Abstract
Purpose
Our understanding of diffuse midline glioma (DMG) biology inclusive of diffuse intrinsic pontine glioma has been revolutionized by the discovery of novel mutations on the tails of histone 3, leading to the reclassification in 2016 of ‘diffuse midline glioma, H3 K27M-mutant.’ Given the abundance of basic, translational, and clinical information put forth in recent years, a review of the epigenetics of diffuse midline glioma is warranted.
Methods
Literature for the epigenetics of diffuse midline glioma published from 1989 to 2019 was reviewed by searching PubMed using the terms “diffuse intrinsic pontine glioma”, “pontine glioma”, or “midline glioma”. The final references list was generated on the basis of originality and relevance to the broad scope of our review.
Results
The effects of H3K27M-mutation, while better understood, suggest multiple consequences on the chromatin landscape and DNA modification states, contributed to the progression of DMG. A rapid pace of translational development is occurring for epigenetic modifiers, and several classes of inhibitors have already made their way into clinical trial testing. As more agents become clinically accessible, immense effort is underway to understand the target effects, tumor penetration, and immune microenvironmental changes of epigenetic modification.
Conclusion
We continue to seek a comprehensive understanding of the mechanisms that govern chromatin dysregulation and DNA modification in DMG, and in parallel we forge ahead with clinical testing of epigenetic modifiers. The determined efforts from bench to bedside, along with collaborative mindset and unified mission, will ultimately result in improved outcomes for DMG.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Genetics/Epigenetics of diffuse midline glioma (DMG)
Discovery of histone 3 mutations and emergence of new disease classification
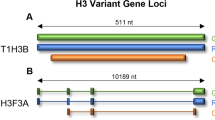
Diffuse intrinsic pontine glioma (DIPG) is an aggressive tumor of the brainstem, and accounts for 75% of brain stem tumors in children [1]. Median survival of DIPG remains only 11.2 months despite decades of clinical trials [2]. Fortunately, our understanding of DIPG biology has been revolutionized by the discovery of novel mutations on the tails of histone 3 (H3) [3]. Histones are nuclear proteins that play an essential role in condensing DNA into nucleosomes and regulating chromatin [4]. The H3 family of proteins includes canonical H3.1 and variant H3.3, each with a protruding amino-acid tail which is subject to a number of post-translational modifications (PTMs) [5]. These PTMs, such as methylation and acetylation, have a range of effects on the epigenetic regulation of transcription and chromatin structure [6]. One such PTM, the trimethylation of lysine 27 on histone 3 (H3K27me3), is usually associated with gene repression [7]. Trimethylation of H3K27 is catalyzed by the enhancer of zeste homolog (EZH1 or 2) subunit of the Polycomb Repressive Complex (PRC2), and has been shown to maintain transcriptional silencing, allowing the preservation of cell identity [8].
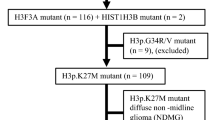
Mutations in H3, specifically the lysine to methionine substitution at position 27 (H3K27M), were initially reported almost a decade ago and represented the first description of histone mutations in cancer [3, 9, 10]. These highly recurrent, somatic, non-synonymous mutations occur not only in ~ 80% of DIPGs (65% in H3F3A, and 15% in HIST1H3B), but in gliomas arising from other midline structures such as the thalamus or spinal cord, and confer a worse prognosis than wildtype cases [11,12,13]. Thus, in 2016 the World Health Organization (WHO) recognized a new entity of diffuse midline glioma, H3 K27M-mutant [12]. Functional studies have since shown H3K27M mutation to drive DMG growth in vitro and in vivo, suggesting that epigenetic misregulation promotes DMG tumorigenesis [14,15,16]. DMG H3K27M mutant tumors, despite carrying the H3K27M mutation on merely 5–17% of H3 histones, have a global loss of H3K27me3 and DNA methylation, ultimately leading to a loss of gene repression [10, 17]. However, the mechanism by which H3K27M confers these effects is still debated.
The role of polycomb repressive complex 2
Following the discovery of H3K27M, it was shown that H3K27M is a gain of function mutation which enables the sequestration and inhibition of PRC2 [10]. The lysine-to-methionine mutation targets the active site of SET-domain containing methyltransferases, such as that in EZH2, and effectively competes with native lysine. By doing so, it prevents the hydrolysis of the methyl- donor S-adenosylmethionine (SAM) [18]. This, as well as methionine’s increased binding affinity for the active site of EZH2 relative to that of lysine, results in EZH2 being sequestered at the H3K27M site, inhibiting the complex and preventing global methylation at cognate residues. In support of this, a recent report by Diehl et al. demonstrated that the presence of bivalent dinucleotides containing native H3K27me3 plays a key role in the inhibition of PRC2 [19]. Within the PRC2 complex, the binding of H3K27M to the catalytic subunit EZH2, and of H3K27me3 to the allosteric activator EED, work cooperatively to inhibit the PRC2 complex. The binding of H3K27me3 to EED increases the affinity of PRC2 to the adjacent nucleosome (H3K27M), and thus further enables the sequestration of PRC2 to the chromatin site [19].
Although this was the first proposed hypothesis of H3K27M mediated affects, recent in vivo studies such as those by Piunti et al. have shown that, in contrast to the hypothesis that H3K27M sequesters PRC2, the localization of H3K27M is inversely correlated with PRC2 occupancy [20]. Specifically, H3K27M is often deposited at transcriptionally active regions of chromatin, and its expression is associated with increased levels of H3K27 acetylation at specific genomic regions [20, 21]. Additionally, the PRC2 subunits EZH2 and SUZ12 are often excluded from chromatin on sites containing H3K27M. Together, this suggests that the formation of locus specific H3K27M/H3K27ac heterotypic nucleosomes results in the exclusion of PRC2 from chromatin binding, and subsequently a loss of H3K27me3. This demonstrates that, in contrast to previous reports, H3K27M and PRC2 do not have strong interactions in all chromatin contexts. As well, the p16 protein (p16) is a well-known negative cell cycle regulator, tumour suppressor, and target of PRC2 mediated transcriptional repression [22]. However, upon PRC2 repression, p16 was not shown to be upregulated in H3K27M cells. This supports mechanisms proposed in other models, which suggest that the pro-proliferative activity of PRC2 inhibited cells occurs through a p16-independent mechanism [23].
In addition to the development of these hypotheses, a recent study by Stafford et al. proposed a third hypothesis to try and resolve the mechanism by which both PRC2 and the overall chromatin landscape of DIPG are affected by H3K27M [24]. Bridging the gap between the two-prior propositions, this study demonstrated that the allosterically activated form of PRC2 binds more strongly to H3K27M, as shown by Lewis et al. [10, 24]. This binding occurs aberrantly at loci that are not typical of PRC2 recruitment and occurs early after H3K27M expression. However, the H3K27M-PRC2 binding is transient; upon binding, EZH2 undergoes a conformational change which renders it catalytically non-functional, and the PRC2 complex inhibited. At this point, PRC2 is released from the chromatin site, and the inhibition of PRC2 persists, without lasting deposition at H3K27M (Fig. 1).
a In wild-type cells, PRC2 complex is recruited to target genes and catalyzes the trimethylation of lysine 27 on the protruding histone H3 amino acid tail through the EZH2 catalytic subunit, maintaining gene repression. Following deposition, PRC2 spreads the trimethyl mark to additional target genes, or dissociates from the histone b Presence of K27M mutation on histone H3 inhibits the PRC2 complex, preventing the deposition of the trimethyl mark and subsequent spreading of PRC2
Histone Effects Beyond PRC2
As well as provide an additional mechanism of PRC2 inhibition, Stafford et al. also demonstrated that H3K27M also results in the global increase of other histone activating marks [24]. This includes additional histone H3 lysine methylation marks such as the PRC2 antagonist H3K26me2/3, as well as increased H3K36me2/3, both of which are typically associated with areas of active transcription. These compliment the well-established effect of DNA hypomethylation, which causes increased expression as a result of loss of gene silencing [17]. Taken together, these alterations reveal the effects of H3K27M surpass that of the H3K27 site and suggest multiple consequences of H3K27M on the chromatin landscape and DNA modification states, suggested to contribute to the progression of DMG [24, 33].
Histone H3.1 and H3.3 Distinctions
The global effects of H3K27M in DMG can be assessed within the context of their specific H3K27M subgroups, H3.1K27M and H3.3K27M. Canonical H3.1 and variant H3.3 can be distinguished by their patterns of deposition throughout chromatin. The incorporation of H3.1 into chromatin is DNA replication dependent as H3.1 is only expressed during S-phase of the cell cycle, where it is incorporated throughout the genome with relative uniformity [25]. However, variant H3.3 is expressed throughout the cell cycle, and thus incorporation is DNA-independent. In contrast to H3.1, H3.3 is incorporated into actively transcribed regions of chromatin, as well as pericentric heterochromatin and telomeres [25]. These differences suggest that the means by which these histones influence tumorigenesis may, in fact, not be identical.
The possibility that tumorigenesis may vary by type of H3 mutation is supported by the distinct secondary mutations associated with H3.1K27M and H3.3K27M tumours [26]. Within H3.1K27M tumours, there is a well-recognized co-occurrence with mutations in activin-receptor type 1 (ACVR1) and phosphoinositide 3-kinase (PI3K), whereas H3.3K27M tumours commonly co-occur with mutations or deletions in tumour suppressor 53 (TP53), and amplification of platelet-derived growth factor alpha (PDGFRa) [26]. Additionally, H3.3K27M tumours have shown enrichment for genes involved in the Ras homolog guanosine triphosphatase (Rho GTPase) activity over H3.1K27M tumours, promoting both tumour cell migration and invasion, and consistent with H3.3K27M DIPG primary tumours being more invasive, more aggressive, and less differentiated than H3.1K27M tumours [27,28,29].
Epigenetic modifier clinical experience for DMG
Given the mounting evidence that epigenetic alterations are associated with DMG development and progression, a rapid pace of translational development is occurring for epigenetic modifiers, and several classes of inhibitors have already made their way into clinical trial testing.
Histone deacetylase inhibition
Histone deacetylase (HDAC) inhibitors were the first class of epigenetic modifiers to be brought to DIPG clinical trial testing. Well before the discovery of histone mutations in DIPG, the Children’s Oncology Group (COG) conducted a phase I/II clinical trial testing suberoylanilide hydroxamic acid (SAHA, Vorinostat), a histone deacetylase inhibitor, and local irradiation, followed by maintenance SAHA in children with newly diagnosed DIPG (NCT01189266). The study is no longer recruiting, and the results are yet to be published.
Following preclinical work described by Grass and colleagues [30], a multi-institutional phase 1 trial of the pan-HDAC inhibitor panobinostat for children with DIPG (NCT02717455) was launched through the Pediatric Brain Tumor Consortium (PBTC) in 2016. The maximum tolerated dose (MTD) of panobinostat administered to children with progressive DIPG on a schedule of 3 weeks on, 1 week off, Monday/Wednesday/Friday was 10 mg/m2/dose, much lower than anticipated [31]. The trial was amended to evaluate the MTD of panobinostat in children with non-progressive DIPG, treated on an alternate week dosing schedule after completion of radiation therapy. The second stratum has been enrolling since October 2017 and is on-going.
Around similar time, a multi-institutional trial using molecularly targeted therapy for DIPG (NCT02274987) opened through the Pacific Pediatric Neuro-Oncology Consortium (PNOC). Patients enrolled from September 2014 through January 2016, and up to four FDA-approved drugs were used for every patient, dependent upon the tumor’s molecular profile based on RNA sequencing as well as whole exome sequencing results [32]. Panobinostat was administered in doses ranging from 15–37 mg/m2/dose, at varying schedule and combination with other agents [33]. While the study achieved feasibility, the optimal dose and schedule of panobinostat remains in question. Furthermore, since it’s FDA approval in 2015, oral panobinostat has been used off label for pediatric DMG in a variety of doses and schedules. One published example has been its use concomitantly with reirradiation in children with progressive DIPG, at doses ranging from 22–25 mg/m2/dose, 3 times weekly for 2 weeks in 3-week cycles [34].
As the PBTC phase 1 trial nears completion, questions remain for oral panobinostat’s ability to penetrate the blood tumor barrier effectively, even at the highest tolerated doses. Answers will come through a well powered efficacy trial, still several years away. In the mean-time, the clinical trial landscape is opening to alternative HDAC inhibitors as well as alternative routes of HDAC inhibitor delivery. Following promising small and large animal model data, PNOC’s single institution phase I/II trial of MTX110 for children with DIPG opened in 2018 and delivers the water-soluble form of panobinostat intratumorally through convection enhanced delivery (CED) (NCT03566199) [35]. PNOC has also opened a multi-center target validation study of Fimepinostat, a novel oral, dual pan-HDAC and PI3K inhibitor for children with newly diagnosed DIPG and recurrent HGG in 2019 (NCT03893487). Further work is needed to define the optimal route, dose, schedule, and timing of HDAC inhibitors for children with DMG.
BMI-1 inhibition
The DMG trial landscape has already expanded beyond HDAC inhibitors to include targeted inhibition of BMI-1, a key regulatory component of the human PRC1 complex. Drawing from their preclinical work showing DIPG cell sensitization to radiomimetic drug-induced DNA damage by BMI-1 inhibition, the CONNECT consortium opened a multi-institutional phase 1b study of the BMI-1 inhibitor PTC596 (NCT03605550) in 2018 for children with newly diagnosed DIPG and HGG [36]. Enrollment is ongoing, providing PTC596 concomitantly with radiotherapy, followed by single-agent maintenance dosing.
Future epigenetic modifier strategies for DMG
Combination epigenetic therapy
We are years away from the results of a well powered efficacy study, but it is well presumed that agents such as HDACi will not succeed as monotherapy, and combinatorial approaches are needed to improve outcomes. To that extent, pre-clinical efforts are ongoing to test novel combinations. A recent comprehensive high-throughput drug screen followed by DMG xenograft testing identified potent synergy of panobinostat with the proteasome inhibitor marizomib. The models used were representative of H3WT, H3.3K27M, and H3.1K27M, and the authors propose the combination’s efficacy is driven by a metabolic crisis ubiquitous within all DMG cells [37]. A multi-institutional phase 1 combination trial of panobinostat and marizomib for children with DMG is expected to open in early 2020.
Another recent study used a CRISPR screen to reveal the sensitization of DIPG cells to HDAC inhibition by LSD-1 knockout, and that Corin, a bifunctional inhibitor of HDACs and LSD1, potently inhibited DIPG growth in vitro and in xenografts. Mechanistically, Corin increased H3K27me3 levels suppressed by H3K27M histones, and simultaneously increased HDAC-targeted H3K27ac and LSD1-targted H3K4me1 at differentiation-associated genes [38]. Although further identification of relevant clinical agents and the associated preclinical testing is needed, the combination of LSD1 inhibition with HDAC inhibition will likely be brought to clinical trial testing for DMG in coming years.
Effective penetrance
For epigenetic modification to reach clinical success, the chosen agents must achieve adequate exposure to the tumor and its sites of spread. Focal drug delivery methods such as CED will continue, as well as intra-arterial delivery methods such as the recently completed trial at Johns Hopkins (NCT01688401). Whether epigenetic modifiers may be delivered safely and effectively by focal delivery remains to be seen. In general, it is presumed that focal treatment will not be curative for DMG unless the therapeutic(s) reaches and eradicates all tumor cells prior to disease spread [39]. Most likely, systemic therapy will be a part of future success. Ongoing pre-clinical efforts to study systemically delivered epigenetic modifiers are placing importance on tumor penetrance. The HDAC inhibitor quisinostat has been identified as a possible superior option due to its increased selectivity for HDAC1, and is currently undergoing further preclinical testing [40]. Regardless of which epigenetic agents are chosen for future clinical trials, we will likely see importance placed on trial design to determine intratumoral target validation.
Correlative studies for epigenetic modifier trials
As more agents become clinically accessible, a simultaneous effort is underway to understand the target effects of epigenetic modification.
Meaningful Biomarkers
An urgent need exists within DMG research to identify predictive biomarkers and validate them through clinical trials. Our understanding of the posttranslational modifications that occur with H3K27M mutation is growing—not only H3K27me3 and H3K27me2 reduction, but disruption of H3K4me3 and H3K36me2, histone acetylation, and DNA methylation [14, 17, 41]. How these aberrations lead to tumorigenesis, and how broadly reprogramming the epigenome through the aforementioned inhibitors leads to tumor reduction, is still not understood. Preclinical work such as that previously described using panobinostat single agent or in combination with marizomib and corin demonstrated efficacy not only for H3K27M mutant tumors, but H3 wild type as well [30, 37, 38]. More thorough mechanistic understanding will allow us to refine (or expand) our targeted populations. Currently, chromatin immunoprecipitation sequencing (ChIP-seq) and RNA sequencing are allowing for broader preclinical analysis of the transcriptome leading to a fuller contextual understanding. Alternatively, partial or total efficacy of epigenetic modifiers may lie in their effects beyond that of epigenome reprogramming.
Immune correlates
We are becoming increasingly aware of the immunologic effects of epigenetic modification. For example, DNA methylation inhibitors (DNMTi) and HDAC inhibitors (HDACi), in addition to broadly reprogramming the epigenome, alter the function of immune cells relevant to acquired immunity in various malignancies. HDACi have led to immunosuppressive effects such as upregulation of PDL-1 in adult melanoma, and DNMTi have upregulated PD1 in T cells of adult patients with MDS and AML [42, 43]. In other contexts, epigenetic therapy has enhanced immune activation. The combination of DNMTi and HDACi induced the secretion of T helper 1 (TH1) cell-type response cytokines, an immune signature associated with effective antitumour immunity in both pretreated lung cancers and those treated with immune checkpoint blockade [44]. These findings highly the need to evaluate changes of the DMG immune microenvironment with epigenetic modifier treatment, and we will see immune correlates included more often within clinical trials.
Conclusion
We continue to seek a comprehensive understanding of the mechanisms that govern chromatin dysregulation and DNA modification in DMG, and their roles in tumorigenesis. In parallel, we forge ahead with clinical testing of epigenetic modifiers. Straddled between mechanistic discovery and clinical development lies preclinical testing, for which we are iteratively evaluating the appropriate models and thresholds of efficacy. The determined efforts from individuals working bench to bedside, along with collaborative mindset and unified mission, will ultimately result in improved outcomes for DMG.
References
Ostrom QT, Cioffi G, Gittleman H, Patil N, Waite K, Kruchko C, Barnholtz-Sloan JS (2019) CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2012–2016. Neuro Oncol 21(5):v1–v100. https://doi.org/10.1093/neuonc/noz150
Cooney T, Lane A, Bartels U, Bouffet E, Goldman S, Leary SES, Foreman NK, Packer RJ, Broniscer A, Minturn JE, Shih CS, Chintagumpala M, Hassall T, Gottardo NG, Dholaria H, Hoffman L, Chaney B, Baugh J, Doughman R, Leach JL, Jones BV, Fouladi M, Warren KE, Monje M (2017) Contemporary survival endpoints: an international diffuse intrinsic pontine glioma registry study. Neuro Oncol 19(9):1279–1280. https://doi.org/10.1093/neuonc/nox107
Khuong-Quang DA, Buczkowicz P, Rakopoulos P, Liu XY, Fontebasso AM, Bouffet E, Bartels U, Albrecht S, Schwartzentruber J, Letourneau L, Bourgey M, Bourque G, Montpetit A, Bourret G, Lepage P, Fleming A, Lichter P, Kool M, von Deimling A, Sturm D, Korshunov A, Faury D, Jones DT, Majewski J, Pfister SM, Jabado N, Hawkins C (2012) K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol 124(3):439–447. https://doi.org/10.1007/s00401-012-0998-0
Marino-Ramirez L, Kann MG, Shoemaker BA, Landsman D (2005) Histone structure and nucleosome stability. Expert Rev Proteomics 2(5):719–729. https://doi.org/10.1586/14789450.2.5.719
Hacques MF, Muller S, De Murcia G, Van Regenmortel MH, Marion C (1990) Use of an immobilized enzyme and specific antibodies to analyse the accessibility and role of histone tails in chromatin structure. Biochem Biophys Res Commun 168(2):637–643. https://doi.org/10.1016/0006-291x(90)92368-a
Campos EI, Reinberg D (2009) Histones: annotating chromatin. Annu Rev Genet 43:559–599. https://doi.org/10.1146/annurev.genet.032608.103928
Plass C, Pfister SM, Lindroth AM, Bogatyrova O, Claus R, Lichter P (2013) Mutations in regulators of the epigenome and their connections to global chromatin patterns in cancer. Nat Rev Genet 14(11):765–780. https://doi.org/10.1038/nrg3554
Comet I, Riising EM, Leblanc B, Helin K (2016) Maintaining cell identity: PRC2-mediated regulation of transcription and cancer. Nat Rev Cancer 16(12):803–810. https://doi.org/10.1038/nrc.2016.83
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K, Sturm D, Fontebasso AM, Quang DA, Tonjes M, Hovestadt V, Albrecht S, Kool M, Nantel A, Konermann C, Lindroth A, Jager N, Rausch T, Ryzhova M, Korbel JO, Hielscher T, Hauser P, Garami M, Klekner A, Bognar L, Ebinger M, Schuhmann MU, Scheurlen W, Pekrun A, Fruhwald MC, Roggendorf W, Kramm C, Durken M, Atkinson J, Lepage P, Montpetit A, Zakrzewska M, Zakrzewski K, Liberski PP, Dong Z, Siegel P, Kulozik AE, Zapatka M, Guha A, Malkin D, Felsberg J, Reifenberger G, von Deimling A, Ichimura K, Collins VP, Witt H, Milde T, Witt O, Zhang C, Castelo-Branco P, Lichter P, Faury D, Tabori U, Plass C, Majewski J, Pfister SM, Jabado N (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231. https://doi.org/10.1038/nature10833
Lewis PW, Muller MM, Koletsky MS, Cordero F, Lin S, Banaszynski LA, Garcia BA, Muir TW, Becher OJ, Allis CD (2013) Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340(6134):857–861. https://doi.org/10.1126/science.1232245
Jones C, Karajannis MA, Jones DTW, Kieran MW, Monje M, Baker SJ, Becher OJ, Cho YJ, Gupta N, Hawkins C, Hargrave D, Haas-Kogan DA, Jabado N, Li XN, Mueller S, Nicolaides T, Packer RJ, Persson AI, Phillips JJ, Simonds EF, Stafford JM, Tang Y, Pfister SM, Weiss WA (2017) Pediatric high-grade glioma: biologically and clinically in need of new thinking. Neuro Oncol 19(2):153–161. https://doi.org/10.1093/neuonc/now101
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Buczkowicz P, Hoeman C, Rakopoulos P, Pajovic S, Letourneau L, Dzamba M, Morrison A, Lewis P, Bouffet E, Bartels U, Zuccaro J, Agnihotri S, Ryall S, Barszczyk M, Chornenkyy Y, Bourgey M, Bourque G, Montpetit A, Cordero F, Castelo-Branco P, Mangerel J, Tabori U, Ho KC, Huang A, Taylor KR, Mackay A, Bendel AE, Nazarian J, Fangusaro JR, Karajannis MA, Zagzag D, Foreman NK, Donson A, Hegert JV, Smith A, Chan J, Lafay-Cousin L, Dunn S, Hukin J, Dunham C, Scheinemann K, Michaud J, Zelcer S, Ramsay D, Cain J, Brennan C, Souweidane MM, Jones C, Allis CD, Brudno M, Becher O, Hawkins C (2014) Genomic analysis of diffuse intrinsic pontine gliomas identifies three molecular subgroups and recurrent activating ACVR1 mutations. Nat Genet 46(5):451–456. https://doi.org/10.1038/ng.2936
Larson JD, Kasper LH, Paugh BS, Jin H, Wu G, Kwon CH, Fan Y, Shaw TI, Silveira AB, Qu C, Xu R, Zhu X, Zhang J, Russell HR, Peters JL, Finkelstein D, Xu B, Lin T, Tinkle CL, Patay Z, Onar-Thomas A, Pounds SB, McKinnon PJ, Ellison DW, Zhang J, Baker SJ (2019) Histone H3.3 K27M Accelerates Spontaneous Brainstem Glioma and Drives Restricted Changes in Bivalent Gene Expression. Cancer Cell 35(1):140–155. https://doi.org/10.1016/j.ccell.2018.11.015
Cordero FJ, Huang Z, Grenier C, He X, Hu G, McLendon RE, Murphy SK, Hashizume R, Becher OJ (2017) Histone H33K27M Represses p16 to Accelerate Gliomagenesis in a Murine Model of DIPG. Mol Cancer Res 15(9):1243–1254. https://doi.org/10.1158/1541-7786.MCR-16-0389
Pathania M, De Jay N, Maestro N, Harutyunyan AS, Nitarska J, Pahlavan P, Henderson S, Mikael LG, Richard-Londt A, Zhang Y, Costa JR, Hebert S, Khazaei S, Ibrahim NS, Herrero J, Riccio A, Albrecht S, Ketteler R, Brandner S, Kleinman CL, Jabado N, Salomoni P (2017) H3.3(K27M) Cooperates with Trp53 Loss and PDGFRA Gain in Mouse Embryonic Neural Progenitor Cells to Induce Invasive High-Grade Gliomas. Cancer Cell 32(5):684–700. https://doi.org/10.1016/j.ccell.2017.09.014
Bender S, Tang Y, Lindroth AM, Hovestadt V, Jones DT, Kool M, Zapatka M, Northcott PA, Sturm D, Wang W, Radlwimmer B, Hojfeldt JW, Truffaux N, Castel D, Schubert S, Ryzhova M, Seker-Cin H, Gronych J, Johann PD, Stark S, Meyer J, Milde T, Schuhmann M, Ebinger M, Monoranu CM, Ponnuswami A, Chen S, Jones C, Witt O, Collins VP, von Deimling A, Jabado N, Puget S, Grill J, Helin K, Korshunov A, Lichter P, Monje M, Plass C, Cho YJ, Pfister SM (2013) Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24(5):660–672. https://doi.org/10.1016/j.ccr.2013.10.006
Justin N, Zhang Y, Tarricone C, Martin SR, Chen S, Underwood E, De Marco V, Haire LF, Walker PA, Reinberg D, Wilson JR, Gamblin SJ (2016) Structural basis of oncogenic histone H3K27M inhibition of human polycomb repressive complex 2. Nat Commun 7:11316. https://doi.org/10.1038/ncomms11316
Diehl KL, Ge EJ, Weinberg DN, Jani KS, Allis CD, Muir TW (2019) PRC2 engages a bivalent H3K27M-H3K27me3 dinucleosome inhibitor. Proc Natl Acad Sci U S A 116(44):22152–22157. https://doi.org/10.1073/pnas.1911775116
Piunti A, Hashizume R, Morgan MA, Bartom ET, Horbinski CM, Marshall SA, Rendleman EJ, Ma Q, Takahashi YH, Woodfin AR, Misharin AV, Abshiru NA, Lulla RR, Saratsis AM, Kelleher NL, James CD, Shilatifard A (2017) Therapeutic targeting of polycomb and BET bromodomain proteins in diffuse intrinsic pontine gliomas. Nat Med 23(4):493–500. https://doi.org/10.1038/nm.4296
Herz HM, Morgan M, Gao X, Jackson J, Rickels R, Swanson SK, Florens L, Washburn MP, Eissenberg JC, Shilatifard A (2014) Histone H3 lysine-to-methionine mutants as a paradigm to study chromatin signaling. Science 345(6200):1065–1070. https://doi.org/10.1126/science.1255104
Bracken AP, Kleine-Kohlbrecher D, Dietrich N, Pasini D, Gargiulo G, Beekman C, Theilgaard-Monch K, Minucci S, Porse BT, Marine JC, Hansen KH, Helin K (2007) The Polycomb group proteins bind throughout the INK4A-ARF locus and are disassociated in senescent cells. Genes Dev 21(5):525–530. https://doi.org/10.1101/gad.415507
Piunti A, Rossi A, Cerutti A, Albert M, Jammula S, Scelfo A, Cedrone L, Fragola G, Olsson L, Koseki H, Testa G, Casola S, Helin K, d'Adda di Fagagna F, Pasini D (2014) Polycomb proteins control proliferation and transformation independently of cell cycle checkpoints by regulating DNA replication. Nat Commun 5:3649. https://doi.org/10.1038/ncomms4649
Stafford JM, Lee CH, Voigt P, Descostes N, Saldana-Meyer R, Yu JR, Leroy G, Oksuz O, Chapman JR, Suarez F, Modrek AS, Bayin NS, Placantonakis DG, Karajannis MA, Snuderl M, Ueberheide B, Reinberg D (2018) Multiple modes of PRC2 inhibition elicit global chromatin alterations in H3K27M pediatric glioma. Sci Adv 4(10):eaau5935. https://doi.org/10.1126/sciadv.aau5935
Goldberg AD, Banaszynski LA, Noh KM, Lewis PW, Elsaesser SJ, Stadler S, Dewell S, Law M, Guo X, Li X, Wen D, Chapgier A, DeKelver RC, Miller JC, Lee YL, Boydston EA, Holmes MC, Gregory PD, Greally JM, Rafii S, Yang C, Scambler PJ, Garrick D, Gibbons RJ, Higgs DR, Cristea IM, Urnov FD, Zheng D, Allis CD (2010) Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140(5):678–691. https://doi.org/10.1016/j.cell.2010.01.003
Mackay A, Burford A, Carvalho D, Izquierdo E, Fazal-Salom J, Taylor KR, Bjerke L, Clarke M, Vinci M, Nandhabalan M, Temelso S, Popov S, Molinari V, Raman P, Waanders AJ, Han HJ, Gupta S, Marshall L, Zacharoulis S, Vaidya S, Mandeville HC, Bridges LR, Martin AJ, Al-Sarraj S, Chandler C, Ng HK, Li X, Mu K, Trabelsi S, Brahim DH, Kisljakov AN, Konovalov DM, Moore AS, Carcaboso AM, Sunol M, de Torres C, Cruz O, Mora J, Shats LI, Stavale JN, Bidinotto LT, Reis RM, Entz-Werle N, Farrell M, Cryan J, Crimmins D, Caird J, Pears J, Monje M, Debily MA, Castel D, Grill J, Hawkins C, Nikbakht H, Jabado N, Baker SJ, Pfister SM, Jones DTW, Fouladi M, von Bueren AO, Baudis M, Resnick A, Jones C (2017) Integrated Molecular Meta-Analysis of 1000 Pediatric High-Grade and Diffuse Intrinsic Pontine Glioma. Cancer Cell 32(4):520–553. https://doi.org/10.1016/j.ccell.2017.08.017
Qin EY, Cooper DD, Abbott KL, Lennon J, Nagaraja S, Mackay A, Jones C, Vogel H, Jackson PK, Monje M (2017) Neural Precursor-Derived Pleiotrophin Mediates Subventricular Zone Invasion by Glioma. Cell 170(5):845–859. https://doi.org/10.1016/j.cell.2017.07.016
Castel D, Philippe C, Calmon R, Le Dret L, Truffaux N, Boddaert N, Pages M, Taylor KR, Saulnier P, Lacroix L, Mackay A, Jones C, Sainte-Rose C, Blauwblomme T, Andreiuolo F, Puget S, Grill J, Varlet P, Debily MA (2015) Histone H3F3A and HIST1H3B K27M mutations define two subgroups of diffuse intrinsic pontine gliomas with different prognosis and phenotypes. Acta Neuropathol 130(6):815–827. https://doi.org/10.1007/s00401-015-1478-0
Nagaraja S, Quezada MA, Gillespie SM, Arzt M, Lennon JJ, Woo PJ, Hovestadt V, Kambhampati M, Filbin MG, Suva ML, Nazarian J, Monje M (2019) Histone Variant and Cell Context Determine H3K27M Reprogramming of the Enhancer Landscape and Oncogenic State. Mol Cell 76(6):965–980. https://doi.org/10.1016/j.molcel.2019.08.030
Grasso CS, Tang Y, Truffaux N, Berlow NE, Liu L, Debily MA, Quist MJ, Davis LE, Huang EC, Woo PJ, Ponnuswami A, Chen S, Johung TB, Sun W, Kogiso M, Du Y, Qi L, Huang Y, Hutt-Cabezas M, Warren KE, Le Dret L, Meltzer PS, Mao H, Quezado M, van Vuurden DG, Abraham J, Fouladi M, Svalina MN, Wang N, Hawkins C, Nazarian J, Alonso MM, Raabe EH, Hulleman E, Spellman PT, Li XN, Keller C, Pal R, Grill J, Monje M (2015) Functionally defined therapeutic targets in diffuse intrinsic pontine glioma. Nat Med 21(7):827. https://doi.org/10.1038/nm0715-827a
Cooney T, Onar-Thomas A, Huang J, Lulla R, Fangusaro J, Kramer K, Baxter P, Fouladi M, Dunkel IJ, Warren KE, Monje M (2018) DIPG-22 A Phase 1 Trial Of The Histone Deacetylase Inhibitor Panobinostat In Pediatric Patients With Recurrent Or Refractory Diffuse Intrinsic Pontine Glioma: A Pediatric Brain Tumor Consortium (Pbtc) Study. Neuro Oncol 20:i53. https://doi.org/10.1093/neuonc/noy059.115
Mueller S, Jain P, Liang WS, Kilburn L, Kline C, Gupta N, Panditharatna E, Magge SN, Zhang B, Zhu Y, Crawford JR, Banerjee A, Nazemi K, Packer RJ, Petritsch CK, Truffaux N, Roos A, Nasser S, Phillips JJ, Solomon D, Molinaro A, Waanders AJ, Byron SA, Berens ME, Kuhn J, Nazarian J, Prados M, Resnick AC (2019) A pilot precision medicine trial for children with diffuse intrinsic pontine glioma-PNOC003: A report from the Pacific Pediatric Neuro-Oncology Consortium. Int J Cancer 145(7):1889–1901. https://doi.org/10.1002/ijc.32258
Kilburn L, Kuhn J, Nazemi K, Molinaro A, Liang W, Banerjee A, Hwang E, Crawford J, Nicolaides T, Packer R, Nazarian J, Gupta N, Resnick AC, Byron S, Berens M, Prados M, Mueller S (2018) DIPG-76 PNOC-003: Precision medicine trial for children with diffuses intrinsic pontine glioma: preliminary experience with multi-agent personalized therapy recommendations. Neuro Oncol 20:i64. https://doi.org/10.1093/neuonc/noy059.168
Wang ZJ, Ge Y, Altinok D, Poulik J, Sood S, Taub JW, Edwards H, Kieran MW, Steven M (2017) Concomitant Use of Panobinostat and Reirradiation in Progressive DIPG: Report of 2 Cases. J Pediatr Hematol Oncol 39(6):e332–e335. https://doi.org/10.1097/MPH.0000000000000806
Singleton WGB, Bienemann AS, Woolley M, Johnson D, Lewis O, Wyatt MJ, Damment SJP, Boulter LJ, Killick-Cole CL, Asby DJ, Gill SS (2018) The distribution, clearance, and brainstem toxicity of panobinostat administered by convection-enhanced delivery. J Neurosurg Pediatr 22(3):288–296. https://doi.org/10.3171/2018.2.PEDS17663
Kumar SS, Sengupta S, Lee K, Hura N, Fuller C, DeWire M, Stevenson CB, Fouladi M, Drissi R (2017) BMI-1 is a potential therapeutic target in diffuse intrinsic pontine glioma. Oncotarget 8(38):62962–62975. https://doi.org/10.18632/oncotarget.18002
Lin GL, Wilson KM, Ceribelli M, Stanton BZ, Woo PJ, Kreimer S, Qin EY, Zhang X, Lennon J, Nagaraja S, Morris PJ, Quezada M, Gillespie SM, Duveau DY, Michalowski AM, Shinn P, Guha R, Ferrer M, Klumpp-Thomas C, Michael S, McKnight C, Minhas P, Itkin Z, Raabe EH, Chen L, Ghanem R, Geraghty AC, Ni L, Andreasson KI, Vitanza NA, Warren KE, Thomas CJ, Monje M (2019) Therapeutic strategies for diffuse midline glioma from high-throughput combination drug screening. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aaw0064
Anastas JN, Zee BM, Kalin JH, Kim M, Guo R, Alexandrescu S, Blanco MA, Giera S, Gillespie SM, Das J, Wu M, Nocco S, Bonal DM, Nguyen QD, Suva ML, Bernstein BE, Alani R, Golub TR, Cole PA, Filbin MG, Shi Y (2019) Re-programing Chromatin with a Bifunctional LSD1/HDAC Inhibitor Induces Therapeutic Differentiation in DIPG. Cancer Cell 36(5):528–544. https://doi.org/10.1016/j.ccell.2019.09.005
Caretti V, Bugiani M, Freret M, Schellen P, Jansen M, van Vuurden D, Kaspers G, Fisher PG, Hulleman E, Wesseling P, Vogel H, Monje M (2014) Subventricular spread of diffuse intrinsic pontine glioma. Acta Neuropathol 128(4):605–607. https://doi.org/10.1007/s00401-014-1307-x
Biery M, Myers, C., Girard, E., Morris, S., Carmack, S., Noll, A., Sarthy, J., Ferguson, E., Mhyre, A., Strand, A., Olson, J., Vitanza, N. (2018) DIPG-35. A novel HDAC inhibitor in new patient-derived diffuse intrinsic pontine glioma (DIPG) model Neuro Oncol 20:i56
Nagaraja S, Vitanza NA, Woo PJ, Taylor KR, Liu F, Zhang L, Li M, Meng W, Ponnuswami A, Sun W, Ma J, Hulleman E, Swigut T, Wysocka J, Tang Y, Monje M (2017) Transcriptional Dependencies in Diffuse Intrinsic Pontine Glioma. Cancer Cell 31(5):635–652. https://doi.org/10.1016/j.ccell.2017.03.011
Orskov AD, Treppendahl MB, Skovbo A, Holm MS, Friis LS, Hokland M, Gronbaek K (2015) Hypomethylation and up-regulation of PD-1 in T cells by azacytidine in MDS/AML patients: A rationale for combined targeting of PD-1 and DNA methylation. Oncotarget 6(11):9612–9626. https://doi.org/10.18632/oncotarget.3324
Booth L, Roberts JL, Poklepovic A, Kirkwood J, Dent P (2017) HDAC inhibitors enhance the immunotherapy response of melanoma cells. Oncotarget 8(47):83155–83170. https://doi.org/10.18632/oncotarget.17950
Topper MJ, Vaz M, Chiappinelli KB, DeStefano Shields CE, Niknafs N, Yen RC, Wenzel A, Hicks J, Ballew M, Stone M, Tran PT, Zahnow CA, Hellmann MD, Anagnostou V, Strissel PL, Strick R, Velculescu VE, Baylin SB (2017) Epigenetic Therapy Ties MYC Depletion to Reversing Immune Evasion and Treating Lung Cancer. Cell 171(6):1284–1300. https://doi.org/10.1016/j.cell.2017.10.022
Funding
This review was not funded.
Author information
Authors and Affiliations
Contributions
All authors contributed to the analysis and interpretation of reviewed data, contributed to the writing of the manuscript, and reviewed and approved the final version.
Corresponding author
Ethics declarations
Conflicts of interest
All authors declare no competing interests or conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cooney, T.M., Lubanszky, E., Prasad, R. et al. Diffuse midline glioma: review of epigenetics. J Neurooncol 150, 27–34 (2020). https://doi.org/10.1007/s11060-020-03553-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03553-1