Abstract
Although yeasts belonging to the genus Candida are frequently seen as commensals in the oral cavity, they possess virulence attributes that contribute for pathogenicity. The aims of the present study were to study the prevalence of Candida spp. isolated from the oral cavity of renal transplant recipients and to analyze strains virulence factors. We isolated a total of 70 Candida strains from 111 transplant recipients, and Candida albicans was the most prevalent species (82.86 %). Oral candidiasis was diagnosed in 14.4 % kidney transplant patients, while 11 isolates (15.7 %) corresponded to non-Candida albicans Candida (NCAC) species. C. albicans adhered to a higher extension than NCAC strains. Some isolates of Candida tropicalis were markedly adherent to human buccal epithelial cells and highly biofilm-forming strains. Regarding proteinase activity, Candida orthopsilosis was more proteolytic than Candida metapsilosis. Candida glabrata and Candida dubliniensis showed very low ability to form biofilm on polystyrene microtiter plates. We have demonstrated here diverse peculiarities of different Candida species regarding the ability to express virulence factors. This study will contribute for the understanding of the natural history and pathogenesis of yeasts belonging to the genus Candida in the oral cavity of patients who were submitted to kidney transplant and are under immunosuppressive therapies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Candida comprises yeasts that colonize the human oral cavity as commensal organisms in 20–80 % of adults without evidences of infection [1]. Nevertheless, under certain circumstances, specifically among immunosuppressed individuals, such as AIDS patients and transplant organ recipients, they may cause opportunistic diseases ranging from superficial infections of mucous membrane to systemic infections leading to life-threatening diseases [2].
Oral candidiasis is an opportunistic infection commonly observed among renal transplant recipients [3]. The prevalence of this disease for this group of patients ranges from 9.4 to 46.7 % [3]. Candida albicans is the most frequent species associated with oral lesions, but other less virulent Candida species such as Candida glabrata, Candida tropicalis, Candida parapsilosis, Candida krusei and Candida dubliniensis have also been isolated from the saliva of patients with or without oral candidiasis [4].
Kidney transplant is the treatment of choice for treating patients with chronic renal insufficiency. The main oral manifestations cited in the literature of kidney transplant patients under immunosuppressive therapy are the following: oral candidiasis, gingival hyperplasia, herpetic infections and pilous leukoplakia [5].
The transition of opportunistic yeasts from commensalism to infection is related to the virulence factors as well as to host susceptibility [6]. Several putative virulence factors contribute to Candida spp. pathogenicity, including the ability to adhere to epithelial and endothelial cells, yeast-to-hypha transition, phenotypic switching and the production of hydrolytic enzymes, specifically phospholipases and proteinases and biofilm formation [7–9].
Although several factors may contribute for the development of oral candidiasis among kidney transplant recipients, virulence factors of Candida species isolated from the oral cavity of this group of patients were not yet determined in this specific clinical scenario. Therefore, the objective of the present study was to evaluate the prevalence of yeasts isolated from the oral cavity of kidney transplant recipients and some in vitro virulence factors of the isolates, including the ability to adhere to human buccal epithelial cells (HBEC), biofilm formation and proteinase production. This study will contribute to the discussion of the essential pathogenicity factors important for the transition from commensalism to infection of Candida spp. during oral candidiasis in renal transplant patients.
Materials and Methods
Patients
We evaluated a total of 111 kidney transplant recipients followed up at the Onofre Lopes University Hospital, Natal, Brazil, from April to July 2008. The patients were subjected to clinical examination of the oral cavity for candidiasis diagnosis, according to the criteria established for oral lesions in HIV patients recommended by the EC-Clearinghouse and World Health Organization classifications [10]. Only patients who agreed to take part on a surveillance confidential study, in accordance with the Local Research Ethics committee from the referred Hospital, were enrolled in this study. Patients were under immunosuppressive therapy during the whole period of the study.
Samples Collection and Yeasts Identification
Samples containing 2 mL of saliva were collected from all patients, by previous stimulation with chewing gums [11]. Subsequently, 100 μL of cell suspensions was inoculated on the surface of Sabouraud Dextrose Agar (SDA; Oxoid, UK) added 300 μg/mL of chloramphenicol (Park–Davis), using a Drigalsky loop. The plates were incubated at 37 °C for 48 h. Yeast colonies were plated on CHROMagar Candida® (CHROMagar Microbiology, Paris, France) to check for purity and screening for different color colonies. Species identification was based on the characteristics of the cells observed microscopically after cultivation on cornmeal agar added Tween 80, as well as assimilation and fermentation testing and ID32C System (bioMérieux Marcy l’Etoile, France), whenever it was necessary [12]. C. dubliniensis presumptive identification was based on its inability to grow on Sabouraud dextrose broth with 6.5 % NaCl (Sigma-Aldrich, Brazil), after 72 h of incubation at 37 °C [13].
Molecular Identification
We performed molecular methods to identify the cryptic species C. dubliniensis and the species of the C. parapsilosis species complex (C. parapsilosis, Candida orthopsilosis and Candida metapsilosis).
DNA Extraction
Candida albicans cells were grown overnight in YPD liquid medium (dextrose 20 g/L, peptone 20 g/L, yeast extract 10 g/L) incubated at 30 °C and rotated at 200 rpm in a gyratory shaker (TE-420, Tecnal® Piracicaba, Brazil). DNA was extracted using the PrepMan Ultra sample preparation reagent (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s instructions. Genomic DNA concentration and purity were checked with a NanoDrop instrument (Thermo Scientific; Amersham Pharmacia Biotech, Wilmington, DE, USA).
Differentiation Among C. albicans and C. dubliniensis
In order to double-check the C. dubliniensis presumptive identification, the isolates previously identified by phenotypic methods as C. dubliniensis were identified with ABC genotyping, which is based on the amplification of an intron present in the 26 S rDNA, where C. dubliniensis is considered to belong to the former C. albicans genotype D [14]. C. albicans ATCC 90028 and C. dubliniensis CBS 7987 were used as control strains. The ABC identification was performed with the following primers: CA–INT–L (5′-ATA AGG GAA GTC GGC AAA ATA GAT CCG TAA-3′) and CA–INT–R (5′-CCT TGG CTG TGG TTT CGC TAG ATA GTA GAT-3′; [14]. Briefly, 1.0 μL of DNA 40 ng/μL was added to 2× PCR Master Mix (Promega) to a final volume of 25 μL. The samples were amplified in a Thermocycler (Amplitherm TX 96, USA) using the following cycling parameters: one initial cycle of 94 °C for 3 min followed by 30 cycles of 1 min at 94 °C, 1 min at 57 °C, 1 min at 72 °C and a final cycle of 5 min at 72 °C. PCR products were size-separated by agarose gel electrophoresis, and the gel was stained in a 0.5 μg/mL ethidium bromide buffer solution (1× TAE).
Differentiation of the Species Belonging to C. parapsilosis Species Complex
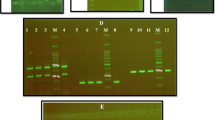
To differentiate the cryptic species belonging to the C. parapsilosis species complex, we have used the RAPD primer RPO2 validated in combination with DNA sequencing by Ge et al. [15]. For RAPD reactions, 1.0 μL of DNA 40 ng/μL, 2.5 μL of 10× PCR buffer (100 mM Tris–HCl, pH 8.3, 500 mM KCl, 3.5 mM MgCl2), 5 μL of dNTPmix (100 mM each dNTP), 1.0 μL of primer (50 pmol/μL), 0.13 μL of Tween 20 and 1.0 unit of Taq DNA polymerase were added to a final volume of 25 μL. Amplifications were performed in a thermal circler Amplitherm TX 96 (USA) with the following three-step program: 2 min at 94 °C followed by 45 cycles of 1 min at 94 °C, 1 min at 36 °C, 75 s at 72 °C and then 10 min at 72 °C. PCR products were size-separated by agarose gel electrophoresis at 100 V for 30 min followed by 55 V for 5.5 h in a Tris–acetate buffer (TAE). The gel was stained in a 0.5 μg/mL ethidium bromide buffer solution (1× TAE) for 10 min and distained twice for 15 min in 400 mL of water.
Candida spp. Virulence Factors
For the Candida spp. virulence factors tested in vitro, C. albicans ATCC90028, C. tropicalis 13803, C. dubliniensis CBS797, C. glabrata ATCC2001, C. orthopsilosis ATCC96143 and C. metapsilosis ATCC96139 were used as reference controls. For these experiments, we randomly selected 26 isolates of C. albicans and included all non-Candida albicans Candida (NCAC) strains isolated (n = 11).
Candida spp. Adherence to Human Buccal Epithelial Cells
Candida cells were grown overnight to stationary phase in NGY 0.1 % (Neopeptone [Difco], 0.4 % glucose and 0.1 % yeast extract [Difco]) at 37 °C and were mixed with HBEC from healthy volunteers at a ratio of 10 yeast cells per HBEC. The mixtures were incubated at 37 °C for 45 min with shaking; then cells were vortexed, formalin-fixed and transferred to a microscope slide. The number of Candida cells adhering to 150 HBEC was determined with the operator blinded to the nature of the material on the slide. Tests were done in triplicate [16].
Candida spp. Proteinase Activity
Proteinase activity was determined by a method of Macdonald and Odds [17]. Fifty microliter samples from NGY cultures were grown in 5 mL YCB + BSA medium (11.7 g/L Yeast Carbon Base [Difco]; 10 g/L glucose; 5 g/L bovine serum albumin, fraction V, Batch 08k0560 [Sigma]) rotated in a rotator shaker at 30 °C for 72 h, 200 rpm. Proteolytic activity was determined by measuring the increase in trichloroacetic acid-soluble products absorbing at 280 nm in triplicate after 1-h incubation of the culture supernatant with BSA substrate at 37 °C. Specific activity was expressed as OD280/OD600 of the culture.
Candida spp. Biofilm Formation
Biofilms were performed according to Melo et al. [18]. At first, 100 μL aliquots of a standardized cell suspension (107 cells/mL) were transferred to flat-bottom 96-well microtiter plates and incubated for 1.5 h at 37 °C in a shaker at 75 rpm. As controls, eight wells of each microtiter plate were handled in an identical fashion, except that no Candida suspensions were added. Following the adhesion phase, the cell suspensions were aspirated and each well was washed twice with 150 μL of PBS to remove loosely adherent cells. A total of 100 μL of RPMI-1640 medium buffered with MOPS was added to each of the washed wells and incubated at 37 °C in a shaker at 75 rpm. The biofilms were allowed to develop for 72 h, and then the yeasts were quantified by the crystal violet assay. Briefly, the biofilm-coated wells of microtiter plates were washed twice with 150 μL of PBS and then air-dried for 45 min. Then, each of the washed wells was stained with 110 μL of 0.4 % aqueous crystal violet solution for 45 min. Afterward, each well was washed four times with 350 μL of sterile distilled water and immediately destained with 200 μL of 95 % ethanol. After 45 min of destaining, 100 μL of destaining solution was transferred to a new well and the amount of the crystal violet stain in the destaining solution was measured with a microtiter plate reader (SpectraMAX 340 Tunable Microplate Reader; Molecular Devices Ltd.) at 595 nm. The absorbance values for the controls were subtracted from the values for the test wells to minimize background interference.
Statistical Analysis
Data were analyzed using the statistical software “GraphPad,” version 3.0. Results were presented as mean ± standard deviation, and differences were analyzed by the Mann–Whitney test. For all the analyses, P was considered a default value of 0.05 and the confidence interval of 95 %.
Results
Patients Demographic Data
Patient’s clinical and demographic data are summarized in Table 1.
Microbiological Profiling
Yeasts were isolated from the oral cavity of 70 out of 111 kidney transplant patients (63.1 %). C. albicans was the most prevalent species with 58 isolates (82.86 %), while 11 isolates (15.7 %) corresponded to NCAC, such as C. tropicalis (3 isolates; 4.35 %), C. orthopsilosis (2 isolates; 2.89 %), C. metapsilosis (2 isolates; 2.89 %), C. glabrata (2 isolates; 2.89 %) and C. dubliniensis (2 isolates; 2.89 %).
From the 111 kidney transplant recipients, we were able to diagnose oral candidiasis in 16 (14.4 %) of them, while 54 (48.6 %) individuals were only colonized by Candida spp. On the other hand, 41 (36.9 %) patients had negative Candida culture. In the patients diagnosed with oral candidiasis, atrophic candidiasis was the most prevalent clinical symptom (supplementary material). Interestingly, from the 70 kidney transplant patients whose yeasts were isolated from the oral cavity (colonization or infection), 33 had denture prosthesis. Twelve out of 16 patients with oral candidiasis had dental prosthesis. Therefore, the relationship between oral candidiasis and dentures prosthesis wearing was considered significant (P = 0.01). Patients only colonized by Candida spp. did not show any symptoms compatible to oral candidiasis. It seems that the specific immunosuppressive regimen adopted did not influence the positivity of cultures, because the percentage of patients with positive cultures ranged from 55.5 to 68 % independently of the immunosuppressors combination adopted (Table 2).
Adhesion to HBEC
Due to our limited number of NCAC strains representative for each species, we compared in vitro attributes of virulence among two groups: 26 C. albicans isolates versus 11 NCAC isolates. The ability of the Candida spp. isolates to adhere to HBEC was expressed by the number of blastoconidia adhered to 150 HBEC, determined with optical microscopy. In general, C. albicans isolates were highly capable to adhere to HBEC with an average of 237.38 blastoconidia/150 HBEC (Table 4). Nevertheless, an enormous variation among the isolates was observed. The number of C. albicans cells adhered to buccal epithelia ranged from 60 ± 3 to 379 ± 29/150 HBEC (Tables 3, 4). NCAC strains were statistically significant less adherent to HBEC then C. albicans isolates (P = 0.01; Table 4). Interestingly, an isolate of C. tropicalis was highly adherent to HBEC (335 ± 20/150 HBEC; Table 3).
Proteinase Activity
All the strains evaluated were proteinase producers. Proteinase activity was determined by growing cells in the presence of albumin as a sole nitrogen source. The enzymatic activity ranged from 0.01 to 0.08 (Table 3). We could not detect differences for proteinase production between C. albicans versus NCAC strains. However, a comparison within C. parapsilosis species complex showed that all C. orthopsilosis strains produced higher amounts of proteinase than C. metapsilosis isolates, including reference strains (mean of 0.06 ± 0.01 and 0.03 ± 0.01, respectively; P = 0.02.)
Biofilm Formation
Biofilms were formed when cells were grown on microtiter plates at 37 °C for 72 h of incubation. Once again, we could not detect a statistically significant difference between the two groups evaluated. Nevertheless, we noticed a C. tropicalis strain that was a high biofilm producer (1.82 ± 0.21); C. glabrata and C. dubliniensis were in general low biofilm producers both with a mean equal to 0.02 ± 0.01).
Virulence Factors of Candida Species Versus the Presence of Oral Candidiasis
We could not find a statistically significant correlation for any of the virulence factors evaluated and patient’s clinical condition (colonization × infection) when all the strains for both groups were evaluated.
Discussion
Here, we evaluated the prevalence of oral candidiasis as well as colonization rates among patients submitted to renal transplant. We observed that 14.4 % of our patients showed clinical signs and symptoms compatible with oral candidiasis, while 48.6 % of them were colonized by Candida spp. The high number of patients colonized in the oral cavity with Candida spp. is threatening, because previous colonization may lead to infection. The prevalence of oral candidiasis in this specific group of patients ranges from 9.4 to 46.7 % [3, 5]. This noticeable variation may be related to the different procedures applied in the different studies, regarding the collection and processing of saliva samples and the different levels of immunosuppression.
Several studies have demonstrated that erythematous candidiasis is the major clinical manifestation of oral candidiasis [3, 19–21]. Nevertheless, our investigation showed chronic atrophic candidiasis as the main clinical manifestation. This fact might be related to the high frequency of our denture-wearing population associated with poor hygiene habits favoring the development of this manifestation of oral candidiasis. We also found a statistically significant association between wearing dentures and the presence of oral candidiasis (P = 0.01), corroborating the investigation of Darwazeh et al. [22], who reported that the presence of prosthetic dentures is a risk factor for developing oral candidiasis.
We found that C. albicans was the main species isolated from the oral cavity of our patients, corresponding to 82.86 %. C. albicans is still the most frequently isolated species from the oral cavity among immunosuppressed patients [23, 24]. Nevertheless, other Candida species have been implicated as etiological agents of oral candidiasis, such as C. glabrata [23, 25].
Most of the studies have demonstrated that C. albicans is the most adherent Candida species to HBEC [26, 27]. This trend was also observed in our study. In fact, we also observed a highly adherent C. tropicalis strain. C. tropicalis isolates’ adherence to laminin and fibronectin was described as significantly higher than that obtained for the C. albicans isolates [28].
Regarding proteinase production, we have found that C. orthopsilosis isolates were lower proteinase producers than C. metapsilosis strains. In fact, Sabino et al. [29] showed that C. orthopsilosis were SAP producers, whereas C. metapsilosis were not.
We found a tremendous variation regarding the ability of the strains to form biofilm. An interesting finding is that we could detect a very strong C. tropicalis biofilm-forming strain. C. tropicalis clinical isolates have been classified as abundant biofilm formers [21], and their mature biofilms consist of a dense network of yeast cells in addition to evident filamentous morphologies [30].
In the present study, C. dubliniensis showed lower ability to form biofilm on polystyrene microtiter plates, suggesting clear differences between the two phenotypically undistinguishable species C. albicans and C. dubliniensis. Biofilm formation by C. albicans isolates has been reported as statistically significant higher than by C. dubliniensis [31]. In addition, all C. glabrata strains were low biofilm producers. This fact is also in agreement with the findings of Silva et al. [21] and Shin et al. [32] who reported that biofilm formation by C. glabrata is lower than in other NCAC species.
Studies regarding oral candidiasis in kidney transplant recipients are mostly concerning the epidemiology of Candida spp. and clinical manifestations. In respect of that, we could find that chronic atrophic candidiasis is the main clinical manifestation in our specific clinical scenario. Nevertheless, we also describe here some peculiarities of Candida spp. virulence factors. For instance, C. albicans was very adherent to HBEC, but some strains of C. tropicalis may be also highly adherent to buccal epithelia and biofilm producer. On the other hand, C. glabrata and C. dubliniensis may be less effective in biofilm formation. Further investigations with a higher number of NCAC strains should be performed in order to confirm whether our findings follow the same trend in larger population studies.
References
Mendonça FH, Santos SS, Faria Ida S, Gonçalves e Silva CR, Jorge AO, Leão MV. Effects of probiotic bacteria on Candida presence and IgA anti-Candida in the oral cavity of elderly. Braz Dent J. 2012;23:534–8.
Jin Y, Samaranayake LP, Samaranayake Y, Yip HK. Biofilm formation of Candida albicans is variably affected by saliva and dietary sugars. Arch Oral Biol. 2004;49:789–98.
Anees MM, Reich A, Hirschberg L, Watorek E, El-Shinnawi UM, Ibrahiem TM, et al. Enhanced enzymatic activity of Candida species responsible for oral candidiasis in renal transplant recipients. Mycoses. 2011;54:337–44.
Zahir RA, Himratul-Aznita WH. Distribution of Candida in the oral cavity and its differentiation based on the internally transcribed spacer (ITS) regions of rDNA. Yeast. 2013;30:13–23.
de la Rosa-Garcia E, Mondragon-Padilla A, Irigoyen-Camacho ME, Bustamante-Ramirez MA. Oral lesions in a group of kidney transplant patients. Med Oral Patol Oral Cir Bucal. 2005;10:196–204.
Silva S, Negri M, Henriques M, Oliveira R, Williams DW, Azeredo J. Adherence and biofilm formation of non-Candida albicans Candida species. Trends Microbiol. 2011;19:241–7.
Mayer FL, Wilson D, Hube B. Candida albicans pathogenicity mechanisms. Virulence. 2013; 9:4 [Epub ahead of print].
Lyon JP, de Resende MA. Correlation between adhesion, enzyme production, and susceptibility to fluconazole in Candida albicans obtained from denture wearers. Oral Surg Oral Med Oral Pathol Oral Radiol Endod. 2006;102:632–8.
Nikawa H, Egusa H, Makihira S, Okamoto T, Kurihara H, Shiba H, et al. An in vitro evaluation of the adhesion of Candida species to oral and lung tissue cells. Mycoses. 2006;49:14–7.
Classification and diagnostic criteria for oral lesions in HIV infection. EC-Clearinghouse on oral problems related to HIV infection and WHO Collaborating Centre on oral manifestations of the immunodeficiency virus. J Oral Pathol Med. 1993;22:289–91.
Torres SR, Peixoto CB, Caldas DM, Silva EB, Akiti T, Nucci M, et al. Relationship between salivary flow rates and Candida counts in subjects with xerostomia. Oral Surg Oral Med Oral Pathol Oral Radiol Endod. 2002;93:149–54.
Pincus DH, Orenga S, Chatellier S. Yeast identification—past, present, and future methods. Med Mycol. 2007;45:97–121.
Alves SH, Milan EP, de Laet Sant’Ana P, Oliveira LO, Santurio JM, Colombo AL. Hypertonic sabouraud broth as a simple and powerful test for Candida dubliniensis screening. Diagn Microbiol Infect Dis. 2002;43:85–6.
McCullough MJ, Clemons KV, Stevens DA. Molecular and phenotypic characterization of genotypic Candida albicans subgroups and comparison with Candida dubliniensis and Candida stellatoidea. J Clin Microbiol. 1999;37:417–21.
Ge YP, Boekhout T, Zhan P, Lu GX, Shen YN, Li M, et al. Characterization of the Candida parapsilosis complex in East China: species distribution differs among cities. Med Mycol. 2012;50:56–66.
Chaves GM, Bates S, Maccallum DM, Odds FC. Candida albicans GRX2, encoding a putative glutaredoxin, is required for virulence in a murine model. Genet Mol Res. 2007;6:1051–63.
Macdonald F, Odds FC. Inducible proteinase of Candida albicans in diagnostic serology and in the pathogenesis of systemic candidosis. J Med Microbiol. 1980;13:423–35.
Melo AS, Bizerra FC, Freymüller E, Arthington-Skaggs BA, Colombo AL. Biofilm production and evaluation of antifungal susceptibility amongst clinical Candida spp. isolates, including strains of the Candida parapsilosis complex. Med Mycol. 2011;49:253–62.
Dunne WM Jr. Bacterial adhesion: seen any good biofilms lately? Clin Microbiol Rev. 2002;15:155–66.
Trofa D, Gacser A, Nosanchuk JD. Candida parapsilosis, an emerging fungal pathogen. Clin Microbiol Rev. 2008;21:606–25.
Silva S, Henriques M, Martins A, Oliveira R, Williams D, Azeredo J. Biofilms of non-Candida albicans Candida species: quantification, structure and matrix composition. Med Mycol. 2009;47:681–9.
Darwazeh AM, Al-Refai S, Al-Mojaiwel S. Isolation of Candida species from the oral cavity and fingertips of complete denture wearers. J Prosthet Dent. 2001;86:420–3.
Dongari-Bagtzoglou A, Dwivedi P, Ioannidou E, Shaqman M, Hull D, Burleson J. Oral Candida infection and colonization in solid organ transplant recipients. Oral Microbiol Immunol. 2009;24:249–54.
Golecka M, Oldakowska-Jedynak U, Mierzwinska-Nastalska E, Adamczyk-Sosinska E. Candida-associated denture stomatitis in patients after immunosuppression therapy. Transplant Proc. 2006;38:155–6.
Li L, Redding S, Dongari-Bagtzoglou A. Candida glabrata: an emerging oral opportunistic pathogen. J Dent Res. 2007;86:204–15.
Biasoli MS, Tosello ME, Magaro HM. Adherence of Candida strains isolated from the human gastrointestinal tract. Mycoses. 2002;45:465–9.
Lyon JP, de Resende MA. Evaluation of adhesion to buccal epithelial cells in Candida species obtained from denture wearers after exposure to fluconazole. Mycoses. 2007;50:21–4.
Costa KR, Ferreira JC, Lavrador MA, Baruffi MD, Candido RC. Virulence attributes and genetic variability of oral Candida albicans and Candida tropicalis isolates. Mycoses. 2012;55:e97–105.
Sabino R, Sampaio P, Carneiro C, Rosado L, Pais C. Isolates from hospital environments are the most virulent of the Candida parapsilosis complex. BMC Microbiol. 2011;8:11–80.
Silva S, Negri M, Henriques M, Oliveira R, Williams DW, Azeredo J. Candida glabrata, Candida parapsilosis and Candida tropicalis: biology, epidemiology, pathogenicity and antifungal resistance. FEMS Microbiol Rev. 2012;36:288–305.
Villar-Vidal M, Marcos-Arias C, Eraso E, Quindos G. Variation in biofilm formation among blood and oral isolates of Candida albicans and Candida dubliniensis. Enferm Infecc Microbiol Clin. 2011;29:660–5.
Shin JH, Kee SJ, Shin MG, Kim SH, Shin DH, Lee SK, et al. Biofilm production by isolates of Candida species recovered from nonneutropenic patients: comparison of bloodstream isolates with isolates from other sources. J Clin Microbiol. 2002;40:1244–8.
Acknowledgments
We thank the Brazilian Ministry of Education, on behalf of CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior), grant Procad-NF N° 08/2008. This work was also supported by the Postgraduate Programme of Pharmaceutical Sicences, from the Federal University of Rio Grande do Norte, Brazil. We are very grateful to Prof. Arnaldo Colombo for the donation of Candida spp. reference strains.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chaves, G.M., Diniz, M.G., da Silva-Rocha, W.P. et al. Species Distribution and Virulence Factors of Candida spp. Isolated from the Oral Cavity of Kidney Transplant Recipients in Brazil. Mycopathologia 175, 255–263 (2013). https://doi.org/10.1007/s11046-013-9640-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-013-9640-5