Abstract
Tripartite motif (TRIM) proteins are involved in tumorigenesis. Here, we examined the expression, biological function, and clinical significance of tripartite motif containing 28 (TRIM28) in glioma, a locally aggressive brain tumor. First, TRIM28 expression was significantly higher in glioma (n = 138) than in non-glioma controls (n = 6). TRIM28 expression was positively correlated with tumor malignancy, and associated with poor overall survival (OS) and progression-free survival (PFS). Notably, TRIM28 expression was negatively correlated with p21 expression in patients with glioblastoma multiforme (GBM). A multivariate analysis that included relevant measures indicated that high TRIM28 expression is an independent prognostic factor for poor OS and PFS in GBM patients. In experiments with cultured glioma cells, down-regulating TRIM28 with shRNA increased p21 expression, and induced cell cycle arrest at the G1 phase. In a xenograft model, down-regulating TRIM28 suppressed tumor growth. These results indicate that over-expression of TRIM28 is associated with poor outcome in glioma patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gliomas, and glioblastoma multiforme (GBM) in particular, are locally invasive brain tumors with poor prognosis, even after treatment with temozolomide and/or radiation [1, 2].
E3 ligase, a member of the ubiquitin–proteasome system, has been implicated in tumorigenesis on the basis of regulation of oncoproteins and tumor inhibitors [3–6]. Tripartite motif (TRIM) family of proteins function as E3 ligases via their RING-finger domain [7]. More than 70 members of the TRIM family have been identified so far. Several of these factors have been associated with oncogenic processes, such as apoptosis, cell proliferation and transcriptional regulation [8]. Some of the TRIM gene family members (such as TRIM19 and TRIM24) are transported to other genes and play important roles in tumorigenesis and cancer progression [9, 10]. Other TRIM proteins, including TRIM25 and TRIM68, regulate nuclear receptor activation [8]. TRIM28 expression is elevated in a variety of tumors, including breast cancer, gastric cancer and lung cancer [11–16]. In a clinical study in patients with early-stage lung tumor, TRIM28 expression was correlated with overall survival (OS) [17]. In a mouse model for aging, TRIM28 knockdown could reverse senescence [18].
In the current study, we compared TRIM28 gene expression in glioma tissues acquired from surgical resection versus non-glioma tissues. Functional significance of TRIM28 was examined in cultured glioma cells and a xenograft model using shRNA gene silencing.
Materials and methods
Cell lines and culture
Four representative glioma cell lines (SHG44, GL261, U87 and U251: from the Chinese Academy of Sciences Shanghai Branch Bank of Cells) were cultured in DMEM medium containing 10 % fetal bovine serum (GIBCO; Carlsbad, CA, USA) as previously described [19].
Patients and follow-up
A total of 138 glioma tissue microarray (TMA) samples were obtained from patients who underwent curative resection between 2002 and 2009 in the Department of Neurosurgery, Huashan Hospital, Fudan University. The patient characteristics are shown in Table S1. Of them, 70 cases were GBM (Table S2). Six control brain specimens were randomly chosen from patients undergoing trauma/epilepsy surgery. Ethical approval was provided by the Ethics Committee of Huashan Hospital. Written informed consent was obtained from each patient.
Construction of the TMAs and immunohistochemistry (IHC)
Detailed description of the construction of the TMA and IHC are available in the Supplementary Materials and Methods.
Transfection, quantitative real-time polymerase chain reaction (qRT-PCR) and Western blotting
Detailed description of the transfection experiments with shRNA for TRIM28, qRT-PCR and Western blotting analysis are available in the Supplementary Materials and Methods.
Cell proliferation, colony formation, flow cytometry and xenograft experiments
Detailed description of the cell proliferation assay, colony formation assay, flow cytometry assay of the cell cycle and xenograft experiments are available in the Supplementary Materials and Methods.
Statistical analysis
Statistical analysis was performed with the SPSS 19.0 software program (SPSS, Chicago, IL, USA). Data are presented as the mean ± standard deviation. Student’s t-test and Fisher’s exact probability were used for comparisons between groups. OS and progression-free survival (PFS) were evaluated using the Kaplan–Meier method for univariate analysis, and differences were assessed using the log-rank test. For the multivariate analysis, Cox proportional hazards regression model was used. Values of p < 0.05 were considered statistically significant.
Results
Bioinformatics analysis of TRIM family expression
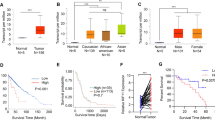
We searched the Cancer Genome Atlas (TCGA) for differential expression of TRIM family proteins in GBM tissue (n = 483) vs. normal brain tissue (n = 10) (Fig. 1a). The analysis using a nanobody-based reverse proteomics approach [20] revealed high TRIM28 expression in GBM stem-like cells [20]. Such a result was confirmed by an analysis of the Murat database (Fig. 1b).
TRIM28 is overexpressed in glioma and correlates with poor prognosis
Examination of TRIM28 expression in 11 cases of glioma (n = 4, 3, and 4 for grade II, III and IV) and 3 control tissues by qRT-PCR and Western blotting revealed significantly higher TRIM28 mRNA (p = 0.009) and protein (p = 0.034) in glioma (Fig. 2a, b). TRIM28 IHC and counterstaining with hematoxylin and eosin (H&E) revealed localization of TRIM28 in cell nuclei (Fig. 2c). Upon IHC analysis, the TRIM28 protein was substantially higher in glioma (n = 138) than in control tissues (n = 6) (p = 0.002, Fig. 2d). Higher TRIM28 expression was also evident in high-grade versus low-grade glioma (p = 0.009, Fig. 2d).
TRIM28 is overexpressed in glioma tissues and associated with poor prognosis. a, b A total of 11 cases of glioma (including 4 grade II, 3 grade III and 4 grade IV) and 3 control tissues evaluated by qRT-PCR (p = 0.009) and immunoblotting (p = 0.034). c Representative photomicrographs of the tumor sections following IHC analysis for TRIM28. d Elevated TRIM28 expression is correlated with the histological malignancy. Expression of TRIM28 was higher in glioma tissues than in control tissue (p = 0.002), and in high-grade glioma compared with low-grade glioma (p = 0.009). e Kaplan–Meier survival curves indicating the cumulative survival as a function of time for patients with high TRIM28 expression versus those with low expression (OS p = 0.009, PFS p = 0.001). The patients with high TRIM28 expression experience significantly worse outcomes. (*p < 0.05, **p < 0.01)
In the 70 GBM cases, high and low TRIM28 expression was noted in 32 and 38 cases, respectively. Both the OS (p = 0.009) and PFS (p = 0.001) differed between gliomas that expression high versus low levels of TRIM28. High TRIM28 expression was associated with poor prognosis (Fig. 2e). The 1- and 2-year OS was 76.3 and 56.3 %, respectively for GBM with low TRIM 28 expression, and 26.3 and 12.5 %, respectively for GBM with high TRIM28 expression. The 1- and 2-year PFS was 42.1 and 25.0 %, respectively for GBM with low TRIM28 expression, and 13.2 and 3.1 %, respectively for GBM with high TRIM28 expression.
The prognosis did not correlate with gender, sex, extent of resection, tumor location, or expression of Mind bomb E3 ubiquitin protein ligase 1 (MIB-1) and TP53 (Table 1).
A univariate analysis indicated better prognosis in patients with a Karnofsky performance status (KPS) at 70 or above (OS: p = 0.006; PFS: p = 0.035) (Table 2). In addition, reduced TRIM28 expression was associated with longer OS (p = 0.009) and PFS (p = 0.002) (Table 2). A multivariate analysis using Cox proportional hazard regression model confirmed poor OS (p = 0.026, HR = 1.776, 95 % CI 1.069–2.950) and PFS (p = 0.004, HR = 2.120, 95 % CI 1.263–2.350) in glioma with high TRIM28 expression. KPS at above 70 was also a prognostic factor for OS (p = 0.044, HR = 1.811 95 % CI 1.016–3.227) but not for PFS (p = 0.171).
TRIM28 knockdown decreases cell proliferation and induces G1 arrest
Examination of TRIM28 expression in 4 representative glioma cell lines by qRT-PCR and Western blot revealed more robust increase in TRIM28 expression in U251 and U87 cells versus in SHG44 and GL261 cells versus normal brain tissues (Fig. 3a).
TRIM28 down-regulation inhibited glioma cell proliferation. a qRT-PCR and immunoblotting analyses of the expression of TRIM28 in 4 glioma cell lines and control brain tissue. b Stably transfected glioma cells with decreased TRIM28 expression were established. Cell proliferation was assessed by an MTT assay. c The clone formation ability was impaired after TRIM28 interference. d TRIM28 down-regulation induces G1 arrest. e P53, p27, p21 and p16 were assessed by immunoblotting after the down-regulation of TRIM28 in glioma cells. f The level of TRIM28 expression affected the growth abilities of glioma cells in the xenograft model of nude mice, as determined by the tumor size. The error bars represent the s.d. (*p < 0.05, **p < 0.01, ***p < 0.001)
To functional significance of TRIM28 expression, we knocked down the expression of TRIM28 in U87 and U251 cells via transfection with pGMLV-GFP-vshRNA-TRIM28 (Fig. 3b). TRIM28 knockdown decreased cell proliferation (Fig. 3b) and clone formation (Fig. 3c) in U87 and U251 cells, and led to G1 arrest in glioma cells (Fig. 3d). TRIM28 knockdown significantly increased the expression of p21, slightly altered p53 and p16, but did not affect p27 expression (Fig. 3e). In a xenograft model using U87 cells, a decrease in TRIM28 expression inhibited tumor growth (Fig. 3f).
TRIM28 and p21 expression levels are independent prognostic markers in glioma patients
To examine the relationship between TRIM28 and p21, we examined the expression of TRIM28 and p21 in 70 GBM tissue samples (representative staining in Fig. 4a). The results indicated a negative correlation between the expression of TRIM28 and p21 (r = −0.2601, p = 0.0296, Fig. 4b). Kaplan–Meier survival curves failed to show significant difference in survival time between patients with positive versus negative p21 expression (Fig. 4c), but did reveal poor prognosis in patients with high level of TRIM28 and negative p21 expression (Fig. 4d).
Combination of TRIM28 overexpression and p21 down-regulation is associated with poor survival in glioma patients. a Representative immunostaining images of TRIM28 and p21. b The relationship between the protein levels of TRIM28 and those of p21 in glioma tissues. c Kaplan–Meier survival curves indicating no significance for survival time between patients with positive p21 expression versus those with negative expression (OS p = 0.087, PFS p = 0.251). d Kaplan–Meier survival curves indicating the cumulative survival as a function of time for patients with TRIM28 high/p21 negative expression versus those with TRIM28 low/p21 positive expression (OS p < 0.001, PFS p < 0.001). Glioma patients co-expressing TRIM28 high/p21 negative had the worst prognosis
Discussion
TRIM proteins could affect (either promote or inhibit) oncogenesis and tumor progression by affecting cellular physiological processes, such as DNA repair, cell proliferation and apoptosis. The biological functions of TRIM family members as oncogenes or tumor suppressor genes have been extensively studied. In the present study, we analyzed TCGA data and found that 18 TRIM genes were up-regulated and 16 were down-regulated in 483 GBM tissue samples compared with 10 control brain tissues. Among the 18 genes with elevated expression, we focused on TRIM28, a molecule previously known to play important roles in the oncogenesis of several tumors [11, 13].
The clinical part of the current study showed that TRIM28 expression is positively associated with malignancy of glioma and poor patient prognosis. Next, we selected 4 representative glioma cell lines with a range of TRIM28 expression levels to investigate the role played by TRIM28. Inhibiting TRIM28 expression in the 2 cell lines with high TRIM28 expression (e.g., U87 and U251 cells) with lentivirus-mediated pGMLV-GFP-shRNAs decreased cell proliferation and colony formation, and arrested cell cycle at the G1 phase. These results suggest that down-regulation of the TRIM28 protein plays an inhibitory role in the development of glioblastoma and may be a key regulator of the G1-S transition in glioblastoma cells. Extrapolation of the findings from cultured cells and xenograft models back into in vivo situations, however, must be attempted with caution.
TRIM28 is a member of the TRIM family of E3 ligases, and could activate or suppress transcription via various mechanisms under different contexts [21–24]. TRIM28 is known to be associated with the histone deacetylase complex NuRD and the histone methyltransferase SETDB1 and leads to the silencing of specific genes [25, 26]. Via other mechanisms, TRIM28 causes transcriptional activation through its recruitment to certain response elements, such as the Nur response element (NuRE) [23]. Santos et al. showed that in human embryonic lung fibroblast cell line IMR90, TRIM28 expression inhibit p16 induction without affecting p53 or p21 expression [27]. However, in our study, TRIM28 regulated the expression of p21 in glioma cells; such a finding was consistent with the negative correlation of TRIM28 with p21 in GBM tissues. P21 is a tumor suppressor gene and a cell cycle inhibitor. The down-regulation of p21 plays a vital role in the development of many cancers. The relationship between TRIM28 and p21 observed in the current study may partly explain the biological functions of TRM28. Lee et al. demonstrated that sumoylated TRIM28 decreased H3-K9 and H3-K14 acetylation and augments H3-K9 methylation at the p21 promoter [28]. Also a result, the downregulation of p21 may also be related to the sumoylation of TRIM28 via a chromatin-silencing process.
In our experiments, TRIM28 was detected in cell nuclei but not in cytoplasm. E3 ligases, including TRIM28, catalyzes ubiquitination of the proteins destined for degradation, and by doing so, serves as an important post-translational mechanism to regulate various cell functions, such as transcriptional regulation, protein quality control, DNA repair and cell cycle regulation. Protein degradation indeed occurs in cytoplasm, but ubiquitination may occur in cell nuclei. [29].
In conclusion, the results from the current study highlighted the biological and clinical significance of TRIM28 expression in glioma. Importantly, the combination of negative p21 expression and high TRIM28 expression could predict a poor prognosis in GBM. Decreasing TRIM28 expression could inhibit the growth of glioma cells both in vitro and in vivo, suggesting that targeting TRIM28 could be a promising therapeutic approach to manage gliomas.
Abbreviations
- TRIM:
-
Tripartite motif
- TRIM28:
-
Tripartite motif containing 28
- OS:
-
Overall survival
- PFS:
-
Progression-free survival
- GBM:
-
Glioblastoma multiforme
- TMA:
-
Tissue microarray
- IHC:
-
Immunohistochemistry
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- TCGA:
-
The Cancer Genome Atlas
- MIB-1:
-
Mind bomb E3 ubiquitin protein ligase 1
- KPS:
-
Karnofsky performance status
- MGMT:
-
O-6-methylguanine-DNA methyltransferase
- STR:
-
Subtotal resection
- GTR:
-
Gross total resection
References
Dolecek TA, Propp JM, Stroup NE, Kruchko C (2012) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2005–2009. Neuro-oncology 14(Suppl 5):v1–v49
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, Gorlia T, Allgeier A, Lacombe D et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996
Weissman AM (1997) Regulating protein degradation by ubiquitination. Immunol Today 18(4):189–198
Koepp DM, Schaefer LK, Ye X, Keyomarsi K, Chu C, Harper JW, Elledge SJ (2001) Phosphorylation-dependent ubiquitination of cyclin E by the SCFFbw7 ubiquitin ligase. Science 294(5540):173–177
Onoyama I, Tsunematsu R, Matsumoto A, Kimura T, de Alboran IM, Nakayama K, Nakayama KI (2007) Conditional inactivation of Fbxw7 impairs cell-cycle exit during T cell differentiation and results in lymphomatogenesis. J Exp Med 204(12):2875–2888
Tsunematsu R, Nakayama K, Oike Y, Nishiyama M, Ishida N, Hatakeyama S, Bessho Y, Kageyama R, Suda T, Nakayama KI (2004) Mouse Fbw7/Sel-10/Cdc4 is required for notch degradation during vascular development. J Biol Chem 279(10):9417–9423
Reymond A, Meroni G, Fantozzi A, Merla G, Cairo S, Luzi L, Riganelli D, Zanaria E, Messali S, Cainarca S, Guffanti A, Minucci S, Pelicci PG, Ballabio A (2001) The tripartite motif family identifies cell compartments. EMBO J 20(9):2140–2151
Hatakeyama S (2011) TRIM proteins and cancer. Nat Rev Cancer 11(11):792–804
de The H, Lavau C, Marchio A, Chomienne C, Degos L, Dejean A (1991) The PML-RAR alpha fusion mRNA generated by the t(15;17) translocation in acute promyelocytic leukemia encodes a functionally altered RAR. Cell 66(4):675–684
Le Douarin B, Zechel C, Garnier JM, Lutz Y, Tora L, Pierrat P, Heery D, Gronemeyer H, Chambon P, Losson R (1995) The N-terminal part of TIF1, a putative mediator of the ligand-dependent activation function (AF-2) of nuclear receptors, is fused to B-raf in the oncogenic protein T18. EMBO J 14(9):2020–2033
Beer DG, Kardia SL, Huang CC, Giordano TJ, Levin AM, Misek DE, Lin L, Chen G, Gharib TG, Thomas DG, Lizyness ML, Kuick R, Hayasaka S, Taylor JM, Iannettoni MD, Orringer MB et al (2002) Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat Med 8(8):816–824
Landi MT, Dracheva T, Rotunno M, Figueroa JD, Liu H, Dasgupta A, Mann FE, Fukuoka J, Hames M, Bergen AW, Murphy SE, Yang P, Pesatori AC, Consonni D, Bertazzi PA, Wacholder S et al (2008) Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS One 3(2):e1651
Ho J, Kong JW, Choong LY, Loh MC, Toy W, Chong PK, Wong CH, Wong CY, Shah N, Lim YP (2009) Novel breast cancer metastasis-associated proteins. J Proteome Res 8(2):583–594
Su LJ, Chang CW, Wu YC, Chen KC, Lin CJ, Liang SC, Lin CH, Whang-Peng J, Hsu SL, Chen CH, Huang CY (2007) Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics 8:140
Hou J, Aerts J, den Hamer B, van Ijcken W, den Bakker M, Riegman P, van der Leest C, van der Spek P, Foekens JA, Hoogsteden HC, Grosveld F, Philipsen S (2010) Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS One 5(4):e10312
Yokoe T, Toiyama Y, Okugawa Y, Tanaka K, Ohi M, Inoue Y, Mohri Y, Miki C, Kusunoki M (2010) KAP1 is associated with peritoneal carcinomatosis in gastric cancer. Ann Surg Oncol 17(3):821–828
Chen L, Chen DT, Kurtyka C, Rawal B, Fulp WJ, Haura EB, Cress WD (2012) Tripartite motif containing 28 (Trim28) can regulate cell proliferation by bridging HDAC1/E2F interactions. J Biol Chem 287(48):40106–40118
Liu B, Wang Z, Ghosh S, Zhou Z (2013) Defective ATM-Kap-1-mediated chromatin remodeling impairs DNA repair and accelerates senescence in progeria mouse model. Aging Cell 12(2):316–318
Mo LJ, Ye HX, Mao Y, Yao Y, Zhang JM (2013) B7-H4 expression is elevated in human U251 glioma stem-like cells and is inducible in monocytes cultured with U251 stem-like cell conditioned medium. Chin J Cancer 32(12):653–660
Jovcevska I, Zupanec N, Kocevar N, Cesselli D, Podergajs N, Stokin CL, Myers MP, Muyldermans S, Ghassabeh GH, Motaln H, Ruaro ME, Bourkoula E, Turnsek TL, Komel R (2014) TRIM28 and beta-actin identified via nanobody-based reverse proteomics approach as possible human glioblastoma biomarkers. PLoS One 9(11):e113688
Friedman JR, Fredericks WJ, Jensen DE, Speicher DW, Huang XP, Neilson EG, Rauscher FJ III (1996) KAP-1, a novel corepressor for the highly conserved KRAB repression domain. Genes Dev 10(16):2067–2078
Wang C, Rauscher FJ III, Cress WD, Chen J (2007) Regulation of E2F1 function by the nuclear corepressor KAP1. J Biol Chem 282(41):29902–29909
Rambaud J, Desroches J, Balsalobre A, Drouin J (2009) TIF1beta/KAP-1 is a coactivator of the orphan nuclear receptor NGFI-B/Nur77. J Biol Chem 284(21):14147–14156
Chang CJ, Chen YL, Lee SC (1998) Coactivator TIF1beta interacts with transcription factor C/EBPbeta and glucocorticoid receptor to induce alpha1-acid glycoprotein gene expression. Mol Cell Biol 18(10):5880–5887
Schultz DC, Ayyanathan K, Negorev D, Maul GG, Rauscher FJ III (2002) SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16(8):919–932
Schultz DC, Friedman JR, Rauscher FJ III (2001) Targeting histone deacetylase complexes via KRAB-zinc finger proteins: the PHD and bromodomains of KAP-1 form a cooperative unit that recruits a novel isoform of the Mi-2alpha subunit of NuRD. Genes Dev 15(4):428–443
Santos J, Gil J (2014) TRIM28/KAP1 regulates senescence. Immunol Lett 162(1 Pt B):281–289
Lee YK, Thomas SN, Yang AJ, Ann DK (2007) Doxorubicin down-regulates Kruppel-associated box domain-associated protein 1 sumoylation that relieves its transcription repression on p21WAF1/CIP1 in breast cancer MCF-7 cells. J Biol Chem 282(3):1595–1606
Hoeller D, Hecker CM, Dikic I (2006) Ubiquitin and ubiquitin-like proteins in cancer pathogenesis. Nat Rev Cancer 6(10):776–788
Acknowledgments
This work was supported by China National Funds for Distinguished Young Scientists (81025013 to Ying Mao), National Natural and Science Foundation of China (81402053) and International Scientific and Technological Cooperation Projects (2014DFA31470 to Wei Zhu).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None.
Additional information
Zeng-Xin Qi and Jia-Jun Cai have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qi, ZX., Cai, JJ., Chen, LC. et al. TRIM28 as an independent prognostic marker plays critical roles in glioma progression. J Neurooncol 126, 19–26 (2016). https://doi.org/10.1007/s11060-015-1897-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-015-1897-8