Abstract
The identification of MR images of the brain with tumours is one of the most critical tasks of any brain tumour (BT) detection system. Interestingly, because of its non-invasive image properties, the field of BT research has used the principles of medical image treatment, in particular in MR images. Diagnostic or detection systems assisted by a computer are becoming more complex and are still an open issue because of variations in tumour types, areas, and sizes. This study investigates the use of deep classification methods with deep features to identify the tumorous brain MR images. The proposed scheme involves three pre-trained networks, namely alexnet, vgg16 & vgg19, and their deep features. Again, a deep fusion approach is implemented to increase classification model performance. Also, the performance of classifiers is examined with the introduction of principal component analysis (PCA) to reduce the feature vector dimensions. The uniqueness of the proposed approach; it avoids the reproduction of MR images. The reproduction techniques generate anatomically incorrect images and are under investigation. In this research, small dataset in its original form is used. So, prejudice diagnosis is avoided. So, even if the proposed approach is data constrained but competent enough with the state-of-art. The proposed approach, i.e., vgg16 with the fused features of fc6 and fc7 without any dimension reduction techniques with linear SVM achieved the maximum value of accuracy is 0.9789, sensitivity is 1, specificity is 1, precision is 1, and F1 Score 97.92.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Due to the growing technology in medical image processing, Brain tumour (BT) and its study are of great interest. According to the National Brain Tumour Foundation (NBTF) survey, the production of BT among masses and the death rate due to brain tumours were successful globally in the previous year’s statistics [32, 49]. In addition, many researchers have suggested structures or methods in recent decades to illustrate the BT region that may or may not be accompanied by phases such as classification, therapy preparation, and outcome predictions. “Medical image BT segmentation is critical and is normally governed by variables such as poor contrast, noise, and missing boundaries [16]“. Diagnostic techniques such as magnetic resonance (MR) imaging, positron emission tomography (PET), and computed tomography (CT) scans are used to monitor these variables accurately. In detecting various types of diseases, these imaging processes are beneficial.
“The use of harmless magnetic fields and radio waves in the effective diagnosis and treatment of BT makes MR images very common [30]“. For diagnosis, accurate identification and precise localization of abnormal tissues are essential. This fact is completely supported by successful segmentation or classification methods or their combination, both quantitatively and qualitatively, for brain characterization. MR images can be processed based on human interactivity using manual [26, 50, 88], semi-automatic [13, 22, 86], and fully automatic [27, 31, 35, 82] techniques. Segmentation/classification should be precise in medical image processing, which is usually done manually by experts and hence time-consuming. “At the same time, developing a fully automated and effective approach to segmentation is still far from fact. In addition, such structures often require a second opinion, as human life is the main object here [83]“. The efficiency of automated techniques depends solely on the knowledge base, even in the absence of experts. Researchers have proposed several approaches to develop these knowledge bases and thus the potential of tumour detection systems. The clinical and pathological identification of all strategies depends solely on the ease of measurement and the degree of consumer supervision [29].
AlexNet and VGGNet are the milestones in the CNN development and basement of various computer vision tasks. In addition, the performance of CNNs is publicly validated on the annual ImageNet Large Scale Image Recognition Challenge (ILSIRC), between the year 2010 to 2015 [66]. It reveals besides shallow networks, the AlexNet and VGGNet achieved less Top-5 error. Further, those CNN models developed after 2015 having more than 100 layers, which increases the computational complexity. So, AlexNet, VGG16, and VGG 19 are considered for this research, which has 8,16 and 19 layers, respectively.
The work in this paper aimed to evaluate the performances of the pre-trained network, such as AlexNet, VGG16, and VGG19, combined with SVM for detecting the BT using the 2D brain MRI slices. The SVM is introduced for classification instead of SoftMax as in the original network. The detection process was performed: (1) using individual deep features such as fc6, fc7, and fc8, (2) using the fusion of two deep feature, i.e., fc6 & fc7, (3) using fusion of three deep features, i.e., fc6, fc7 & fc8, (4) using three deep features with PCA. After finding the best suitable feature or fused features of the pre-trained network, the top two features or fused features are again considered for fusion for performing the task. In this work, the performance of the proposed system was confirmed by computing the accuracy, precision, sensitivity, specificity, FPR, and F1-Score.
The significant contribution of this article is as follows:
-
1.
A data constraint approach for detection of BT based on the deep convolutional network.
-
2.
A deep feature based extensive comparison is carried out with the fusion technique.
-
3.
The proposed model avoids the reproduction of MR images. The reproduction techniques generate anatomically incorrect images and are under investigation. It uses small dataset in its original form. So, prejudice diagnosis is avoided.
-
4.
Achieved results are evaluated with recent methods, which prove that the proposed model performed better than existing techniques.
The remaining article is set out as follows. Section 2 presents related study. The theoretical background is discussed in section 3. A detail of the proposed method is presented in section 4. Experimental results are given in Section 5, which include findings and comparison with the existing methods. Finally, the research is concluded in section 6.
2 Related study
Researchers with a chosen machine learning (ML) or deep learning (DL) technique are proposing and implementing a large number of traditional and current BT detection procedures in the literature [36, 41, 48, 63, 84, 87]. “The main goal of the new automated and semi-automated disease assessment technique is to establish an effective method of disease identification to support the doctor during the process of diagnosis and treatment planning [19]“. “Due to its superiority and detection precision, most of the new disease diagnostic systems incorporate the DL technique [59]“. A transfer-learning-based deep learning architecture (DLA) was introduced in the work of Talo et al. [78] to detect tumours using 2D MRI slices and achieved 98% accuracy in classification. Also, Talo et al. [79] provided a thorough review of the current DLA in the literature and reported that during the BT detection process, the ResNet50 had a better classification accuracy of 95%. Amin et al. [10] used BRATS2013, 2015, and the clinical database to introduce a BT assessment protocol and achieved an accuracy of 98%. Improved binomial thresholding and multi-feature selection-based approach were implemented in the work of Sharif et al. [69] to identify the BT and achieve an improved result. A DLA to detect glioblastoma using hyperspectral 3D and 2D brain images was introduced in the work of Fabelo et al. [28]. Sajid et al. [67] introduced a BT detection protocol based on DL and achieved better sensitivity and specificity values. Acharya et al. [2] addressed the higher-order spectrum feature-based identification and classification of the abnormal segment in a brain MRI. A significant number of methods to improve the detection accuracy of a class of brain MRI images ranging from benchmark datasets and clinical images have been proposed and implemented by a significant number of researchers [14, 37, 52, 55, 70, 81]. Several researchers have documented different aspects of BT diagnosis, such as tumour visualisation [53], classification [7, 11, 43] and segmentation [15, 21, 47, 62, 71,72,73, 77, 84].
For effective brain tumour detection, segmentation and classification using MR images, several deep learning techniques, namely deep neural networks (DNN) [37, 85], convolutional neural networks (CNNs) [60, 61], deep convolutional neural networks (DCNNs) [25, 39], auto-encoders [58], stacked auto-encoders [26, 82], have been developed. In order to achieve the highest outcomes, research at a great pace continues to explore more in the deep learning approach. “To increase the performance of the classification, deep learning combined with other techniques is also observed. Acting differently, one study focused on proper planning of diagnosis and proposed a hybrid method that predicts low-grade 1p/19q status of glioma (LGG) [5].” “ANT’s open-source image registration software library is used to register multimodal CE-T1W and T2-W images (a total of 159 MR images of LGG with non-deleted status for 57 images and co-deleted for 102 images) [12], followed by tumour segmentation using semi-automatic software [4], and then CNN classification.” Finally, 87.7% accuracy, 93.3% sensitivity, and 97.7% specificity were recorded from cross-domain outcomes.
“A three-stage automatic approach for segmentation, referred to as WMMFCM, is proposed to address the constraints of FCM. Multi-resolution wavelet (WM), morphological pyramid (M), and Fuzzy C-means (FCM) clustering are used in the three stages concerned. Two datasets are used to verify performance: BrainWeb (152 MR images with T1-W, T2-W, and PD modalities) and BRATSS (81 images from multi-modal brain tumor segmentation having glioma with T1-W, T2-W, FLAIR, CE-T1W modalities). For BrainWeb, an accuracy of 97.05% is recorded, and with BRATS, 95.853% accuracy is achieved [6]”. In the following year, the idea of small kernels based on CNN was used in another work [61]. The paper provides a novel way of dealing with overfitting, given less weights in the network are presented. Starting with unusual strength and patch normalization, the analysis demonstrated the combination’s efficacy along with data augmentation. Further preparation for patches is achieved by artificially spinning them. Finally, the specified threshold is set to enforce volumetric constraints, i.e., the exclusion of small clusters that are essentially erroneous and categorized as small tumours. The accuracy rate for a baseline network turned to 84% during the brain tumour experimentation process, while for U-net, 88% accuracy is achieved.
An automated system based on deep convolutional neural networks (DCNN) is designed to focus on the same problem of over-fitting with sparse data [38]. “It incorporates DCNN with the implementation of layers of max-out and drop-out. On the BRATS 2013 dataset with T1-W, T2-W, CE-T1W, and FLAIR MR image modalities, the acceptability of the methodology is assessed. Experiments are conducted on a system using training is to the testing ratio of 80:20 based on three parameters, namely, dice similarity coefficient (whole tumor - 80%, core - 67%, and enhancing - 85%), sensitivity (whole tumor - 82%, core - 63%, and enhancing - 83%), and specificity (whole tumor - 85%, core - 82%, and enhancing - 88%). Several approaches are designed to strengthen and outdo CNN performance in terms of hardware requirements, accuracy, and processing time, especially when handling large-size images [56].” The method uses Fuzzy c-means for MR image segmentation to classify 66 T2-W MR images into four classes: regular, glioblastoma, sarcoma, and metastatic bronchogenic carcinoma tumour, followed by discrete wavelet transformation (DWT) integrated with the Deep Neural Network (DNN). The algorithm’s success resulted in a classification rate of 96.97%.
The extended version of the DCNN was proposed in the same year to address segmentation problems [3, 60, 75]. The incidence of multiple tumours is another common problem studied by a large group of researchers [56, 61]. “Tumor multiplicity demands more precision and thus increases complexity; in such a scenario, MR image input type and its features matter. In order to work with both MR and diffusion tensor imaging (DTI), a novel multimodal super voxel-based segmentation technique integrated with random forest (RF) is introduced [76].” A range of Gabor characteristics is extracted to train the RF classifier for each super voxel. Using multimodal images from BRATS and clinical databases, each super voxel is categorized as healthy and tumorous (core or edoema) (30 images). Performance is reported in terms of sensitivity and dice score. The respective values for the clinical dataset are 86% and 0.84. In comparison, better values are reported for BRATS, 96% and 0.89, respectively. Three separate network models have been proposed in one study: the Interpolated Network (IntNet), Skip-Net, and SE-Net for brain tumour segmentation [40]. IntNet’s dominance over SE-Net and Skip-Net is illustrated by experiments on BRATS 2015 MR images with four modalities (T1-W, T2-W, CE-T1W, and FLAIR). IntNet achieves the highest values on a full dataset for all the three parameters considered: dice coefficient (90%), sensitivity (88%), and specificity (73%). In the present situation, CNN enhancement is used to overcome the lengthy manual method of diagnosis. An efficient automatic segmentation approach has been developed that combines enhanced CNN (ECNN) with the BAT algorithm [80]. BAT works on the loss of functions, while ECNN’s small kernel characteristics allow networks with less weight allocation to manage over-fitting.
In contrast to conventional CNN, the output of ECNN is found to be 3% more accurate. “An end-to-end incremental DNN based model known as “EnsembleNet” is proposed for the segmentation of glioblastomas (high as well as low) [68].” It aggregates with incremental XCNet on parallel instances that generate model CNNs (2CNet and 3CNet) using the non-parametric fusion technique. On the BRATS 2017 dataset, the dice score turns out to be 0.88.
3 Theoretical background
3.1 Convolutional neural network
Convolutional Neural Network is multi-layered architectures based on deep learning, the popular technique of recent times. CNN shows the latest technology performance in many areas where it is applied, and its use is increasing day by day. Convolution neural networks, especially in image classification problems, achieve good results. “The CNN network structure is simple with few training parameters. Due to CNN weight sharing, the complex structure of the network model and the number of weights is reduced. The network includes layers called convolution, activation, pooling, and fully connected layer. CNN contains a loss function like softmax in its last layer [51].” The convolution layer aims to extract characteristics from the image of the input. By learning image properties using small squares of input data, convolution maintains the spatial relationship between pixels. The function of the pooling layer is to gradually reduce the spatial dimension of the representation to reduce the number of parameters and computation in the network and thus control overfitting. It is common in convolutional network architecture to add a pooling layer between successive layers of convolution periodically. “In convolutional neural network architectures, convolution, activation, and pooling layers are followed by a fully connected layer. This layer is connected to all the neurons of the previous layer. The fully connected layer is connected with all neurons in the previous layer. It is used to classify the extracted features into various input image classes based on the training data set. The last layer of convolutional neural networks can make an ultra-specific classification by combining all the specific features extracted from input data in previous layers [34].”
Convolutional neural networks first attracted attention by participating in the ImageNet ILSVRC competition held in 2012 with the CNN architecture called Alexnet by Kizevsky and his friends [45]. The network has a very similar architecture to LeNet [46] but is deeper and broader. Unlike the previous architectures in which a pooling layer is stacked right after just a single convolution layer [42], it is also distinct that it includes overlapping convolutional layers.
“The AlexNet architecture is made up of 8 layers. The first two layers are convolution + max + norm; the third and fourth layers are convolution; the fifth layer is convolution + max, the sixth and seventh layers are fully connected, and the last layer is softmax. The architecture uses an image of 227 × 227 pixels as an input. During the convolution process, 96 11 × 11 filters are used in the first layer. As a result of the convolution phase in the network, the number of steps is 4, and the image size is 55 × 55. 3 × 3 dimensional filters are used in the first pooling layer of the Alexnet model. The size of the image after the process is 27 × 27. In the following layers, the same processes are repeated [45]”.
The output of the convolution and pooling layers represents the high-level features of the input image. Convolution and pooling layers do not make classification predictions. The fully connected layer aims to use these features to classify the input image based on the training data set. Each neuron in a fully connected layer is a class. Since the model is designed to classify 1000 images, the last layer contains 1000 neurons. On the last layer, softmax is allocated to perform the task of classification.
VGGNet was the ILSVRC 2014 runner-up, network from Karen Simonyan and Andrew Zisserman’, known as VGGNet [74]. Its key contribution was to demonstrate that the breadth of the network is a critical component of successful results. “VGG 16 is a 16-layer architecture with a pair of convolution layers, a pooling layer, and a fully connected layer at the top. The VGG network is the concept of much deeper networks and much narrower filters. VGGNet increased the number of layers in AlexNet from eight layers. Right now, there were versions with 16 to 19 layers of the VGGNet version. One key point is that these models have very small 3 × 3 Conv filters all the way, which is essentially the smallest conv filter size that looks at a little bit of the adjacent pixels. And they’ve only retained this very simple 3 × 3 conv structure with periodic pooling all the way through the network. VGG used small filters because of fewer parameters and stack more of them instead of having larger filters. VGG has smaller filters with more depth instead of having large filters. It has ended up having the same effective receptive field as if you only have one 7 × 7 convolutional layers [74].”
3.2 Principal component analysis
In many places, large data sets are increasingly common. In order to analyze such data sets, methods are required to minimize their dimensionality with the retention of most of the information in the data. The principal component analysis is one of the most commonly used approaches. Principal components analysis is a mathematical technique to describe information in a multivariate numerical data set with fewer variables but limited information loss. The overall variability clarifies the knowledge in the data set. PCA decreases the size of large data sets [1]. PCA is the transform of data from one coordinate system to another. After implementation, the first dimension in our new coordinate system has the maximum variance it can make. The second dimension has the most variance it can take, and so on. The dimension reduction of PCA is based on converting the correlated variables in the dataset into variables that are not correlated with some linear transformations. These new variables are a linear combination of existing ones and are referred to as Prime Components. This technique works well when there is an excessive correlation between variables in datasets, and data can contain high errors. The high correlation between variables indicates that the set carries unnecessary information [57].
The stages of the PCA dimension reduction technique are as follows [64]:
-
1)
Prepare the data: Here, the data is centered by subtracting the average from each variable. Thus, a data set with a mean of 0 is obtained. The mean of the dataset a is a=0
-
2)
The covariance matrix (C) for the dataset features is calculated as in formula (1).
-
3)
The eigenvalue V eigenvector E values of the covariance matrix are calculated as in Eq. (2).
-
4)
List the eigenvalues and their corresponding eigenvectors: The highest eigenvector is the fundamental component of the data. The eigenvectors are ordered according to eigenvalues from highest to lowest.
-
5)
Choose K eigenvalues and build an eigenvectors matrix.
-
6)
Transform the original matrix.
3.3 Support vector machine
“SVM is one of the most chosen, very simple, and efficient methods for solving classification problems with supervised learning. The purpose is to find a hyperplane in an m-dimensional space that divides data points into potential groups. The sub-plane should be located at the maximum distance from the data points. Due to the proximity of data points with a minimum distance to the hyperplane, known as support vectors, their effect on the hyperplane’s exact position is greater than other data points [20].” SVM algorithms use a set of mathematical functions that are defined as the kernel. The kernel tricks are used to solve non-linear problems using linear classifier. “The function of the kernel is to take data as input and transform it into the required form. Different SVM algorithms use different types of kernel functions. These functions can be of various types—for example, linear, nonlinear, polynomial, radial basis function (RBF), and sigmoid. The kernel functions return the inner product between two points in a suitable feature space [8].”
4 Materials and methods
In the literature, several CNN models are proposed to detect the abnormalities in the medical images using pre-trained CNN and customized CNN [36, 59, 84]. Most of the earlier work for BT detection uses thousands of MR images like BRATS [18, 54] and TCIA [23] dataset. So, an open challenge exists for the detection of BT based on minimal data. Here, a total number of 253 MR images are collected from the Kaggle repository [17]. The datasets consist of 155 tumorous brain MR images and 98 non-tumorous brain MR images. Figure 1 depicts the samples of MR images considered in this work.
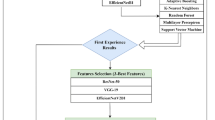
The proposed work is executed in six approaches to feeding deep features to SVM for BT detection. In the first approach, the deep features fc6 is considered, secondly fc7, thirdly fc8, fourthly the fusion of fc6 & fc7, fifthly fusion of fc6, fc7 & fc8 and sixthly fc6, fc7 & fc8 with dimension reduction technique namely PCA is employed. The feature fusion process is done by concatenating the two features or more to form a high dimensional feature to compensate for a single feature’s inadequacy. Again, we adopt PCA for the selection and dimensionality reduction of features and analyze classifiers’ performance. As three number of pre-trained network with six approaches of deep feature feedings are taken into consideration so, a total of 18 classifiers are framed. The proposed approach that selects the best classifier for BT detection is illustrated in Fig. 2.
The SVM is a well-known and most preferable classifier in the machine learning approach, especially image classification. The SVM uses a hyperplane for dataset labeling based on features obtained during the training process. One of the most commonly used for categorizing MRI images is SVM. For BT MR image classification based on deep features, the linear polynomial kernel (SVM-Linear) SVM is adapted. The 18 classifiers with their respective pre-trained network, deep features, fusion technique, feature dimension reduction technique, and classifier are framed in Table 1.
5 Results and discussion
The experimental studies were implemented using the MATLAB 2020a deep learning toolbox. “All applications were run on a laptop, i.e., Acer Predator Helios 300 Core i5 8th Gen - (8 GB/1 TB HDD/128 GB SSD/Windows 10 Home/4 GB Graphics) and equipped with NVIDIA GeForce GTX 1050Ti.” In terms of accuracy, sensitivity, specificity, precision, false-positive rate (FPR), and F1 Score, each classifier’s output was calculated. Measures of the confusion matrix are expressed in (4) to (9) equations.
Where TP = true positive, TN = true negative, FP = false positive, FN = false negative.
The hyperparameters used in all of the experiments in these approaches are: “solver type: stochastic gradient descent, the initial learning rate is 0.001, learning rate policy: Step (decreases by a factor of 10 every 50/5 epochs), momentum: 0.9, drop out is 0.2, Number of Epochs is 50 and minibatch size:64”. “The adaptive learning rate is good compared to the fixed learning rate. An adaptive algorithm usually converges much faster than simple back-propagation with a poorly chosen fixed learning rate [24, 33]“. The performance measures are given in Tables 2, 3 and 4. Note that all the results recorded in Tables 2, 3, and 4 are based on 30 independent runs.
It was observed from Table 2; VGG16 achieves the highest accuracy with the fused feature of fc6 and fc7. The vgg16 provides better results compared to alexnet and vgg19 concerning all features and combinations of features. In comparison to fc6, fc7, and fc8, fc6 contributes better results thanfc7 and fc8. Again, fc7 contribute better results than fc8 irrespective of the pre-trained network. The fused features of fc6 and fc7 improve the performance of classifiers. But, the fused features offc6, fc7, and fc8 reduced the performance of classifiers. Again, by the introduction of PCA, the performances of classifiers are further reduced. Hence, the fused features of fc6 and fc7 without any dimension reduction techniques with linear SVM achieved the best results, such as the maximum value of accuracy is 0.9789, sensitivity is 1, specificity is 1, precision is 1, and F1 Score 97.92. Also, the VGG16 with fused features of fc6 and fc7 have significantly less FPR, i.e., the mean value of 0.1133 and maximum value of 0.3, which is less compared to the other classifiers (Table 5).
It is observed from the literature that most of the work BRATS dataset is used with augmentation to increase the volume of the dataset. The BRATS dataset contains thousands of MR images in all of its version. The introduction of augmentation techniques to increase the dataset has many disadvantages, such as (1) easy generation of anatomically incorrect samples (2) not trivial to implement (3) mode collapse problem. The real impact of incorporating unrealistic samples into training sets still needs investigation. Hence, the VGG16 with fused future of fc6 andfc7 with linear SVM is the appropriate classifier to detect BT using MR images with a small dataset without duplicates.
6 Conclusion
The main objective of this research is to identify the best deep features or combination of deep features for the detection of BT using a small dataset. Here, we are avoiding reproduction techniques to make the duplicate of MR images as the reproduction techniques generate anatomically incorrect images and are under investigation. The VGG16 with the fused feature of fc6 and fc7 with a linear SVM classifier is competent enough with the state-of-art even if using a small dataset.
References
Abdi H, Williams LJ (2010) Principal component analysis. Wiley interdisciplinary reviews: computational statistics 2(4):433–459
Acharya UR, Meiburger KM, Faust O, Koh JE, Oh SL, Ciaccio EJ, Subudhi A, Jahmunah V, Sabut S (2019) Automatic detection of ischemic stroke using higher order spectra features in brain MRI images. Cogn Syst Res 58:134–142
Ahmed KB, Hall LO, Goldgof DB, Liu R, Gatenby RA (2017) Fine-tuning convolutional deep features for MRI based brain tumor classification. In: Medical imaging 2017: computer-aided diagnosis, vol 10134, p 101342E. International Society for Optics and Photonics
Akkus Z, Sedlar J, Coufalova L, Korfiatis P, Kline TL, Warner JD, Agrawal J, Erickson BJ (2015) Semiautomated segmentation of pre-operative low grade gliomas in magnetic resonance imaging. Cancer Imaging 15(1):12
Akkus Z, Ali I, Sedlar J, Kline TL, Agrawal JP, Parney IF, Giannini C, Erickson BJ (2016) Predicting 1p19q chromosomal deletion of low-grade gliomas from MR images using deep learning. arXiv:1611.06939
Ali H, Elmogy M, El-Daydamony E, Atwan A (2015) Multi-resolution MRI brain image segmentation based on morphological pyramid and fuzzy c-mean clustering. Arab J Sci Eng 40(11):3173–3185
Amarapur B (2020) Computer-aided diagnosis applied to MRI images of brain tumor using cognition based modified level set and optimized ANN classifier. Multimed Tools Appl 79(5):3571–3599
Amari SI, Wu S (1999) Improving support vector machine classifiers by modifying kernel functions. Neural Netw 12(6):783–789
Amin J, Sharif M, Yasmin M, Fernandes SL (2018) Big data analysis for brain tumor detection: deep convolutional neural networks. Futur Gener Comput Syst 87:290–297. https://doi.org/10.1016/j.future.2018.04.065
Amin J, Sharif M, Raza M, Saba T, Anjum MA (2019) Brain tumor detection using statistical and machine learning method. Comput Methods Prog Biomed 177:69–79
Anitha V, Murugavalli SJ (2016) Brain tumour classification using two-tier classifier with adaptive segmentation technique. IET Comput Vis 10(1):9–17
Avants BB, Tustison NJ, Song G, Cook PA, Klein A, Gee JC (2011) A reproducible evaluation of ants similarity metric performance in brain image registration. Neuroimage 54(3):2033–2044
Bakas S, Zeng K, Sotiras A, Rathore S, Akbari H, Gaonkar B, Rozycki M, Pati S, Davatzikos C (2015) GLISTRboost: combining multimodal MRI segmentation, registration, and biophysical tumor growth modeling with gradient boosting machines for glioma segmentation. In: BrainLes 2015. Springer, Berlin, pp. 144–155
Bakator M, Radosav D (2018) Deep learning and medical diagnosis: a review of literature. Multimodal Technologies and Interaction 2(3):47
Banday SA, Mir AH (2017) Statistical textural feature and deformable model based brain tumor segmentation and volume estimation. Multimed Tools Appl 76(3):3809–3828
Bhandarkar SM, Koh J, Suk M (1997) Multiscale image segmentation using a hierarchical self-organizing map. Neurocomputing 14(3):241–272
Brain MRI Images for brain tumor detection. http://kaggle.com
Brain Tumour Database (BraTS-MICCAI). Available online: http://hal.inria.fr/hal-00935640.
Buda M, Saha A, Mazurowski MA (2019) Association of genomic subtypes of lower-grade gliomas with shape features automatically extracted by a deep learning algorithm. Comput Biol Med 109:218–225
Chamasemani, F. F., & Singh, Y. P. (2011, September). Multiclass support vector machine (SVM) classifiers--an application in hypothyroid detection and classification. In 2011 Sixth International Conference on Bio-Inspired Computing: Theories and Applications (pp. 351-356). IEEE.
Chandra SK, Bajpai MK (2020) Brain tumor detection and segmentation using mesh-free super-diffusive model. Multimed Tools Appl 79(3):2653–2670. https://doi.org/10.1007/s11042-019-08374-7
Chow D, Qi J, Guo X, Miloushev V, Iwamoto F, Bruce J, Lassman A, Schwartz L, Lignelli A, Zhao B et al (2014) Semi-automated volumetric measurement on postcontrast MR imaging for analysis of recurrent and residual disease in glioblastoma multiforme. Am J Neuroradiol 35(3):498–503
Clark K, Vendt B, Smith K, Freymann J, Kirby J, Koppel P, Moore S, Phillips S, Maffitt D, Pringle M, Tarbox L (2013) The cancer imaging archive (TCIA): maintaining and operating a public information repository. J Digit Imaging 26(6):1045–1057
Cubuk ED, Zoph B, Mane D, Vasudevan V, Le QV (2019) Autoaugment: learning augmentation strategies from data. InProceedings of the IEEE conference on computer vision and pattern recognition, pp. 113-123).
David DS, Saravanan D, Jayachandran A (2020) Deep convolutional neural network based early diagnosis of multi class brain tumour classification system. Solid State Technology 63(6):3599–3623
Dolz J, Betrouni N, Quidet M, Kharroubi D, Leroy HA, Reyns N, Massoptier L, Vermandel M (2016) Stacking denoising auto-encoders in a deep network to segment the brainstem on MRI in brain cancer patients: a clinical study. Comput Med Imaging Graph 52:8–18
Drozdzal M, Chartrand G, Vorontsov E, Shakeri M, Di Jorio L, Tang A, Romero A, Bengio Y, Pal C, Kadoury S (2018) Learning normalized inputs for iterative estimation in medical image segmentation. Med Image Anal 44:1–13
Fabelo H, Halicek M, Ortega S, Shahedi M, Szolna A, Piñeiro JF, Sosa C, O’Shanahan AJ, Bisshopp S, Espino C, Márquez M (2019) Deep learning-based framework for in vivo identification of glioblastoma tumor using hyperspectral images of human brain. Sensors 19(4):920
Foo JL (2006) A survey of user interaction and automation in medical image segmentation methods. Tech rep ISUHCI 20062, Human-Computer Interaction Department, Iowa State University
Gaillard AF (2020) Brain tumors. [online]. Available: https://radiopaedia.org/articles/brain-tumours.
Georgiadis P, Cavouras D, Kalatzis I, Daskalakis A, Kagadis GC, Sifaki K, Malamas M, Nikiforidis G, Solomou E (2008) Improving brain tumor characterization on MRI by probabilistic neural networks and non-linear transformation of textural features. Comput Methods Prog Biomed 89(1):24–32
Gibbs P, Buckley DL, Blackband SJ, Horsman A (1996) Tumour volume determination from MR images by morphological segmentation. Phys Med Biol 41(11):2437–2446
Goodfellow I, Bengio Y, Courville A, Bengio Y (2016 Nov 18) Deep learning. MIT press, Cambridge http://www.deeplearningbook.org
Ian Goodfellow Yoshua Bengio and AAron Courville (2016) Deep learning, page 526
Gordillo N, Montseny E, Sobrevilla P (2013) State of the art survey on MRI brain tumor segmentation. Magn Reson Imaging 31(8):1426–1438
Gudigar A, Raghavendra U, San TR, Ciaccio EJ, Acharya UR (2019) Application of multiresolution analysis for automated detection of brain abnormality using MR images: a comparative study. Futur Gener Comput Syst 90:359–367
Havaei M, Davy A, Warde-Farley D, Biard A, Courville A, Bengio Y, Pal C, Jodoin PM, Larochelle H (2017) Brain tumor segmentation with deep neural networks. Med Image Anal 35:18–31
Hussain S, Anwar SM, Majid M (2017) Brain tumor segmentation using cascaded deep convolutional neural network. In: IEEE 39th annual international conference of the IEEE engineering in medicine and biology society (EMBC), pp 1998–2001
Hussain S, Anwar SM, Majid M (2018) Segmentation of glioma tumors in brain using deep convolutional neural network. Neurocomputing 282:248–261
Iqbal S, Ghani MU, Saba T, Rehman A (2018) Brain tumor segmentation in multi-spectral MRI using convolutional neural networks (CNN). Microsc Res Tech 81(4):419–427
Kanmani P, Marikkannu P (2018) MRI brain images classification: a multi-level threshold based region optimization technique. J Med Syst 42(4):62
Karpathy A (2016) Cs231n convolutional neural networks for visual recognition. Neural Netw 1(1)
Kaur T, Saini BS, Gupta S (2019) An adaptive fuzzy K-nearest neighbor approach for MR brain tumor image classification using parameter free bat optimization algorithm. Multimed Tools Appl 78(15):21853–21890
Khawaldeh S, Pervaiz U, Rafiq A, Alkhawaldeh RS (2018) Noninvasive grading of glioma tumor using magnetic resonance imaging with convolutional neural networks. Appl Sci 8(1):27
A. Krizhevsky, I. Sutskever, and G. E. Hinton (2012) ImageNet classification with deep convolutional neural networks. In NIPS, pp. 1106–1114
LeCun Y, Bottou L, Bengio Y, Haffner P (1998) Gradient-based learning applied to document recognition. Proc IEEE 86(11):2278–2324
Lin F, Wu Q, Liu J, Wang D, Kong X (2020) Path aggregation U-net model for brain tumor segmentation. Multimed Tools Appl. https://doi.org/10.1007/s11042-020-08795-9
Liu M, Zhang J, Nie D (2018) Anatomical landmark based deep feature representation for MR images in brain disease diagnosis. IEEE J Biomed Health Inform 22:1476–1485
Logeswari T, Karnan M (2010) An improved implementation of brain tumor detection using segmentation based on hierarchical self-organizing map. Int J Comput Theory Eng 2(4):591
Luo S, Li R, Ourselin S (2003) A new deformable model using dynamic gradient vector flow and adaptive balloon forces. APRS workshop on digital image computing, Brisbane, Australia, In
Ma W, Lu J (2017) An equivalence of fully connected layer and convolutional layer. arXiv preprint arXiv:1712.01252
Mallick PK, Ryu SH, Satapathy SK, Mishra S, Nguyen GN, Tiwari P (2019) Brain MRI image classification for cancer detection using deep wavelet autoencoder-based deep neural network. IEEE Access 7:46278–46287
Mehmood I, Sajjad M, Muhammad K, Shah SI, Sangaiah AK, Shoaib M, Baik SW (2019) An efficient computerized decision support system for the analysis and 3D visualization of brain tumor. Multimed Tools Appl 78(10):12723–12748
Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby J, Burren Y, Porz N, Slotboom J, Wiest R, Lanczi L (2014) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imaging 34(10):1993–2024
Mohsen H (2018) Classification using deep learning neural networks for brain tumors. Future Comput Inform J 3:68–71
Mohsen H, El Dahshan ESA, El Horbaty ESM, Salem A-BM (2018) Classification using deep learning neural networks for brain tumors. Future Computing and Informatics Journal 3(1):68–71
Morchid M, Dufour R, Bousquet P-M, Linarès G, Torres-Moreno J-M (2014) Feature selection using principal component analysis for massive retweet detection. Pattern Recogn Lett 49:33–39
Myronenko A (2018) 3D MRI brain tumor segmentation using autoencoder regularization. In: International MICCAI brain lesion workshop. Springer, Berlin, pp 311–320
Nadeem MW, Ghamdi MA, Hussain M, Khan MA, Khan KM, Almotiri SH, Butt SA (2020) Brain tumor analysis empowered with deep learning: a review, taxonomy, and future challenges. Brain Sciences 10(2):118
Pan Y, Huang W, Lin Z, Zhu W, Zhou J, Wong J, Ding Z (2015) Brain tumor grading based on neural networks and convolutional neural networks. In: IEEE 37th annual international conference of the IEEE engineering in medicine and biology society (EMBC), pp 699–702
Pereira S, Pinto A, Alves V, Silva CA (2016) Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans Medical Imaging 35(5):1240–1251
Punn NS, Agarwal S (2020) Multi-modality encoded fusion with 3D inception U-net and decoder model for brain tumor segmentation. Multimed Tools Appl. https://doi.org/10.1007/s11042-020-09271-0
Raja NSM, Fernandes SL, Dey N, Satapathy SC (2018) Contrast enhanced medical MRI evaluation using Tsallis entropy and region growing segmentation. J Ambient Intell Humaniz Comput 1–12. https://doi.org/10.1007/s12652-018-0854-8
Rajagopal R, Ranganathan V (2017) Evaluation of effect of unsupervised dimensionality reduction techniques on automated arrhythmia classification. Biomedical Signal Processing and Control 34:1–8
Rajinikanth V, Joseph Raj AN, Thanaraj KP, Naik GR (2020) A customized VGG19 network with concatenation of deep and handcrafted features for brain tumor detection. Appl Sci 10(10):3429
Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, Huang Z, Karpathy A, Khosla A, Bernstein M, Berg AC (2015) Imagenet large scale visual recognition challenge. Int J Comput Vis 115(3):211–252
Sajid S, Hussain S, Sarwar A (2019) Brain tumor detection and segmentation in MR images using deep learning. Arab J Sci Eng 44(11):9249–9261
Saouli R, Akil M, Retal K (2018) Fully automatic brain tumor segmentation using end-to-end incremental deep neural networks in MRI images. Comput Methods Prog Biomed 166:39–49
Sharif M, Tanvir U, Munir EU, Khan MA, Yasmin M (2018) Brain tumor segmentation and classification by improved binomial thresholding and multi-features selection. J Ambient Intell Humaniz Comput. https://doi.org/10.1007/s12652-018-1075-x
Sharif MI, Li JP, Khan MA, Saleem MA (2020) Active deep neural network features selection for segmentation and recognition of brain tumors using MRI images. Pattern Recogn Lett 129:181–189
Sheela CJ, Suganthi G (2020) Brain tumor segmentation with radius contraction and expansion based initial contour detection for active contour model. Multimed Tools Appl 79:23793–23819. https://doi.org/10.1007/s11042-020-09006-1
Sheela CJ, Suganthi G (2020) Morphological edge detection and brain tumor segmentation in magnetic resonance (MR) images based on region growing and performance evaluation of modified fuzzy C-means (FCM) algorithm. Multimed Tools Appl 79:17483–17496. https://doi.org/10.1007/s11042-020-08636-9
Shivhare SN, Kumar N, Singh N (2019) A hybrid of active contour model and convex hull for automated brain tumor segmentation in multimodal MRI. Multimed Tools Appl 78(24):34207–34229
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556.
Singh L, Chetty G, Sharma D (2012) A novel machine learning approach for detecting the brain abnormalities from MRI structural images. In: IAPR international conference on pattern recognition in bioinformatics. Springer, Berlin, pp 94–105
Soltaninejad M, Yang G, Lambrou T, Allinson N, Jones TL, Barrick TR, Howe FA, Ye X (2018) Supervised learning based multimodal MRI brain tumour segmentation using texture features from supervoxels. Comput Methods Prog Biomed 157:69–84
Sun J, Li J, Liu L (2020) Semantic segmentation of brain tumor with nested residual attention networks. Multimed Tools Appl. https://doi.org/10.1007/s11042-020-09840-3
Talo M, Baloglu UB, Yıldırım Ö, Acharya UR (2019) Application of deep transfer learning for automated brain abnormality classification using MR images. Cogn Syst Res 54:176–188
Talo M, Yildirim O, Baloglu UB, Aydin G, Acharya UR (2019) Convolutional neural networks for multi-class brain disease detection using MRI images. Comput Med Imaging Graph 78:101673
Thaha MM, Kumar KPM, Murugan B, Dhanasekeran S, Vijayakarthick P, Selvi AS (2019) Brain tumor segmentation using convolutional neural networks in MRI images. J Med Syst 43(9):294
Tiwari A, Srivastava S, Pant M (2020) Brain tumor segmentation and classification from magnetic resonance images: review of selected methods from 2014 to 2019. Pattern Recogn Lett 131:244–260
Vaidhya K, Thirunavukkarasu S, Alex V, Krishnamurthi G (2015) Multimodal brain tumor segmentation using stacked denoising autoencoders. In: BrainLes. Springer, Berlin, pp 181–194
Vishnuvarthanan G, Rajasekaran MP, Subbaraj P, Vishnuvarthanan A (2016) An unsupervised learning method with a clustering approach for tumor identification and tissue segmentation in magnetic resonance brain images. Appl Soft Comput 38:190–212
Wang G, Li W, Zuluaga MA, Pratt R, Patel PA, Aertsen M, Doel T, David AL, Deprest J, Ourselin S, Vercauteren T (2018) Interactive medical image segmentation using deep learning with image-specific fine tuning. IEEE Trans Med Imaging 37(7):1562–1573
Xiao Z, Huang R, Ding Y, Lan T, Dong R, Qin Z, Zhang X, Wang W (2016) A deep learning based segmentation method for brain tumor in MR images. In: IEEE 6th international conference on computational advances in bio and medical sciences (ICCABS), pp 1–6
Xue X, Xue Z, Cao F, Zhu Y, Young GS, Li Y, Yang J, Wong ST (2010) Pice: prior information constrained evolution for 3-d and 4-d brain tumor segmentation. In: IEEE international symposium on biomedical imaging: from nano to macro, pp 840–843
Yao J (2006) Image processing in tumor imaging. New Techniques in Oncologic Imaging, pp 79–102
Zhu Y, Young GS, Xue Z, Huang RY, You H, Setayesh K, Hatabu H, Cao F, Wong ST (2012) Semiautomatic segmentation software for quantitative clinical brain glioblastoma evaluation. Acad Radiol 19(8):977–985
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sethy, P.K., Behera, S.K. A data constrained approach for brain tumour detection using fused deep features and SVM. Multimed Tools Appl 80, 28745–28760 (2021). https://doi.org/10.1007/s11042-021-11098-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-021-11098-2