Abstract
Long non-coding RNAs (lncRNAs), a recently discovered class of non-coding genes, are transcribed throughout the genome. Emerging evidence suggests that lncRNAs may be involved in modulating various aspects of tumor biology, including regulating gene activity in response to external stimuli or DNA damage. No data are available regarding the expression of lncRNAs during genotoxic stress-induced apoptosis and/or necrosis in human glioma cells. In this study, we detected a change in the expression of specific candidate lncRNAs (neat1, GAS5, TUG1, BC200, Malat1, MEG3, MIR155HG, PAR5, and ST7OT1) during DNA damage-induced apoptosis in human glioma cell lines (U251 and U87) using doxorubicin (DOX) and resveratrol (RES). We also detected the expression pattern of these lncRNAs in human glioma cell lines under necrosis induced using an increased dose of DOX. Our results reveal that the lncRNA expression patterns are distinct between genotoxic stress-induced apoptosis and necrosis in human glioma cells. The sets of lncRNA expressed during genotoxic stress-induced apoptosis were DNA-damaging agent-specific. Generally, MEG3 and ST7OT1 are up-regulated in both cell lines under apoptosis induced using both agents. The induction of GAS5 is only clearly detected during DOX-induced apoptosis, whereas the up-regulation of neat1 and MIR155HG is only found during RES-induced apoptosis in both cell lines. However, TUG1, BC200 and MIR155HG are down regulated when necrosis is induced using a high dose of DOX in both cell lines. In conclusion, our findings suggest that the distinct regulation of lncRNAs may possibly involve in the process of cellular defense against genotoxic agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioma is the most common primary brain tumor in humans, and glioblastoma multiforme (GBM) is one of the most aggressive and lethal brain tumors. Five-year survival rates are less than 5 % for glioblastoma [1]. Identifying the genes that may be critically associated with oncogenesis and their function, as demonstrated by further examination, will contribute to the discovery of a therapeutic target for glioma.
RNA is not only a messenger between DNA and proteins. Only 10 % of the transcribed RNA encodes for proteins. The other 90 % is described as non-coding RNA,in which transcripts that are longer than 200 bp are defined as long non-coding RNA (lncRNA) [2]. According to the proximity between neighboring transcripts, lncRNAs can be classified as sense, antisense, bidirectional, intronic, and intergenic [3]. Gene expression profiling indicates that certain lncRNAs influence several biological aspects, including the nuclear architecture, regulation of gene expression, and embryonic stem cell pluripotency, although the relevant mechanisms remain poorly understood [4, 5]. Increasing evidence links mutations and the dysregulation of lncRNAs to diverse human diseases, including a variety of human cancers [6, 7].
Recently, emerging evidence reveals that the transcription of lncRNAs can modulate gene activity in response to external stimuli or DNA damage [8, 9]. Other experiments demonstrate that some genotoxic-responsive lncRNAs function as tumor suppressors or oncogenes. For example, doxorubicin (DOX) increased the level of the lincRNA RoR (lincRNA-Regulator of Reprogramming) in HCT-116 cells expressing wild type p53 (HCT-116 WT cells). Further evidence demonstrate that RoR plays an important role in the p53-mediated tumor suppression network [10]. The lncRNA loc285194 is induced in response to DOX in HCT-116 WT cells. Further results suggest that loc285194 is a p53-regulated tumor suppressor [11].
However, the differential expression of lncRNAs during genotoxic induced stress in human glioma cells remains to be demonstrated. In this study, we aimed to investigate the change in the expression of 9 lncRNAs during genotoxic stress in human glioma cells. Although there are many lncRNAs that can be transcribed from the human genome, a relatively small number of lncRNAs have been shown to be associated with human cancer. Therefore, we focused on these tumor-related and/or brain disease-related lncRNA that have been validated in human cancer tissues and/or in the human central nervous system. PubMed and lncRNAdb were searched to retrieve these 9 candidate lncRNAs. Our findings provide novel information on the lncRNA expression profiles, which may be altered by genotoxic stress-induced cell death.
Materials and methods
Reagents
Resveratrol (RES) and DOX were purchased from Sigma Chemical Co. (St. Louis, MO, USA) and were dissolved in DMSO at a stock concentration of 100 mmol/L and 1 mmol/L, respectively. They were further diluted in DMEM containing 10 % fetal bovine serum to the appropriate final concentrations. Control experiments utilized similar volumes of DMSO alone.
Study design
In this study, we aimed to detect the differential expression of 9 candidate lncRNAs in response to genotoxic stress in human glioma cells. For this purpose, we chose two human glioma cell lines (U251 and U87) and two different genotoxic agents (DOX and RES). DOX is one of the most widely used chemotherapeutic agents, and it can induce cell death via DNA damage [12]. RES can induce DNA damage and inhibit different stages of carcinogenesis, including tumor initiation, promotion and progression [13]. Treatment of human glioma cells with DOX or RES has been confirmed to induce apoptosis [14, 15]. Cell death can occur via a spectrum of morphologically and biochemically distinct pathways, including apoptosis and necrosis. A specific concentration of a drug or different intensity of the stimulus may induce a particular pathway of cell death. We hypothesized that these different concentrations of DOX and RES induce different pathways of cell death. lncRNAs in these cell lines may exhibit distinct responses during different pathways of cell death.
Cell culture
The U251 and U87 cells were generous gifts from Dr. Lv (Department of Neurosurgery, Xinqiao Hospital, The Third Military Medical University, Chongqing, P.R. China), and these cells were grown in DMEM culture medium (Gibco). All culture media were supplemented with 10 % FBS (Gibco), 2 mM glutamine, and 100 U/ml penicillin. The cells were incubated at 37 °C in a humidified chamber supplemented with 5 % CO2.
Apoptosis detection via Annexin V/propidium iodide (PI) staining
Apoptosis was analyzed by double-staining with FITC-labeled Annexin V and PI, using an Annexin V-FITC apoptosis detection kit (BD Biosciences, San Jose, CA, USA) according to the manufacturer’s instructions. Briefly, 2 × 106 cells were harvested using EDTA-free trypsin, washed twice with cold PBS and resuspended in 100 µL of binding buffer. Annexin V-FITC (5 µL) and PI (5 µL) were added to each sample. Then, the samples were incubated for 15 min at room temperature in the dark. Immediately afterwards, flow cytometry analysis was performed (FACSCalibur,BD Biosciences).
In situ terminal-deoxytransferase-mediated dUTP nick end labeling (TUNEL) assay
The TUNEL assay was performed to detect apoptosis using the In Situ Cell Death Detection Kit (Roche Molecular Biochemicals, Indianapolis, IN, USA). Briefly, U251 and U87 cells on coverslips were treated with RES (100 μM) for 48 h or DOX (1 μM) for 24 h, fixed in 4 % phosphate-buffered paraformalin for 60 min and then washed with PBS. The cells were incubated in blocking solution for 10 min at room temperature and incubated in permeabilisation solution for 2 min. Then, the cells were washed twice with PBS and incubated in 50 µL of the TUNEL reaction mixture for 60 min at 37 °C. After incubation, the sections were washed with PBS. The cells were counterstained with DAPI in the dark for 30 min and visualized using an Olympus microscope (Tokyo, Japan). The percentage of TUNEL-positive cells was determined by counting at least 200 cells in 5 randomly selected visual fields (0.4 mm2) on each coverslip.
Hoechst 33342 and PI double-staining
Apoptosis and necrosis were distinguished via PI and Hoechst 33342 double staining using the Apoptotic Cell Hoechst 33342/PI Detection Kit (KeyGENE Biotech, Nanjing, China) according to the manufacturer’s instructions. The U251 and U87 cells were seeded on 12-well plates (1 × 10 4 cells/well) and treated with different concentrations of DOX. After incubation, the cells were washed with PBS and then stained with Hoechst 33342 (10 µL) for 15 min at room temperature. The cells were washed with PBS and then incubated in 400 µL of 1 × buffer A and PI (5 µL) for 10 min at room temperature. The presence of fluorescence was immediately verified.
Quantitative real-time polymerase chain reaction (PCR)
Total RNA was isolated from 5–10 × 106 cells using an RNAiso Plus Kit (Takara, Japan). Real time PCR was performed using a BioRad CFX96 system according to the protocol for SYBR® Premix Ex Taq™ (Perfect Real Time) (Takara, Japan). The thermal cycling conditions included an initial denaturation step at 95 °C for 30 s and 35 cycles at 95 °C for 10 s followed by 60 °C for 30 s. Primers for neat1, GAS5, TUG1, BC200, Malat1, MEG3, MIR155HG, PAR5, ST7OT1 and GAPDH were obtained from the Human Disease-Related LncRNA Profiler [CAT# RA920D, System Biosciences (SBI, USA)]. The results of three independent experiments were evaluated to calculate the average value and were standardized to the housekeeping gene GAPDH. All samples were examined in triplicate. Comparative quantification was performed according to the 2−ΔΔCt method.
Statistical analysis
Statistical analysis was performed using the statistical program SPSS 13.0. The quantitative results were expressed as the mean ± standard error (SE) and were analyzed using Student’s t test. P values of <0.05 were considered to be statistically significant.
Results
DOX and RES induce apoptosis in U251 and U87 cells
We performed an Annexin V-FITC/PI double-staining experiment to determine the levels of apoptosis induced by DOX and RES in human glioma cell lines. As shown in Fig. 1, after treatment with RES (100 µM) for 48 h, the percentage of early apoptotic cells significantly increased in both U251 and U87 cells. However, a considerable increase in the number of late apoptotic cells and necrotic cells was detected in U251 and U87 cells treated with DOX (1 µM) for 24 h. These results indicate that RES primarily induced early apoptosis and DOX caused more cells to enter into a late apoptotic or necrotic phase.
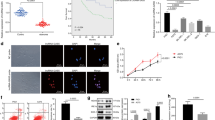
DOX and RES induce apoptosis in U251 and U87 cells based on flow cytometric analysis. A The Annexin V-FITC positive cells in the top (PI positive) and bottom (PI negative) right quadrants are shown. The U87 cells were used as control cells (a) and were treated with 100 µM (b) RES for 48 h or 1 µM (c) DOX for 24 h. The U251 cells were used control cells (d) and were treated with 100 µM (e) RES or 1 µM (f) DOX. B The histogram displays the number of early apoptotic, late apoptotic and necrotic cells induced using RES or DOX. **P < 0.01
Validation of the FACS results via TUNEL staining assays
The TUNEL assay is an invaluable technique for examining late-stage apoptosis. The apoptotic cells display red fluorescently stained nuclei, whereas the non-apoptotic cells are not stained. As shown in Fig. 2, after treatment of the U87 cells with 1 μM DOX for 24 h or 100 μM RES for 48 h, the number of DNA fragmentation-dependent apoptotic cells relative to that of the control cells was significantly increased (Fig. 2A). The experiment was repeated using U251 cells in vitro, with similar results (Fig. 2B). Moreover, the number of TUNEL-positive apoptotic cells after DOX treatment was higher than that after RES treatment in both the U251 and U87 cells, demonstrating that DOX induces more late-stage apoptosis in human glioma cells, which is consistent with the flow cytometry results.
TUNEL assay using U251 and U87 cells to detect DOX- and RES-induced apoptosis. A, B Both cell lines were treated with 100 µM RES for 48 h or 1 µM DOX for 24 h. The representative red fluorescence images were captured under a microscope after the treatment. C The percentage of red fluorescent cells after DOX or RES treatment compared to the control treatment. **P < 0.01. (Color figure online)
Apoptosis and necrosis detection via Hoechst 33342/PI double-staining
To determine whether DOX displays a necrosis-inducing activity in glioma cell lines, Hoechst 33342 and PI double-staining assays were performed. U251 and U87 cells were treated with DOX at various doses and durations and then double-stained with Hoechst 33342 and PI. Hoechst stains the nuclei of cells containing either a disrupted or intact cell membrane, whereas PI only stains the nuclei of cells containing a disrupted cell membrane. Figure 3 shows the nuclear morphology of cell death visualized via fluorescence microscopy using Hoechst 33342/PI staining following exposure to DOX at 1 µM, or 2 µM for 24 h or at 1 µM for 48 h. The nuclei of normal control cells displayed round blue staining. However, at the increased exposure time, shrunken, condensed nuclei, a marker of apoptosis, displayed brighter blue fluorescence in both U251 and U87 cells (Fig. 3A). Moreover, the percentage of necrotic cells increased with the increasing of concentration of DOX in both cell lines (Fig. 3A). These morphologic findings suggested that treatment of U251 and U87 cells with DOX resulted in apoptosis in a time-dependent manner, and a higher concentration of DOX caused necrosis in both cell lines. Therefore, we evaluated the effects of 100 µM RES or 1 µM DOX for 24 h or 48 h on the induction of apoptosis in subsequent experiments. Moreover, 2 µM DOX for 24 h was selected as an inducer of necrosis for the profiling experiments.
Nuclear morphology of various types of cell death visualized via fluorescence microscopy using Hoechst33342/PI staining following exposure to DOX. A Both cell lines were treated with 1 µM or 2 µM DOX for 24 h or 1 µM DOX for 48 h. B The percentage of apoptotic cells and necrotic cells after treatment compared to the control. **P < 0.01
Genotoxic stress-induced expression of lncRNAs in human glioma cells
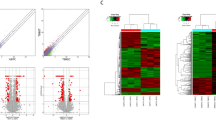
We determined the expression of nine candidate lncRNAs following apoptosis-inducing treatment in U251 and U87 cells using DOX or RES. According to the above results, we used 100 µM RES or 1 µM DOX for 24 h or 48 h for apoptosis induction. Under the apoptotic conditions, MEG3 and ST7OT1 were up-regulated in both cell types in response to both DNA-damaging agents in a time-dependent manner, whereas TUG1, BC200, Malat1, and PAR5 displayed no significant changes in expression (Fig. 4). The expression of both ST7OT1 and MEG3 was more pronounced in the U251 cells than in the U87 cells following treatment with DOX. A similar pattern was observed when both cell lines were treated with RES (Fig. 4).
The expression level of 9 lncRNAs during genotoxic stress-induced apoptosis in U87 and U251 cells. The examined genes include neat1, GAS5, TUG1, BC200, Malat1, PAR5, MIR155HG, MEG3, and ST7OT1. A, B Both cells were treated with doxo at 1 µM for 24 h or 48 h before extraction of total RNA for qRT-PCR. C, D Both cells were treated with RES at 100 µM for 24 h or 48 h before extraction of total RNA for qRT-PCR.**P < 0.01
Interestingly, the response of some lncRNAs to genotoxic stress was DNA-damaging agent-specific. The up-regulation of GAS5 was only detected in both cell lines treated with DOX. The increase in GAS5 expression was more pronounced in the U251 cells than in the U87 cells (Fig. 4A, B). For neat1 and MIR155HG, a statistically significant increase was detected in a time-dependent manner in both cell lines treated with RES (Fig. 4C, D), but not those treated with DOX.
To investigate whether these lncRNAs respond differentially during necrotic stress compared with apoptotic stress, we also analyzed the expression level of these lncRNAs in both cell lines treated with a high concentration of DOX. The down-regulation of TUG1, BC200 and MIR155HG was detected when necrosis was induced using a high dose of DOX in both cell lines, but no change in expression was detected for the other genes (Fig. 5).
The expression level of 9 lncRNAs during genotoxic stress-induced necrosis in U87 and U251 cells. The examined genes include neat1, GAS5, TUG1, BC200, Malat1, PAR5, MIR155HG, MEG3, and ST7OT1. A U87 cells were treated with doxo at 1 µM or 2 µM for 24 h before extraction of total RNA for qRT-PCR. B U251 cells were treated with doxo at 1 µM or 2 µM for 24 h before extraction of total RNA for qRT-PCR. **P < 0.01
DISCUSSION
LncRNAs have recently been found to be pervasively transcribed throughout the genome. Alterations in the primary structure, secondary structure and expression levels of lncRNAs are often associated with human diseases, particularly cancer [6]. Numerous reports have identified a specific set of lncRNAs that function in the DNA damage response [8, 16]. However, no data are available on the alteration of the expression of these lncRNAs during genotoxic stress-induced cell death in human glioma cells. In this report, we demonstrated that the expression patterns of lncRNAs are altered during two different types of cell death induced using genotoxic agents in human glioma cells. Additionally, we found not only universally altered lncRNAs, but also DNA-damaging agent-specific lncRNAs in human glioma cells during apoptosis or necrosis induced using DOX or RES.
Although both drugs exert genotoxic effects, their specific mechanisms are different. Resveratrol is a polyphenolic compound highly enriched in a wide variety of plant products, including red grapes, exhibits anticancer [17], antioxidant [18], and anti-inflammatory properties [19]. Resveratrol has been reported to elicit many cellular responses including cell cycle arrest, differentiation, and apoptosis in various cancer cell lines, including human U251 and U87 glioma cells [15, 20]. The biological effects of DOX include the induction of DNA damage, the impairment of mitochondrial function, and cell death. DOX is involved in the activation of the apoptotic pathway in malignant glioma cells in vitro [21]. Different stresses or stimuli can exert distinct effects on the same cell line or tissue. At high concentrations, intense stimuli may induce necrosis rather than apoptosis. We investigated the effects of DOX and RES on human glioma cells to determine which stage of apoptosis or necrosis was induced by these genotoxic agents. We confirmed that both damaging agents induce apoptosis in human glioma cells. However, DOX causes significant late-stage apoptosis,whereas RES primarily induces early apoptosis. Moreover, at higher concentrations, DOX primarily causes necrosis in both cell lines.
The pattern of expression of these lncRNAs during genotoxic stress-induced apoptosis was DNA-damaging agent-specific. Under apoptotic conditions, we found that some lncRNAs (MEG3, ST7OT1, GAS5, neat1, and MIR155HG) were up-regulated, whereas others (TUG1, BC200, MALAT1, and PAR5) were not affected. The up-regulated lncRNAs may be involved in apoptosis-mediated biological processes that regulate other genes required for apoptosis. Interestingly, both MEG3 and ST7OT1 were induced during apoptosis in both cell lines treated with DOX or RES. These data suggest that ST7OT1 and MEG3 are involved in the cellular responses to DNA-damaging agents in human glioma cells. MEG3 acts as a tumor suppressor gene in a variety of human tumors. Ectopic expression of MEG3 inhibits cell proliferation and promotes apoptosis in human glioma cell lines [22]. Recent data demonstrated the clear induction of MEG3 by genotoxic stress in MCF-7 cells [8]. Our findings corroborate the possibility that genotoxic responsive-lncRNAs may function as tumor suppressors or oncogenes. ST7OT1 overlaps with exon 1 and the promoter of RAY1/ST7, which has been reported to be a suppressor of tumorigenicity [23]. Because the trend of the altered expression of ST7OT1 was similar to that of MEG3, we speculate that ST7OT1 may function as a tumor suppressor in human glioma cells. Although MEG3 and ST7OT1 are both up-regulated under apoptotic conditions, no changes were detected under necrotic conditions induced using a high dose of DOX. Potentially, these lncRNAs are involved in an apoptotic signaling pathway rather than a necrotic signaling pathway.
The up-regulation of GAS5 was restricted to DOX-treated cells, whereas the up-regulation of neat1 and MIR155HG was confined to RES-treated cells. The differential expression of lncRNAs induced by different drugs is likely related to the particular stage of apoptosis or the distinct pathway induced by each drug. Emerging evidence suggests that GAS5 regulates cell proliferation and promotes apoptosis [24–26]. Our findings are consistent with the induction of GAS5 using certain apoptotic stimuli in HeLa cells [8]. These results suggest that GAS5 may be involved in a specific process that mediates the mechanism of apoptosis. Neat1 is induced during the differentiation of various cell lineages, and its transcription is necessary for the formation of a nuclear structure termed the paraspeckle [27, 28]. MIR155HG encompasses the pre-miR155 sequence, and elevated levels of MIR155HG are associated with elevated levels of miR-155, which is considered to be an onco-miRNA [29, 30]. The MIR155HG transcript, which accumulates in lymphoma cells, participates in oncogenesis [31]. Neat1 and MIR155HG may play roles in maintaining the integrity of DNA during apoptotic stress. However, we detected no remarkable change in GAS5 or neat1 expression, whereas MIR155HG was down-regulated under necrotic conditions in both cell lines. Increased stress may have initiated the necrosis-related machinery, thus down-regulating the expression of MIR155HG.
Despite the altered expression of MIR155HG, we still detected the down-regulation of TUG1 and BC200. Evidence shows that TUG1 binds to the PRC2 epigenetic regulatory complex and plays a role in repressing specific genes involved in cell-cycle regulation [32]. BC200 is selectively expressed in the central nervous system and targets eIF4A activity, resulting in the decoupling of ATP hydrolysis from RNA duplex unwinding [33]. However, BC200 is up-regulated in breast, ovarian, cervical, lung and other cancers compared with corresponding normal tissues [34]. In our study, we detected a decrease in TUG1 and BC200 expression in response to high stress despite no change in the expression of these lncRNAs during apoptosis. It is plausible that TUG1 and BC200 are only involved in necrosis rather than apoptosis. Considering the oncogenic potential of MIR155HG, TUG1, and BC200, the down-regulation of these molecules under stress conditions may contribute to the maintenance of chromatin stability and may helpfully avoid genomic instability.
In summary, our data suggest that these lncRNA expression levels are altered during genotoxic stress, indicating that these lncRNAs are involved in the genetic regulation of cellular stress responses. These lncRNAs may play roles in cellular defense mechanisms against individual genotoxic agents. These lncRNAs may cause cells to be more flexible in their responses to external stimuli. Although the functions of these identified lncRNAs in human glioma cells remain unknown, their ability to respond to genotoxic stress suggests the need for further investigation. Future studies are necessary to elucidate the function and mechanisms of the regulation of individual lncRNAs in human glioma cells.
References
Ostrom QT, Gittleman H, Farah P, Ondracek A, Chen Y, Wolinsky Y, Stroup NE, Kruchko C, Barnholtz-Sloan JS (2013) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2006-2010. Neuro-oncol 15(suppl 2):ii1–ii56. doi:10.1093/neuonc/not151
Costa FF (2010) Non-coding RNAs: meet thy masters. BioEssays 32(7):599–608. doi:10.1002/bies.200900112
Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. doi:10.1016/j.cell.2009.02.006
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10:155–159. doi:10.1038/nrg2521
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227. doi:10.1038/nature07672
Wapinski O, Chang HY (2011) Long noncoding RNAs and human disease. Trends Cell Biol 21:354–361. doi:10.1016/j.tcb.2011.04.001
Huarte M (2010) Large non-coding RNAs: missing links in cancer? Hum Mol Genet 19:152–161
Ozgur E, Mert U, Isin M, Okutan M, Dalay N, Gezer U (2013) Differential expression of long non-coding RNAs during genotoxic stress-induced apoptosis in HeLa and MCF-7 cells. Clin Exp Med 13:119–126. doi:10.1007/s10238-012-0181-x
Mizutani R, Wakamatsu A, Tanaka N, Yoshida H, Tochigi N, Suzuki Y, Oonishi T, Tani H, Tano K, Ijiri K, Isogai T, Akimitsu N (2012) Identification and characterization of novel genotoxic stress-inducible nuclear long noncoding RNAs in mammalian cells. PLoS One 7(4):e34949. doi:10.1371/journal.pone.0034949
Zhang A, Zhou N, Huang J, Liu Q, Fukuda K, Ma D, Lu Z, Bai C, Watabe K, Mo Y-Y (2013) The human long non-coding RNA-RoR is a p53 repressor in response to DNA damage. Cell Res 23:340–350. doi:10.1038/cr.2012.164
Liu Q, Huang J, Zhou N, Zhang Z, Zhang A, Lu Z, Wu F, Mo Y-Y (2013) LncRNA loc285194 is a p53-regulated tumor suppressor. Nucleic Acids Res 41:4976–4987. doi:10.1093/nar/gkt182
Hurley LH (2002) DNA and its associated processes as targets for cancer therapy. Nat Rev Cancer 2:188–200. doi:10.1038/nrc749
Delmas D, Lancon A, Colin D, Jannin B, Latruffe N (2006) Resveratrol as a chemopreventive agent: a promising molecule for fighting cancer. Curr Drug Targ 7:423–442. doi:10.2174/138945006776359331
Gopinath S, Vanamala SK, Gujrati M, Klopfenstein JD, Dinh DH, Rao JS (2009) Doxorubicin-mediated apoptosis in glioma cells requires NFAT3. Cell Mol Life Sci 66:3967–3978. doi:10.1007/s00018-009-0157-5
Jiang H, Zhang L, Kuo J, Kuo K, Gautam SC, Groc L, Rodriguez AI, Koubi D, Hunter TJ, Corcoran GB, Seidman MD, Levine RA (2005) Resveratrol-induced apoptotic death in human U251 glioma cells. Mol Cancer Ther 4:554–561. doi:10.1158/1535-7163.mct-04-0056
Hung T, Wang Y, Lin MF, Koegel AK, Kotake Y, Grant GD, Horlings HM, Shah N, Umbricht C, Wang P, Wang Y, Kong B, Langerod A, Borresen-Dale A-L, Kim SK, van de Vijver M, Sukumar S, Whitfield ML, Kellis M, Xiong Y, Wong DJ, Chang HY (2011) Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet 43:621. doi:10.1038/ng.848
Atten MJ, Godoy-Romero E, Attar BM, Milson T, Zopel M, Holian O (2005) Resveratrol regulates cellular PKC alpha and delta to inhibit growth and induce apoptosis in gastric cancer cells. Invest New Drugs 23:111–119. doi:10.1007/s10637-005-5855-8
Ovesna Z, Kozics K, Bader Y, Saiko P, Handler N, Erker T, Szekere T (2006) Antioxidant activity of resveratrol, piceatannol and 3,3’,4,4’,5,5’-hexahydroxy-trans-stilbene in three leukemia cell lines. Oncol Rep 16:617–624
Notas G, Nifli A-P, Kampa M, Vercauteren J, Kouroumalis E, Castanas E (2006) Resveratrol exerts its antiproliferative effect on HepG2 hepatocellular carcinoma cells, by inducing cell cycle arrest, and NOS activation. Biochim Biophys Acta 1760:1657–1666. doi:10.1016/j.bbagen.2006.09.010
Joe AK, Liu H, Suzui M, Vural ME, Xiao D, Weinstein IB (2002) Resveratrol induces growth inhibition, S-phase arrest, apoptosis, and changes in biomarker expression in several human cancer cell lines. Clin Cancer Res 8:893–903
Wolff JE, Trilling T, Molenkamp G, Egeler RM, Jurgens H (1999) Chemosensitivity of glioma cells in vitro: a meta analysis. J Cancer Res Clin Oncol 125:481–486. doi:10.1007/s004320050305
Wang P, Ren Z, Sun P (2012) Overexpression of the long non-coding RNA MEG3 impairs in vitro glioma cell proliferation. J Cell Biochem 113:1868–1874. doi:10.1002/jcb.24055
Vincent JB, Petek E, Thevarkunnel S, Kolozsvari D, Cheung J, Patel M, Scherer SW (2002) The RAY1/ST7 tumor-suppressor locus on chromosome 7q31 represents a complex multi-transcript system. Genomics 80:283–294. doi:10.1006/geno.2002.6835
Kino T, Hurt DE, Ichijo T, Nader N, Chrousos GP (2010) Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci Signal 3:ra8. doi:10.1126/scisignal.2000568
Mourtada-Maarabouni M, Pickard MR, Hedge VL, Farzaneh F, Williams GT (2009) GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28:195–208. doi:10.1038/onc.2008.373
Pickard MR, Mourtada-Maarabouni M, Williams GT (2013) Long non-coding RNA GAS5 regulates apoptosis in prostate cancer cell lines. Biochim Biophys Acta 1832:1613–1623. doi:10.1016/j.bbadis.2013.05.005
Mercer TR, Qureshi IA, Gokhan S, Dinger ME, Li G, Mattick JS, Mehler MF (2010) Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci 11:14. doi:10.1186/1471-2202-11-14
Sunwoo H, Dinger ME, Wilusz JE, Amaral PP, Mattick JS, Spector DL (2009) MEN epsilon/beta nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res 19:347–359. doi:10.1101/gr.087775.108
Eis PS, Tam W, Sun L, Chadburn A, Li Z, Gomez MF, Lund E, Dahlberg JE (2005) Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci USA 102:3627–3632. doi:10.1073/pnas.0500613102
Kluiver J, Poppema S, de Jong D, Blokzijl T, Harms G, Jacobs S, Kroesen B-J, van den Berg A (2005) BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J Pathol 207:243–249. doi:10.1002/path.1825
Elton TS, Selemon H, Elton SM, Parinandi NL (2013) Regulation of the MIR155 host gene in physiological and pathological processes. Gene 532:1–12. doi:10.1016/j.gene.2012.12.009
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Morales DR, Thomas K, Presser A, Bernstein BE, van Oudenaarden A, Regev A, Lander ES, Rinn JL (2009) Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA 106:11667–11672. doi:10.1073/pnas.0904715106
Lin D, Pestova TV, Hellen CUT, Tiedge H (2008) Translational control by a small RNA: dendritic BC1 RNA targets the eukaryotic initiation factor 4A helicase mechanism. Mol Cell Biol 28:3008–3019. doi:10.1128/mcb.01800-07
Chen W, Bocker W, Brosius J, Tiedge H (1997) Expression of neural BC200 RNA in human tumours. J Pathol 183:345–351. doi:10.1002/(sici)1096-9896(199711)183:3<345:aid-path930>3.0.co;2-8
Acknowledgments
This work was supported by grants from the Basic Medical Science Faculty of Chongqing Medical University (Projects # JC201307) and Chongqing Science and Technology Commission (Projects # cstc2014jcyjA10028).
Conflict of interest
None of the authors has a conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, Q., Sun, S., Yu, W. et al. Altered expression of long non-coding RNAs during genotoxic stress-induced cell death in human glioma cells. J Neurooncol 122, 283–292 (2015). https://doi.org/10.1007/s11060-015-1718-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-015-1718-0