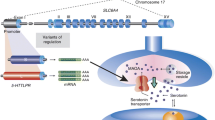
Abstract
We have demonstrated in both the human serotonin transporter gene (5HTT) and the dopamine transporter gene (DAT1) that specific polymorphic variants termed Variable Number Tandem Repeats (VNTRs), which correlate with predisposition to a number of neurological and psychiatric disorders, act as transcriptional regulatory domains. We have demonstrated that these domains can act as both tissue-specific and stimulus-inducible regulators of gene expression. As such they can act to be mechanistically associated with the progression or initiation of a behavioural disorder by altering the level of transporter mRNA, which in turn regulates the concentration of transporter in specific cells or in response to a challenge; chemical, environmental or physiological. The synergistic actions of such transcriptional domains will modulate gene expression. Our hypothesis is that these VNTR variants are one mechanism by which nurture can modify concentrations of neurotransmitters in a differential manner.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although we are all genetically individual humans share 99.9% sequence similarity between their genomes. It is estimated that the human genome has about 10 million polymorphisms or allelic variations. It is this variation between our genomes, and the mechanisms employed to regulate expression of our genes, that lead in part to individuality between us. A different combination of these polymorphisms creates the “genetic fingerprint”, which is used to distinguish one person’s DNA from another.
Genetic factors have been implicated in the aetiology of mental illness and many studies have determined that changes in protein structure correlate with a predisposition to specific conditions. There are, however, a number of polymorphisms found in non-coding sequences, which do not affect protein structure, that have been identified as risk factors for behavioural and affective disorders including Alzheimer’s disease [1], schizophrenia [2], anxiety [3], obsessive compulsive disorder (OCD) [4], unipolar depression [5], bipolar depression [6–8], migraine [9, 10], and Parkinson’s disease [11]. Having a genetic variant described as predisposing for a certain disorder does not mean that an individual with that variant will develop that disorder, however these polymorphisms might act as “markers” indicating a predisposition to a disorder. We have demonstrated that polymorphisms have functional effects on gene expression, displaying both tissue-specific and stimulus-inducible regulation [12–16]. The functional significance of a polymorphism may be relevant to an individual’s response to pharmacological treatment or for modulation in response to environmental factors. This suggests that individuals with a particular combination of polymorphisms may respond differently to the same medications or environmental stresses.
Identification of specific polymorphisms which associate with specific disorders, or respond to specific environmental stimuli, could lead to tailored treatments or bespoke medication for individuals based on genome variation. This review briefly describes types of allelic variation that exist, with particular emphasis on the Variable Number Tandem Repeats (VNTR) found within non-coding sequences in the serotonin transporter (5-HTT also termed SLC6A4) and dopamine transporter (DAT1 also termed SLC6A3) genes. It will also consider the possibility that these VNTR represent a class of cis-regulatory element to which a number of trans-acting factors may bind to modulate gene expression and it will focus on the modulatory nature of VNTR function in response to illicit drugs.
Allelic variation: Single Nucleotide Polymorphism (SNP)
Genetic variation is manifest in a number of modes. The most widely investigated polymorphisms and the first type that were identified, are single nucleotide polymorphisms (SNPs). SNPs constitute a major source of variation in the genome and occur about every 1,000 bp in the entire human sequence. There is even variability in the type of SNP that can occur. The simplest form of SNP is a base pair substitution where one nucleotide in the DNA sequence is exchanged for another, for example, an adenine (A) may be substituted for one of the other nucleotides such as a guanine (G). Other SNPs include; omissions, where a nucleotide is absent in one sequence but not in another or insertions, where an extra nucleotide is found in a DNA sequence. The SNP Consortium’s Allele Frequency project or the International HapMap project is a collaboration of numerous international research groups working together to produce a freely available, extensive analysis of the variation in the human genome across different populations (http://www.snp.cshl.org/). HapMap, which estimates the occurrence of 10 million SNPs within the genome, has currently identified more than 1.5 million SNPs. Many of these SNPs occur together, which is termed linked, for example as in Fig. 1a, a SNP at locus A might always be present with a SNP at locus B, or even with another polymorphism at loci C or D. This linkage occurs as a result of inheritance and SNPs which are always inherited linked together are known as haplotypes (Fig. 2), so a particular haplotype, rather than an individual SNP, may be associated with predisposition to a specific condition. Haplotypes may be identified by a common tagging SNP (tSNP) reducing the need to examine all individual SNP loci when performing an association study. A tSNP is defined as a change in a particular nucleotide at a given position associated with a specific haplotype (Fig. 2).
Definition of polymorphisms. The polymorphisms at locus A and locus B represent Single Nucleotide Polymorphism(s) or SNP(s), where one nucleotide has been substituted for another. Deletion or insertion of a single nucleotide is another example of a SNP. Locus C shows a micro-satellite polymorphism, or Short Tandem Repeat (STR). Common STRs are composed of di-, tri- or tetra-nucleotide repeats. Locus D is representative of a Variable Number Tandem Repeat (VNTR). These are a subclass of mini-satellite composed of repeats that are 10–100 nucleotides in length. They are variable as each allele may have a different copy number of these repeats. For example, here one allele has 3.5 copies of the repeat while the other has only two copies
Haplotypes and tagging SNPs. Linked polymorphisms that are inherited together are termed haplotypes. Here, for example, the first bold polymorphic position is always present with the other two bold polymorphic positions. At any of these positions the polymorphism is a substitution SNP where one nucleotide is substituted for another e.g., g for a, or t for c. In this example this would lead to the possible existence of nine different haplotypes. As genotyping these SNPs allows identification of each haplotype without having to analyse the whole sequence they are know as tagging SNPs or tSNPs. tSNP are particularly useful if one is always linked with a larger polymorphism as genotyping a SNP is more cost effective and less time consuming than genotyping a mini- or micro-satellite
Allelic variation: micro-, mini-satellites, repeats and Variable Number Tandem Repeats (VNTR)
There are over 500,000 identified micro- or mini-satellites in the human genome. Mini-satellites consist of 10–100 nucleotides repeated several times in tandem, which are bordered by unique DNA sequences. Micro-satellites also called Short Tandem Repeats (STRs), are similar to mini-satellites but the sequence repeats are smaller e.g., tetra- and di-nucleotide sequence repeats (Fig. 1). Most repeats are not comprised of “perfect” repeating units and show some degree of degeneration, where one repeat may be slightly different to the next but overall the core consensus sequence is maintained. Genotyping is employed to determine if a repeating element found within the genome is variable i.e., exists as a different allelic copy number between individuals in a population, for example at the same allelic loci, one individual may have 10 copies of a repeat, whilst another may have 12 copies. In this review we will concentrate on a subclass of mini-satellite repeats commonly referred to in the literature as Variable Number Tandem Repeats (VNTR). We define these VNTR as having sufficient DNA sequence in the repeat, for example greater than 6 bp, to act as a sequence specific DNA binding site for proteins such as transcription factors, and therefore have the potential to act as transcriptional regulatory domains (Fig. 1). This situation is similar to the regulatory repeat domains seen in retroviruses or herpesviruses [17–19].
These potential regulatory classes of VNTR are many and varied but can share some common features. Firstly, the majority of these VNTRs are located in non-coding regions of the genome. Secondly, many are present in the genome as a feature of an emerging evolution. VNTRs display evolutionary conservation between humans and non-human primates but are often not found in lower mammals [20, 21]. Indeed, many appear to be constantly evolving in modern primates by dramatic changes in copy number or large variation in primary sequence of the repeat (personal observations). Thirdly, many repeats contain potential cis-regulatory elements or transcription factor binding sites producing localised clusters of one or more particular binding site which may have implications for gene expression [14, 15, 21]. Lastly, in support of the prediction that these VNTR may act as modulators of gene expression many are found at higher density in gene-enriched areas compared to non-genic regions (G. Breen SDGP, IOP, Kings College, London, personal communication). Potential regulatory VNTRs can be identified by analysis of variation using a genome browser such as the UCSC Genome Bioinformatics Site, http://www.genome.ucsc.edu, and have been identified for example, in the following human genes: the serotonin transporter (SERT/5HTT), dopamine transporter (DAT1), N-methyl-d-aspartame receptor 1 (GRIN1/NMDAR), dopamine D4 receptor (DRD4), and monoamine oxidase A (MAOA).
Since they do not act to alter protein structure, these VNTRs would be predicted to act as endogenous modulators of transcription or alter post transcriptional properties (of the gene) such as mRNA stability. We propose, based on our published data, that this subclass of VNTR would function in both a tissue-specific and stimulus-inducible manner to fine-tune gene expression. This fine tuning could be correlated, mechanistically, not only with normal physiological function and variation between individuals, but also with a predisposition to behavioural disorders by altering neurotransmitter signalling in response to challenges and stress. Furthermore, if stimulus-inducible expression varies dependant on a specific polymorphism associated with a disorder then that may have similar implications in the response of an individual to pharmacological treatment of that disorder [12–15, 22].
Serotonin transporter
5-HT and the 5-HT transporter (sert/5-HTT/SLC6A4)
Clinical abnormalities in serotonin (5-HT) metabolism have been implicated in the pathophysiology of many CNS-related disorders. Early indications that 5-HT played a role in depression were based on the biogenic amine hypothesis which states that depression is caused by a deficiency of monoamines [23]. It was demonstrated that drugs which increased monoamines alleviated depression and visa-versa. It is now known that the monoamine 5-HT (5-hydroxytryptamine) is a major neuromodulator of cognitive and emotional behaviours involved in regulating diverse processes such as appetite [24, 25], memory and learning [26], aggression and antisocial behaviour [27], anxiety and depression [28, 29].
In the CNS, 5-HT and 5-HTT are mostly localized in the neurons of the raphe nuclei whose processes innervate many areas of the brain thought to be involved in cognition and behaviour regulation [30, 31]. Decreased serotonergic signaling as a result of low levels of 5-HT in the synaptic cleft, desensitization of 5-HT receptors, decreased expression of the 5-HT receptor, or changes in the expression of monoaminergic transporters can lead to the development of mood disorders such as depression. Conversely, increasing the concentration of 5-HT in the synaptic cleft can lead to the development of affective disorders such as schizophrenia. It is not surprising then, that the serotonergic signaling pathway is often a target for treating these complex disorders. In particular, changing the bioavailability of 5-HT in the synaptic cleft is a major mechanism of regulating 5-HT signaling. Abundance of 5-HT in the cleft is regulated principally via re-uptake by the monoaminergic, sodium and chloride dependant, 5-HT transporter (SERT/5HTT/SLC6A4) [32]. Following re-uptake, 5-HT is degraded by monoamine oxidases for recycling [33].
Since 5-HTT is a key regulator of the bioavailability of 5-HT, any modulation in the expression or action of 5-HTT would be expected to have consequences on behaviour and mood and has been strongly linked with depression [34, 35]. As such 5-HTT is a major target for pharmaceutical intervention in mood disorders. The family of antidepressants known as selective 5-HT reuptake inhibitors or SSRIs (such as Prozac) target only 5-HTT [36]. SSRIs block the transporter effectively raising, or at least maintaining, the concentration of 5-HT in the synaptic cleft. Tricyclic antidepressants (TCA) such as amtriptyline also target the 5-HTT and are still in widespread use despite having a number of undesirable side effects because they also target other norepinephrin transporters [36]. 5-HTT is also a target for illicit drugs such as ecstasy (MDMA) [37, 38] and cocaine [39–41].
Variation in the clinical response of individuals to various antidepressant treatments including the SSRIs, in part, will correlate with variation in their ability to process the drug and in their “normal” physiological control of the serotonergic system. For the latter, polymorphisms in the coding regions of the 5-HTT gene cannot account for these differences, and have not been associated with any psychiatric conditions. The identification of a number of VNTR in the non-coding regions of the 5-HTT gene that may have functional effects on the expression of 5-HTT are currently being investigated by a number of research groups, including our own, for their correlation with behavioral and psychiatric conditions.
5-HTTLPR VNTR: association studies and functionality
The human 5-HTT gene consists of 14 exons and spans 37.8 kb on chromosome 17q11.2. It encodes a 630 amino acid protein, containing 12 putative transmembrane domains and it shares 92% sequence homology with the rat 5-HTT gene [42, 43]. Mutation or inappropriate expression of the gene has been postulated as a possible cause of affective disorders. Although there are SNPs in the coding sequence of 5-HTT, thus far, no differences in amino acid sequence have been found between affective disorder patients and healthy controls, this finding therefore does not support a role for alteration of the primary structure of the coding region of the 5-HTT gene in the pathogenesis of affective disorders. However, possible associations between two VNTR polymorphisms in non-coding regions of the 5-HTT gene and susceptibility to affective disorder have been analysed.
The first of these to be identified was a biallelic insertion/deletion found in the 5′ promoter region of the gene 1.2 kb upstream of the transcriptional start site [44, 45]. This VNTR, termed, 5-HTTLPR was initially identified as two variants containing either, 14 (short/deletion) or 16 (long/insertion) copies of a 22 bp repeat (Fig. 3). More recently, Nakamura et al., verified the existence of subgroups within these variants to uncover as many as 14 allelic variations of this locus [46]. One particularly interesting variant of the long polymorphism contains an A to G (la or lg) substitution, tSNP (rs25531) (Fig. 3) [47–49]. Inclusion of this tSNP (lg) has conferred clinical properties normally associated with the short allele onto the long allele. Unless otherwise stated this review examines the common l and s variants referred to elsewhere as 14A (s) and 16A (l) [46]. Individuals can be heterozygous (l/s) or homozygous (l/l or s/s) for the VNTR. The distribution of these genotypes varies, with heterozygous and homozygous l/l individuals appearing with considerably more frequency than homozygous s/s individuals in the Caucasian population [44, 45]. However, allelic frequencies vary between global populations or different ethnic groups. In a review of the literature, Smits et al. determined that the frequency of the alleles for the Caucasian population was 27.4–28.3% (l/l), 43.4–51.0% (s/l) and 21.6–28.3% (s/s), whilst in Asian populations the allelic frequency was 4.2–10.0% (l/l), 30.0–39.2% (s/l) and 55.6–60.0% (s/s) [50].
Sequence and repeat alignment of the 5-HTTLPR VNTR. Schematic representation of the human 5-HTT gene and position of the LPR VNTR (top). On the left is the sequence of the long variant of the VNTR also known as the insertion VNTR due to the presence of four extra copies of the repeat (underlined). The VNTR is not composed of perfect repeating units as can be seen by the designation of Greek symbols to each different repeat (we have arranged the repeat to highlight this core). The bold underlined nucleotide “a” in the first ϕ repeat represents the a/g SNP (rs25531) which confers the functional properties associated with the short variant onto the long variant. On the right is the sequence of the short variant also known as the deletion
Association studies compare the genotype or haplotype of loci between patients and healthy individuals and analyse the data obtained to identify possible correlations between the occurrence of a specific variant and a disorder or disease. Many association studies have indicated that the 5-HTTLPR variants have been linked with a number of affective disorders. The homozygous s/s genotype has been associated with an increased risk to depression [51], unipolar depression [7], bipolar depression [52], anxiety [53, 54], substance abuse [55], and a predisposition to suicide or depression following stressful life events [56, 57]. The effect of stress may be important in these association studies as would be suggested by recent evidence demonstrating an altered amygdala response to stress depending on variable 5-HT signaling resulting from 5-HTTLPR genotype [54, 58]. Conversely, the homozygous l/l genotype is associated with a predisposition to OCD [59] and increased intensity of hallucinations in individuals with schizophrenia [60].
Although one can correlate the function of a specific VNTR with endogenous gene expression, more often the ability of a VNTR to modulate gene expression is analysed in a recombinant DNA construct termed a reporter gene construct (Fig. 4). In such analysis the isolated VNTR is tested for its ability to modulate the activity of a minimal promoter in supporting expression of a heterologous gene such as luciferase which can be easily quantified. The 5-HTTLPR VNTR have been shown to be functional transcriptional regulators that will affect gene expression in vitro. Differential expression supported by the 5-HTTLPR variants in the context of their own promoter has been demonstrated in the human placental choriocarcinoma cell line—JAr [45] and in lymphoblast cell lines [53], with the level of expression supported by the long allele being twice that supported by the short allele. In contrast, in a neuronal cell line, RN46A, Sakai et al., did not find any significant difference in activity between the l and s variants when isolated from the promoter and cloned into a luciferase reporter vector [61]. Conversely, Mortensen et al., did not find any difference in the activity of the l and s variants in JAr cells, but did see differential activity in RN46A cells [62]. In lymphoblast cell lines, the 5-HTT promoter fragments containing the long or short variant demonstrated differential stimulus-inducible response to stimuli such as forskolin or phorbol ester treatment, again with the long allele displaying proportionally greater responses [53]. In addition, having an s allele, as either s/l or s/s resulted in 40–70% less mRNA production than when genotype for the allele is l/l [53, 63, 64]. The same study also found greater uptake rates and an increased binding for 5-HT in lymphoblast cells with the l/l genotype compared with either s/l or s/s [53]. In contrast, a study employing allelic expression imbalance (AEI) techniques did not observe a correlation between 5-HTTLPR variants and 5-HTT mRNA levels in B-lymphocytes [65].
Schematic example of a reporter gene construct. The isolated VNTR sequence is cloned upstream of a minimal promoter such as the SV40 minimal promoter, driving a reporter gene such as firefly or renilla luciferase, or a marker gene, for example, green fluorescent protein (GFP). These constructs are transfected into cells by varying methods. The activity measured as reporter or marker gene read out is compared a construct with only the minimal promoter driving expression the reporter/marker gene (no VNTR control). This comparison provides a measure of the modulatory, and therefore functional, ability of the VNTR to regulate gene expression
Investigations into the functionality of the 5-HTT VNTR in vivo have also been undertaken. Heinz et al., examined in vivo, human 5-HTT binding in the midbrain using single photon emission computed tomography (SPECT) [66]. They found a difference in binding capacity between l/l and −s individuals which was consistent with the in vitro data of Lesch et al. [53], which states that individuals with the short allele show less binding [66]. Contrastingly, later SPECT studies did not show any correlation between genotype and functional regulation of the 5-HTT in the midbrain [67]. Supporting a lack of function in vivo, quantitative autoradiography in the prefrontal cortex of post-mortem brains of suicide victims did not show any correlation between 5-HTT binding and 5-HTTLPR genotype [68]. Further in vivo evidence against a role for this polymorphism regulating 5-HTT function exists from recent positron emission topography (PET) studies which fail to show any association between genotype and 5-HTT binding in the brain [69, 70]. To further compound matters, analysis of post-mortem brains found no difference in binding across the dorsal raphe, median raphe or substantia nigra in homozygous samples (of either s/s or l/l) but found lower levels of binding in heterozygous individuals [40]. As the authors of this study point out, the differences in binding between this study and that of Lesch et al. [53] may be due to the relatively small sample size employed and the same may be the case in other in vivo studies. However in the same study, a correlation between decreased mRNA expression and individuals with at least a single copy of the s allele was reported, which is in agreement with Lesch et al. [53]. More recently, a study investigating the affects of this polymorphism in cognitive error processing in the prefrontal cortex has demonstrated that the 5-HTTLPR short variant is associated with enhanced responsiveness of the brain [71].
Transgenic animal studies have also hinted at the possible functionality of the 5-HTTLPR by over-expressing human 5-HTT in mice to mimic the higher mRNA expression levels, increased uptake and binding observed with the l/l genotype. The l/l genotype is associated with non-anxious controls in healthy human volunteers, and in the transgenic mice results in a low-anxiety phenotype [72].
STIN2 VNTR association studies and functionality
A second widely studied VNTR found in the non-coding region of the 5-HTT gene is located in intron 2 and comprises 9, 10 or 12 copies of a 16–17 bp repeat termed, STin2.9, STin2.10 and STin2.12 respectively [5, 73] (Fig. 5). Individuals can be hetero- or homozygous for the STin2 VNTR giving rise to six possible genotypes 9/9, 10/10, 12/12, 9/10, 9/12 and 10/12. Association studies have also linked this polymorphism with a number of cognitive and affective disorders including; unipolar depression and bipolar disorder to which the STin2.9 allele seems to be a predisposing factor [5, 6]; OCD in which the STin2.12 was seen at a higher frequency in affected individuals compared to healthy controls [4, 74]; migraine, where an increased incidence of the STin2.12 homozygous genotype was observed in migraine without aura, and an increased incidence of the STin2.9 allele was observed in migraine with aura, when compared to non-migraine controls [9]; anxiety which correlated with the presence of at least one copy of STin2.9 [3]; the STin2.12 allele has been associated with varying aspects of schizophrenia [2, 63, 75]; individuals homozygous for the 10 allele of STin2 have recently been shown to be predisposed to suicide [76, 77].
Sequence and repeat alignment of the 5-HTT STin2 VNTR. Schematic representation of the human 5-HTT gene and position of the STin2 VNTR (top). The STin2 repeats exist as 9–12 copies of a 16–17 bp repeat. The repeats are not perfect repeating units. Variation in repeat sequence is represented by a Greek symbol. As can be seen the 10 repeat lacks a γ and η repeats present in the 12 copy while the nine lacks a further ϕ copy
While association of the STin2 VNTR with affective disorders has been noted, the results of meta-analysis of a number of published studies cast doubt on the solidity of these findings [78–80]. For every study reporting a positive association between a genotype and a disorder there are contradictory papers demonstrating a lack of association.
Nevertheless, the STin2 VNTR can display functionality both in vitro [12, 14, 15] and in vivo [13]. In cell lines, we demonstrated that the STin2 VNTR can support differential levels of reporter gene expression, acting as enhancers of transcription, with STin2.12 showing the most robust activity [12]. In addition, we verified that copy number of the repeat unit is not the only factor functioning in the observed regulation of transcription, but that primary sequence variation between the individual repeating units can also affect enhancer activity [14]. We have also demonstrated that in vivo, in transient transgenic mice expressing the human 5-HTT STin2 VNTRs driving a β-galactosidase marker gene, that the STin2 VNTRs may have an important regulatory role in development of the serotonergic system with genotype affecting 5-HTT expression in a temporal and tissue-specific manner [13]. This developmental role for the VNTRs may be particularly significant considering recent implications that transient alterations in 5-HT homeostasis in embryogenesis modify the fine wiring of brain connections leading to permanent changes in adult behaviour [81, 82]. In light of these findings it is possible that the effects of the 5-HTT polymorphisms are more pronounced during embryogenesis and development, a view also expressed more recently by Parsey et al. [70].
5-HTT variants and anti-depressant efficacy
The 5-HTT polymorphic genotypes have been functionally associated with differential efficacy of SSRI treatment, with individuals varying in their response. For different global populations the same response to treatment with SSRI is seen across different genotypes, suggesting that combinatorial interactions of polymorphisms unique to each population may be producing distinct effects e.g., Caucasians with 5-HTTLPR s/s genotype were found to display a poor response to SSRI, but in contrast, Asian individuals with the 5-HTTLPR l/l genotype responded poorly to treatment (for detailed review and meta-analysis see [50, 83–85]). As is the case with the 5-HTTLPR polymorphisms, individuals with different STin2 VNTRs may respond differently to treatments for depression such as TCA or SSRI. A systematic review of the current literature indicated that while Caucasians with the 5-HTTLPR s/s genotype showed the least favourable response to SSRI, in Asian populations those with the STin2 10/12 genotype showed the least favourable response [50].
Dopamine (DA) and the DAT1 transporter (DAT1/SLC6A3)
The dopaminergic system is well known to be physiologically involved in motivation and reward (reviewed in [86]), and memory and learning (reviewed in [87]). In addition an involvement of DA has been strongly implicated in a number of addictive behaviours such as drug-taking (reviewed in [88]), smoking (reviewed in [89]), overeating [90] and gambling [91]. A number of studies utilizing DA agonists and antagonists have demonstrated that DA is involved in long term potentiation (LTP); the process by which behaviour is learnt and memories are stored [92–95]. In the CNS, DA acts to fine tune neuronal firing by altering signal-to-noise ratios to produce more focused activity [96, 97]. In the CNS, DA is localised at high synaptic concentrations in the striatum and lower concentrations in the cortex, and is involved in modulating thalmo-cortical signalling pathways [98, 99]. Unlike 5-HT, which mediates its effect by acting solely on receptors located synaptically, DA also activates receptors which are found in extrasynaptic locations by DA spillover [87]. Elevation of DA concentration and the duration of DA exposure in the synaptic and extrasynaptic space are major determinants of DA receptor activation. In common with the 5-HT system, DA availability is regulated by uptake via a member of the sodium and chloride dependant NET family of proteins, the human DA transporter, DAT1 [100]. DA availability is also regulated via methylation by catechol-O-methyltransferase (COMT), which itself contains a VNTR but this is beyond the scope of this review (for review [101]). The DAT1 is a target for illicit drugs such as cocaine and methamphetamine, but also for therapeutic agents such as those used to treat Attention Deficit Hyperactivity Disorder (ADHD) e.g., Ritalin [102, 103].
3′ UTR VNTR: association and functionality
The human DAT1 gene consists of 15 exons and spans 60 kb on chromosome 5p15.3 [104] (Fig. 6). The human DAT1 protein is 620 amino acids in length and has 12 membrane spanning domains and shares over 90% homology with rodent DAT [103, 105]. Although there are SNPs in the coding sequence of DAT1 they do not affect the protein structure of the transporter [106]. Like 5-HTT, the DAT1 is a member of the Na/Cl-dependant NET family of transporters [107] and like 5-HTT, DAT1 also has a number of VNTRs in non-coding sequences. The first of these to be identified and by far the most extensively studied is located in the 3′ untranslated region (3′UTR) of the DAT1 gene [104, 108]. This VNTR, termed the 3′UTR VNTR, comprises 3–11 repeats of a 40 bp unit, with the 9 and 10 repeats being the most common forms. Individuals can be heterozygous or homozygous for the VNTR. As with all VNTRs the distribution of these genotypes varies in the population with the 10 allele appearing more frequently [109]. However, again as with the polymorphisms described earlier in this review differences in allelic frequency are observed between global populations or different ethnic groups [109].
Sequence and repeat alignment of the DAT 3′UTR VNTR. Schematic representation of the human DAT1 gene and position of the 3′UTR VNTR (top). The common 3′UTR repeats exist as 9 and 10 copies of a 40 bp repeat. The repeats are not perfect repeating units. Bold type lettering in the sequence represents degeneration between repeats and variation is denoted by defining each repeat with a Greek symbol. The nine repeat allele VNTR lacks the γ repeat seen in the 10 repeat allele, and in this example a SNP can be seen in the ε repeat of the 10 allele which produces the ι repeat in the nine allele
Association studies have linked the 3′UTR VNTR not only with a number of disorders or affective behaviours but also with differences in normal physiological functioning between individuals e.g., focused neuronal activity [110]. Bertolino et al., demonstrated that individuals with the homozygous 10 allele (10/10) perform with a more focused response to memory related tasks when compared to heterozygous individuals (9/10) [110]. One of the most extensively studied associations for the 3′UTR VNTR is with ADHD. Similarly to the association of 5-HTT polymorphisms with depression, the data is conflicting. For example, a recent study has demonstrated an association of the 9/10 genotype with more severe symptoms of ADHD when compared to the 10/10 genotype [111], whilst others pinpoint association with susceptibility to ADHD with the presence of a 10 repeat allele in family transmission studies [112–117]. In contrast, some studies found no association between DAT1 polymorphisms and ADHD using either family based [118, 119] or population based studies [120, 121]. However, a role for the DAT1 transporter in ADHD is strongly supported by the facts that DAT1 knockout mice are extremely hyperactive [122] and that current therapies for ADHD include drugs that act on the transporter e.g., methylphenidate (Ritalin). Strong conclusive evidence that polymorphisms in DAT1 are associated with ADHD has not yet been found and once again conflicting evidence may have arisen due to differences in the types of association being made e.g., familial versus population studies, ethnic variation, different test parameters, etc. However, it is possible that when acting in concert with other polymorphisms, via an epistatic interaction, the 3′UTR VNTR of DAT1 may display stronger associations with disorders. For example, Carrasco et al., have very recently demonstrated, in a Chilean population, individuals that are heterozygous for a seven repeat VNTR in the dopamine 4 receptor (DRD4) and homozygous for the DAT1 3′UTR VNTR 10 allele polymorphism are likely to suffer from ADHD, but in the presence of only one of these specific polymorphisms the same association can not be made [123].
In addition the DAT1 gene contains several other VNTRs that meet our criteria for acting as potential regulatory domains (personal observations and preliminary data not shown). They could therefore, act synergistically to modulate transporter expression and studies should consider multiple polymorphisms which are both intra- and intergeneic if they target the same neural transmission pathway.
The 3′UTR VNTR has also been linked with a number of other disorders or addictive behaviours. The presence of a nine allele has been linked with a reduced risk for addiction to smoking [124–126] which may be due to decreased DA release in individuals with this genotype [127]. The 3′UTR has also been associated with susceptibility to Parkinson’s disease although the evidence is highly variable. The rare 11 allele has been associated with Parkinson’s in a limited number of studies using different populations [128–130] but not in others [131]. A further study examined the nine allele in Parkinson’s and found an age-related association: those over the age of 60 with a 9-repeat show greater incidence than those younger than 60 with a 9-repeat [132]. In contrast, a further study found no association between DAT1 3UTR polymorphisms and Parkinson’s disease in a Chinese population [133].
Most studies disprove an association of the 3′UTR VNTR and alcoholism [134, 135] although genotype may play a part in severity of withdrawal symptoms [135]. However, one study implies that an A/G SNP in the 10 allele may be associated with vulnerability to alcoholism [136]. To a lesser extent other associations with this VNTR have been examined but most find no correlation, these include; body weight, body mass index or obesity [137], Tourette’s [138], personality traits (novelty seeking, harm avoidance, reward dependence and persistence [139]), cocaine dependence [140]; and schizophrenia [141].
A number of functional studies on the common DAT1 3′UTR VNTR have been conducted in vitro and in vivo with varying results. Transient transfections of recombinant reporter and marker gene plasmids containing a VNTR show conflicting results. However, when we examined the isolated nine allele 3′UTR VNTR linked to a green fluorescent protein marker gene, it could act as an enhancer of transcription in dopaminergic neurons within a murine midbrain slice and in a murine dopaminergic cell line, SN4741 [142]. Furthermore, higher levels of reporter gene expression have been demonstrated for the 3′UTR nine allele in human neuroblastoma SK-N-SH cells [143] and in HEK293 cells when transfected as part of a larger 3′UTR fragment of DNA [144]. In contrast, in a non-human primate cell line, cos-7, a fragment of the 3′UTR containing the 10 repeat was demonstrated to support higher transcription levels than that of the 9 [145]. Furthermore, another study found no difference in transcriptional activity supported by either the 9 or the 10 allele when plasmids were transiently transfected into the human neuroblastoma cell line, SHSY-5Y, or HEK293 cells [146]. Nevertheless, QPCR on cerebellum neurons and lymphocytes demonstrates that the 10 allele is associated with higher levels of endogenous DAT mRNA [147]. In addition, in HEK293 cells with targeted stable integration of recombinant DNA containing the DAT1 coding sequence with different polymorphisms, the 10 allele has been associated with increased DAT1 protein density [148]. In our opinion, the contrasting reporter gene activity demonstrated for this and other VNTRs when tested is that these domains demonstrate tissue specificity as exemplified by our transgenic data for the 5HTT intron 2 VNTRs [13] and that variation in tissue culture reflects this property.
In vivo SPECT studies have demonstrated that having a 9/10 genotype results in lower levels of DAT1 expression in the striatal putamen when compared to individuals homozygous for the 10 allele [149]. In complete contrast, two other studies report higher levels of striatal DAT1 availability in individuals with at least one 9 allele [150, 151]. Three more in vivo studies found no association between 3′UTR polymorphism genotype and DAT1 density [152] or protein availability [153] and function [154].
Intron 8 VNTR: association and functionality
We have identified a 5–6 copy 30 bp VNTR located in intron 8 (Int8) of the human DAT1 gene and were able to demonstrate that it had transcriptional properties [16] (Fig. 7). This VNTR is termed either DAT1Int8.2 (five copies) or DAT1Int8.3 (six copies). As this VNTR has only recently been characterised much less work has focused on association studies with the DATInt8 VNTRs than for instance with that of the 3′UTR. We have described an association between the DATInt8.3 VNTR and predisposition towards cocaine addiction in a Brazilian population [16]. Recently, Brookes et al., have identified an association between the DATInt8 VNTRs and predisposition to ADHD in both English and Taiwanese populations [155]. O’Gara et al., have very recently demonstrated an association between the DATInt8 polymorphism and the ability to quit smoking, but only in the early stages of the attempt [156].
Sequence and repeat alignment of the DAT Int8 VNTR. Schematic representation of the human DAT1 gene and position of the Int8 VNTR (top). The common repeats exist as five (termed two allele) and six (termed three allele) copies of a 25–30 bp repeat. The repeats are not perfect repeating units. Bold type lettering in the sequence represents degeneration between repeats and variation is denoted by defining each repeat with a Greek symbol. The two repeat allele lacks the χ repeat seen in the three repeat allele
Transient transfection of the dopaminergic murine cell line, SN4741, with renillin luciferase reporter plasmids containing an isolated VNTR cloned into an intronic position, demonstrated the ability of the Int8.2 VNTR to support higher basal levels of gene expression than the Int8.3 VNTR [16]. In addition, these Int8 VNTR demonstrated differential responses to stimuli. Namely, the Int8.3 allele displayed more pronounced sensitivity to stimuli such as cocaine than the Int8.2 allele [16].
DAT1 variants and therapeutics
DAT1 is a site of action for illicit drugs but also for therapeutic drugs. The DAT1Int8 VNTRs have been associated with differential response and susceptibility to illicit drugs such as cocaine and nicotine as outlined above. In addition there have been a number of reports indicating an association between the response of individuals to treatment for ADHD (namely methylphenidate), and genotype at the DAT1 3′UTR VNTR. Methylphenidate is a DAT1 re-uptake inhibitor that acts to prolong the duration of action of DA at it’s receptors. In children with ADHD undergoing treatment with methylphenidate homozygosity for the 3′UTR 10 repeat was associated with poorer performance in tests and a different pattern of EEG response when compared to children with a copy of the nine repeat [157]. Furthermore, in a sample of African-American children undergoing treatment for ADHD, Winsburg and Comings correlated homozygosity of the 10 repeat allele with non-responsiveness to the drug [158]. In contrast, others found that poor response to methylphenidate was evident in those individuals homozygous for the nine repeat allele [159, 160]. Discrepancies between these studies are likely to arise from differences in test parameters; however they do demonstrate that the allelic variation at the 3′UTR can be associated with differential responses to treatment. These clinical data also support our theory that regulatory polymorphic domains must be considered together in assessing the response of a particular genotype. In future studies, attempting to correlate clinical data with genotype, we believe attempts must be made to determine which allele the polymorphisms are located on to fully elucidate their summative action in modulating gene expression in specific tissues or in response to a particular challenge.
Transcription regulation by polymorphisms based on primary sequence similarities
Our analysis of the VNTRs within the transporters, 5HTT and DAT1, has focused on comparison of the functional characteristics of different copy number VNTRs [12–15]. However variation in the sequence of individual repeat elements might be as relevant in furthering our understanding of VNTR function. One or more base changes in the sequence at the binding site for a transcription factor, as found within the VNTR elements, could alter the affinity or specificity of a specific transcription factor binding to that site [14]. The synergistic or additive action of these individual elements in the whole VNTR domain may result in differential regulatory properties. Consistent with this we have analysed the transcriptional characteristics of individual repeat elements of the 5-HTT VNTR and demonstrated that they act as functionally distinct regulatory elements [12]. Whilst copy number variation can have effects at the level of gross human phenotype [5, 7, 161] and in in vitro and in vivo model systems, our data indicates that there is an additional layer of transcriptional complexity based on the primary sequence of the VNTR [14]. For example, as stated previously there is debate as to the correlation of VNTR copy number with predisposition to affective disorders, perhaps reanalysing this data taking into account the primary sequence of the VNTR could resolve the conflicting literature and resolve the debate as to the clinical significance of these VNTR domains.
We have demonstrated that the intron 2 VNTR domain of the 5-HTT gene is bound and regulated by the transcription factor Y box binding protein 1 (YB1). This factor was identified by a yeast one hybrid screen, followed by demonstration of it’s ability to transactivate in a 5-HTT STIn2 VNTR construct in a cell line model [15]. YB1 has been implicated as an important protein during development. Interestingly, a transcription factor termed CTCF can also regulate the function of YB1 directly [15]. We have demonstrated that YB1 will bind specifically to the 5-HTT STin2 VNTR elements and this binding can be antagonised by CTCF [15]. We have also recently demonstrated that CTCF can bind the 5-HTT STin2 VNTRs [22]. This action of CTCF is of interest because this protein in addition to direct transcriptional properties, has epigenetic functions such as controlling imprinting and has recently been associated with diseases ranging from Alzheimer’s to cancer [162–165].
In general, transcription of a gene will therefore be determined by the combinatorial action of multiple positive and negative promoter domains that specify the tissue-specific and stimulus-inducible expression of a gene [166]. We postulate that VNTRs could have a direct effect on the expression of a gene in different cell types and in response to various physiological or pharmacological stimuli. This may correlate with the aetiology or progression of a disorder. We therefore predict that studies to better define the function of the VNTRs to modulate gene expression will not only be of importance for normal and abnormal behaviour but be of general importance for understanding gene expression, as related VNTRs are present in many other genes and are often proposed to correlate with susceptibility to various disorders.
Summary
Contradictory reports in the literature linking or associating specific polymorphisms or genes with specific disorders may arise from inter-experimental variation, mixed populations, and inappropriate comparisons between subjects. As most association studies demonstrate a lack of power the conclusion of most studies and of recent meta-analysis is that larger sample sizes and samples that are controlled for the variations above are needed to provide more accurate associations. In addition, variation in function as measured by ability to modulate gene expression (endogenous or reporter gene) can also arise due to differences between experimental paradigms e.g., cell type and growth conditions used, length of “VNTR”; some groups include flanking sequences which may have binding sites for additional transacting factors. The use of different transfection systems, a multitude of different cell types and protocol differences such as time spent in culture may all affect reporter gene expression in transient analysis. One group even reports variation depending on the minimal promoter present in the reporter plasmid used for cloning, possibly due to interaction with a SNP in the repeat under investigation [144]. An interesting point raised by Brookes et al., concerning the contradictory reports surrounding the DAT1 3′UTR VNTR function in ADHD, is that this may be due to the 3′UTR VNTR acting as a marker for an associated polymorphism rather than being involved directly in the aetiology [167]. This could also be true for other VNTR that appear to be weakly associated with a disorder but do not appear to display differential function, or show conflicting associations. What has until recently been overlooked, is the possibility that functional polymorphisms on the same gene (or haplotypes) or even on different genes (epistatic interaction) may increase (or decrease) the propensity towards the development of specific disorders or characteristics. In this case having two or three polymorphisms or haplotypes that might predispose to a specific condition is more of a risk factor for the development of that condition than having a single polymorphism. However, one must take into account that some polymorphisms show a stronger correlation with a disorder than others. A number of haplotype association studies have already been undertaken [64, 115, 132, 155, 167]. In addition, Hranilovic et al., recently examined combinatorial effects of the two VNTR 5-HTT polymorphisms, we have discussed, on mRNA expression by quantitative PCR on lymphoblast cells from schizophrenic patients [168]. They determined that the polymorphisms could be banded, based on expression, into high expressing (l/l and 12/12) and low expressing (l/s, s/s and 10/10 10/12). They found a dominant effect of the low expressing alleles on the high expressing alleles resulting in significantly decreased transporter expression [168]. We are currently investigating the ability of the 5-HTT and DAT1 polymorphisms reviewed here to act in a combinatorial or synergistic manner in regulating transcriptional control of gene expression in stably expressing cell lines, and in response to stimuli such as lithium chloride or cocaine hydrochloride, in both cell lines and primary neuronal cells. The integration of variation in expression of a number of genes with related VNTRs (containing the consensus sequence for binding specific transcription factors) is consistent with recent attempts to take a more global analysis of polymorphic variation and correlation with disease reflected in whole genome analysis [169].
References
Brookes AJ, Prince JA (2005) Genetic association analysis: lessons from the study of Alzheimers disease. Mutat Res 573:152–159
Liu W, Gu N, Feng G, Li S, Bai S, Zhang J, Shen T, Xue H, Breen G, St Clair D, He L (1999) Tentative association of the serotonin transporter with schizophrenia and unipolar depression but not with bipolar disorder in Han Chinese. Pharmacogenetics 9:491–495
Evans J, Battersby S, Ogilvie AD, Smith CA, Harmar AJ, Nutt DJ, Goodwin GM (1997) Association of short alleles of a VNTR of the serotonin transporter gene with anxiety symptoms in patients presenting after deliberate self harm. Neuropharmacology 36:439–443
Baca-Garcia E, Vaquero-Lorenzo C, Diaz-Hernandez M, Rodriguez-Salgado B, Dolengevich-Segal H, Arrojo-Romero M, Botillo-Martin C, Ceverino A, Piqueras JF, Perez-Rodriguez MM, Saiz-Ruiz J (2007) Association between obsessive-compulsive disorder and a variable number of tandem repeats polymorphism in intron 2 of the serotonin transporter gene. Prog Neuropsychopharmacol Biol Psychiatry 31(2):416–420
Ogilvie AD, Battersby S, Bubb VJ, Fink G, Harmar AJ, Goodwim GM, Smith CA (1996) Polymorphism in serotonin transporter gene associated with susceptibility to major depression. Lancet 347:731–733
Battersby S, Ogilvie AD, Smith CA, Blackwood DH, Muir WJ, Quinn JP, Fink G, Goodwin GM, Harmar AJ (1996) Structure of a variable number tandem repeat of the serotonin transporter gene and association with affective disorder. Psychiatr Genet 6:177–181
Collier DA, Arranz MJ, Sham P, Battersby S, Vallada H, Gill P, Aitchison KJ, Sodhi M, Li T, Roberts GW, Smith B, Morton J, Murray RM, Smith D, Kirov G (1996) The serotonin transporter is a potential susceptibility factor for bipolar affective disorder. Neuroreport 7:1675–1679
Bellivier F, Leroux M, Henry C, Rayah F, Rouillon F, Laplanche JL, Leboyer M (2002) Serotonin transporter gene polymorphism influences age at onset in patients with bipolar affective disorder. Neurosci Lett 334:17–20
Ogilvie AD, Russell MB, Dhall P, Battersby S, Ulrich V, Smith CA, Goodwin GM, Harmar AJ, Olesen J (1998) Altered allelic distributions of the serotonin transporter gene in migraine without aura and migraine with aura. Cephalalgia 18:23–26
Yilmaz M, Erdal ME, Herken H, Cataloluk O, Barlas O, Bayazit YA (2001) Significance of serotonin transporter gene polymorphism in migraine. J Neurol Sci 186:27–30
Skipper L, Liu JJ, Tan EK (2006) Polymorphisms in candidate genes: implications for the current treatment of Parkinson’s disease. Expert Opin Pharmacother 7:849–855
Fiskerstrand CE, Lovejoy EA, Quinn JP (1999) An intronic polymorphic domain often associated with susceptibility to affective disorders has allele dependent differential enhancer activity in embryonic stem cells. FEBS Lett 458:171–174
MacKenzie A, Quinn J (1999) A serotonin transporter gene intron 2 polymorphic region, correlated with affective disorders, has allele-dependent differential enhancer-like properties in the mouse embryo. Proc Natl Acad Sci USA 96:15251–15255
Lovejoy EA, Scott AC, Fiskerstrand CE, Bubb VJ, Quinn JP (2003) The serotonin transporter intronic VNTR enhancer correlated with a predisposition to affective disorders has distinct regulatory elements within the domain based on the primary DNA sequence of the repeat unit. Eur J Neurosci 17:417–420
Klenova E, Scott AC, Roberts J, Shamsuddin S, Lovejoy EA, Bergmann S, Bubb VJ, Royer HD, Quinn JP (2004) YB-1 and CTCF differentially regulate the 5-HTT polymorphic intron 2 enhancer which predisposes to a variety of neurological disorders. J Neurosci 24:5966–5973
Guindalini C, Howard M, Haddley K, Laranjeira R, Collier D, Ammar N, Craig I, O’Gara C, Bubb VJ, Greenwood T, Kelsoe J, Asherson P, Murray RM, Castelo A, Quinn JP, Vallada H, Breen G (2006) A dopamine transporter gene functional variant associated with cocaine abuse in a Brazilian sample. Proc Natl Acad Sci USA 103:4552–4557
Quinn JP, Holbrook N, Levens D (1987) Binding of a cellular protein to the gibbon ape leukemia virus enhancer. Mol Cell Biol 7:2735–2744
Quinn JP, McGregor RA, Fiskerstrand CE, Davey C, Allan J, Dalziel RG (1998) Identification of a novel multifunctional structural domain in the herpes simplex virus type 1 genome: implications for virus latency. J Gen Virol 79(Pt 10):2529–2532
Quinn JP, Takimoto M, Iadarola M, Holbrook N, Levens D (1989) Distinct factors bind the AP-1 consensus sites in gibbon ape leukemia virus and simian virus 40 enhancers. J Virol 63:1737–1742
Lesch KP, Meyer J, Glatz K, Flugge G, Hinney A, Hebebrand J, Klauck SM, Poustka A, Poustka F, Bengel D, Mossner R, Riederer P, Heils A (1997) The 5-HT transporter gene-linked polymorphic region (5-HTTLPR) in evolutionary perspective: alternative biallelic variation in rhesus monkeys. Rapid communication. J Neural Transm 104:1259–1266
Soeby K, Larsen SA, Olsen L, Rasmussen HB, Werge T (2005) Serotonin transporter: evolution and impact of polymorphic transcriptional regulation. Am J Med Genet B Neuropsychiatr Genet 136:53–57
Roberts JC, Scott AM, Howard MR, Breen G, Bubb VJ, Klenova E, Quinn JP (2007) Differential regulation of the serotonin transporter gene by lithium is mediated by transcription factors, CCTC binding protein and Y-box binding protein 1, through the polymorphic intron 2 variable number tandem repeat. J Neurosci 27(11):2793–2801
Zis AP, Goodwin FK (1979) Novel antidepressants and the biogenic amine hypothesis of depression. The case for iprindole and mianserin. Arch Gen Psychiatry 36:1097–1107
Noach EL (1994) Appetite regulation by serotoninergic mechanisms and effects of d-fenfluramine. Neth J Med 45:123–133
Garattini S (1995) Biological actions of drugs affecting serotonin and eating. Obes Res 3(Suppl 4):463S–470S
Brunelli M, Garcia-Gil M, Mozzachiodi R, Scuri R, Zaccardi ML (1997) Neurobiological principles of learning and memory. Arch Ital Biol 135:15–36
Lesch KP, Merschdorf U (2000) Impulsivity, aggression, and serotonin: a molecular psychobiological perspective. Behav Sci Law 18:581–604
Lesch KP, Zeng Y, Reif A, Gutknecht L (2003) Anxiety-related traits in mice with modified genes of the serotonergic pathway. Eur J Pharmacol 480:185–204
Ressler KJ, Nemeroff CB (2000) Role of serotonergic and noradrenergic systems in the pathophysiology of depression and anxiety disorders. Depress Anxiety 12(Suppl 1):2–19
McLaughlin DP, Little KY, Lopez JF, Watson SJ (1996) Expression of serotonin transporter mRNA in human brainstem raphe nuclei. Neuropsychopharmacology 15:523–529
Zhou FC, Sari Y, Zhang JK (2000) Expression of serotonin transporter protein in developing rat brain. Brain Res Dev Brain Res 119:33–45
Blakely RD, Berson HE, Fremeau RT Jr, Caron MG, Peek MM, Prince HK, Bradley CC (1991) Cloning and expression of a functional serotonin transporter from rat brain. Nature 354:66–70
Rudnick G, Clark J (1993) From synapse to vesicle: the reuptake and storage of biogenic amine neurotransmitters. Biochim Biophys Acta 1144:249–263
Owens MJ, Nemeroff CB (1994) Role of serotonin in the pathophysiology of depression: focus on the serotonin transporter. Clin Chem 40:288–295
Owens MJ, Nemeroff CB (1998) The serotonin transporter and depression. Depress Anxiety 8(Suppl 1):5–12
Roman DL, Walline CC, Rodriguez GJ, Barker EL (2003) Interactions of antidepressants with the serotonin transporter: a contemporary molecular analysis. Eur J Pharmacol 479:53–63
McCann UD, Szabo Z, Seckin E, Rosenblatt P, Mathews WB, Ravert HT, Dannals RF, Ricaurte GA (2005) Quantitative PET studies of the serotonin transporter in MDMA users and controls using [11C]McN5652 and [11C]DASB. Neuropsychopharmacology 30:1741–1750
Mlinar B, Corradetti R (2003) Endogenous 5-HT, released by MDMA through serotonin transporter- and secretory vesicle-dependent mechanisms, reduces hippocampal excitatory synaptic transmission by preferential activation of 5-HT1B receptors located on CA1 pyramidal neurons. Eur J Neurosci 18:1559–1571
Ramamoorthy S, Bauman AL, Moore KR, Han H, Yang-Feng T, Chang AS, Ganapathy V, Blakely RD (1993) Antidepressant- and cocaine-sensitive human serotonin transporter: molecular cloning, expression, and chromosomal localization. Proc Natl Acad Sci USA 90:2542–2546
Little KY, McLaughlin DP, Zhang L, Livermore CS, Dalack GW, McFinton PR, DelProposto ZS, Hill E, Cassin BJ, Watson SJ, Cook EH (1998) Cocaine, ethanol, and genotype effects on human midbrain serotonin transporter binding sites and mRNA levels. Am J Psychiatry 155:207–213
Sandhu SK, Ross LS, Gill SS (2002) A cocaine insensitive chimeric insect serotonin transporter reveals domains critical for cocaine interaction. Eur J Biochem 269:3934–3944
Lesch KP, Wolozin BL, Estler HC, Murphy DL, Riederer P (1993) Isolation of a cDNA encoding the human brain serotonin transporter. J Neural Transm Gen Sect 91:67–72
Lesch KP, Wolozin BL, Murphy DL, Reiderer P (1993) Primary structure of the human platelet serotonin uptake site: identity with the brain serotonin transporter. J Neurochem 60:2319–2322
Heils A, Teufel A, Petri S, Seemann M, Bengel D, Balling U, Riederer P, Lesch KP (1995) Functional promoter and polyadenylation site mapping of the human serotonin (5-HT) transporter gene. J Neural Transm Gen Sect 102:247–254
Heils A, Teufel A, Petri S, Stober G, Riederer P, Bengel D, Lesch KP (1996) Allelic variation of human serotonin transporter gene expression. J Neurochem 66:2621–2624
Nakamura M, Ueno S, Sano A, Tanabe H (2000) The human serotonin transporter gene linked polymorphism (5-HTTLPR) shows ten novel allelic variants. Mol Psychiatry 5:32–38
Hu X, Schrodi SJ, Ross DA, Cargill M (2004) Selecting tagging SNPs for association studies using power calculations from genotype data. Hum Hered 57:156–170
Kraft JB, Slager SL, McGrath PJ, Hamilton SP (2005) Sequence analysis of the serotonin transporter and associations with antidepressant response. Biol Psychiatry 58:374–381
Wendland JR, Martin BJ, Kruse MR, Lesch KP, Murphy DL (2006) Simultaneous genotyping of four functional loci of human SLC6A4, with a reappraisal of 5-HTTLPR and rs25531. Mol Psychiatry 11:224–226
Smits KM, Smits LJ, Schouten JS, Stelma FF, Nelemans P, Prins MH (2004) Influence of SERTPR and STin2 in the serotonin transporter gene on the effect of selective serotonin reuptake inhibitors in depression: a systematic review. Mol Psychiatry 9:433–441
Cervilla JA, Rivera M, Molina E, Torres-Gonzalez F, Bellon JA, Moreno B, de Dios Luna J, Lorente JA, de Diego-Otero Y, King M, Nazareth I, Gutierrez B (2006) The 5-HTTLPR s/s genotype at the serotonin transporter gene (SLC6A4) increases the risk for depression in a large cohort of primary care attendees: the PREDICT-gene study. Am J Med Genet B Neuropsychiatr Genet 141:912–917
Cho HJ, Meira-Lima I, Cordeiro Q, Michelon L, Sham P, Vallada H, Collier DA (2005) Population-based and family-based studies on the serotonin transporter gene polymorphisms and bipolar disorder: a systematic review and meta-analysis. Mol Psychiatry 10:771–781
Lesch KP, Bengel D, Heils A, Sabol SZ, Greenberg BD, Petri S, Benjamin J, Muller CR, Hamer DH, Murphy DL (1996) Association of anxiety-related traits with a polymorphism in the serotonin transporter gene regulatory region. Science 274:1527–1531
Hariri AR, Mattay VS, Tessitore A, Kolachana B, Fera F, Goldman D, Egan MF, Weinberger DR (2002) Serotonin transporter genetic variation and the response of the human amygdala. Science 297:400–403
Gerra G, Garofano L, Pellegrini C, Bosari S, Zaimovic A, Moi G, Avanzini P, Talarico E, Gardini F, Donnini C (2005) Allelic association of a dopamine transporter gene polymorphism with antisocial behaviour in heroin-dependent patients. Addict Biol 10:275–281
Caspi A, Sugden K, Moffitt TE, Taylor A, Craig IW, Harrington H, McClay J, Mill J, Martin J, Braithwaite A, Poulton R (2003) Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science 301:386–389
Zalsman G, Huang YY, Oquendo MA, Burke AK, Hu XZ, Brent DA, Ellis SP, Goldman D, Mann JJ (2006) Association of a triallelic serotonin transporter gene promoter region (5-HTTLPR) polymorphism with stressful life events and severity of depression. Am J Psychiatry 163:1588–1593
Hariri AR, Brown SM (2006) Serotonin. Am J Psychiatry 163:12
Bengel D, Greenberg BD, Cora-Locatelli G, Altemus M, Heils A, Li Q, Murphy DL (1999) Association of the serotonin transporter promoter regulatory region polymorphism and obsessive-compulsive disorder. Mol Psychiatry 4:463–466
Malhotra AK, Goldman D, Mazzanti C, Clifton A, Breier A, Pickar D (1998) A functional serotonin transporter (5-HTT) polymorphism is associated with psychosis in neuroleptic-free schizophrenics. Mol Psychiatry 3:328–332
Sakai K, Nakamura M, Ueno S, Sano A, Sakai N, Shirai Y, Saito N (2002) The silencer activity of the novel human serotonin transporter linked polymorphic regions. Neurosci Lett 327:13–16
Mortensen OV, Thomassen M, Larsen MB, Whittemore SR, Wiborg O (1999) Functional analysis of a novel human serotonin transporter gene promoter in immortalized raphe cells. Brain Res Mol Brain Res 68:141–148
Hranilovic D, Schwab SG, Jernej B, Knapp M, Lerer B, Albus M, Rietschel M, Kanyas K, Borrmann M, Lichtermann D, Maier W, Wildenauer DB (2000) Serotonin transporter gene and schizophrenia: evidence for association/linkage disequilibrium in families with affected siblings. Mol Psychiatry 5:91–95
Bradley SL, Dodelzon K, Sandhu HK, Philibert RA (2005) Relationship of serotonin transporter gene polymorphisms and haplotypes to mRNA transcription. Am J Med Genet B Neuropsychiatr Genet 136:58–61
Lim JE, Papp A, Pinsonneault J, Sadee W, Saffen D (2006) Allelic expression of serotonin transporter (SERT) mRNA in human pons: lack of correlation with the polymorphism SERTLPR. Mol Psychiatry 11:649–662
Heinz A, Goldman D (2000) Genotype effects on neurodegeneration and neuroadaptation in monoaminergic neurotransmitter systems. Neurochem Int 37:425–432
Willeit M, Stastny J, Pirker W, Praschak-Rieder N, Neumeister A, Asenbaum S, Tauscher J, Fuchs K, Sieghart W, Hornik K, Aschauer HN, Brucke T, Kasper S (2001) No evidence for in vivo regulation of midbrain serotonin transporter availability by serotonin transporter promoter gene polymorphism. Biol Psychiatry 50:8–12
Mann JJ, Huang YY, Underwood MD, Kassir SA, Oppenheim S, Kelly TM, Dwork AJ, Arango V (2000) A serotonin transporter gene promoter polymorphism (5-HTTLPR) and prefrontal cortical binding in major depression and suicide. Arch Gen Psychiatry 57:729–738
Shioe K, Ichimiya T, Suhara T, Takano A, Sudo Y, Yasuno F, Hirano M, Shinohara M, Kagami M, Okubo Y, Nankai M, Kanba S (2003) No association between genotype of the promoter region of serotonin transporter gene and serotonin transporter binding in human brain measured by PET. Synapse 48:184–188
Parsey RV, Hastings RS, Oquendo MA, Hu X, Goldman D, Huang YY, Simpson N, Arcement J, Huang Y, Ogden RT, Van Heertum RL, Arango V, Mann JJ (2006) Effect of a triallelic functional polymorphism of the serotonin-transporter-linked promoter region on expression of serotonin transporter in the human brain. Am J Psychiatry 163:48–51
Fallgatter AJ, Herrmann MJ, Roemmler J, Ehlis AC, Wagener A, Heidrich A, Ortega G, Zeng Y, Lesch KP (2004) Allelic variation of serotonin transporter function modulates the brain electrical response for error processing. Neuropsychopharmacology 29:1506–1511
Jennings KA, Loder MK, Sheward WJ, Pei Q, Deacon RM, Benson MA, Olverman HJ, Hastie ND, Harmar AJ, Shen S, Sharp T (2006) Increased expression of the 5-HT transporter confers a low-anxiety phenotype linked to decreased 5-HT transmission. J Neurosci 26:8955–8964
Lesch KP, Balling U, Gross J, Strauss K, Wolozin BL, Murphy DL, Riederer P (1994) Organization of the human serotonin transporter gene. J Neural Transm Gen Sect 95:157–162
Ohara K, Suzuki Y, Ochiai M, Tsukamoto T, Tani K, Ohara K (1999) A variable-number-tandem-repeat of the serotonin transporter gene and anxiety disorders. Prog Neuropsychopharmacol Biol Psychiatry 23:55–65
Kaiser R, Tremblay PB, Schmider J, Henneken M, Dettling M, Muller-Oerlinghausen B, Uebelhack R, Roots I, Brockmoller J (2001) Serotonin transporter polymorphisms: no association with response to antipsychotic treatment, but associations with the schizoparanoid and residual subtypes of schizophrenia. Mol Psychiatry 6:179–185
Hranilovic D, Stefulj J, Furac I, Kubat M, Balija M, Jernej B (2003) Serotonin transporter gene promoter (5-HTTLPR) and intron 2 (VNTR) polymorphisms in Croatian suicide victims. Biol Psychiatry 54:884–889
Jernej B, Stefulj J, Hranilovic D, Balija M, Skavic J, Kubat M (2004) Intronic polymorphism of tryptophan hydroxylase and serotonin transporter: indication for combined effect in predisposition to suicide. J Neural Transm 111:733–738
Furlong RA, Ho L, Walsh C, Rubinsztein JS, Jain S, Paykel ES, Easton DF, Rubinsztein DC (1998) Analysis and meta-analysis of two serotonin transporter gene polymorphisms in bipolar and unipolar affective disorders. Am J Med Genet 81:58–63
Anguelova M, Benkelfat C, Turecki G (2003) A systematic review of association studies investigating genes coding for serotonin receptors and the serotonin transporter: II. Suicidal behavior. Mol Psychiatry 8:646–653
Lotrich FE, Pollock BG (2004) Meta-analysis of serotonin transporter polymorphisms and affective disorders. Psychiatr Genet 14:121–129
Whitaker-Azmitia PM (2001) Serotonin and brain development: role in human developmental diseases. Brain Res Bull 56:479–485
Gaspar P, Cases O, Maroteaux L (2003) The developmental role of serotonin: news from mouse molecular genetics. Nat Rev Neurosci 4(12):1002–1012
Malhotra AK, Murphy GM Jr, Kennedy JL (2004) Pharmacogenetics of psychotropic drug response. Am J Psychiatry 161:780–796
Lesch KP, Gutknecht L (2005) Pharmacogenetics of the serotonin transporter. Prog Neuropsychopharmacol Biol Psychiatry 29:1062–1073
Serretti A, Kato M, De Ronchi D, Kinoshita T (2006) Meta-analysis of serotonin transporter gene promoter polymorphism (5-HTTLPR) association with selective serotonin reuptake inhibitor efficacy in depressed patients. Mol Psychiatry
Franken IH, Booij J, van den Brink W (2005) The role of dopamine in human addiction: from reward to motivated attention. Eur J Pharmacol 526:199–206
Cragg SJ, Rice ME (2004) DAncing past the DAT at a DA synapse. Trends Neurosci 27:270–277
Haile CN, Kosten TR, Kosten TA (2007) Genetics of dopamine and its contribution to cocaine addiction. Behav Genet 37:119–145
Dani JA (2003) Roles of dopamine signaling in nicotine addiction. Mol Psychiatry 8:255–256
Nirenberg MJ, Waters C (2006) Compulsive eating and weight gain related to dopamine agonist use. Mov Disord 21:524–529
Grosset KA, Macphee G, Pal G, Stewart D, Watt A, Davie J, Grosset DG (2006) Problematic gambling on dopamine agonists: not such a rarity. Mov Disord 21:2206–2208
Otmakhova NA, Lisman JE (1996) D1/D5 dopamine receptor activation increases the magnitude of early long-term potentiation at CA1 hippocampal synapses. J Neurosci 16:7478–7486
Swanson-Park JL, Coussens CM, Mason-Parker SE, Raymond CR, Hargreaves EL, Dragunow M, Cohen AS, Abraham WC (1999) A double dissociation within the hippocampus of dopamine D1/D5 receptor and beta-adrenergic receptor contributions to the persistence of long-term potentiation. Neuroscience 92:485–497
Li S, Cullen WK, Anwyl R, Rowan MJ (2003) Dopamine-dependent facilitation of LTP induction in hippocampal CA1 by exposure to spatial novelty. Nat Neurosci 6:526–531
Swant J, Wagner JJ (2006) Dopamine transporter blockade increases LTP in the CA1 region of the rat hippocampus via activation of the D3 dopamine receptor. Learn Mem 13:161–167
Newman J, Grace AA (1999) Binding across time: the selective gating of frontal and hippocampal systems modulating working memory and attentional states. Conscious Cogn 8:196–212
Seamans JK, Yang CR (2004) The principal features and mechanisms of dopamine modulation in the prefrontal cortex. Prog Neurobiol 74:1–58
Sesack SR, Hawrylak VA, Guido MA, Levey AI (1998) Cellular and subcellular localization of the dopamine transporter in rat cortex. Adv Pharmacol 42:171–174
Lewis DA, Melchitzky DS, Sesack SR, Whitehead RE, Auh S, Sampson A (2001) Dopamine transporter immunoreactivity in monkey cerebral cortex: regional, laminar, and ultrastructural localization. J Comp Neurol 432:119–136
Giros B, Caron MG (1993) Molecular characterization of the dopamine transporter. Trends Pharmacol Sci 14:43–49
Diaz-Asper CM, Weinberger DR, Goldberg TE (2006) Catechol-O-methyltransferase polymorphisms and some implications for cognitive therapeutics. NeuroRx 3:97–105
Seeman P, Madras BK (1998) Anti-hyperactivity medication: methylphenidate and amphetamine. Mol Psychiatry 3:386–396
Bannon MJ, Michelhaugh SK, Wang J, Sacchetti P (2001) The human dopamine transporter gene: gene organization, transcriptional regulation, and potential involvement in neuropsychiatric disorders. Eur Neuropsychopharmacol 11:449–455
Vandenbergh DJ, Persico AM, Hawkins AL, Griffin CA, Li X, Jabs EW, Uhl GR (1992) Human dopamine transporter gene (DAT1) maps to chromosome 5p15.3 and displays a VNTR. Genomics 14:1104–1106
Bannon MJ, Poosch MS, Xia Y, Goebel DJ, Cassin B, Kapatos G (1992) Dopamine transporter mRNA content in human substantia nigra decreases precipitously with age. Proc Natl Acad Sci USA 89:7095–7099
Vandenbergh DJ, Thompson MD, Cook EH, Bendahhou E, Nguyen T, Krasowski MD, Zarrabian D, Comings D, Sellers EM, Tyndale RF, George SR, O’Dowd BF, Uhl GR (2000) Human dopamine transporter gene: coding region conservation among normal, Tourette’s disorder, alcohol dependence and attention-deficit hyperactivity disorder populations. Mol Psychiatry 5:283–292
Amara SG, Sonders MS, Zahniser NR, Povlock SL, Daniels GM (1998) Molecular physiology and regulation of catecholamine transporters. Adv Pharmacol 42:164–168
Sano A, Kondoh K, Kakimoto Y, Kondo I (1993) A 40-nucleotide repeat polymorphism in the human dopamine transporter gene. Hum Genet 91:405–406
Mitchell RJ, Howlett S, Earl L, White NG, McComb J, Schanfield MS, Briceno I, Papiha SS, Osipova L, Livshits G, Leonard WR, Crawford MH (2000) Distribution of the 3′ VNTR polymorphism in the human dopamine transporter gene in world populations. Hum Biol 72:295–304
Bertolino A, Blasi G, Latorre V, Rubino V, Rampino A, Sinibaldi L, Caforio G, Petruzzella V, Pizzuti A, Scarabino T, Nardini M, Weinberger DR, Dallapiccola B (2006) Additive effects of genetic variation in dopamine regulating genes on working memory cortical activity in human brain. J Neurosci 26:3918–3922
Barkley RA, Smith KM, Fischer M, Navia B (2006) An examination of the behavioral and neuropsychological correlates of three ADHD candidate gene polymorphisms (DRD4 7+, DBH TaqI A2, and DAT1 40 bp VNTR) in hyperactive and normal children followed to adulthood. Am J Med Genet B Neuropsychiatr Genet 141:487–498
Cook EH Jr, Stein MA, Krasowski MD, Cox NJ, Olkon DM, Kieffer JE, Leventhal BL (1995) Association of attention-deficit disorder and the dopamine transporter gene. Am J Hum Genet 56:993–998
Gill M, Daly G, Heron S, Hawi Z, Fitzgerald M (1997) Confirmation of association between attention deficit hyperactivity disorder and a dopamine transporter polymorphism. Mol Psychiatry 2:311–313
Daly G, Hawi Z, Fitzgerald M, Gill M (1999) Mapping susceptibility loci in attention deficit hyperactivity disorder: preferential transmission of parental alleles at DAT1, DBH and DRD5 to affected children. Mol Psychiatry 4:192–196
Barr CL, Xu C, Kroft J, Feng Y, Wigg K, Zai G, Tannock R, Schachar R, Malone M, Roberts W, Nothen MM, Grunhage F, Vandenbergh DJ, Uhl G, Sunohara G, King N, Kennedy JL (2001) Haplotype study of three polymorphisms at the dopamine transporter locus confirm linkage to attention-deficit/hyperactivity disorder. Biol Psychiatry 49:333–339
Curran S, Mill J, Tahir E, Kent L, Richards S, Gould A, Huckett L, Sharp J, Batten C, Fernando S, Ozbay F, Yazgan Y, Simonoff E, Thompson M, Taylor E, Asherson P (2001) Association study of a dopamine transporter polymorphism and attention deficit hyperactivity disorder in UK and Turkish samples. Mol Psychiatry 6:425–428
Chen CK, Chen SL, Mill J, Huang YS, Lin SK, Curran S, Purcell S, Sham P, Asherson P (2003) The dopamine transporter gene is associated with attention deficit hyperactivity disorder in a Taiwanese sample. Mol Psychiatry 8:393–396
Swanson JM, Flodman P, Kennedy J, Spence MA, Moyzis R, Schuck S, Murias M, Moriarity J, Barr C, Smith M, Posner M (2000) Dopamine genes and ADHD. Neurosci Biobehav Rev 24:21–25
Holmes J, Payton A, Barrett JH, Hever T, Fitzpatrick H, Trumper AL, Harrington R, McGuffin P, Owen M, Ollier W, Worthington J, Thapar A (2000) A family-based and case–control association study of the dopamine D4 receptor gene and dopamine transporter gene in attention deficit hyperactivity disorder. Mol Psychiatry 5:523–530
Feng Y, Wigg KG, Makkar R, Ickowicz A, Pathare T, Tannock R, Roberts W, Malone M, Kennedy JL, Schachar R, Barr CL (2005) Sequence variation in the 3′-untranslated region of the dopamine transporter gene and attention-deficit hyperactivity disorder (ADHD). Am J Med Genet B Neuropsychiatr Genet 139:1–6
Cheuk DK, Li SY, Wong V (2006) No association between VNTR polymorphisms of dopamine transporter gene and attention deficit hyperactivity disorder in Chinese children. Am J Med Genet B Neuropsychiatr Genet 141:123–125
Giros B, Jaber M, Jones SR, Wightman RM, Caron MG (1996) Hyperlocomotion and indifference to cocaine and amphetamine in mice lacking the dopamine transporter. Nature 379:606–612
Carrasco X, Rothhammer P, Moraga M, Henriquez H, Chakraborty R, Aboitiz F, Rothhammer F (2006) Genotypic interaction between DRD4 and DAT1 loci is a high risk factor for attention-deficit/hyperactivity disorder in Chilean families. Am J Med Genet B Neuropsychiatr Genet 141:51–54
Sabol SZ, Nelson ML, Fisher C, Gunzerath L, Brody CL, Hu S, Sirota LA, Marcus SE, Greenberg BD, Lucas FR 4th, Benjamin J, Murphy DL, Hamer DH (1999) A genetic association for cigarette smoking behavior. Health Psychol 18:7–13
Lerman C, Caporaso NE, Audrain J, Main D, Bowman ED, Lockshin B, Boyd NR, Shields PG (1999) Evidence suggesting the role of specific genetic factors in cigarette smoking. Health Psychol 18:14–20
Timberlake DS, Haberstick BC, Lessem JM, Smolen A, Ehringer M, Hewitt JK, Hopfer C (2006) An association between the DAT1 polymorphism and smoking behavior in young adults from the National Longitudinal Study of Adolescent Health. Health Psychol 25:190–197
Brody AL, Mandelkern MA, Olmstead RE, Scheibal D, Hahn E, Shiraga S, Zamora-Paja E, Farahi J, Saxena S, London ED, McCracken JT (2006) Gene variants of brain dopamine pathways and smoking-induced dopamine release in the ventral caudate/nucleus accumbens. Arch Gen Psychiatry 63:808–816
Le Couteur DG, Leighton PW, McCann SJ, Pond S (1997) Association of a polymorphism in the dopamine-transporter gene with Parkinson’s disease. Mov Disord 12:760–763
Kim JW, Kim DH, Kim SH, Cha JK (2000) Association of the dopamine transporter gene with Parkinson’s disease in Korean patients. J Korean Med Sci 15:449–451
Lim MH, Kim HW, Paik KC, Cho SC, Yoon DY, Lee HJ (2006) Association of the DAT1 polymorphism with attention deficit hyperactivity disorder (ADHD): a family-based approach. Am J Med Genet B Neuropsychiatr Genet 141:309–311
Mercier G, Turpin JC, Lucotte G (1999) Variable number tandem repeat dopamine transporter gene polymorphism and Parkinson’s disease: no association found. J Neurol 246:45–47
Kelada SN, Costa-Mallen P, Checkoway H, Carlson CS, Weller TS, Swanson PD, Franklin GM, Longstreth WT Jr, Afsharinejad Z, Costa LG (2005) Dopamine transporter (SLC6A3) 5′ region haplotypes significantly affect transcriptional activity in vitro but are not associated with Parkinson’s disease. Pharmacogenet Genomics 15:659–668
Leighton PW, Le Couteur DG, Pang CC, McCann SJ, Chan D, Law LK, Kay R, Pond SM, Woo J (1997) The dopamine transporter gene and Parkinson’s disease in a Chinese population. Neurology 49:1577–1579
Sander T, Harms H, Podschus J, Finckh U, Nickel B, Rolfs A, Rommelspacher H, Schmidt LG (1997) Allelic association of a dopamine transporter gene polymorphism in alcohol dependence with withdrawal seizures or delirium. Biol Psychiatry 41:299–304
Schmidt LG, Harms H, Kuhn S, Rommelspacher H, Sander T (1998) Modification of alcohol withdrawal by the A9 allele of the dopamine transporter gene. Am J Psychiatry 155(4):474–478
Ueno S, Nakamura M, Mikami M, Kondoh K, Ishiguro H, Arinami T, Komiyama T, Mitsushio H, Sano A, Tanabe H (1999) Identification of a novel polymorphism of the human dopamine transporter (DAT1) gene and the significant association with alcoholism. Mol Psychiatry 4:552–557
Need AC, Ahmadi KR, Spector TD, Goldstein DB (2006) Obesity is associated with genetic variants that alter dopamine availability. Ann Hum Genet 70:293–303
Yoon DY, Rippel CA, Kobets AJ, Morris CM, Lee JE, Williams PN, Bridges DD, Vandenbergh DJ, Shugart YY, Singer HS (2006) Dopaminergic polymorphisms in Tourette syndrome: association with the DAT gene (SLC6A3). Am J Med Genet B Neuropsychiatr Genet
Kim SJ, Kim YS, Kim CH, Lee HS (2006) Lack of association between polymorphisms of the dopamine receptor D4 and dopamine transporter genes and personality traits in a Korean population. Yonsei Med J 47:787–792
Gelernter J, Kranzler HR, Satel SL, Rao PA (1994) Genetic association between dopamine transporter protein alleles and cocaine-induced paranoia. Neuropsychopharmacology 11:195–200
Byerley W, Hoff M, Holik J, Caron MG, Giros B (1993) VNTR polymorphism for the human dopamine transporter gene (DAT1). Hum Mol Genet 2:335
Michelhaugh SK, Fiskerstrand C, Lovejoy E, Bannon MJ, Quinn JP (2001) The dopamine transporter gene (SLC6A3) variable number of tandem repeats domain enhances transcription in dopamine neurons. J Neurochem 79:1033–1038
Inoue-Murayama M, Adachi S, Mishima N, Mitani H, Takenaka O, Terao K, Hayasaka I, Ito S, Murayama Y (2002) Variation of variable number of tandem repeat sequences in the 3′-untranslated region of primate dopamine transporter genes that affects reporter gene expression. Neurosci Lett 334:206–210
Miller GM, Madras BK (2002) Polymorphisms in the 3′-untranslated region of human and monkey dopamine transporter genes affect reporter gene expression. Mol Psychiatry 7:44–55
Fuke S, Suo S, Takahashi N, Koike H, Sasagawa N, Ishiura S (2001) The VNTR polymorphism of the human dopamine transporter (DAT1) gene affects gene expression. Pharmacogenomics J 1:152–156
Mill J, Asherson P, Craig I, D’Souza UM (2005) Transient expression analysis of allelic variants of a VNTR in the dopamine transporter gene (DAT1). BMC Genet 6:3
Mill J, Asherson P, Browes C, D’Souza U, Craig I (2002) Expression of the dopamine transporter gene is regulated by the 3′ UTR VNTR: evidence from brain and lymphocytes using quantitative RT-PCR. Am J Med Genet 114:975–979
VanNess SH, Owens MJ, Kilts CD (2005) The variable number of tandem repeats element in DAT1 regulates in vitro dopamine transporter density. BMC Genet 6:55
Heinz A, Goldman D, Jones DW, Palmour R, Hommer D, Gorey JG, Lee KS, Linnoila M, Weinberger DR (2000) Genotype influences in vivo dopamine transporter availability in human striatum. Neuropsychopharmacology 22:133–139
Jacobsen LK, Staley JK, Zoghbi SS, Seibyl JP, Kosten TR, Innis RB, Gelernter J (2000) Prediction of dopamine transporter binding availability by genotype: a preliminary report. Am J Psychiatry 157:1700–1703
van Dyck CH, Malison RT, Jacobsen LK, Seibyl JP, Staley JK, Laruelle M, Baldwin RM, Innis RB, Gelernter J (2005) Increased dopamine transporter availability associated with the 9-repeat allele of the SLC6A3 gene. J Nucl Med 46:745–751
Martinez D, Gelernter J, Abi-Dargham A, van Dyck CH, Kegeles L, Innis RB, Laruelle M (2001) The variable number of tandem repeats polymorphism of the dopamine transporter gene is not associated with significant change in dopamine transporter phenotype in humans. Neuropsychopharmacology 24:553–560
Lynch DR, Mozley PD, Sokol S, Maas NM, Balcer LJ, Siderowf AD (2003) Lack of effect of polymorphisms in dopamine metabolism related genes on imaging of TRODAT-1 in striatum of asymptomatic volunteers and patients with Parkinson’s disease. Mov Disord 18:804–812
Contin M, Martinelli P, Mochi M, Albani F, Riva R, Scaglione C, Dondi M, Fanti S, Pettinato C, Baruzzi A (2004) Dopamine transporter gene polymorphism, spect imaging, and levodopa response in patients with Parkinson disease. Clin Neuropharmacol 27:111–115
Brookes KJ, Mill J, Guindalini C, Curran S, Xu X, Knight J, Chen CK, Huang YS, Sethna V, Taylor E, Chen W, Breen G, Asherson P (2006) A common haplotype of the dopamine transporter gene associated with attention-deficit/hyperactivity disorder and interacting with maternal use of alcohol during pregnancy. Arch Gen Psychiatry 63:74–81
O’Gara C, Stapleton J, Sutherland G, Guindalini C, Neale B, Breen G, Ball D (2007) Dopamine transporter polymorphisms are associated with short-term response to smoking cessation treatment. Pharmacogenet Genomics 17:61–67
Loo SK, Specter E, Smolen A, Hopfer C, Teale PD, Reite ML (2003) Functional effects of the DAT1 polymorphism on EEG measures in ADHD. J Am Acad Child Adolesc Psychiatry 42(8):986–993
Winsberg BG, Comings DE (1999) Association of the dopamine transporter gene (DAT1) with poor methylphenidate response. J Am Acad Child Adolesc Psychiatry 38:1474–1477
Stein MA, Waldman ID, Sarampote CS, Seymour KE, Robb AS, Conlon C, Kim SJ, Cook EH (2005) Dopamine transporter genotype and methylphenidate dose response in children with ADHD. Neuropsychopharmacology 30:1374–1382
Joober R, Grizenko N, Sengupta S, Amor LB, Schmitz N, Schwartz G, Karama S, Lageix P, Fathalli F, Torkaman-Zehi A, Stepanian MT (2006) Dopamine transporter 3′-UTR VNTR genotype and ADHD: a pharmaco-behavioural genetic study with methylphenidate. Neuropsychopharmacology
Collier DA, Stober G, Li T, Heils A, Catalano M, Di Bella D, Arranz MJ, Murray RM, Vallada HP, Bengel D, Muller CR, Roberts GW, Smeraldi E, Kirov G, Sham P, Lesch KP (1996) A novel functional polymorphism within the promoter of the serotonin transporter gene: possible role in susceptibility to affective disorders. Mol Psychiatry 1:453–460
Vostrov AA, Quitschke WW (1997) The zinc finger protein CTCF binds to the APBbeta domain of the amyloid beta-protein precursor promoter. Evidence for a role in transcriptional activation. J Biol Chem 272:33353–33359
Quitschke WW, Taheny MJ, Fochtmann LJ, Vostrov AA (2000) Differential effect of zinc finger deletions on the binding of CTCF to the promoter of the amyloid precursor protein gene. Nucleic Acids Res 28:3370–3378
Ohlsson R, Renkawitz R, Lobanenkov V (2001) CTCF is a uniquely versatile transcription regulator linked to epigenetics and disease. Trends Genet 17:520–527
Docquier F, Farrar D, D’Arcy V, Chernukhin I, Robinson AF, Loukinov D, Vatolin S, Pack S, Mackay A, Harris RA, Dorricott H, O’Hare MJ, Lobanenkov V, Klenova E (2005) Heightened expression of CTCF in breast cancer cells is associated with resistance to apoptosis. Cancer Res 65:5112–5122
Quinn JP (1996) Neuronal-specific gene expression—the interaction of both positive and negative transcriptional regulators. Prog Neurobiol 50:363–379
Brookes K, Xu X, Chen W, Zhou K, Neale B, Lowe N, Anney R, Franke B, Gill M, Ebstein R, Buitelaar J, Sham P, Campbell D, Knight J, Andreou P, Altink M, Arnold R, Boer F, Buschgens C, Butler L, Christiansen H, Feldman L, Fleischman K, Fliers E, Howe-Forbes R, Goldfarb A, Heise A, Gabriels I, Korn-Lubetzki I, Johansson L, Marco R, Medad S, Minderaa R, Mulas F, Muller U, Mulligan A, Rabin K, Rommelse N, Sethna V, Sorohan J, Uebel H, Psychogiou L, Weeks A, Barrett R, Craig I, Banaschewski T, Sonuga-Barke E, Eisenberg J, Kuntsi J, Manor I, McGuffin P, Miranda A, Oades RD, Plomin R, Roeyers H, Rothenberger A, Sergeant J, Steinhausen HC, Taylor E, Thompson M, Faraone SV, Asherson P (2006) The analysis of 51 genes in DSM-IV combined type attention deficit hyperactivity disorder: association signals in DRD4, DAT1 and 16 other genes. Mol Psychiatry 11:934–953
Hranilovic D, Stefulj J, Schwab S, Borrmann-Hassenbach M, Albus M, Jernej B, Wildenauer D (2004) Serotonin transporter promoter and intron 2 polymorphisms: relationship between allelic variants and gene expression. Biol Psychiatry 55:1090–1094
Magi R, Kaplinski L, Remm M (2006) The whole genome tagSNP selection and transferability among HapMap populations. Pac Symp Biocomput 11:535–543
Author information
Authors and Affiliations
Corresponding author
Additional information
Special issue article in honor of George Fink.
Rights and permissions
About this article
Cite this article
Haddley, K., Vasiliou, A.S., Ali, F.R. et al. Molecular Genetics of Monoamine Transporters: Relevance to Brain Disorders. Neurochem Res 33, 652–667 (2008). https://doi.org/10.1007/s11064-007-9521-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-007-9521-8