Abstract
Heat shock factors (HSFs) play crucial roles in various plant stress responses. However, the current knowledge about HSFs in cassava, an important crop, is still insufficient. In this research, we identified 32 cassava HSF genes (MeHSFs) and clustered them into three groups (A, B, C) based on phylogenetic analysis and structural characteristics. Conserved motif analyses showed that MeHSFs display domains characteristic to HSF transcription factors. Gene structure analyses suggested that 29 MeHSFs contained only two exons. All identified 32 cassava MeHSFs were distributed on 13 chromosomes. Their expression profiles revealed that the different MeHSFs were expressed differentially in different tissues, most high expression genes belonged to group A. The similar MeHSFs were up-regulated after treatment with both PEG and abscisic acid (ABA), which implied that these MeHSFs may participate in resistance to simulated drought stress associated with the ABA signaling pathway. In addition, several MeHSFs were induced during postharvest physiological deterioration (PPD) in cassava. Our results provided basic but important knowledge for future gene function analysis of MeHSFs toward efforts in improving tolerance to abiotic stress and PPD in cassava.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants routinely experience diverse stress conditions, such as abiotic or biotic stresses, due to their sessile nature. They are therefore required to adapt for survival by developing a range of defense mechanisms [1, 2]. Heat shock response is an important conserved defense mechanism for adapting to environmental stresses. Heat shock proteins (HSPs) play essential roles in defense from environment stresses by folding/unfolding and degrading proteins [3, 4]. Several researches showed that HSFs, the key regulators of HSPs, could increase heat, high salinity and oxidative stress tolerance in plants [5, 6]. Although HSFs vary in their size and sequences, their promoter recognition pattern and basic structures are conserved in plants [7]. Based on their conserved region, plant HSFs can be divided into three classes namely, HSFA, B, and C [8]. Group A HSFs are major members with function as transcription activators, whereas most HSFs that belong to Group B and C do not have a self-activation function [9, 10].
Analyses of different HSF mutants have presented extraordinary functional differences that could not be restored by other HSF types. The precise function of different HSFs in plants is still unclear. Several studies showed that HSFs partake in signal transduction pathways and regulate genes expression responsive for a variety of abiotic/biotic stresses [6, 11, 12]. Evidences showed that HSFA1a was the primary regulator to resist heat stress. HSFA1a could trigger a heat stress response and form a protein complex with HSFA2 and HSFB1 to adjust physiological metabolism during heat stress [8]. Arabidopsis showed an improved heat, salt/osmotic, and oxidative stress tolerance via overexpression of HSFA2 [13, 14]. HSFA3 was shown to be functionally similar to HSFA1a and HSFA2 and since its expression was up-regulated by drought and heat stress, it was regarded as a part of drought stress signaling [15]. The expression of HSFA6a and HSFA6b was also significant up-regulated by salt and cold stress [16]. HSF also showed resistance to heavy metal stress. For example, rice and yeast strains presented better tolerance for cadmium stress by overexpressing TaHSFA4a [17]. HSFA9 regulated the expression of HSP during seed development and showed a unique function to verify the functional diversification of HSF [18, 19]. While HSFAs seemingly displayed the majority of HSF functions, most members belonging to HSFBs did not have an activator function and were usually regarded as repressors of gene expression [9, 10, 20]. A large proportion of group HSFC genes were found in monocots compared to eudicots. However, the exact role of HSFCs in monocots is still unclear [8].
HSFs are pervasive in eukaryotes and function as transcription factors. Plant HSFs comprise of a larger family of proteins compared with vertebrate HSFs and those found in Drosophila [21]. With the development of sequencing technology, several HSF families have been identified in different plants and these are composed of 25 members in Zea mays, more than 56 members in Triticum aestivum, 25 members in Oryza sativa, 30 members in Populus trichocarpa, 19 members in Ricinus communis, and 21 members in Arabidopsis thaliana [8, 22,23,24,25,26,27]. Cassava is a major source of dietary carbohydrate, industrial starch, and bioethanol due to its high starch production [28, 29]. After harvest, its tuberous roots undergo rapid PPD, restricting its use as a raw material in food industry [30, 31]. Physiological and biochemical analysis showed that the production of ROS is the first event during development of PPD. Lower ROS accumulation results in delayed PPD development by manipulating ROS-scavenging-related genes or exogenous application of chemicals [11, 30,31,32]. Cassava presents excellent drought resistance during the growth process [33]. HSFs could increase the tolerance to drought and salinity stress in plants. However, the mechanism underlying resistance to abiotic stress in cassava is unclear.
In this study, HSFs present in cassava were identified and analyzed with regards to their phylogenetic relationships, gene structure, and protein motifs. The expression profiles of the various identified HSFs in different tissues were analyzed. HSFs responded to simulated drought and ABA. The change in HSF expression during the process of PPD was also investigated. Our results may prove meaningful for the analysis of HSFs function in cassava. Our findings also expanded knowledge regarding simulated drought tolerance and the PPD process, and offered novel implications for extending the shelf life and improving the quality of cassava tuberous roots.
Materials and methods
Plant materials and treatments
Arg7 (Manihot esculenta cv. Arg7) is an elite cassava cultivar in Argentina adapted to moderate drought stress. Arg7 were cultured in growth chamber conditions (35 °C/20 °C day/night, 16/8-h light/dark cycle under 70% relative humidity, 200 μmol m−2 s−1 photosynthetic photon flux density). After cultivation for 90 days, cassava seedlings were subjected to 100 μM ABA and PEG6000 (20%) treatment, respectively. Each sample was pooled from five plants with three replicates. Subsequently, two replicates of these samples were chosen for transcriptomic analysis. For analyzing PPD, the tuberous roots of 10-month-old cassava (Manihot esculenta cv. sc124) were cut into 5-mm thick slices and placed on a wet filter paper in Petri dishes. The samples were incubated in the dark for different time periods (0, 6, 12, and 48 h) at 28 °C. RNA was extracted from the tuberous root slices at different time periods (0, 6, 12, and 48 h) and used for transcriptomic analysis (three replicates of each sample).
Identification and phylogenetic analyses
The HSF protein sequences in Arabidopsis and rice were obtained from TAIR (http://www.arabidopsis.org/) and RGAP (http://rice.plantbiology.msu.edu/) databases, respectively. The whole genome sequence of cassava was acquired from a publicly available database (https://phytozome.jgi.doe.gov/pz/portal.html). In order to identify HSFs in the cassava protein sequence library, hidden Markov models (HMM) (http://www.hmmer.org/) were constructed using known HSF sequences [34]. PtHSFs, RcHSFs, AtHSFs, and OsHSFs were used to confirm the recognized cassava HSFs by BLAST analysis. The conserved domains of cassava HSFs were validated by the PFAM (http://pfam.sanger.ac.uk/) and conserved domains database (http://www.ncbi.nlm.nih.gov/cdd/). An evolutionary tree was constructed using cassava, poplar, castor bean, Arabidopsis, and rice HSFs using MEGA 5.0 and Clustal X2.0 softwares [35].
Protein properties and sequence analyses
The molecular weight and isoelectric points of MeHSFs were predicted by proteomics server (ExPASy) (http://expasy.org/). The MEME (http://meme.nbcr.net/meme/cgi-bin/meme.cgi) and InterProScan (http://www.ebi.ac.uk/Tools/pfa/iprscan/) databases were employed to identify the conserved motifs of MeHSFs. MeHSF gene structures and chromosomal location were analyzed through gene structure display server (GSDS) and Phytozome cassava database, respectively [36].
Transcriptomic analysis
RNA extraction, library preparation, and sequencing were performed by Majorbio BioTech Co., Ltd. (Shanghai, China). The sequencing platform was Illumina GAII (Illumina, San Diego, CA, USA). For reliability, the FASTX-toolkit (http://hannonlab.cshl.edu/fastx_toolkit/) and FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) were used to remove adapter sequences and low-quality sequences, respectively. Subsequently, the cassava genome version 4.1 (https://phytozome.jgi.doe.gov/pz/portal.html) was used as the reference for aligning the clean reads through Tophat 2.0 software (http://tophat.cbcb.umd.edu/) [37]. Based on the alignment files, the transcriptomic data were assembled using Cufflinks [38]. Genes were scored as not expressed if the corresponding RNAseq reads could not align to the genome. Expression levels were calculated and normalized as fragments per kilobase per million mapped reads. FPKM values were calculated to create heat map with MeV 4.9 software (CCCB, Boston, MA, USA). DEGseq was used to identify differentially expressed genes (Log2 based fold changes > 1; Log2 based fold changes < − 1) in response to treatment. The expression profiles of cassava housekeeping genes and phylogeny of raw RNA-Seq data were added in the supplement (Table S1–S2, Fig. S1). The expression datum of MeHSFs in various tissues were downloaded from database (shiny.danforthcenter.org/cassava_atlas/) [39]. The generated sequence data were deposited in NCBI’s SRA database under the accession of SRP182603.
Results
Identification and evolutionary analysis of HSFs in Manihot esculenta
The AtHSF and OsHSF sequences were used as queries for HMM and BLAST searches and 32 HSF proteins were identified from the cassava genome, which were designated as MeHSF1–MeHSF32. Their lengths ranged from 217 to 576 amino acid residues and pIs and relative molecular masses varied between 4.63 to 9.06 and 24.8 to 64.2 kDa, respectively (Table S3).
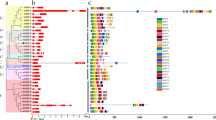
The evolutionary relationship of MeHSFs with AtHSFs, OsHSFs, PtHSF, and RcHSF was investigated by phylogenetic analysis (Fig. 1). The MeHSF family could be classified into 10 groups, Group A1 included MeHSF1, -30; Group A2 included MeHSF9, -23; Group A3 included MeHSF25, -29; Group A4 included MeHSF5, -6, -7 and -28; Group A5 included MeHSF3; Group A6 included MeHSF12, -26; Group A7 included MeHSF14, -15; Group A8 included MeHSF8; Group A9 included MeHSF2, -4; Group B included MeHSF11, -13, -16, -18 -19, -20, -21, -22, -24,-27, -31 and -32; Group C included MeHSF10, -17. MeHSFs presented a closer relationship with RcHSFs and PtHSFs compared to OsHSFs and AtHSFs via orthologous genes.
Conserved motif and gene structure analyses of MeHSFs in Manihot esculenta
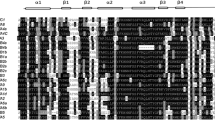
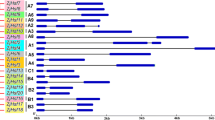
Overall 10 conserved MeHSF motifs were identified by searching in the MEME database. The conserved motifs were annotated using the InterPro database, which presented essential features of the HSF family (Fig. 2). Figure 2 indicated that all MeHSFs contained motifs 3 and 4. In group A1–A9, all MeHSFs contained at least six motifs, except in group A3, whereas only four and at most six motifs were present in group B and C, respectively. All groups presented a similar or identical motif composition, except group A3. These results suggested that all MeHSFs contain essential features of the HSF family and similar motif characters in different groups, further supporting the results of evolutionary relationships.
The exon–intron organization of the MeHSFs were analyzed using the GSDS database (Fig. 3). Interestingly, all MeHSFs contained two exons, except MeHSF9, MeHSF25, and MeHSF29, and those in the same group generally exhibited similar exon–intron organizations.
Chromosomal distribution of MeHSFs in M. esculenta
In order to analyze the distribution of MeHSFs, the chromosomal location of the identified MeHSF1-32 were analyzed. MeHSFs were mapped to be present on chr1, 2, 3, 5, 8, 9, 12, 13, 14, 15, 16, 17, and 18 (Fig. 4). Group A5 and A8 contained only one member located on chr15. MeHSFs in group A2 and A3 were all located on chr3, 8, 9, and 16. Group A4 contained four members located on chr1, 2, and 17; however, MeHSF28 from this group could not be exact located on the chromosome. By using the cassava v7.1 to analyze the information of MeHSF28, it could only be located in chr16. Group B, a large subfamily contained 12 members located on chr1, 2, 5, 8, 9, 12, 13, 14, 16, 17, and 18. The locations of group C were chr3, and 16. Thus, most of MeHSFs were located on Chr1, 2, 3, 9, 15, and 16, while four chromosomes were only distributed for one gene, respectively.
Expression profiles of MeHSFs in different cassava tissues
In order to analyze the expression profiles of MeHSFs in different cassava tissues, the expression data of 11 cassava tissue/organ types were downloaded from a database [39]. Here 11 tissues that were included were leaf, midvein, petiole, stem, lateral bud, shoot apical meristem (SAM), storage root (SR), fibrous root (FR), root apical meristem (RAM), organized embryogenic structure (OES), and friable embryogenic calli (FEC). As shown in Fig. 5, all MeHSFs showed a corresponding expression based on transcriptomic data, except for MeHSF19 (Table S4). Approximately 50% of MeHSFs presented a low transcript abundance, which coincided with its blocking status under normal conditions. Five MeHSFs (MeHSF1, 3, 8, 28, and 30), belonging to Group A, presented high expression levels in all the 11 analyzed tissues, three MeHSFs (MeHSF24, 18, and 10), belonging to Group B and C, presented a high expression in nine tissues, and the other MeHSFs were mainly highly expressed in lateral buds, OES, and FEC.
Expression profiles of MeHSFs in response to PEG and ABA treatment
A PEG treatment was performed in order to analyze the expression profiles of MeHSFs responsive to drought stress. All MeHSFs, except MeHSF12, showed corresponding expression on the basis of transcriptome data. As shown in Fig. 6, MeHSF-5, -7, -8, -9, -10, 18 and -29 showed induction under PEG treatment, MeHSFs-15, 17, 20, 22, and 23 showed suppression under PEG treatment. ABA played an important role in signal transduction pathways that respond to drought stress, and plant HSFs also regulated the expression of genes responsive to various abiotic stresses [8]. The expression profiles of MeHSFs upon ABA treatment were also studied. Similar to treatment with PEG, the expression data of MeHSF21 was not covered. MeHSF-8, -9, -10, -13, -18 and -29 showed induction after ABA treatment, MeHSF22 and MeHSF 23 showed a down-regulation. The up-regulated MeHSFs were identical after treatment with PEG or ABA, which suggested that these MeHSFs might be involved in ABA mediated osmotic response.
Expression profiles of MeHSFs during PPD
PPD is one of the major factors that restricts the use of cassava as a raw material in the food industry. HSFs are strongly associated with oxidative stress, and physiological and biochemical analyses show that production of ROS is the first step in PPD development. The expression profiles of MeHSFs during PPD in cassava were analyzed in the cultivar sc124. All MeHSFs showed a corresponding expression data based on transcriptome data, except MeHSF12 and MeHSF20. Most of the MeHSFs showed induction during PPD and only MeHSF23 and MeHSF26 showed suppression, which suggested these to be a part of the MeHSFs that participate in this process (Fig. 7).
Discussion
Cassava is one of the important crops in tropical and sub-tropical regions, and it provides staple food for over 750 million people around the world [33]. HSFs have many functions involved in different circumstances, such as oxidative stress, high temperatures, and drought stress [5, 6]. Thus, it is necessary to systematically analyze the potential roles of MeHSFs in cassava. In this research, 32 HSFs were identified from the cassava genome, which were classified into three groups (A, B, C) according to their evolutionary relationship. MeHSFA contained nine subclasses (A1–A9). The results of this classification coincided with the classification in rice and Arabidopsis [25, 40]. The cassava genome (742 Mb) is larger than that of Arabidopsis (125 Mb), castor bean (320 Mb), poplar (410 Mb), and rice (430 Mb) [26, 27, 41,42,43]. The number of HSFs in cassava (32 MeHSFs) was roughly similar with that in rice (25 OsHSFs), castor bean (19 RcHSFs), poplar (30 PtHSFs) and Arabidopsis (21 AtHSFs). MeHSFs showed a closer relationship with RcHSFs and PtHSFs. In all species, the HSFs are classified within different subclasses and these might be correlated with the different growth conditions [6]. Although their sequence and sizes are considerably diverse, the fundamental structure of HSFs is conserved in eukaryotes. Our results showed that almost all MeHSFs contained conserved motifs and different groups had similar motif characters. Exon/intron organization analysis showed that all MeHSFs contained only one intron, except MeHSF9, MeHSF25, and MeHSF29, and presented a prominently conservation in all family members. This phenomenon also existed in other species [6]. The extensive MeHSF distribution on chromosomes in cassava showed similarity to those from poplar, rice, and Arabidopsis [25, 40, 44]. Taken together, all results suggested that the classification of MeHSFs was reliable and the HSFs family of proteins was well conserved among different species.
Tissue-specific expression may be related with the function of genes. In Arabidopsis and rice, HSFs are differentially expressed in a tissue-specific manner [40]. In order to explore the possible roles of MeHSFs in different tissues, their expression profiles in 11 tissues were studied [2]. Most MeHSFs presented a low transcript abundance, which coincided with its blocking status under normal conditions. Five MeHSFs (-1, -3, -8, -28 and -30) presented high expression in all the analyzed 11 tissues, which might be involved in wide heat shock functions in various tissues [8]. The tissues that were analyzed here could be divided into three types; the first class consists of tissues that are subjected to air such as SAM, lateral bud, stem, leaf, petiole, and midvein. The second class consists of tissues that grow underground and include the SR, FR, and RAM, whereas the last class consisted of the embryogenic tissues including OES and FEC. The expression profiles of MeHSFs in different tissue classes were similar suggesting that these MeHSFs might exhibit a similar function. More MeHSFs showed a high expression in the lateral bud, OES, and FEC, which might be related to less differentiation of these tissues. None of the MeHSFs were expressed only in one specific tissue, which was similar to that observed in rice. However, AtHSF9 was specifically expressed in seeds [18]. Collectively, the expression profiles revealed that different MeHSFs presented a differential expression pattern in different tissues. Several MeHSFs presented a constitutively high expression in all cassava tissues, indicating their crucial function in cassava development.
Large experimental data has demonstrated that HSFs can increase the tolerance to abiotic stresses in plants including Arabidopsis and rice [8, 45]. Results from this research showed that few MeHSFs including MeHSF-5, -7, -8, 9, 10, -18, and -29, were up-regulated after PEG treatment and the same MeHSFs, such as MeHSF-8, -9, -10, -18, and -29, were up-regulated after ABA treatment. Thus, MeHSFs might resist drought stress associated with the ABA signaling pathway. AtHsfA9 was considered to be related with the ABA signal network [46]. MeHSF8 is a homolog of AtHsfA9, which present high expression after ABA treatment. Thus, this suggests that MeHSF8 might improve the tolerance to drought stress through the ABA signaling pathway. MeHSF9 and MeHSF8 are homologs of AtHSFA2 and AtHSFA8, respectively [25]. AtHSFA2 and AtHSFA8 improved the tolerance to salt/osmotic stress in Arabidopsis. MeHSF9 and MeHSF27 are homologs of OsHSF17 and OsHSF29, respectively. OsHSF17 and OsHSF29 also improved the tolerance to salt/osmotic stress in rice [40]. The expression analysis of these MeHSFs were consistent with Arabidopsis and rice under similar abiotic stress treatments, providing clues for the function of MeHSFs under abiotic stress. HSFs are associated with oxidative stress, and lower ROS accumulation leads to a delayed PPD process [30,31,32, 47, 48]. In this research, we observed that MeHSFs were induced during PPD in the analyzed cultivar. OsHsfC2a and OsHsfA5 seem to be the major players related to ROS sensing and accumulation [48, 49], which are homologs of MeHSF10/17 and MeHSF3, respectively, and that show high expression during PPD. These MeHSFs may be regarded as candidate genes for genetic improvement of cassava toward resistance to PPD.
Conclusion
In this research, 32 MeHSFs were identified from cassava and their classification, protein motifs, and gene structures were analyzed in detail. All identified MeHSFs were distributed on 13 different chromosomes. Tissue expression analysis showed that none of the MeHSFs were expressed only in one specific tissue. Transcriptomic analysis suggested that the MeHSFs were involved in response to simulated drought and ABA treatments. MeHSFs were also related with PPD and may operate mainly through ROS-regulated gene networks. In conclusion, our results offer critical basic knowledge for future gene function analysis of MeHSFs in cassava.
Supplementary materials
Table S1 The expression profiles of cassava housekeeping genes (PPD), Table S2 The expression profiles of cassava housekeeping genes (ABA, PEG), Table S3 The list of MeHSF members identified, Table S4 Expression data of MeHSF genes in various tissues/organs.
References
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24(1):23–58. https://doi.org/10.1080/07352680590910410
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K (2017) Transcriptional regulatory network of plant heat stress response. Trends Plant Sci 22(1):53–65
Hartl FU, Hayer-Hartl M (2002) Protein folding—molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295(5561):1852–1858. https://doi.org/10.1126/science.1068408
Lindquist S (1986) The heat-shock response. Annu Rev Biochem 55:1151–1191. https://doi.org/10.1146/annurev.bi.55.070186.005443
Jiang Y, Zheng Q, Chen L, Liang Y, Wu J (2018) Ectopic overexpression of maize heat shock transcription factor gene ZmHsf04 confers increased thermo and salt-stress tolerance in transgenic Arabidopsis. Acta Physiol Plant 40(1):9
Guo M, Liu JH, Ma X, Luo DX, Gong ZH, Lu MH (2016) The plant heat stress transcription factors (HSFs): structure, regulation, and function in response to abiotic stresses. Front Plant Sci 7:114. https://doi.org/10.3389/fpls.2016.00114
Baniwal SK, Bharti K, Chan KY, Fauth M, Koskull-Döring PV (2004) Heat stress response in plants: a complex game with chaperones and more than twenty heat stress transcription factors. J Biosci 29(4):471–487
Scharf KD, Berberich T, Ebersberger I, Nover L (2012) The plant heat stress transcription factor (Hsf) family: structure, function and evolution. Biochim Biophys Acta 1819(2):104–119
Kotak S, Port M, Ganguli A, Bicker F, von Koskull-Doring P (2004) Characterization of C-terminal domains of Arabidopsis heat stress transcription factors (Hsfs) and identification of a new signature combination of plant class A Hsfs with AHA and NES motifs essential for activator function and intracellular localization. Plant J 39(1):98–112. https://doi.org/10.1111/j.1365-313X.2004.02111.x
Czarnecka-Verner E, Yuan CX, Scharf KD, Englich G, Gurley WB (2000) Plants contain a novel multi-member class of heat shock factors without transcriptional activator potential. Plant Mol Biol 43(4):459–471. https://doi.org/10.1023/a:1006448607740
Zhou J, Xu X, Cao J, Yin L, Yu J (2018) HsfA1a is essential for Mi-1.2-mediated resistance to Meloidogyne incognita, and regulates Wfi1 transcription and H2O2 production. Plant Physiol 176(3):2456–2471
Zhang X, Xu W, Ni D, Wang M, Guo G (2020) Genome-wide characterization of tea plant (Camellia sinensis) Hsf transcription factor family and role of CsHsfA2 in heat tolerance. BMC Plant Biol. https://doi.org/10.1186/s12870-020-02462-9
Banti V, Mafessoni F, Loreti E, Alpi A, Perata P (2010) The heat-inducible transcription factor HsfA2 enhances anoxia tolerance in Arabidopsis. Plant Physiol 152(3):1471–1483. https://doi.org/10.1104/pp.109.149815
Ogawa D, Yamaguchi K, Nishiuchi T (2007) High-level overexpression of the Arabidopsis HsfA2 gene confers not only increased themotolerance but also salt/osmotic stress tolerance and enhanced callus growth. J Exp Bot 58(12):3373–3383. https://doi.org/10.1093/jxb/erm184
Schramm F, Larkindale J, Kiehlmann E, Ganguli A, Englich G, Vierling E, von Koskull-Doering P (2008) A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J 53(2):264–274. https://doi.org/10.1111/j.1365-313X.2007.03334.x
Miller G, Mittler R (2006) Could heat shock transcription factors function as hydrogen peroxide sensors in plants? Ann Bot-london 98(2):279–288. https://doi.org/10.1093/aob/mc/mc1107
Shim D, Hwang J-U, Lee J, Lee S, Choi Y, An G, Martinoia E, Lee Y (2009) Orthologs of the class A4 heat shock transcription factor HsfA4a confer cadmium tolerance in wheat and rice. Plant Cell 21(12):4031–4043
Almoguera C, Rojas A, Diaz-Martin J, Prieto-Dapena P, Carranco R, Jordano J (2002) A seed-specific heat-shock transcription factor involved in developmental regulation during embryogenesis in sunflower. J Biol Chem 277(46):43866–43872. https://doi.org/10.1074/jbc.M207330200
Prieto-Dapena P, Almoguera C, Personat J-M, Merchan F, Jordano J (2017) Seed-specific transcription factor HSFA9 links late embryogenesis and early photomorphogenesis. J Exp Bot 68(5):1097–1108
Bharti K, von Koskull-Doring P, Bharti S, Kumar P, Tintschl-Korbitzer A, Treuter E, Nover L (2004) Tomato heat stress transcription factor HsfB1 represents a novel type of general transcription coactivator with a histone-like motif interacting with the plant CREB binding protein ortholog HAC1. Plant Cell 16(6):1521–1535. https://doi.org/10.1105/tpc.019927
Nover L, Scharf KD, Gagliardi D, Vergne P, Czarnecka-Verner E, Gurley WB (1996) The Hsf world: classification and properties of plant heat stress transcription factors. Cell Stress Chaperones 1(4):215–223
Xue GP, Sadat S, Drenth J, McIntyre CL (2014) The heat shock factor family from Triticum aestivum in response to heat and other major abiotic stresses and their role in regulation of heat shock protein genes. J Exp Bot 65(2):539–557. https://doi.org/10.1093/jxb/ert399
Wang F, Dong Q, Jiang H, Zhu S, Chen B, Xiang Y (2012) Genome-wide analysis of the heat shock transcription factors in Populus trichocarpa and Medicago truncatula. Mol Biol Rep 39(2):1877–1886. https://doi.org/10.1007/s11033-011-0933-9
Lin YX, Jiang HY, Chu ZX, Tang XL, Zhu SW, Cheng BJ (2011) Genome-wide identification, classification and analysis of heat shock transcription factor family in maize. BMC Genom. https://doi.org/10.1186/1471-2164-12-76
Guo JK, Wu J, Ji Q, Wang C, Luo L, Yuan Y, Wang YH, Wang J (2008) Genome-wide analysis of heat shock transcription factor families in rice and Arabidopsis. J Genet Genom 35(2):105–118. https://doi.org/10.1016/s1673-8527(08)60016-8
Chan AP, Crabtree J, Zhao Q, Lorenzi H, Orvis J, Puiu D, Melake-Berhan A, Jones KM, Redman J, Chen G, Cahoon EB, Gedil M, Stanke M, Haas BJ, Wortman JR, Fraser-Liggett CM, Ravel J, Rabinowicz PD (2010) Draft genome sequence of the oilseed species Ricinus communis. Nat Biotechnol 28(9):951–956. https://doi.org/10.1038/nbt.1674
Tuskan G, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313(5793):1596–1604
Hu W, Xia Z, Yan Y, Ding Z, Tie W, Wang L, Zou M, Wei Y, Lu C, Hou X, Wang W, Peng M (2015) Genome-wide gene phylogeny of CIPK family in cassava and expression analysis of partial drought-induced genes. Front Plant Sci. https://doi.org/10.3389/fpls.2015.00914
Hu W, Wei Y, Xia Z, Yan Y, Hou X, Zou M, Lu C, Wang W, Peng M (2015) Genome-wide identification and expression analysis of the NAC transcription factor family in cassava. PLoS ONE. https://doi.org/10.1371/journal.pone.0136993
Vanderschuren H, Nyaboga E, Poon JS, Baerenfaller K, Grossmann J, Hirsch-Hoffmann M, Kirchgessner N, Nanni P, Gruissem W (2014) Large-scale proteomics of the cassava storage root and identification of a target gene to reduce postharvest deterioration. Plant Cell 26(5):1913–1924. https://doi.org/10.1105/tpc.114.123927
Zidenga T, Leyva-Guerrero E, Moon H, Siritunga D, Sayre R (2012) Extending cassava root shelf life via reduction of reactive oxygen species production. Plant Physiol 159(4):1396–1407. https://doi.org/10.1104/pp.112.200345
Xu J, Duan XG, Yang J, Beeching JR, Zhang P (2013) Enhanced reactive oxygen species scavenging by overproduction of superoxide dismutase and catalase delays postharvest physiological deterioration of cassava storage roots. Plant Physiol 161(3):1517–1528
Okogbenin E, Setter TL, Ferguson M, Mutegi R, Ceballos H, Olasanmi B, Fregene M (2013) Phenotypic approaches to drought in cassava: review. Front Physiol 4(93):93
Eddy SR (2011) Accelerated profile HMM searches. PLoS Comput Biol. https://doi.org/10.1371/journal.pcbi.1002195
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. https://doi.org/10.1093/molbev/msr121
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25(9):1105–1111
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7(3):562–578. https://doi.org/10.1038/nprot.2012.016
Wilson MC, Mutka AM, Hummel AW, Berry J, Chauhan RD, Vijayaraghavan A, Taylor NJ, Voytas DF, Chitwood DH, Bart RS (2017) Gene expression atlas for the food security crop cassava. New Phytol 213(4):1632–1641. https://doi.org/10.1111/nph.14443
Jin G, Gho H, Jung K (2013) A systematic view of rice heat shock transcription factor family using phylogenomic analysis. J Plant Physiol 170(3):321–329. https://doi.org/10.1016/j.jplph.2012.09.008
Yu J, Hu SN, Wang J, Wong GKS, Li SG, Liu B, Deng YJ, Dai L, Zhou Y, Zhang XQ, Cao ML, Liu J, Sun JD, Tang JB, Chen YJ, Huang XB, Lin W, Ye C, Tong W, Cong LJ, Geng JN, Han YJ, Li L, Li W, Hu GQ, Huang XG, Li WJ, Li J, Liu ZW, Li L, Liu JP, Qi QH, Liu JS, Li L, Li T, Wang XG, Lu H, Wu TT, Zhu M, Ni PX, Han H, Dong W, Ren XY, Feng XL, Cui P, Li XR, Wang H, Xu X, Zhai WX, Xu Z, Zhang JS, He SJ, Zhang JG, Xu JC, Zhang KL, Zheng XW, Dong JH, Zeng WY, Tao L, Ye J, Tan J, Ren XD, Chen XW, He J, Liu DF, Tian W, Tian CG, Xia HG, Bao QY, Li G, Gao H, Cao T, Wang J, Zhao WM, Li P, Chen W, Wang XD, Zhang Y, Hu JF, Wang J, Liu S, Yang J, Zhang GY, Xiong YQ, Li ZJ, Mao L, Zhou CS, Zhu Z, Chen RS, Hao BL, Zheng WM, Chen SY, Guo W, Li GJ, Liu SQ, Tao M, Wang J, Zhu LH, Yuan LP, Yang HM (2002) A draft sequence of the rice genome (Oryza sativa L. ssp indica). Science 296(5565):79–92. https://doi.org/10.1126/science.1068037
Goff SA, Ricke D, Lan TH, Presting G, Wang RL, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D, Hutchinson D, Martin C, Katagiri F, Lange BM, Moughamer T, Xia Y, Budworth P, Zhong JP, Miguel T, Paszkowski U, Zhang SP, Colbert M, Sun WL, Chen LL, Cooper B, Park S, Wood TC, Mao L, Quail P, Wing R, Dean R, Yu YS, Zharkikh A, Shen R, Sahasrabudhe S, Thomas A, Cannings R, Gutin A, Pruss D, Reid J, Tavtigian S, Mitchell J, Eldredge G, Scholl T, Miller RM, Bhatnagar S, Adey N, Rubano T, Tusneem N, Robinson R, Feldhaus J, Macalma T, Oliphant A, Briggs S (2002) A draft sequence of the rice genome (Oryza sativa L. ssp japonica). Science 296(5565):92–100. https://doi.org/10.1126/science.1068275
Arabidopsis Genome I (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408(6814):796–815
Zhu Y, Yan H, Wang Y, Feng L, Chen Z, Xiang Y (2016) Genome duplication and evolution of heat shock transcription factor (HSF) gene family in four model angiosperms. J Plant Growth Regul 35(4):903–920. https://doi.org/10.1007/s00344-016-9590-5
Albihlal WS, Irabonosi O, Blein T, Persad R, Chernukhin I, Crespi M, Bechtold U, Mullineaux PM (2018) Arabidopsis heat shock transcription FactorA1b regulates multiple developmental genes under benign and stress conditions. J Exp Bot 69(11):2847–2862
Carranco R, Manuel Espinosa J, Prieto-Dapena P, Almoguera C, Jordano J (2010) Repression by an auxin/indole acetic acid protein connects auxin signaling with heat shock factor-mediated seed longevity. Proc Natl Acad Sci USA 107(50):21908–21913. https://doi.org/10.1073/pnas.1014856107
Reilly K, Gomez-Vasquez R, Buschmann H, Tohme J, Beeching JR (2004) Oxidative stress responses during cassava post-harvest physiological deterioration. Plant Mol Biol 56(4):625–641. https://doi.org/10.1007/s11103-005-2271-6
Mittal D, Chakrabarti S, Sarkar A, Singh A, Grover A (2009) Heat shock factor gene family in rice: genomic organization and transcript expression profiling in response to high temperature, low temperature and oxidative stresses. Plant Physiol Biochem 47(9):785–795. https://doi.org/10.1016/j.plaphy.2009.05.003
Miller G, Shulaev V, Mittler R (2008) Reactive oxygen signaling and abiotic stress. Physiol Plant 133(3):481–489. https://doi.org/10.1111/j.1399-3054.2008.01090.x
Acknowledgements
This work was supported by the Key Research and Development Project of Hainan Province (ZDYF2018220), Key Projects of Shaoguan University (Grant No. SZ2018KJ05), Foundation for Young Talents in Higher Education of Guangdong (Natural Science) (Grant No. 2018KQNCX234), Science and Technology Planning Project of Shaoguan (Grant No. 2019sn087), Doctoral Scientific Research Startup Foundation of Shaoguan University (Grant No. 99000615), National Natural Science Foundation of China (31901537, 31801419), Central Public-interest Scientific Institution Basal Research Fund for Chinese Academy of Tropical Agricultural Sciences (CATAS) (1630052019023), the Central Public-interest Scientific Institution Basal Research Fund for Innovative Research Team Program of CATAS (17CXTD-28; 1630052017017). We thank Dandan Li for editorial assistance and helpful discussions.
Author information
Authors and Affiliations
Contributions
Conceptualization, WH; Methodology and software, ZHD, JZ; Validation, JZ; Formal analysis, FFL, JJM, WWT, and YY; Writing original draft preparation, JZ, CLW, CW; Supervision, WH.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zeng, J., Wu, C., Wang, C. et al. Genomic analyses of heat stress transcription factors (HSFs) in simulated drought stress response and storage root deterioration after harvest in cassava. Mol Biol Rep 47, 5997–6007 (2020). https://doi.org/10.1007/s11033-020-05673-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05673-3