk B, van der Spek H, Grivell LA, Simons G (2003) Quantitative comparison of cDNA-AFLP, microarray, and GeneChip expression data in Saccharomyces cerevisiae. Genomics 82:606–618
Schulze-Lefert P, Vogel J (2000) Closing the ranks to attack by powdery mildew. Trends Plant Sci 5:343–348
Tao Y, Xie Z, Chen W, Glazebrook J, Chang HS, Han B, Zhu T, Zou G, Katagiri F (2003) Quantitative nature of Arabidopsis responses during compatible and incompatible interactions with bacterial pathogen Pseudomonas syringae. Plant Cell 15:317–330
Van der Hoeven R, Ronning C, Giovannoni J, Martin G, Tanksley S (2002) Deductions about the number, organization, and evolution of genes in the tomato genome based on analysis of a large expressed sequence tag collection and selective genomic sequencing. Plant Cell 14:1441–1456
Vos P, Hogers R, Bleek M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new concept for DNA fingerprinting. Nuclei Acids Res 23:4965–4970
Acknowledgements
We would like to thank Dr C. Bachem for the advice and help on cDNA-AFLP, Dr X. Wang for helpful discussion and Ms J. Tang for the help on developing computer program RE-predictor. This work was supported by the Joint PhD program between Wageningen University and Chinese Academy of Agricultural Sciences and by the grants of International Foundation for Science (C/3395–1), the Laboratory of Plant Breeding of Wageningen University, and by the opening Key Laboratory of Vegetable Genetics and Physiology of Chinese Ministry of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, C., Bai, Y., Jacobsen, E. et al. Tomato defense to the powdery mildew fungus: differences in expression of genes in susceptible, monogenic- and polygenic resistance responses are mainly in timing. Plant Mol Biol 62, 127–140 (2006). https://doi.org/10.1007/s11103-006-9008-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-006-9008-z