Abstract
Glioblastomas (GBMs) are the highest grade of primary brain tumors with astrocytic similarity and are characterized dispersal of tumor cell. Metastasis suppressor 1 (MTSS1) play an important role in cancer metastasis. Recent studies indicating that MTSS1 as a potential tumor suppressor and its reduced expression associated with poor prognosis in many cancer types. However, the relationship with the prognosis of patients and the molecular mechanism of MTSS1 renders a tumor suppressor effect in GBM is unknown. Here, we showed that low MTSS1 gene expression is associated with poor outcomes in patients with GBM. Overexpression of MTSS1 in U-87 MG cells exhibited inhibited glioma cell growth, colony formation, migration and invasion. Mechanistically, we found that high MTSS1 expression in U-87 MG reduced expression of CTTN. These results implicate that the role of MTSS1 suppresses cell migration and invasion by inhibiting expression of CTTN and as a prognosis biomarker in GBM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastomas (GBMs) are the most common and malignant primary brain tumors and are aggressively treated with surgery, chemotherapy, and radiotherapy, have very poor survival, remain the most pervasive and lethal of all brain malignancies [1], although many improve therapeutic strategies had been made to prolong the average survival of patients with GBM [2]. Nonetheless, the discovery of new potential targets oncogenes or tumor suppressor genes for anticancer drug development remains a major goal of cancer research.
Metastasis suppressor 1 (MTSS1) as a potential metastasis suppressor gene was originally identified in non-metastatic bladder cancer cell line [3]. Subsequent studies have been demonstrated that decreased expression of MTSS1 in many cancers [4–6], Ma et al. suggests that high MTSS1 expression has been observed in hepatocellular carcinoma [7]. And also studies founded MTSS1 expression was maintained in metastatic cell lines [8], basal cell carcinomas [9] and Colorectal Cancer [10]. Although these studies cited above suggested MTSS1 as a promising candidate biomarker and playing an important role in tumorigenesis, little is known about the role and biochemical mechanisms for MTSS1 in GBM. In our study, we sought to determine the expression of MTSS1 in GBM and patients with survival, in an attempt to discover the potential influence of MTSS1 on the development of GBM.
Here we found that low expression of MTSS1 was associated with poor prognosis. Low MTSS1 in patients with GBM and three glioma cell lines in vitro. Furthermore, overexpression of MTSS1 in U-87 MG cells inhibited the cell growth, colony formation, migration, and invasion in vivo. The molecular mechanism investigation demonstrates high level MTSS1 could inhibit expression of cortactin (CTTN), which closely related to tumor migration and invasion. These data suggest that MTSS1 as a potential tumor suppressor and prognostic biomarker in patients with GBM.
Materials and methods
Cell culture
The U-87 MG and U-251 MG human glioma cell lines were obtained from American type culture collection (ATCC). U-373 MG was purchased from xiangf bio company. Normal human astrocytes (NHA) were obtained from Reynolds JL. All cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; invitrogen) supplemented with either 10 % fetal bovine serum (FBS; HyClone) and antibiotic–antimycotic (invitrogen) in a humidified atmosphere of 5 % CO2 at 37 °C.
Patient samples and data
GBM patient tissues were obtained from surgical resections in accordance with an approved protocol from The First Affiliated Hospital of Liaoning Medical University. Tissues from adjacent to tumors were used as normal tissue. We obtained gene expression and Kaplan–Meier survival analysis results of patient with brain tumor samples from the National Cancer Institute (NCI) oncogenomics gene expression databases (available at: http://pob.abcc.ncifcrf.gov/cgi-bin/JK) and NCI REMBRANDT website (available at: http://rembrandt.nci.nih.gov). These databases include patients with all tumor information and all patient data from these databases were included in our analysis. The IntAct (available at: http://www.ebi.ac.uk/intact/) was performed for prediction molecular interaction.
RNA extraction and quantitative real-time RT-PCR
The total RNA from normal and tumor tissue and cells were extracted using TRIzol reagent (TaKaRa) according to the manufacturer’s protocol. cDNA was synthesized by using the PrimeScript RT reagent Kit (TaKaRa). The primer sequences were listed as follows: MTSS1 forward, 5′-CACATCCTGGTGAGAGCAGA-3′, and reverse, 5′-CTTCTTGGACGCCTTTCAGA-3′; CTTN forward, 5′-TAATCC AATGAGGAAT TTCCAG-3′, and reverse, 5′-TAGAGCCTGGTGCCTGGG-3′; β-actin forward, 5′-CATTGTAAGAGCCAA-3′, and reverse, 5′-TGAGGTTCGTTGTC-3′; Quantitative RT-PCR (qRT-PCR) analyses was performed on CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad) to detect mRNA expression using SYBR Premix Ex Taq™ (TaKaRa), and β-actin was used as an internal control, and all qRT-PCR data was normalized to β-actin expression.
Western blotting
Normal and tumor tissue or Cells were lysed by RIPA buffer supplemented with protease inhibitor (PMSF) on ice, and total proteins (10 mg/lane) were separated on a 10 % SDS-PAGE gel (TaKaRa) and transferred to a PVDF membrane (Bio-Rad), after blocked with 5 % nonfat milk, the membrane was incubated overnight at 4 °C with each primary antibodies against MTSS1 (Abcam), CTTN (Santa Cruz Biotechnology) and β-actin (Zhongshan, Bei-jing, china) as the loading control. Membranes were then incubated with horseradish peroxidase-conjugated secondary antibody at 1:1,000 dilution for 2 h at room temperature. Finally, Signal detection was performed using the ImageJ program (National Institutes of Health) by using ECL reagents (Pierce). The specific bands were subjected to densitometry analysis using Quantity One software.
Lentiviral infection
For production of MTSS1 overexpression lentivirus, the lentiviral with an empty control vector or MTSS1 were purchased from HANBIO (Shanghai, China). Glioma cell U-87 MG was infected with recombinant lentivirus units plus 6 μg/ml polybrene (Sigma, USA) with a multiplicity of infection (MOI) of 30. The stable lentivirus infection cells were selected and enriched by flow cytometry (BD).
Cell proliferation assay and colony formation assay
All cells were seeded 1,000 cells per well into 96-well plates and cultured for 1, 2, 3, 4 and 5 days. Cell proliferation assay was performed with the Cell Counting Kit-8 (CCK-8; Dojindo) according to the manufacturer’s instruction. For the colony formation assay, 1,000 cells were placed into each well of six-well plate. Colonies were fixed with methanol and stained with 0.1 % crystal violet in 20 % methanol for 15 min. The number of colonies was counted using an inverted microscope.
Wound healing assay
The cells were seeded into six-well plates and when cells were grown to confluence, wound healing was made by manually scraping the cells with a pipette tip. Debris was removed from the culture by washing it twice with PBS. Then the cells were incubated with serum-free medium to exclude the effect of cell growth facilitated by serum and allowed to migrate into the wounded area at 37 °C. Images were immediately acquired at 0, 24, and 48 h after wounding. Migratory ability of cells was expressed as the percentage of the area covered by cells measured using Olympus DP72 software.
In vitro invasion assays
For invasion assays, transwell chambers (8 mm pore size; Millipore, USA) containing membrane filter inserted in 24-well tissue culture plates were used. Cells (2.5 × 105) were suspended in medium without serum and seeded into the upper segment of the chambers that coated with matrigel were placed in 24-well plates, and under the chambers was the 15 % fetal bovine serum, the cells were incubated at 37 °C for 48 h. Cells will grow from low level serum to higher-level serum. After 48 h of incubation, the upper side of the filter was scraped with a cotton tip to eliminate cells that had not migrated through it. The invaded cells on the lower membrane surface were fixed in 100 % methanol for 10 min. Cells that pass through the membrane were stained with crystal violet. The cells that invaded were counted with a light microscope and expressed as the average cell number with six visual fields were randomly determined in each chamber.
Statistical analysis
All values are expressed as mean ± SD. Patients survival was analyzed with Kaplan–Meier method, using the log-rank test for comparison. The statistical significantly of differences between two groups was evaluated using Student’s t test. Analysis was performed using JSTAT statistical software (version 8.2). Statistical significance was accepted at p < 0.05(*), or p < 0.01 (**).
Results
MTSS1 gene expression is negatively correlated to the prognosis of brain cancer patients
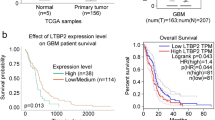
To evaluate the prognostic relevance of MTSS1 expression in patients with brain tumor, we analyzed the association of MTSS1 gene expression patients with brain cancer outcomes using results from microarray analyses of brain cancer obtained from the NCI as detailed above. Low expression of MTSS1 was significantly associated with poor outcomes of with neuroblastoma (p = 2.7 × 10−8, n = 252) (Fig. 1a). We independently evaluated the association of MTSS1 expression with patients survival in the NCI database, in these 57 patients, low MTSS1 expression was also significantly associated with lower survival rates (p = 1.6 × 10−4) (Fig. 1b). These results suggest that low MTSS1 expression is associated with the outcomes of patients with brain tumor.
The outcomes of patients with glioblastoma are based on MTSS1 gene expression. The NCI oncogenomics gene expression databases and The NCI REMBRANDT databases were evaluated for outcomes of patients with brain tumor, and Kaplan–Meier survival curves were generated. a Compared to the Kaplan–Meier survival for numbers (n = 252) of patients with high (red) or low (blue) MTSS1 gene expression (log-rank test; p = 2.7 × 10−8). b Estimated Kaplan–Meier survival for patients with all glioma who have high (red; n = 34) and low (blue; n = 23) MTSS1 gene expression (log-rank test; p = 1.6 × 10−4)
MTSS1 expression is reduced in gliomas
In an effort to understand the role of MTSS1 in glioma malignancy, we examined the MTSS1 mRNA levels and protein levels among human glioma tumors compared to control normal tissue and cell lines. We assessed the expression of MTSS1 in human glioma tumors compared to control normal tissue, and expression was strikingly reduced in glioma tissues (Fig. 2a), and the results from three glioma cell lines U-87 MG, U-373 MG, U-251 MG and NHA showed also low of MTSS1 expression in three glioma cells (Fig. 2b). In contrast to GBMs, NHA cell maintained some high level of MTSS1 expression. Next, we examined the presence of MTSS1 protein in tumor and normal tissue, and the commonly three human glioma cell lines through western blot. Notably, MTSS1 was found to be expressed at high levels in normal tissue from patients 245 and 474, tumor tissue exhibit low expression of MTSS1 (Fig. 2c). The MTSS1 in three human glioma cells examined also reduced expression compared with NHA (Fig. 2d). β-actin expression was used as a control for protein loading among the tissue and cell samples. Densitometry analyses of the bands on the immunoblots indicate that there is a complete reduction of MTSS1 expression in GBMs compared to nontumor tissue following normalization for the β-actin loading control. These results suggest that loss of MTSS1 protein expression may be an important event in glioma progression.
Decreased expression of MTSS1 in glioblastoma tissues and glioma cell lines. a MTSS1 expression in normal human tissues and glioblastoma tissues measured by qRT-PCR, the relative expression of MTSS1 was normalized by β-actin expression. b MTSS1 expression in normal human astrocytes (NHA), three glioma cell lines U-251MG, U-373MG and U-87 MG measured by qRT-PCR, the relative expression of MTSS1 was normalized by β-actin expression. c Western blotting analyses of MTSS1 protein expression in normal tissues and glioblastoma tissues. Biopsies were taken from the tumor center (G) and normal tissue (N). Expression of MTSS1 was quantitative and normalized against β-actin. d MTSS1 protein expression in NHA, U-251MG, U-373 MG and U-87 MG analyses by western blot. Expression of MTSS1 was quantitative and normalized against β-actin. All experiments were performed at least in triplicate and the data are presented as the mean ± SD, ** p < 0.01
Overexpression of MTSS1 suppresses glioma cell proliferation and colony formation in vitro
To determine the impact of MTSS1 on glioma cell proliferation, we established U-87 transfection stably expressing MTSS1 using lentivirus infection. We found that overexpressed MTSS1 reduced GBM cell proliferation (Fig. 3a). The colony formation assays results revealed that overexpressed MTSS1 can significantly suppress U-87 cell clone formation. We multiple clones tested and the results of a single representative clone presented (p < 0.05, Fig. 3b, c).collectively, our data indicated that MTSS1 could suppresses the proliferation and colony formation of GBM cells in vitro.
Tumor suppressive properties of MTSS1. a Overexpression of MTSS1 in U-87 MG results in decreased growth rate. Right shows a growth curve comparing the rate of proliferation of U-87 MG with MTSS1 overexpression versus control vector. Five thousand cells were plated in both cases. All experiments were performed in triplicate. All graphs report mean ± SD. * p < 0.05; ** p < 0.001. b U-87 MG cell expressing MTSS1 or vector was measured to form colonies in vitro. c The numbers of colonies in U-87 MG expressing control or MTSS1 cells. The result shows cultures of U-87 MG with overexpression MTSS1 were statistically significant reduction in colony formation (p < 0.05)
Overexpression of MTSS1 suppresses GBM cell migration and invasion
Previously study showed that regulation of MTSS1 protein expression had an impact on breast cancer cell aggressiveness [5] and the significant low expression of MTSS1 in glioma tumor samples and glioma cell lines. Based on these data, we hypothesized that overexpression of MTSS1 in U-87 MG cells would also suppress glioma migration and invasion. We evaluated this hypothesis using a scratch wound assay and transwell assay, respectively. Confluent monolayers of U-87 MG cells overexpressing either vector or MTSS1 were scratched to form a wound. Cells migration was measured after 24 and 48 h, the migration of cells was observed by optical microscopes (Fig. 4a). The migratory ability of cells was expressed as the percentage of the area covered by cells measured, U-87 MG cells overexpressing MTSS1 had impaired migration significant reduction in the distance migrated (Fig. 4b). Additionally, Transwell invasion assay results observed that overexpression of MTSS1 also inhibited U-87 MG cell invasion (Fig. 4c, d). These results indicated that MTSS1 could influence the migration and invasion ability of GBM cell.
The effect of overexpression MTSS1 protein inhibitor on U-87 MG migration and invasion. a Migration of cells to cover the wound was followed over 48 h. Dashed line mark the wound edge at 0 h. (×20). b The percent reduction of the initial scratch area at 12 and 24 h was compared among groups. * p < 0.05. ** p < 0.01. The data represented the mean values of three measurements, and the error bars represented the SD of the mean. c The representative images of invaded cells by transwell invasion assay. The U-87 MG cells overexpression vector or MTSS1 penetrating the membrane was fixed and were stained with 0.1 % crystal violet after 24 h (×40). d The numbers of invaded U-87 MG cells with overexpression vector or MTSS1. Experiments were performed at least in triplicate and the data are presented as the mean ± SD. * p < 0.05. ** p < 0.01
MTSS1 targets CTTN and inhibits its expression
To further determine the mechanisms that MTSS1 suppresses GBM cell proliferation and invasion, we performed bioinformatic analysis to predict protein interactions and ten potential molecular were found in IntAct (data not shown). We focused on CTTN, which has emerged as a key player in aggressive cancers [11] and whether it was responsible for the cancer invasion led by reduced MTSS1 expression. As showed in Fig. 5a and b, CTTN was significantly reduced in MTSS1 overexpressing cells as compared with control. The results reveal that MTSS1 suppresses and reduces the expression of CTTN, thereby inhibiting proliferation and invasion of U-87 MG GBM cells.
MTSS1 overexpression suppresses the CTTN. a qRT-PCR analysis of MTSS1 and CTTN in U-87 MG with empty vector or MTSS1 overexpressing vector. b Western blot analysis of MTSS1 and CTTN in U-87 MG with empty vector or MTSS1 overexpressing vector. The expression results were normalized against β-actin. All experiments were performed at least in triplicate and the data are presented as the mean ± SD. ** p < 0.01
Discussion
This study demonstrates the decreased expression of MTSS1 in glioma patient tissues and tumor cells. Our data demonstrated a reduced level of MTSS1 expression in glioma tissues compared to the normal tissues and the expression of MTSS1 was associated with the clinical pathology and prognosis of the patients with brain cancer. Cellular function tests further demonstrated that enhanced MTSS1 expression has been associated with inhibition to the glioma cell’s growth, clone formation, migration and invasion. Most importantly, we identified CTTN as a target molecular of MTSS1. To our best knowledge, this study is the first report on the function of MTSS1 in glioma. These results facilitated our understanding about the mechanisms that MTSS1 as a tumor suppressor in GBM progression.
MTSS1 may exert its metastasis suppressor function by acting as a scaffold protein that interacts with actin-associated protein to regulate lamellipodia formation [12]. We believed that other regulatory mechanisms exist in different tumors. The CTTN was first identified as a prominent substrate of the Src nonreceptor tyrosine kinase [13] and also as a key player in aggressive cancers [14–16]. Previously reported that down regulation of CTTN could decrease cell migration [17, 18]. It has also been reported that Overexpression of CTTN in colorectal cancer, melanoma, gastric cancer and GBM [11, 19–22]. Consistent with our study, we found that overexpressing MTSS1 could inhibit the expression of CTTN and suppress GBM cell’s migration and invasion.
Our functional studies have demonstrated that the MTSS1 over-expression resulted in a dramatic reduction in tumor cell migration, invasion and growth. But the mechanism for loss of MTSS1 expression in brain tumor was needed to be revealed. More recent publications indicate a role for DNA methylation 3B (DNMT3B) in regulation of MTSS1 expression in hepatocellular carcinoma [23]. MicroRNAs are important in tumorigenesis and effects by post-transcriptionally regulating target genes to induce mRNA degradation and inhibit mRNA translation [24, 25].Whether the expression of MTSS1 was regulated by microRNA need further study. Further analysis of MTSS1 expression or inaction in glioma tissue samples and its association with other human cancers will remain to be investigated.
In summary, low expression of MTSS1 was observed in GBM tissues and glioma cell lines, and loss its expression associated with poor prognosis in patients with brain tumor. We found that MTSS1 is capable to suppress cell growth, clone formation, migration and invasion by reduced expression of CTTN. These findings are useful in understanding the progression of GBM and may be served as a useful biomarker for the prediction of outcome of GBM.
References
Leder K, Pitter K, Laplant Q, Hambardzumyan D, Ross BD, Chan TA et al (2014) Mathematical modeling of PDGF-driven glioblastoma reveals optimized radiation dosing schedules. Cell 156(3):603–616
Tanaka S, Louis DN, Curry WT, Batchelor TT, Dietrich J (2013) Diagnostic and therapeutic avenues for glioblastoma: no longer a dead end? Nat Rev Clin Oncol 10(1):14–26
Lee YG, Macoska JA, Korenchuk S, Pienta KJ (2002) MIM, a potential metastasis suppressor gene in bladder cancer. Neoplasia 4(4):291–294
Utikal J, Gratchev A, Muller-Molinet I, Oerther S, Kzhyshkowska J, Arens N et al (2006) The expression of metastasis suppressor MIM/MTSS1 is regulated by DNA methylation. Int J Cancer 119(10):2287–2293
Parr C, Jiang WG (2009) Metastasis suppressor 1 (MTSS1) demonstrates prognostic value and anti-metastatic properties in breast cancer. Eur J Cancer 45(9):1673–1683
Huang XY, Huang ZL, Xu YH, Huang XY, Zhou J, Ye SL et al (2010) Effects of MIM-B gene on invasive and metastatic potentials of human hepatocellular carcinoma MHCC97H cells. Zhonghua Gan Zang Bing Za Zhi 18(12):915–919
Ma S, Guan XY, Lee TK, Chan KW (2007) Clinicopathological significance of missing in metastasis B expression in hepatocellular carcinoma. Hum Pathol 38(8):1201–1206
Bompard G, Sharp SJ, Freiss G, Machesky LM (2005) Involvement of Rac in actin cytoskeleton rearrangements induced by MIM-B. J Cell Sci 118(Pt 22):5393–5403
Callahan CA, Ofstad T, Horng L, Wang JK, Zhen HH, Coulombe PA et al (2004) MIM/BEG4, a Sonic hedgehog-responsive gene that potentiates Gli-dependent transcription. Genes Dev 18(22):2724–2729
Wang D, Xu MR, Wang T, Li T, Zhu J (2011) MTSS1 overexpression correlates with poor prognosis in colorectal cancer. J Gastrointest Surg 15(7):1205–1212
Weaver AM (2008) Cortactin in tumor invasiveness. Cancer Lett 265(2):157–166
Lin J, Liu J, Wang Y, Zhu J, Zhou K, Smith N, Zhan X (2005) Differential regulation of cortactin and NWASP- mediated actin polymerization by missing in metastasis (MIM) protein. Oncogene 24(12):2059–2066
Wu H, Reynolds AB, Kanner SB, Vines RR, Parsons JT (1991) Identification and characterization of a novel cytoskeleton-associated pp60src substrate. Mol Cell Biol 11(10):5113–5124
Buday L, Downward J (2007) Roles of cortactin in tumor pathogenesis. Biochim Biophys Acta 1775(2):263–273
Stuible M, Dube N, Tremblay ML (2008) PTP1B regulates cortactin tyrosine phosphorylation by targeting Tyr446. J Biol Chem 283(23):15740–15746
MacGrath SM, Koleske AJ (2012) Cortactin in cell migration and cancer at a glance. J Cell Sci 125(Pt 7):1621–1626
Mezi S, Todi L, Orsi E, Angeloni A, Mancini P (2012) Involvement of the Src-cortactin pathway in migration induced by IGF-1 and EGF in human breast cancer cells. Int J Oncol 41(6):2128–2138
Nakane K, Fujita Y, Terazawa R, Atsumi Y, Kato T, Nozawa Y et al (2012) Inhibition of cortactin and SIRT1 expression attenuates migration and invasion of prostate cancer DU145 cells. Int J Urol 19(1):71–79
Hirakawa H, Shibata K, Nakayama T (2009) Localization of cortactin is associated with colorectal cancer development. Int J Oncol 35(6):1271–1276
Hashimoto A, Hashimoto S, Ando R, Noda K, Ogawa E, Kotani H et al (2011) GEP100-Arf6-AMAP1-cortactin pathway frequently used in cancer invasion is activated by VEGFR2 to promote angiogenesis. PLoS ONE 6(8):e23359
Cen G, Ding HH, Liu B, Wu WD (2014) FBXL5 targets cortactin for ubiquitination-mediated destruction to regulate gastric cancer cell migration. Tumour Biol 35(9):8633–82014
Xu XZ, Garcia MV, Li TY, Khor LY, Gajapathy RS, Spittle C et al (2010) Cytoskeleton alterations in melanoma: aberrant expression of cortactin, an actin-binding adapter protein, correlates with melanocytic tumor progression. Mod Pathol 23(2):187–196
Fan H, Chen L, Zhang F, Quan Y, Su X, Qiu X et al (2012) MTSS1, a novel target of DNA methyltransferase 3B, functions as a tumor suppressor in hepatocellular carcinoma. Oncogene 31(18):2298–2308
Gurtan AM, Sharp PA (2013) The role of miRNAs in regulating gene expression networks. J Mol Biol 425(19):3582–3600
Li J, Lu X (2013) The emerging roles of 3′ untranslated regions in cancer. Cancer Lett 337(1):22–25
Acknowledgments
We thank Reynolds JL for providing the NHA cell. Chenyuan For helpful discussion, and Zhangyuan for technical assistance.
Conflict of interest
The authors declared that they have no conflicts of interest to this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Qigui Qi has contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhang, S., Qi, Q. MTSS1 suppresses cell migration and invasion by targeting CTTN in glioblastoma. J Neurooncol 121, 425–431 (2015). https://doi.org/10.1007/s11060-014-1656-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-014-1656-2