Abstract
Hepatocellular carcinoma (HCC) is the most common form of liver cancer, occurring primarily in regions where viral hepatitis infections are common. Unfortunately, most HCC cases remain undiagnosed until late stages of the disease when patient outcome is poor, typically limiting survival from a few months to a year after initial diagnosis. In order to better care for HCC patients, new target-specific approaches are needed to improve early detection and therapeutic intervention. In this work, polymeric nanoparticles functionalized with a HCC-specific aptamer were examined as potential targeted drug delivery vehicles. Specifically, doxorubicin-loaded nanoparticles were prepared via nanoprecipitation of blends of poly(lactic-co-glycolic acid)-b-poly(ethylene glycol). These particles were further functionalized with the HCC-specific TLS11a aptamer. The in vitro interaction and therapeutic efficacy of the aptamer and aptamer-functionalized nanoparticles were characterized in a hepatoma cell line. Nanoparticles were found to be spherical in shape, roughly 100–125 nm in diameter, with a low polydispersity (≤0.2) and slightly negative surface potential. Doxorubicin was encapsulated within the particles at ~40 % efficiency. Drug release was found to occur through anomalous transport influenced by diffusion and polymer relaxation, releasing ~50 % doxorubicin in the first 10 h and full release occurring within 36 h. Confocal microscopy confirmed binding and attachment of aptamer-targeted nanoparticles to the cell surface of cultured HCC cells. Efficacy studies demonstrated a significant improvement in doxorubicin delivery and cell-killing capacity using the aptamer-functionalized, drug-loaded nanoparticles versus controls further supporting use of aptamer nanoparticles as a targeted drug delivery system for HCC tumors.

In this work, polymeric nanoparticles functionalized with a liver cancer-specific aptamer were examined as potential targeted drug delivery vehicles. The aptamer-functionalized nanoparticles were found to significantly improve doxorubicin drug delivery and cell-killing capacity in vitro versus non-targeted controls, supporting their use as a targeted treatment toward liver cancer tumors.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Liver cancer is the second leading cause of cancer-related death worldwide (Ferlay et al. 2015). The most common form of liver cancer is hepatocellular carcinoma (HCC), accounting for roughly 75 % of the 782,000 new cases of liver cancer diagnosed and of the 745,000 liver cancer-related deaths occurring each year (Ferlay et al. 2015). Contributing to the high mortality of this disease is the late stage at initial diagnosis for most patients, high recurrence rate, lack of effective treatments, and underlying liver dysfunction due to chronic diseases such as cirrhosis or viral hepatitis infection (Forner et al. 2012). Improvements in surveillance of high-risk patients with imaging and serum biomarkers have increased the rate of early detection in some parts of the world, yet early HCC detection remains problematic in developing countries where HCC is most prevalent (Attwa 2015; Dhanasekaran et al. 2012; Forner et al. 2012). When detected at an early stage (which is the case for 43 % of patients in the USA), 5-year survival rates for HCC are 31 %; however, survival rates fall to 11 and 3 % for advanced stages of the disease exhibiting regional and distant metastases, respectively (American Cancer Society 2016).
Existing treatment options for HCC remain quite limited. Surgery is the standard treatment for early-stage disease offering potentially curative outcomes through resection, transplantation, or ablation (Forner et al. 2012; Knox et al. 2015). Other localized treatments include radiotherapy and various forms of chemoembolization, which disrupts the tumor blood supply by obstructing the hepatic artery and delivering chemotherapy locally (Forner et al. 2012; Knox et al. 2015). Conventional chemotherapy with doxorubicin alone or in a multi-drug cocktail has previously been used to treat HCC, but there is no clear evidence of a benefit on overall survival and side effects associated with systemic toxicity that limit the usable dosage (Knox et al. 2015). At present, the only FDA-approved drug for treatment of advanced HCC is sorafenib (Nexavar®), which is a targeted therapeutic that blocks angiogenic and proliferative signaling pathways (Llovet et al. 2008); however, sorafenib recently failed to improve outcomes when used as an adjuvant treatment in phase 3 clinical trials (Bruix et al. 2015). Thus, a need remains for the development of new therapeutic strategies that can reduce systemic toxicity and enhance existing drug efficacy in the treatment of HCC. Our targeted drug delivery approach seeks to overcome both these challenges through the use of biodegradable drug-loaded polymeric nanocarriers functionalized with a HCC-specific aptamer.
Aptamers are a class of molecular targeting agents that consist of single-stranded DNA or RNA oligonucleotides that fold into unique secondary structures, enabling recognition of biomolecules with high affinity and specificity (Ellington and Szostak 1990; Stoltenburg et al. 2007). The advantages of aptamers over antibodies include increased stability, low cost, and ease of chemical functionalization or modification (Meng et al. 2015; Sun et al. 2014). The HCC-specific aptamer employed in the current study (designated TLS11a) was developed in the Tan group using a modified whole-cell SELEX procedure (Shangguan et al. 2008; Xu et al. 2015). TLS11a has been shown to exhibit high specificity toward mouse/human HCC cells in vitro over normal liver cells and other cancer cell types (Shangguan et al. 2008; Xu et al. 2015). Specifically, the low affinity of this aptamer toward cell lines from non-liver human tumor types including leukemia, lymphoma, and lung cancer (Jurkat/K562/CCRF-CEM, Ramos, and H23, respectively) and normal mouse/human liver cells (CL.2 and HU 1082, respectively) has been demonstrated (Shangguan et al. 2008; Xu et al. 2015). The utility and effectiveness of TLS11a as an HCC-specific probe have also been demonstrated in recent biosensor applications (Kashefi-Kheyrabadi et al. 2014; Qu et al. 2014; Sun et al. 2015; Sun et al. 2016), although the specific cell surface protein or carbohydrate moiety recognized by the TLS11a aptamer remains unknown. In addition to binding at the cell surface, rapid internalization and localization of the TLS11a aptamer to late endosomes or lysosomes have been described (Meng et al. 2012; Weigum et al. 2014). As such, the TLS11a aptamer is an ideal targeting agent for drug delivery since it selectively binds to the HCC cell surface and elicits cellular uptake (Bareford and Swaan 2007; Meng et al. 2012).
Nanoparticle (NP) drug carriers afford several advantages in the delivery of chemotherapeutics by increasing drug concentration in tumor versus normal tissue via active and passive targeting, thereby improving the pharmacokinetic profile of the drug, and increasing cellular uptake (Danhier et al. 2010). The use of nanocarriers can significantly improve tumor selectivity passively by taking advantage of the enhanced permeability and retention (EPR) effect. The EPR effect is the result of the tumor-induced formation of leaky angiogenic blood vessels that enable the extravasation of macromolecules and nanocarriers into the tumor, as well as of deficient tumor lymphatics that lead to nanocarrier accumulation (McDonald and Baluk 2002). The vascular density and leakiness associated with the EPR effect have been previously utilized for the delivery of several nanocarriers to tumors (Danquah et al. 2011; Hainfeld et al. 2011; Huang and Hainfeld 2013; Maeda et al. 2013; Onda et al. 2013). Similarly, the use of nanocarriers enables concurrent incorporation of targeting moieties with high affinity to cancer cell biomarkers such as antibodies, peptides, or aptamers as in the current study to facilitate cancer cell recognition, binding, and carrier internalization (Byrne et al. 2008; Yu et al. 2012).
The use of aptamer-targeted nanocarriers for the delivery of chemotherapeutic agents to tumors has gained significant interest in recent years. Notable progress has occurred in the development of NP-aptamer conjugates against prostate cancer by Farokhzad and colleagues (Dhar et al. 2008; Farokhzad et al. 2006; Farokhzad et al. 2004). These systems have advanced to phase I and phase II clinical trials through BIND Therapeutics (Cambridge, MA). Despite the potential of aptamer-targeted nanomedicines, there have been relatively few attempts to investigate the efficacy of aptamer-targeted nanocarrier delivery of chemotherapeutic agents to HCC. Among these, examples include cytochrome C-capped mesoporous silica NPs (Zhang et al. 2014) and poly(lactic-co-glycolic acid) (PLGA) NPs (Yu et al. 2013) functionalized with the nucleolin-specific AS1411 aptamer. Similarly, carboxymethyl cellulose-modified magnetic iron oxide NPs have been targeted to HCC cells utilizing an aptamer specific to the epithelial cell adhesion molecule (EpCAM) (Pilapong et al. 2014). Finally, PLGA NPs have been targeted to cancer stem cells in HCC utilizing aptamers specific to both CD133 and the epidermal growth factor receptor (EGFR) (Jiang et al. 2015).
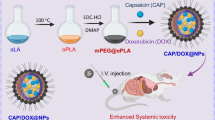
The goal of this study is to target the delivery of the chemotherapeutic agent doxorubicin to HCC utilizing DNA aptamer-functionalized biodegradable polymeric NPs (Fig. 1). These NPs were prepared through the co-nanoprecipitation of blends of two biodegradable and biocompatible amphiphilic block copolymers. The characteristics of the polymers and NPs, as well as the in vitro selectivity and therapeutic efficacy of these nanocarriers in HCC cells, are described.
Materials and methods
Materials
D,L-lactide, tin(II) 2-ethylhexanoate (stannous octoate, Sn(Oct)2), 1-(3-dimethylaminopropyl)-3-ethylcarbodiimide hydrochloride (EDC), and doxorubicin hydrochloride were obtained from Alfa Aesar (Ward Hill, MA). Glycolide and poly(ethylene glycol) methyl ether (MW 2000 Da, mPEG2000) were procured from Sigma-Aldrich (St. Louis, MO). N-hydroxysuccinimide (NHS), sodium bicarbonate, methanol, dimethyl sulfoxide (DMSO), and chloroform-d were obtained from Thermo Scientific (Waltham, MA). The heterofunctional poly(ethylene glycol) polymer OH-PEG2000-COOH (where 2000 represents the molecular weight in Da of the polymer) was obtained from Laysan Bio (Arab, AL). All solvents used were of at least ACS grade and used as received unless otherwise noted. Ultrapure water (DI H2O, 18.2 MΩ cm) was prepared with a Millipore Direct-Q system equipped with a BioPak disposable ultrafilter. Cell culture media and fetal bovine serum (heat-inactivated FBS) were purchased from Mediatech Inc. (Manassas, VA), while all other supplements were obtained from Sigma-Aldrich, unless otherwise noted.
Aptamer sequence
Aptamers were purchased from Integrated DNA Technologies (IDT; Coralville, IA) with a 5′ primary amine modification and an internal fluorescein-modified thymine residue closest to the 3′ end. Aptamers were HPLC purified and resuspended at a concentration of 100 μM in nuclease-free water. TLS11a aptamer was 63 nucleotides in length with sequence of 5′-ACA GCA TCC CCA TGT GAA CAA TCG CAT TGT GAT TGT TAC GGT TTC CGC CTC ATG GAC GTG CT*G-3′. The predicted secondary structure obtained using IDT OligoAnalyzer 3.1 software at 37 °C with 150 mM [Na+] and 0.71 mM [Mg2+], i.e., the temperature and buffer/media conditions used for imaging and cytotoxicity assays, is shown in Fig. 2. At these conditions, the predicted melting temperature was 54.4 °C with a ΔG of −8.94 kcal mol−1.
Synthesis of PLGA-mPEG and PLGA-PEG-COOH
Prior to polymerization, D,L-lactide and glycolide were purified by recrystallization in hot ethyl acetate and dried under vacuum overnight. PEGs were purified and dried by cold ethyl ether precipitation and dried under vacuum overnight. Anhydrous solvents were stored over molecular sieves in nitrogen-purged flasks.
Poly(D,L-lactide-co-glycolide)-b-poly(ethylene glycol) methyl ether (PLGA-mPEG) and poly(D,L-lactide-co-glycolide)-b-poly(ethylene glycol) carboxymethyl (PLGA-PEG-COOH) were synthesized by the tin(II) 2-ethylhexanoate (Sn(Oct)2)-catalyzed ring opening polymerization of D,L-lactide and glycolide onto the hydroxyl end of mPEG2000 or HO-PEG2000-COOH, as previously described (Betancourt et al. 2009). Polymerizations were performed in a three-arm round bottom flask heated to 110 °C in a silicon oil bath under toluene reflux and nitrogen gas injection. D,L-lactide and the respective PEG precursor were dissolved in 10 mL of toluene and added to the three-arm flask. To prevent the formation of insoluble poly(glycolic acid), glycolide was dissolved in 5 mL acetone and added in 0.5-mL aliquots every 30 min through an addition funnel. Sn(Oct)2 was dissolved in 5 mL of toluene and added to the three-arm flask at the beginning of the polymerization process. The reactants were allowed to react for 6 h. The resulting polymer was dissolved in chloroform and centrifuged to remove insoluble portions; the supernatant was then precipitated in cold ethyl ether. The precipitated polymer was dried in a Labconco FreeZone lyophilizer for 48 h. To confirm their composition and determine their molecular weight, the polymers were dissolved in CDCl3 and analyzed by 1H–NMR spectroscopy using a Bruker Avance 400-MHz spectrophotometer.
Nanoparticle preparation and characterization
NPs were prepared by nanoprecipitation (Betancourt et al. 2007; Betancourt et al. 2009), as shown in Fig. 1. Polymer solutions were prepared by dissolving specific quantities of doxorubicin, PLGA-mPEG, and PLGA-PEG-COOH (polymer molar ratio or 19:1) in acetonitrile containing 4.2 % DMSO to a final concentration of 8 mg polymer/mL. The amount of doxorubicin included in this organic phase was varied to result in a targeted loading of 1, 3, or 5 % (w/w) of doxorubicin in the NPs. The resulting organic phase (0.272 mL) was then added dropwise to ultrapure DI water (4 mL) while magnetically stirring. NP suspensions were then centrifuged three times at 28,000 rpm (74,074×g) for 60 min in an Optima L-90 K Ultracentrifuge (Beckman Coulter Inc., USA) and washed with DI water to remove unencapsulated drug. Drug-free NPs were prepared by the same procedure but omitting doxorubicin. All samples were prepared in triplicate. NP samples used for cell studies were purified using autoclaved ultrapure DI water and sterile-filtered using a 0.2-μm syringe filter.
NP size, polydispersity, and zeta potential were determined using a Zetasizer Nano ZS (Malvern Instruments Ltd., UK). For size measurement, samples of NP suspensions were diluted with water in disposable polystyrene cuvettes for dynamic light scattering (DLS) measurements. DLS utilizes changes in scattered light intensity to determine the diffusivity of particles undergoing Brownian motion. The particle diffusivity is then used to calculate the particle size and size distribution using the Stokes-Einstein equation. Zeta potential measurements were obtained using laser Doppler microelectrophoresis. In this technique, NPs exposed to an electric field move with a velocity associated to their zeta potential. This velocity is used to calculate the electrophoretic mobility which then enables the measurement of the zeta potential of the NPs using Smoluchowski’s approximation (Kaszuba et al. 2010). For zeta potential measurements, 0.9 mL of aqueous NP suspension is mixed with 0.1 mL of 100 mM potassium chloride (KCl) to result in a final concentration of 10 mM KCl. Zeta potential measurements were carried out using a dip cell kit (Malvern Instruments Ltd., UK).
NP morphology was determined by transmission electron microscopy (TEM). TEM samples were prepared by drop casting the NP suspensions onto 200-mesh copper/formvar carbon grids (Electron Microscopy Sciences FCF-200Cu). Samples were stained with uranyl acetate (2 % w/v) prior to imaging in a JEM 1200EXII TEM (JEOL Ltd., JP). The percent recovery for each batch of NPs was determined by comparing the mass of recovered, lyophilized NPs to the known mass of polymer, and drug used for the preparation of the NPs.
Aptamer attachment
Aptamers were attached to the surface of the NPs using standard carbodiimide coupling chemistry, as previously reported (Betancourt et al. 2009; Kamaly et al. 2012). Specifically, the carboxylic acid on the surface of the NPs was activated into an amine-reactive N-hydroxysuccinimide ester using EDC and NHS and reacted with the amine-modified aptamer. Specifically, 1 mL of NPs suspended in water was incubated with 10 mM EDC and 6 mM NHS in the presence of 1 μM (fivefold molar excess of aptamer to PLGA-PEG-COOH) of the amine-modified TLS11a oligonucleotide for 1 h. After the reaction, the NPs were centrifuged three times at 28,000 rpm (74,074×g) for 60 min in an Optima L-90 K Ultracentrifuge (Beckman Coulter Inc., USA) and washed with DI H2O three times to remove unbound oligonucleotides. The attachment of the aptamer to the NPs was confirmed by fluorescence spectroscopy.
Doxorubicin drug loading
To calculate final drug loading, a known volume of NP suspensions was lyophilized overnight. A known mass of lyophilized NPs was then dissolved in DMSO and the amount of doxorubicin in the sample was determined from a standard curve of the fluorescence intensity of
where m drug is the mass of the drug found in a specific mass of lyophilized NPs (m NP).
The drug entrapment efficiency was determined by comparing the mass of doxorubicin found within a batch of lyophilized NPs (m drug) to the mass of drug utilized in the preparation of the batch (m drug,prep) according to Eq. 2,
Drug release
To conduct drug release studies, 24 mg of drug-loaded NPs was dispersed into 60 mL of phosphate-buffered saline (PBS; pH 7.4). The resulting drug-loaded NP suspension was then transferred to 1.5-mL microcentrifuge tubes in 1-mL aliquots. The tubes were capped and placed on a shaker (250 rpm) within a 37 °C incubator. At specific times, three microcentrifuge tubes were collected from the incubator and centrifuged at 15,000 rpm (21,130×g) for 30 min in a Eppendorf 5424 centrifuge; 0.750 mL of the resulting supernatant from each tube was then filtered using a 0.02-μm syringe-driven membrane filter (Whatman Anotop 10, GE Healthcare), and 0.400 mL of the filtrate was collected in a new microcentrifuge tube. The microcentrifuge tubes containing the pelleted NPs and the microcentrifuge tubes containing the filtered supernatant were then lyophilized overnight. The lyophilized samples were dissolved in DMSO, and the amount of doxorubicin in the supernatant and NP pellet was determined from a standard curve of the fluorescence intensity of doxorubicin in DMSO. The percent of doxorubicin released from the NPs was determined by comparing the mass of doxorubicin in the filtered supernatant to the mass of doxorubicin retained in the NPs.
The short-time approximation power law model of drug release from spherical, non-swellable polymer particles was applied to the data to investigate the mechanism of drug release (Ritger and Peppas 1987; Siepmann and Peppas 2001). Specifically, release data for the first 60 % of the drug release was fit to Eq. 3,
where M t /M ∞ is the fractional release, t is the release time, k is a constant, and n is a diffusional constant that provides information about the mechanism associated with the release of the drug from the particles. For spherical particles, n = 0.43 represents purely Fickian diffusion, 0.43 < n < 1 represents anomalous (non-Fickian) or case II transport, and n = 1 represents zero-order release (Ritger and Peppas 1987; Siepmann and Peppas 2001).
In vitro cell culture
HCC mouse liver cells (BNL 1ME A.7R.1, #TIB-75, American Type Culture Collection, Manassas, VA), abbreviated MEAR, were seeded from frozen stocks into T-25 tissue culture treated flasks at an initial seeding density 1 × 104 cells/cm2 in culture medium. Complete growth medium was modified from previous reports by Shangguan et al. (2008) and consisted of a 1:1 (v/v) mixture of Dulbecco’s modified Eagle’s medium (DMEM)/Ham’s F-12 medium (Corning® cellgro® #90–090-PB, Mediatech Inc., Manassas, VA) supplemented with 2.5 g/L sodium bicarbonate, 10 % FBS, 2.5 mM L-glutamine, 50 ng/mL epidermal growth factor, 40 ng/mL dexamethasone, and insulin/transferrin/selenium solution (ITS) diluted to a final concentration of 10 μg/mL insulin, 5.5 μg/mL transferrin, and 5 ng/mL selenium. Cells were maintained at 37 °C under a 5 % CO2 atmosphere until they reached ~80 % confluency, at which time they were subcultured or frozen in growth medium containing 10 % DMSO.
Confocal microscopy
For targeted NP imaging studies in live HCC cells, MEAR cells were subcultured from T-25 flasks onto 22 × 22-mm glass coverslips and grown for 48–72 h or until reaching roughly 80 % confluency. The TLS11a aptamer-NP conjugates were prepared as described above but without the doxorubicin drug in order to prevent spectral overlap with the selected cell stains and fluorophores. The aptamer concentration in the bulk NP suspension was calculated from a standard curve of the fluorescence of the TLS11a fluorescein label. The aptamer-NPs were then diluted in complete culture media to a final concentration of 100 nM aptamer. The control NPs without aptamer were diluted using the same v/v ratio. For staining, individual coverslips were transferred to a clean 35-mm-diameter petri dish and incubated with 250 μL of the aptamer-NP solution in media for 30 min at 37 °C in a CO2 incubator. After 30 min, the aptamer-NP solution was removed and the cells were counterstained with 2 drops/mL NucBlue® Live ReadyProbes® (360 nm ex/460 nm em) and 1 μg/mL wheat germ agglutinin-Alexa-Fluor®647 conjugate (WGA) (650 nm ex/668 nm em) (both from Molecular Probes® Life Technologies, Carlsbad, CA) for 5 min at room temperature. NucBlue® Live ReadyProbes® contains Hoechst membrane permeable dye which binds nucleic acids within the nucleus of live cells. WGA is a lectin which binds to sialic acid and N-acetylglucosaminyl residues on cell surface glycoproteins/glycolipids. In live cells, WGA is membrane impermeable and served as a plasma membrane marker. Cells were washed two times in warm Live Cell Imaging Solution (Molecular Probes® Life Technologies), float mounted onto a glass slide, and imaged start-to-finish in less than 30 min on an Olympus FluoView FV1000 laser scanning confocal microscope with a 60X PlanApoN oil immersion objective (1.4 NA) (Olympus Corporation, Tokyo, Japan). All images were collected sequentially using the appropriate fluorophore-matched filter cube (Hoechst 33,258, FITC, AF647) in order to reduce spectral bleed-through across channels.
Determination of therapeutic efficacy of targeted NPs
MEAR cells were maintained in modified DMEM/F12 media as previously described. For efficacy studies, cells were seeded in 96-well plates at a density of 3000 cells/well. After 24 h, culture media was replaced with media containing doxorubicin-loaded NPs, doxorubicin-loaded NPs functionalized with TLS11a, blank NPs, or doxorubicin at a range of concentrations. The cells were exposed to treatment for 4 h at 37 °C and 5 % CO2. After this exposure period, the treatment media was replaced with complete culture media and the cells were incubated for an additional 48 h. Cell viability was determined by measuring cell mitochondrial activity. A freshly prepared solution of 0.5 mg/mL thiazolyl blue tetrazolium bromide (MTT) dissolved in Dulbecco’s phosphate-buffered saline (DPBS) was added to the cells and incubated for 2 h. After incubation, the MTT reagent solution was removed and 100 μL of DMSO containing 16 mM NH4OH (Wang et al. 2012) was added to dissolve formazan crystals. The absorbance of the samples was measured at 555 nm (formazan absorption peak) and 700 nm (background). Six replicates were utilized per condition. Cell viability was determined by normalizing the absorbance of treated cells to that of cells grown in normal media (negative control). Positive control consisted of cells treated with methanol for 15 min prior to exposure to the MTT reagent solution.
Results and discussion
Polymer preparation
PLGA-mPEG and PLGA-PEG-COOH copolymers were successfully prepared by the ring opening polymerization of lactide and glycolide dimers onto the hydroxyl group of poly(ethylene glycol) methyl ether or poly(ethylene glycol) carboxymethyl, respectively. 1H NMR was used to confirm the composition and estimate the molecular weight of the polymers. Figures S1 and S2 of the Supplementary Materials show the NMR spectra and peak assignments for these two copolymers. The peaks associated with PEG protons at 3.6 ppm, lactide protons at 5.2 and 1.6 ppm, and glycolide at 4.8 ppm are clearly present. The areas under the peaks were used to estimate the average molecular weight of the polymers based on the known starting molecular weight of the poly(ethylene glycol) methyl ether or poly(ethylene glycol) carboxymethyl precursors, as well as to determine the molar ratio of lactide to glycolide. Table 1 lists the characteristics of the copolymers.
Nanoparticle preparation and characterization
Drug-free and doxorubicin-loaded NPs prepared by nanoprecipitation were characterized by DLS, electron microscopy, laser Doppler microelectrophoresis, and fluorescence spectroscopy, respectively. Table 2 summarizes the size, zeta potential, drug loading, drug encapsulation efficiency, and percent recovery for each group of NPs produced. It should be noted that the characterization data provided in Table 2 is for NPs that have not been functionalized with an aptamer.
The NPs had a size of approximately 100 to 125 nm in diameter with a nearly monodispersed distribution (polydispersity indexes between 0.1 and 0.2). As can be seen in Table 2, and also in Fig. 2a, the size of the NPs increased linearly with increasing target drug loading. Nonetheless, all NPs were within the appropriate size range for use as intravenously injected formulations. It is well known that to enable injection and long blood circulation time, NPs should be between 20 and 150 nm in order to prevent premature renal clearance if too small or rapid sequestration by the reticuloendothelial system if too large (Durymanov et al. 2015). Figure 1b, c shows the representative transmission electron microscopy images of blank and drug-loaded NPs. As shown, the NPs have a spherical morphology.
The zeta potential of all NP formulations was negative (Table 2), as expected for these NPs due to the presentation of terminal carboxylic acid groups from the PLGA-PEG-COOH polymer on the surface of the particles. Optimal surface charge of NPs should be negative or neutral to increase their circulation time in the bloodstream and ability to transport deeper into tumors (minimum electrostatic interactions with negative extracellular matrix glycoproteins) and to reduce uptake by macrophages (Durymanov et al. 2015). In addition, the presence of poly(ethylene glycol) on the surface of the NPs ensures that they remain as a stable colloidal suspension via steric stabilization as a result of the high flexibility and hydrophilicity of this polymer (Avgoustakis 2004; Gref et al. 2000; Mehvar 2000; Otsuka et al. 2003). PEG is known to increase the residence time of molecules and nanocarriers in the circulation by shielding them against protein adsorption and delaying removal by the reticuloendothelial system (Prencipe et al. 2009; Torchilin and Trubetskoy 1995).
The experimentally determined drug loading also increased linearly with increasing target drug loading (Table 2 and Fig. 3d), suggesting that higher drug loading could be achieved. However, the drug entrapment efficiency was only approximately 40 % in all cases. This is expected as a result of the relatively high water solubility of doxorubicin in its salt form. Finally, the recovery of the NPs was better than 75 % in all cases. Here, recovery signifies the mass of NPs collected after nanoprecipitation and NP purification. Recovery is highly influenced by the centrifugation process utilized for removal of unencapsulated drug, which typically results in the loss of smaller particles that would require extended centrifugation times to collect.
Nanoparticle characterization. a Relationship between NP diameter as determined by DLS analysis to the target drug loading. b TEM image of blank NPs. c TEM image of doxorubicin-loaded NPs prepared with a target drug loading of 5 %. d Relationship between target drug loading and actual drug loading achieved. e DLS size distribution of NPs functionalized with TLS11a aptamer. f Zeta potential distribution of NPs functionalized with TLS11a aptamer. g Fluorescence of NPs functionalized with fluorescein-functionalized TLS11a aptamer and washed, demonstrating successful attachment (λ ex = 490 nm)
Aptamer-functionalized NPs had sizes and zeta potentials similar to those of the aptamer-free NPs. Figure 3e, f shows the representative size and zeta potential distribution profiles for NPs functionalized with the TLS11a aptamer. Aptamer attachment via EDC/NHS coupling chemistry was confirmed by fluorescence spectrophotometry of washed NPs (Fig. 3g). For this study, NPs were excited with 490-nm light, and the emission spectra were recorded between 510 and 700 nm. Fluorescence from the fluorescein-labeled aptamer is clearly observed.
Drug release
The release of doxorubicin from the NPs under physiological conditions was simulated by incubation of the particles in phosphate-buffered saline at pH 7.4. The method utilized enabled the quantification of the drug released to the supernatant, as well as that remaining within the NPs for increased accuracy. Figure 4 shows the doxorubicin release profile observed for these NPs. As shown, approximately 30 % of the drug were released in the first 6 h of the study, while 40, 80, and 90 % were released within 12, 24, and 36 h, respectively. After 36 h, no additional drug release was observed.
Utilizing the short-time approximation power law release model for spherical drug delivery systems (Ritger and Peppas 1987; Siepmann and Peppas 2001), it was determined that the diffusional constant n for this release process was 0.65, as shown in Fig. S3 of the Supplementary Materials. The value of the diffusional constant indicates that doxorubicin was released from the particles mainly through anomalous transport, i.e., through a process likely controlled by both Fickian diffusion and polymer relaxation (Ritger and Peppas 1987; Siepmann and Peppas 2001). It should be noted that in the analysis of drug release through this model, the data is treated as if the release had occurred from a purely monodispersed NP population. As demonstrated by Riger and Peppas, polydisperse size distributions can affect the analysis of the data (1987). Fortunately, the polydispersity of the NPs herein presented is small (Table 2), suggesting that the model is appropriate for this analysis.
Cell surface binding of aptamer-functionalized NPs
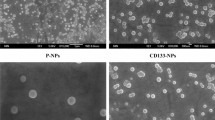
The cell surface binding of the TLS11a aptamer-functionalized NPs was examined in vitro using live MEAR cells and confocal microscopy. The aim of this experiment was to demonstrate adequate cellular binding (i.e., targeting) of the aptamer-functionalized nanoparticles using a previously characterized cell line known to interact with the TLS11a aptamer (Shangguan et al. 2008; Meng et al. 2012). After incubation with the drug-free aptamer-functionalized NPs for 30 min at 37 °C, the fluorescein-tagged aptamer-NP conjugates (green fluorescence channel) were found primarily near the cell surface, where they localized with the WGA/Alexa-Fluor®647 cell membrane marker (red fluorescence channel) (Fig. 5a). Some weak diffuse staining was apparent in the cytoplasm within the green channel; however, negative controls treated with blank NPs (i.e., no fluorescein-tagged aptamer) demonstrated a similar pattern and level of green fluorescence intensity within the cytoplasm, suggesting a low level of background autofluorescence at this 480/40-nm excitation and 535/50-nm emission wavelength range (Fig. 5b). While these results confirm the successful binding of the full aptamer-NP conjugates to the cell surface, time limitations for the cells to be maintained under non-physiological conditions without environmental control prevented tracking of cellular uptake beyond 30 min using the existing microscope setup. As such, functional studies with drug-loaded NPs were critical to assess uptake and drug delivery within the cell.
Cellular interactions of aptamer-functionalized NPs. a Confocal micrograph, single z slice, of live MEAR cells treated with fluorescein-tagged TLS11a-NPs (green) counterstained to visualize nuclei (blue) and the cell membrane (red) along with companion transmitted light micrograph in the right panel. b Negative control image of MEAR cells treated with blank NPs (no aptamer) and counterstained along with companion transmitted light image in the right panel. Brightness and contrast were adjusted for printing purposes and were applied universally to TLS11a-NP and control images
Determination of therapeutic efficacy
Next, the aptamer-functionalized NPs carrying the chemotherapeutic drug doxorubicin were used to treat MEAR HCC cells and an MTT viability assay was used to assess their ability to induce a therapeutic effect (i.e., cause a relative reduction in cell viability) versus unfunctionalized NPs and blank NP controls. As shown in Fig. 6, the TLS11a-functionalized NPs containing doxorubicin exhibited the strongest therapeutic effect on the MEAR cells in vitro with a reduction in cell viability of over 50 % relative to that of control even at the lowest doxorubicin concentration. Cells treated with doxorubicin-loaded NPs without the targeting aptamer consistently had higher cell viability than the TLS11a-targeted NPs, supporting the use of TLS11a as a useful targeting agent for HCC using this NP system. A reduction in the cell viability (~20 %) relative to control was seen in the presence of PLGA-mPEG/PLGA-PEG-COOH NPs alone (NP blank), demonstrating a low-level, albeit, non-trivial, cytotoxicity from the NPs themselves at high concentrations (>1 mg/mL). The free doxorubicin-HCl control yielded cytotoxicity levels comparable to the TLS11a-NPs. Free doxorubicin controls often exhibit an equivalent or even greater cytotoxic effect in vitro over doxorubicin packaged in a NP carrier (Meng et al. 2012; Pilapong et al. 2014; Zhang et al. 2014). This has been attributed to the ability of the free drug to readily permeate the cell membrane relative to the targeted NP carrier which requires additional time to adequately interact with and bind to cell surface markers, undergo endocytosis, and deliver therapeutic payload within the cell (Betancourt et al. 2007; Meng et al. 2012; Pilapong et al. 2014; Zhang et al. 2014). Thus, the true value of targeted NPs for drug delivery is not fully realized until tested in vivo. Separate MTT studies with the TLS11a aptamer alone in MEAR cells showed the inability of the aptamer to reduce cell viability compared to untreated control in the concentration range of 1000–8 nM (data not shown).
It should be also noted that while colorimetric viability assays such as the MTT study utilized in this work enable determination of the extent of therapeutic effect of targeted drug delivery system in vitro, they are not able to differentiate whether the relative reduction in cell viability of the target cancer cells is due to cytostatic or cytotoxic effects. Since the aptamer-targeted nanoparticles act as a carrier for the delivery of doxorubicin to the target cancer cells, the reduction in cell viability observed is most likely associated with the drug’s action. Doxorubicin is known to induce apoptosis and/or cell cycle arrest in cancer cells by intercalating into DNA, inhibiting macromolecule biosynthesis, generating reactive oxygen species, and inhibiting topoisomerase II function (Gewirtz 1999; Minotti et al. 2004). Nonetheless, future work will need to specifically demonstrate the mechanism of cell death caused by these targeted nanoparticles on hepatocellular carcinoma cells.
Conclusions
In this work we have combined a well-characterized, HCC-specific aptamer with a biodegradable polymer NP drug delivery system to demonstrate efficacy in treating HCC tumor cells in vitro. NPs consisting of PLGA-mPEG/PLGA-PEG-COOH polymers were successfully synthesized, loaded with the drug doxorubicin, and functionalized with the TLS11a HCC-specific aptamer. The aptamer-functionalized NPs effectively bound to HCC tumor cells, and functional studies with drug-loaded TLS11a-NPs showed high therapeutic effect versus NPs without the targeting aptamer. Taken together, these results provide a first step toward the development of TLS11a-functionalized NPs as a new therapeutic strategy to treat HCC tumors and support progression to further testing in vivo. Future studies will explore the aptamer-NP drug carrier effects on normal and cancerous liver cell lines at various stages of differentiation, investigate the mechanisms of action of the targeted nanoparticles, and further elucidate the cell surface molecule to which the TLS11a aptamer binds.
References
American Cancer Society (2016) Cancer facts & figures 2016.
Attwa MH (2015) Guide for diagnosis and treatment of hepatocellular carcinoma world. J Hepatol 7:1632–1632. doi:10.4254/wjh.v7.i12.1632
Avgoustakis K (2004) Pegylated poly(lactide) and poly(lactide-co-glycolide) nanoparticles: preparation, properties and possible applications in drug delivery. Curr Drug Deliv 1:321–333
Bareford LM, Swaan PW (2007) Endocytic mechanisms for targeted drug delivery. Adv Drug Deliv Rev 59:748–758. doi:10.1016/j.addr.2007.06.008
Betancourt T, Brown B, Brannon-Peppas L (2007) Doxorubicin-loaded PLGA nanoparticles by nanoprecipitation: preparation, characterization and in vitro evaluation. Nanomedicine (Lond) 2:219–232. doi:10.2217/17435889.2.2.219
Betancourt T et al (2009) PEGylation strategies for active targeting of PLA/PLGA nanoparticles. J Biomed Mater Res A 91:263–276. doi:10.1002/jbm.a.32247
Bruix J et al (2015) Adjuvant sorafenib for hepatocellular carcinoma after resection or ablation (STORM): a phase 3, randomised, double-blind, placebo-controlled trial. The Lancet Oncology 16:1344–1354. doi:10.1016/S1470-2045(15)00198-9
Byrne JD, Betancourt T, Brannon-Peppas L (2008) Active targeting schemes for nanoparticle systems in cancer therapeutics. Adv Drug Deliv Rev 60:1615–1626. doi:10.1016/j.addr.2008.08.005
Danhier F, Feron O, Préat V (2010) To exploit the tumor microenvironment: passive and active tumor targeting of nanocarriers for anti-cancer drug delivery. 148. doi:10.1016/j.jconrel.2010.08.027
Danquah MK, Zhang XA, Mahato RI (2011) Extravasation of polymeric nanomedicines across tumor vasculature. Adv Drug Deliv Rev 63:623–639. doi:10.1016/j.addr.2010.11.005
Dhanasekaran R, Limaye A, Cabrera R (2012) Hepatocellular carcinoma: current trends in worldwide epidemiology, risk factors, diagnosis, and therapeutics. Hepatic medicine : evidence and research 4:19–37. doi:10.2147/HMER.S16316
Dhar S, Gu FX, Langer R, Farokhzad OC, Lippard SJ (2008) Targeted delivery of cisplatin to prostate cancer cells by aptamer functionalized Pt(IV) prodrug-PLGA-PEG nanoparticles. Proc Natl Acad Sci U S A 105:17356–17361. doi:10.1073/pnas.0809154105
Durymanov MO, Rosenkranz AA, Sobolev AS (2015) Current approaches for improving intratumoral accumulation and distribution of nanomedicines. Theranostics 5:1007–1020. doi:10.7150/thno.11742
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346:818–822. doi:10.1038/346818a0
Farokhzad OC et al (2006) Targeted nanoparticle-aptamer bioconjugates for cancer chemotherapy in vivo. Proc Natl Acad Sci U S A 103:6315–6320. doi:10.1073/pnas.0601755103
Farokhzad OC, Jon S, Khademhosseini A, Tran TN, Lavan DA, Langer R (2004) Nanoparticle-aptamer bioconjugates: a new approach for targeting prostate cancer cells. Cancer Res 64:7668–7672. doi:10.1158/0008-5472.CAN-04-2550
Ferlay J et al (2015) Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer 136:E359–E386. doi:10.1002/ijc.29210
Forner A, Llovet JM, Bruix J (2012) Hepatocellular carcinoma. Lancet 379:1245–1255. doi:10.1016/S0140-6736(11)61347-0
Gewirtz DA (1999) A critical evaluation of the mechanisms of action proposed for the antitumor effects of the anthracycline antibiotics adriamycin and daunorubicin. Biochem Pharmacol 57:727–741
Gref R et al (2000) “Stealth” corona-core nanoparticles surface modified by polyethylene glycol (PEG): influences of the corona (PEG chain length and surface density) and of the core composition on phagocytic uptake and plasma protein adsorption. Colloids Surf B Biointerfaces 18:301–313
Hainfeld JF, O'Connor MJ, Dilmanian FA, Slatkin DN, Adams DJ, Smilowitz HM (2011) Micro-CT enables microlocalisation and quantification of Her2-targeted gold nanoparticles within tumour regions. Br J Radiol 84:526–533. doi:10.1259/bjr/42612922
Huang HS, Hainfeld JF (2013) Intravenous magnetic nanoparticle cancer hyperthermia. Int J Nanomedicine 8:2521–2532. doi:10.2147/IJN.S43770
Jiang J, Chen H, Yu C, Zhang Y, Chen M, Tian S, Sun C (2015) The promotion of salinomycin delivery to hepatocellular carcinoma cells through EGFR and CD133 aptamers conjugation by PLGA nanoparticles. Nanomedicine (Lond) 10:1863–1879. doi:10.2217/nnm.15.43
Kamaly N, Xiao Z, Valencia PM, Radovic-Moreno AF, Farokhzad OC (2012) Targeted polymeric therapeutic nanoparticles: design, development and clinical translation. Chem Soc Rev 41:2971–3010. doi:10.1039/c2cs15344k
Kashefi-Kheyrabadi L, Mehrgardi MA, Wiechec E, Turner APF, Tiwari A (2014) Ultrasensitive detection of human liver hepatocellular carcinoma cells using a label-free aptasensor. Anal Chem 86:4956–4960. doi:10.1021/ac500375p
Kaszuba M, Corbett J, Watson FM, Jones A (2010) High-concentration zeta potential measurements using light-scattering techniques. Philos Trans A Math Phys Eng Sci 368:4439–4451. doi:10.1098/rsta.2010.0175
Knox JJ, Cleary SP, Dawson LA (2015) Localized and systemic approaches to treating hepatocellular carcinoma. Journal of clinical oncology : official journal of the American Society of Clinical Oncology 33:1835–1844. doi:10.1200/JCO.2014.60.1153
Llovet JM et al (2008) Sorafenib in advanced hepatocellular carcinoma. N Engl J Med 359:378–390. doi:10.1056/NEJMoa0708857
Maeda H, Nakamura H, Fang J (2013) The EPR effect for macromolecular drug delivery to solid tumors: improvement of tumor uptake, lowering of systemic toxicity, and distinct tumor imaging in vivo. Adv Drug Deliv Rev 65:71–79. doi:10.1016/j.addr.2012.10.002
McDonald DM, Baluk P (2002) Significance of blood vessel leakiness in cancer. Cancer Res 62:5381–5385
Mehvar R (2000) Modulation of the pharmacokinetics and pharmacodynamics of proteins by polyethylene glycol conjugation. J Pharm Pharm Sci 3:125–136
Meng HM, Fu T, Zhang XB, Tan W (2015) Cell-SELEX-based aptamer-conjugated nanomaterials for cancer diagnosis and therapy. National Science Review 2:71–84. doi:10.1093/nsr/nwv001
Meng L, Yang L, Zhao X, Zhang L, Zhu H, Liu C, Tan W (2012) Targeted delivery of chemotherapy agents using a liver cancer-specific aptamer. PLoS One 7:e33434. doi:10.1371/journal.pone.0033434
Minotti G, Menna P, Salvatorelli E, Cairo G, Gianni L (2004) Anthracyclines: molecular advances and pharmacologic developments in antitumor activity and cardiotoxicity. Pharmacol Rev 56:185–229. doi:10.1124/pr.56.2.6
Onda N, Kemmochi S, Morita R, Ishihara Y, Shibutani M (2013) In vivo imaging of tissue-remodeling activity involving infiltration of macrophages by a systemically administered protease-activatable probe in colon cancer tissues. Transl Oncol 6:628–637
Otsuka H, Nagasaki Y, Kataoka K (2003) PEGylated nanoparticles for biological and pharmaceutical applications. Adv Drug Deliv Rev 55:403–419
Pilapong C, Sitthichai S, Thongtem S, Thongtem T (2014) Smart magnetic nanoparticle-aptamer probe for targeted imaging and treatment of hepatocellular carcinoma. Int J Pharm 473:469–474. doi:10.1016/j.ijpharm.2014.07.036
Prencipe G et al (2009) PEG branched polymer for functionalization of nanomaterials with ultralong blood circulation. J Am Chem Soc 131:4783–4787. doi:10.1021/ja809086q
Qu L, Xu J, Tan X, Liu Z, Xu L, Peng R (2014) Dual-aptamer modification generates a unique Interface for highly sensitive and specific electrochemical detection of tumor cells. ACS Appl Mater Interfaces 6:7309–7315. doi:10.1021/am5006783
Ritger PL, Peppas NA (1987) A simple equation for description of solute release I. Fickian and non-fickian release from non-swellable devices in the form of slabs, spheres, cylinders or discs. J Control Release 5:23–36. doi:10.1016/0168-3659(87)90034-4
Shangguan D et al (2008) Identification of liver cancer-specific aptamers using whole live cells. Anal Chem 80:721–728. doi:10.1021/ac701962v
Siepmann J, Peppas NA (2001) Modeling of drug release from delivery systems based on hydroxypropyl methylcellulose (HPMC). Adv Drug Deliv Rev 48:139–157
Stoltenburg R, Reinemann C, Strehlitz B (2007) SELEX—a (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomol Eng 24:381–403. doi:10.1016/j.bioeng.2007.06.001
Sun D, Lu J, Chen Z, Yu Y, Mo M (2015) A repeatable assembling and disassembling electrochemical aptamer cytosensor for ultrasensitive and highly selective detection of human liver cancer cells. Anal Chim Acta 885:166–173. doi:10.1016/j.aca.2015.05.027
Sun D, Lu J, Zhong Y, Yu Y, Wang Y, Zhang B, Chen Z (2016) Sensitive electrochemical aptamer cytosensor for highly specific detection of cancer cells based on the hybrid nanoelectrocatalysts and enzyme for signal amplification. Biosens Bioelectron 75:301–307. doi:10.1016/j.bios.2015.08.056
Sun H, Zhu X, Lu PY, Rosato RR, Tan W, Zu Y (2014) Oligonucleotide aptamers: new tools for targeted cancer therapy. Molecular therapy Nucleic acids 3:e182–e182. doi:10.1038/mtna.2014.32
Torchilin VP, Trubetskoy VS (1995) Which polymers can make nanoparticulate drug carriers long-circulating. Adv Drug Deliv Rev 16:141–155. doi:10.1016/0169-409x(95)00022-Y
Wang H, Wang F, Tao X, Cheng H (2012) Ammonia-containing dimethyl sulfoxide: an improved solvent for the dissolution of formazan crystals in the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) assay. Anal Biochem 421:324–326. doi:10.1016/j.ab.2011.10.043
Weigum SE, Sutton M, Barnes E, Miller S, Betancourt T (2014) Targeting hepatocellular carcinoma with aptamer-functionalized PLGA/PLA-PEG nanoparticles. 9166:916605–916605 doi:10.1117/12.2062283
Xu J et al (2015) Molecular recognition of human liver cancer cells using DNA aptamers generated via cell-SELEX. PLoS One 10:e0125863. doi:10.1371/journal.pone.0125863
Yu D, Zhang Y, Mao Z, Gao C (2013) Study of the selective uptake progress of aptamer-modified PLGA particles by liver cells. Macromol Biosci 13:1413–1421. doi:10.1002/mabi.201300165
Yu MK, Park J, Jon S (2012) Targeting strategies for multifunctional nanoparticles in cancer imaging and therapy. Theranostics 2:3–44. doi:10.7150/thno.3463
Zhang B, Luo Z, Liu J, Ding X, Li J, Cai K (2014) Cytochrome c end-capped mesoporous silica nanoparticles as redox-responsive drug delivery vehicles for liver tumor-targeted triplex therapy in vitro and in vivo. Journal of controlled release : official journal of the Controlled Release Society 192:192–201. doi:10.1016/j.jconrel.2014.06.037
Acknowledgments
This research was supported in part by the Texas Emergent Technology Fund (start-up funds to TB), the Robert A. Welch Foundation (AI-0045), NIGMS South Texas Doctoral Bridge Program (R25-GM102783) fellowship to C. Munoz, and Texas State University (start-up funds to SW).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
ESM 1
(DOCX 1171 kb)
Rights and permissions
About this article
Cite this article
Weigum, S., McIvor, E., Munoz, C. et al. Targeted therapy of hepatocellular carcinoma with aptamer-functionalized biodegradable nanoparticles. J Nanopart Res 18, 341 (2016). https://doi.org/10.1007/s11051-016-3633-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11051-016-3633-5