Abstract
In accordance with the Asian BRCA Consortium data, there is a significant difference in incidence rate of breast cancer depending on age, as well as spectrum and prevalence of BRCA1/2 mutations between Mongoloid (East Asian) and Caucasoid (European) people. However, European strategies to identify familial BC are still applied to the Asian population, including Russian Mongoloids (Khakas, Buryats, Tyvans and Yakuts and others). The main purpose of the study was to identify molecular changes associated with hereditary BC in Russian Mongoloid BC patients (Buryats). Thirty-nine patients were included in the study. Genomic DNA extracted from lymphocytes was used to prepare DNA-libraries. Target sequencing was designed to cover 27 genes, such as ATM, APC, BARD1, BRCA1, BRCA2, BRIP1, CDH1, CHEK2 and others. Paired-end sequencing (2 × 150 bp) was conducted on a NextSeq 500 system (Illumina, USA). Three pathogenic mutations in non-BRCA genes were found (prevalence of 8%). The pathogenic mutations were found in the RAD51D and PTEN genes. The pathogenic variant in the RAD51D gene (rs137886232, NC_000017.10:g.33428366G>A, p.R141X) was observed in two unrelated individuals aged under 40. One of these patients had a family history of late-onset stomach cancer in second-degree relatives. The pathogenic mutation in the PTEN gene (rs786201044, NC_000010.10:g.89692922T>C, p.C136R) was observed in a 38 years old breast cancer patient with no family history. In our study, we first describe pathogenic mutations in RAD51D and PTEN genes found in young Buryat patients.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BC) is the most common female malignancy worldwide. Mutations in BRCA1 and BRCA2 genes are responsible for hereditary BC. Individuals with inherited mutations in BRCA genes should be offered the risk-reduction strategies, such as screening (mammography and breast magnetic resonance imaging), surveillance (clinical breast examination, breast self-examination), chemoprevention, and risk-reduction surgery [1].
BC is also caused by mutations in the TP53, ATM, CDH1, PTEN, and STK11 genes associated with hereditary syndromes. Mutations in the genes mentioned above can inhibit DNA repair pathways [2, 3].
In Russians, mutations of BRCA1/2 genes were found only among Slavic women (newcomers), and were not found in Khakas, Buryats, Tyvans, Yakuts and others women (indigenous) [4, 5]. However, the cancer burden in Khakas, Buryats, Tyvans has risen and the cancer risk assessment has been limited [6,7,8]. The main purpose of the study was to identify molecular changes associated with hereditary BC in Russian Mongoloid BC patients.
Materials and methods
Thirty-nine patients were included in the study. The median age of patients at BC diagnosis was 42 years (range: 26–55). Eighty-one percent of patients were diagnosed with BC before the age of 50. More than one-third of patients under the age of 50 had a family history of BC. Almost all tested women were diagnosed with invasive (ductal) carcinoma of no special type. Information, including age at diagnosis, family history, histological type of cancer, and family origin was obtained. Reported clinical characteristics of the patients are given in Table 1.
Blood samples were collected in ethylenediaminetetra-acetic acid-containing tubes. Genomic DNA from peripheral blood was extracted using the phenol–chloroform method. Purity of the DNA was assessed using a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific, USA) and then quantified using the Qubit 2.0 fluorometer and HS dsDNA Assay Kit (Thermo Fisher Scientific, USA). Integrity of the DNA (DIN) was verified on a 2200 TapeStation system (Agilent, USA). The positive control sample with BRCA1 c.3755_3758delTGTC pathogenic mutation was included as an inner control.
DNA library were prepared using the Hereditary Cancer Solution™ kit (Sophia GENETICS, Switzerland) to cover 27 genes: ATM, APC, BARD1, BRCA1, BRCA2, BRIP1, CDH1, CHEK2, EPCAM, FAM175A, MLH1, MRE11A, MSH2, MSH6, MUTYH, NBN, PALB2, PIK3CA, PMS2, PMS2CL, PTEN, RAD50, RAD51C, RAD51D, STK11, TP53, and XRCC2. Paired-end sequencing (2 × 150 bp) was conducted using NextSeq 500 system (Illumina, USA). The pathogenic variants were validated using Sanger sequencing (SeqStudio, Thermo Fisher Scientific, USA).
Bioinformatics analysis
Sequencing data was analyzed according to the GATK best practice recommendation for Whole Exome Sequencing using GRCh37 as a reference for Burrows-Wheeler alignment. The obtained variants were annotated with ANNOVAR software and ranged according to population frequency (genomic exome, gnomAD genome, and ExAC), ClinVar, CADD, and literature data [9,10,11]. Detected sequence variants were annotated using PolyPhen2, Mutation Taster, and SIFT [12,13,14].
Results
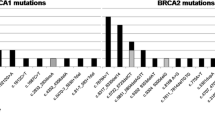
In our study, 8% (3/39) of patients harbored one pathogenic variant and 15% (6/39) of patients harbored likely pathogenic variant. In addition, 8% of patients had VUS, 15% had conflicting variants and 54% had only benign variants (Fig. 1a).
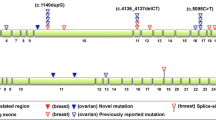
It should be noted that the pathogenic variants were found in two non-BRCA1/2 susceptibility genes and were diagnosed only in BC patients under 45 years old (Figs. 1b, 2). Table 2 illustrates variants that are described as highly pathogenic by dbPubMed, likely pathogenic (possibly/probably damaging by PolyPhen2 or deleterious by SIFT).
The RAD51D variant (rs137886232) was observed in two unrelated individuals. One of these patients had a family history of late-onset stomach cancer in second-degree relatives. Another pathogenic variant was observed in the PTEN gene (rs786201044) in a BC patient aged 38 with no family history of BC.
Variants in the ATM, MSH6 and MLH1 genes (likely pathogenic) were previously described as VUS by dbPubMed, but were predicted as probably damaging by PolyPhen2 and/or deleterious by SIFT. The probably damaging variants (PolyPhen2) in the ATM (rs150757822) and MSH6 (rs142254875) genes were observed in a 49-year-old patient with a burdened family history and in a 48-year-old patient with an unknown family history, respectively. One MLH1 variant (rs4986984) was classified as probably damaging (PolyPhen2) and deleterious (SIFT), and was found in two patients diagnosed with BC prior to 46 years and in a 52-year-old patient with a burdened family history. Another probably damaging/deleterious variant in the MLH1 gene (rs367654552) was found in a 55-year-old patient with a burdened family history.
Rare genetic variants classified by dbPubMed as VUS are given in Table 3. All variants presented in Table 3 were considered benign by PolyPhen2 or tolerated by SIFT. One missense VUS (rs80359254) was identified in the BRCA2 gene. VUS were most commonly encountered in the ATM (n = 3), MSH6 (n = 2) and MLH1 (n = 3) genes.
The rs367654552 of the MLH1 gene was previously described only in the East Asia, whereas the rs80359254 variant of the BRCA2 gene was observed exclusively in Europeans. The rs150757822 and rs1800058 variants of the ATM gene and the rs104894994 variant of the MLH1 gene were found in South Asian populations but not in the East Asian populations. In general, 77% (10 out of 13) of the identified variants were previously found among Europeans (Table 4).
No mutations were found in the MRE11A, PIK3CA, RAD51C or XRCC2 gene.
4. Discussion
Mongoloid population is the most prevalent among all human populations [15]. In accordance with the Asian BRCA Consortium data, there is a significant difference in incidence rate of BC depending on age, as well as spectrum and prevalence of BRCA1/2 mutations and clinical significance of rare VUS between Mongoloid (East Asian) and Caucasoid (European) people [16,17,18,19,20]. However, European strategies to identify familial BC are still applied to the Asian population, including Mongoloids.
There are more than 200 different ethnic groups in Russia. Most of the population in Russia includes Russians (81%), the largest ethnicities are Tatars, Belarusians, Ukrainians, Bashkirs, Chuvash, Chechens, and Armenians (up to 10%), and smallest nationalities include Kazakhs, Yakuts, Buryats, Ingush, Udmurts, Ossetians and others (up to 0.5% of each) [5]. In our study, we continued to search for mutations in BRCA-negative BC women living in Russia (Buryat). Overall, two pathogenic germline variants in RAD51D and PTEN were found in 8% (3/39) of patients under 40. In addition, 8% of patients had VUS, 15% had conflicting variants and 54% had only benign variants.
The pathogenic variant of RAD51D gene (rs137886232) with low minor allele frequency was observed in two young unrelated Buryat patients. The germline mutation of RAD51D gene (rs137886232) was suggested to have a founder effect in Chinese population [21]. This variant was also described in BC families of European ancestry. It was also reported that RAD51D-deficient tumor cells were sensitive to poly-(ADP) ribose polymerase inhibitors [22].
The pathogenic variant of the PTEN gene (rs786201044), which was predicted to be damaging by in silico analysis, was observed in BC patient aged 38 with no family history of BC. Different studies also reported on this variant (rs786201044) of the PTEN gene in families with Cowden syndrome (an autosomal dominant inherited disorder) [23,24,25,26]. Previous studies have also found that this variant (rs786201044) of the PTEN gene may impact on protein stability and lead to increased proteasome activity [27,28,29,30,31].
Interesting, germline variants of RAD51D and PTEN genes were also described in the COSMIC database as somatic mutations, COSM4721157 and COSM5096, respectively.
In 54% of the patients, no clinically significant variants were identified, probably due to some limitations to our study. In particular, we used a panel of only 27 genes that did not include other BC-predisposing genes, such as BLM, ESR1, FANCA and NQO2.
In this study, over 20% of the Buryat BC patients were found to carry a rare VUS. Buryats are characterized by molecular diversity due to the long generation time or the mixed nature of origin compared with other ethnic groups living in Siberia [32,33,34,35]. It is obvious that a more detailed genetic analysis of the Buryats is required.
Conclusion
In this study, we provide the first description of two pathogenic germline variants in the RAD51D (rs137886232) and PTEN (rs786201044) genes in BRCA1/2-negative Mongoloid (Buryat) women with BC.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Dutil J, Teer JK, Golubeva V, Yoder S et al (2019) Germline variants in cancer genes in high-risk non-BRCA patients from Puerto Rico. Sci Rep 9(1):17769
Frey MK, Pothuri B (2017) Homologous recombination deficiency (HRD) testing in ovarian cancer clinical practice: a review of the literature. Gynecol Oncol Res Pract 4:4
Rodríguez-Balada M, Roig B, Melé M, Albacar C et al (2020) Identification of germline pathogenic variants in DNA damage repair genes by a next-generation sequencing multigene panel in BRCAX patients. Clin Biochem 76:17–23
Iyevleva AG, Suspitsin EN, Kroeze K, Gorodnova TV et al (2010) Non-founder BRCA1 mutations in Russian breast cancer patients. Cancer Lett 298:258–263
Sokolenko AP, Preobrazhenskaya EV, Aleksakhina SN, Iyevleva AG et al (2015) Candidate gene analysis of BRCA1/2 mutation-negative high-risk Russian breast cancer patients. Cancer Lett 359:259–261
Cherdyntseva NV, Pisareva LF, Ivanova A, Panferova Y et al (2014) Ethnic aspects of hereditary breast cancer in the region of Siberia. Vestn Ross Akad Med Nauk 11–12:72–79
Cherdyntseva N, Gervas P, Voropaeva E, Denisov E et al (2017) New variants in the BRCA1 gene in Buryat Mongol breast cancer patients: report from two families. Cancer Biomark 18:291–296
Gervas P, Klyuch B, Denisov E, Kiselev A et al (2019) New germline BRCA2 gene variant in the Tuvinian Mongol breast cancer patients. Mol Biol Rep 46(5):5537–5541
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R et al (2013) From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinform 43:11
DePristo MA, Banks E, Poplin R, Garimella KV et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43:491–498
McKenna A, Hanna M, Banks E, Sivachenko A et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249
Schwarz JM, Cooper DN, Schuelke M, Seelow D (2014) MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 11(4):361–362
Kumar P, Henikoff S, Ng PC (2009) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4:1073–1081
Kiseleva OA, Robustova OV, Bessmertny AM, Zakharova EK, Avdeev RV (2013) Prevalence of primary glaucoma in representatives of different races and ethnic groups in Russia and in CIS. Ophthalmology 10(4):11–15
Kwong A, Shin VY, Ho JC, Kang E et al (2016) Comprehensive spectrum of BRCA1 and BRCA2 deleterious mutations in breast cancer in Asian countries. J Med Genet 53:15–23
Nakamura S, Kwong A, Kim SW et al (2016) Current status of the management of hereditary breast and ovarian cancer in Asia: first report by the Asian BRCA Consortium. Public Health Genomics 19:53–60
Khoo US, Chan KY, Cheung AN, Xue WC et al (2002) Recurrent BRCA1 and BRCA2 germline mutations in ovarian cancer: a founder mutation of BRCA1 identified in the Chinese population. Hum Mutat 19:307–308
Ginsburg OM, Dinh NV, To TV, Quang LH et al (2011) Family history, BRCA mutations and breast cancer in Vietnamese women. Clin Gene 80:89–92
Kim YC, Zhao L, Zhang H, Huang Y et al (2016) Prevalence and spectrum of BRCA germline variants in mainland Chinese familial breast and ovarian cancer patients. Oncotarget 7:9600–9612
Sun J, Meng H, Yao L, Lv M, Bai J et al (2017) Germline mutations in cancer susceptibility genes in a large series of unselected breast cancer patients. Clin Cancer Res 23(20):6113–6119
Loveday C, Turnbull C, Ramsay E, Hughes D et al (2011) Germline mutations in RAD51D confer susceptibility to ovarian cancer. Nat Genet 43(9):879–882
Paparo L, Rossi GB, Delrio P, Rega D et al (2013) Differential expression of PTEN gene correlates with phenotypic heterogeneity in three cases of patients showing clinical manifestations of PTEN hamartoma tumour syndrome. Hered Cancer Clin Pract 11(1):8
He X, Arrotta N, Radhakrishnan D, Wang Y et al (2013) Cowden syndrome-related mutations in PTEN associate with enhanced proteasome activity. Cancer Res 73(10):3029–3040
Galatola M, Paparo L, Duraturo F, Turano M et al (2012) Beta catenin and cytokine pathway dysregulation in patients with manifestations of the "PTEN hamartoma tumor syndrome". BMC Med Genet 13:28
Venturini G, Moulin AP, Deprez M, Uffer S et al (2012) Clinicopathologic and molecular analysis of a choroidal pigmented schwannoma in the context of a PTEN hamartoma tumor syndrome. Ophthalmology 119(4):857–864
Pilarski R, Stephens JA, Noss R, Fisher JL, Prior TW (2011) Predicting PTEN mutations: an evaluation of Cowden syndrome and Bannayan-Riley-Ruvalcaba syndrome clinical features. J Med Genet 48(8):505–512
Tan MH, Mester J, Peterson C, Yang Y et al (2011) A clinical scoring system for selection of patients for PTEN mutation testing is proposed on the basis of a prospective study of 3042 probands. Am J Hum Genet 88(1):42–56
Kubo Y, Urano Y, Hida Y, Ikeuchi T et al (2000) A novel PTEN mutation in a Japanese patient with Cowden disease. Br J Dermatol 142(6):1100–1105
Mester J, Eng C (2015) Cowden syndrome: recognizing and managing a not-so-rare hereditary cancer syndrome. J Surg Oncol 111(1):125–130
Cheng J, Peng J, Fu J, Khan A, Tan P, Wei C, Deng X, Chen H, Fu J (2020) Identification of a novel germline BRCA2 variant in a Chinese breast cancer family. J Cell Mol Med 24(2):1676–1683
Stepanov VA, Spiridonova MG, Puzyrev VP (2003) Comparative phylogenetic study of native north Eurasian populations from a panel of autosomal microsatellite loci. Genetika 39(11):1564–1572
Stepanov VA, Spiridonova MG, Tadinova VN, Puzyrev VP (2003) Analysis of genetic diversity of population of Northern Eurasia from autosomal microsatellite loci. Genetika 39(10):1381–1388
Har'kov VN, Hamina KV, Medvedeva OF (2014) Gene pool of Buryats: clinal variability and territorial subdivision based on data of Y-chromosome markers. Genetika 50(2):203–213
Khitrinskaia II, Khar'kov VN, Stepanov VA (2010) Genetic diversity of X-chromosome in populations of aboriginal Siberian ethnic groups: linkage disequilibrium structure and haplotype philogeography of ZFX locus. Mol Biol (Mosk) 44(5):804–815
Acknowledgements
We acknowledge the Tomsk State University for the Competitiveness Improvement Program. The work with DNA sequencing was performed on equipment from the Tomsk regional common use center with the support of the Russian Ministry; Agreement No.14.594.21.0001 (RFMEFI59414X0001).
Funding
The reported study was funded by RFBR according to research project 18-29-09046.
Author information
Authors and Affiliations
Contributions
PG, AM, AS, AK, LP recruited patients, collected samples and conducted experiments. PG wrote the manuscript; and NB revised the manuscript. EC, NC, YT and LP supervised the project. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The current study was approved by the review board of the Cancer Research Institute, and written informed consent was obtained from all patients and in compliance with the recommendations of the Helsinki Declaration.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gervas, P., Molokov, A., Schegoleva, A. et al. New germline mutations in non-BRCA genes among breast cancer women of Mongoloid origin. Mol Biol Rep 47, 5315–5321 (2020). https://doi.org/10.1007/s11033-020-05612-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05612-2