Abstract
Successful pathogenicity often resulted from a complicated association between virulence and antibiotic resistance in Pseudomonas aeruginosa infections. Therefore, the current study aimed to investigate the relationship between the las system and antibiotic resistance. Seventy-three (73) P. aeruginosa isolates were collected from burn wounds (26.02%), blood cultures (30.13%), catheters (12.32%), and urine culture (31.50%). Among the 73 collected isolates, 22 isolates were considered as multi-drug resistant (MDR) and 11 isolates as extensively-drug resistant (XDR). Furthermore, phenazines and LasA protease were detected among 21.91% and 32.87% of isolates, respectively. Quantitative real-time PCR assessment of KPC, MBL, and lasI/R indicated that resistance and virulence factors are more expressed in XDR strains than MDR strains. Also, the expression level of KPC and MBL reduced in non-biofilm forming strains. However, increased expression levels of lasI, lasR, and the KPC genes were observed in LasA and LasB protease producing strains. Interestingly, 16 known sequence types (including ST108, ST260, ST217) and three novel STs (ST2452, ST2427, and ST2542) were characterized among the collected isolates, which are related to the virulence and resistance. In MDR-XDR strains, a strong correlation between lasI/R and the variants of antibiotic resistance genes was found. In conclusion, the pathogenicity of P. aeruginosa may increase the prevalence of antibiotic-resistant strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudomonas aeruginosa possesses an arsenal of virulence factors and a variety of antibiotic resistance mechanisms to cause disease successfully. To regulate the pathogenicity in response to cell density, P. aeruginosa develops three different quorum sensing (QS) systems, including Las, Rhl, and PQS [1]. QS causes organisms to act in synchrony for controlling a variety of functions such as bioluminescence, biofilm formation, virulence factor production, and antibiotic resistance [2]. The systems mentioned above dependently regulate QS- related processes in the organism and are auto-regulated to commence regulatory cascades [3].
The Las is a two-component system which includes a sensor-lasI, and a response regulator- lasR. In response to the growth phase and accumulation of autoinducer peptides (AIPs), such as acyl-homoserine lactones (AHL), LasI expresses, which results in LasR activation [4]. LasR, as a global regulator, controls the production of siderophores, metalloprotease elastase (LasB), metalloprotease (LasA), phenazines, and exotoxin A [5]. Moreover, pyocyanin as a critical virulence factor of P. aeruginosa is regulated and secreted under the control of QS system. This factor is a blue redox-active phenazine, which acts through reactive oxygen species (ROS) production, and plays different roles in disturbing critical organs, including respiratory, cardiovascular, urogenital, and central nervous system (CNS) [6, 7].
QS plays a vital role in the biofilm formation by microorganisms, particularly in chronic infections, and consequently increases antibiotic resistance [8]. Intrinsic or acquired resistance to antibiotics have been evolved through decades and resulted in multi-drug- or extensively drug-resistant (MDR and XDR) strains of P. aeruginosa [9]. Due to high energy-consuming processes of antibiotic resistance, microorganisms neatly fine-tune the resistance and the virulence. During P. aeruginosa infections, QS plays a notable role to control the resistance and the virulence. In order to illustrate, efflux pumps upregulate in exposure to C4HSL (an AIP from Rhl QS system), and as a consequence, antibiotic resistance increases [10]. However, the relationship between virulence and resistance is still unclear, and contradictions are not resolved.
Although resistance to different antibiotics can affect the fitness costs of the organism and results in attenuation of the production of virulence factors [11, 12], some studies demonstrated a contradiction. It has been mentioned that aztreonam-resistance leads to hypervirulent strains of P. aeruginosa in cystic fibrosis (CF) patients [13]. Therefore, the present study aimed to determine the relationship between presence and expression of las regulated virulence factors and resistance genes in MDR and XDR P. aeruginosa strains.
Material and methods
Study design and sample collection
In this descriptive-analytic study, which was performed during six months (April 2015 till August 2016), 73 different clinical specimens, including burn wounds, blood cultures, catheters, and UC (urine culture) samples, were collected from educational Hospitals of Hamadan, Iran. The isolates were characterized and identified by oxidase test (Himedia, India), triple sugar iron (TSI) agar (Merck, Germany), oxidation- fermentation (OF) media (Merck, Germany), and MacConkey agar (Merck, Germany). The oxidase-positive colonies which had a grapelike odor, grown on TSI agar with an alkaline/no change reaction, and consumed glucose only in the aerobic condition in OF media, considered as P. aeruginosa isolates [14].
Antimicrobial agents susceptibility testing
The collected isolates were investigated using antibiotic susceptibility tests for different antibiotic categories including Amikacin (30 µg), Doripenem(10 µg), Meropenem(10 µg), Imipenem(10 µg), Cefoxitin(30 µg), Cefpodoxime(30 µg), Cefotaxime(30 µg), Ceftazidime(30 µg), Ceftriaxone (30 µg), Ciprofloxacin (5 µg), Piperacillin-Tazobactam (100/10 µg), Piperacillin (100 µg),Ticarcillin (75 µg) and Aztreonam (30 µg). The antibiotic susceptibility testing was done based on CLSI 2016. Resistance to at least one antibiotic in more than three antimicrobial categories was considered as MDR. Also, resistance to at least one agent in more than 6 antibiotic families was regarded as XDR. P. aeruginosa ATCC 27,853 was used as the reference strain in each assay. All antibiotic disks were purchased from Mast, UK.
Phenotypic characterization of pyocyanin production
In order to examine pyocyanin production, the chloroform and HCl (Merck, Germany) extraction method based on El Fouly et al. was used [15]. The OD520nm of samples was multiplied to 17.072 to determine the concentration of pyocyanin in µg/mL.
Phenotypic characterization of pyoverdine production
Strains were inoculated to RPMI1640 (Invitrogen, USA) and incubated at 37 °C by shaking 100 rpm overnight. The OD600nm of the cultures was measured. Then the cultures were centrifuged at 200 g for 30 min. The supernatants were collected and filtered by 0.22 µm Millipore filters (Merck, Germany). Using a spectrophotometer, the OD405nm of supernatants was measured, and then Relative Pyoverdine Production (RPP) was calculated by the following formula: RPP: OD405/OD600.
Biofilm formation
To examine the capacity of isolates to produce biofilm, the Crystal violet assay was done according to O'Toole et al. study [16]. To define the biofilm production of the isolates, OD cut off was calculated using a formula as below: ODcut off: ODavg of negative control + 3*SD of ODs of the negative control. If OD of tests were less than ODcut off, the isolates were a non-biofilm producer. The isolates interpreted as weak biofilm producers if the OD lays between ODcut off and 2*ODcut off. And strong biofilm formers showed an OD more than 4*ODcut off.
Phenotypic characterization of LasA and LasB enzymatic activity
LasA enzymatic activity was measured according to Oldak et al. [17]. Briefly, an overnight culture of Staphylococcus aureus ATCC25923 was centrifuged and re-suspended in PBS buffer, and the OD600nm was adjusted to 0.8. Also, overnight cultures of P. aeruginosa strains were centrifuged and re-suspended in the CDMC solution. The CDMC was (pH 7.4) consisted of glucose (30 mM), NaCl (8 mM), K2HPO4 (60 mM), KH2PO4 (35 mM), ZnCl2 (0.025 mM), (NH4)2SO2 (15 mM), l-glutamine (7 mM), CaCl2 (0.05 mM), FeCl3 (0.017 mM), C6H5Na3O7 (35 mM), MgCl2 (1.4 mM), thiamine (0.15 mM), dl-arginine (0.22 mM), uracil (0.2 mM), and nicotinic acid (0.1 mM). This solution stimulates LasA Staphylolytic activity. Then, 100 µL of CDMC was added to 900 µL of Staphylococcal suspension, and the decrease in absorbance of the solution was spectrophotometrically monitored at OD595nm. To determine the protease activity of LasB, 1% skim milk medium was used. Isolates were cultured on the plates, incubated overnight at 37 °C, and the clear zone around colonies was monitored the next day.
Genomic DNA
Strains were inoculated into LB broth (Merck, Germany), and then incubated at 37 °C. In order to extract genomic DNA and plasmid, the Qiagen extraction kit (Germany) was applied using manufacturer instructions.
Virulence factor production and resistance genes detection
PCR method was applied to determine virulence factor production for lasB, lasA, apr, plcH, phzI, phzM. Also, resistance genes, including carbapenemase, ESBLs, MBLs, and AmpC families, were detected using primers, according to Fazeli and et al. study [18, 19]. Multiplex PCR was done in a 50 µL volume reaction containing 2 µL DNA, 1 µL of each primer (5 pmol L−1) and 20 µL of master mix (Amplicon, Denmark) reached 50 µL by adding deionized water. The PCR program consisted of an initial denaturation at 96 °C for 10 min, 30 cycles of 1 min denaturation at 96 °C, annealing 58 °C for 1 min, an extension for 1 min at 72 °C, and a final extension at 72 °C for 10 min.
Sequencing
PCR products were purified and sequenced by Bioneer Co., Korea mediated by Pishgam Co., Iran. The data were analyzed using the Chromas software and compared to the microbial genome using the Basic Local Alignment Search Tool (BLAST) to confirm the sequence authenticity.
Sample selection for gene expression
All strains of MDR and XDR P. aeruginosa were selected to evaluate gene expression. The selected isolates demonstrated variation in Las-regulated virulence production, antibiotic susceptibility patterns, and demographic characteristics.
RNA extraction and qRT-PCR
The strains were inoculated into LB broth (Merck, Germany), and then incubated at 37 °C. RNA was extracted, and cDNA synthesis was performed using the GeneAll RNA extraction kit and GeneAll cDNA synthesis kit (GeneAll, Korea) according to the manufacturer instructions. Quantitive RT-PCR was used to determine the expression of lasR, lasI, amp, mexR. KPC, MBL genes using a syber green master mix (Amplicon, Denmark) and aroC was applied as the reference gene. The primers of Lima et al. [20], Quale et al. [21], Zaman et al. [22], and Geyer et al. [23], studies were used for q-RT PCR. To qualify the qPCR test, the standard validation tests for assessing the sensitivity and Melt curve analysis for determining the specificity of primers were performed. To determine the amplification efficiency (EFF %), 10 dilutions of cDNA were prepared, and the most efficient dilution was determined according to the 10−slope equation. The most suitable amount in this equation is less than 2. To calculate the expression level, the 2−ΔΔCT equation was used based on the Pfaffl study [24]. To normalize expression levels, P. aeruginosa PAO1 was used as a reference strain.
Multilocus sequence typing (MLST)
MLST was performed using the scheme described by Vernez et al., [25]. The primers used to detect seven housekeeping genes (acsA, aroE, guaA, mutL, nuoD, ppsA, and trpE). For each of the seven MLST loci, 50 µL PCR reactions were performed in 96-well plates. The PCR mixture contained 39.75 µL of molecular grade water, 5 µL of 10 × PCR buffer (Qiagen, UK), 1 µL of a 10 µM concentration of each forward and reverse primer, 1 µL of 10 mM deoxynucleoside triphosphate (dNTP) mix (Invitrogen, UK), 0.25 µL of HotStart Taq DNA polymerase (Qiagen, UK) and 2 µL of P. aeruginosagDNA. The MLST PCR was performed using a Bio-Rad C1001 thermocycler (California, United States) and thermocycling conditions were; 35 cycles at 94 °C for 15 s, 50 °C for 1 min, 72 °C for 1 min. This was followed by a hold step of 72 °C for 7 min.
Statistical analysis
All statistical analyses of phenotypic data were carried out in SPSS software (version 16, Chicago, IL, USA), using One-Way and Two-Way analysis of variance (ANOVA) for individual comparisons and Tukey’s for multiple comparisons. A p-value of less than 0.05 was reported as statistically significant (*p < 0.05; **p < 0.01; ***p < 0.001). The statistical significance was defined as P < 0.05. Gene expression analysis was performed using REST® 2009 (Qiagen, Germany) software. For MLST analysis, strain information that includes strain designation, subspecies, serovar, year of isolation, host, continent, and the source was stored in a MEGA 6 (version 6.0,https://www.megasoftware.net). The nucleotide sequences of gene fragments were also stored in the same database.
Results
Isolate characterization and antimicrobial resistance profiles
Totally, 73 isolates of P. aeruginosa were isolated from 19 (26.02%) burn wounds, 22 (30.13%), blood cultures, 9 (12.32%) catheters, and 23 (31.50%) urine culture samples in Hamadan University hospitals. Also, according to the disk diffusion method, resistance to cefoxitin, ciprofloxacin, and cefotaxime reported in 79.45%, 75.34% and 97.26% of isolates. Moreover, more than 49% of the isolates were resistant to cefpodoxime (67.12%) and ceftazidime (49.31%). Besides, 39.73% of the isolates were non-susceptible to ceftriaxone, 36.97% to imipenem, 31.50% to doripenem, 30.13% to piperacillin, and 28.76% to meropenem. Resistance to other antibiotics (except for ticarcillin and amikacin) was observed in more than 20% of the isolates (Table 1).
Pyocyanin and pyoverdine Production
Out of 73 P. aeruginosa isolates, 16 isolates (21.91%) possess phenazines, and pyocyanin was detected in 22 isolates (30.13%) (Table 1).
Biofilm formation
Out of 73 P. aeruginosa isolates, 17 (21.91%) isolates were strong biofilm producers, 1 (21.91%) isolates moderately formed biofilm, and 55 (75.34%) isolates were a weak or non-biofilm producer.
LasA and LasB enzymatic activity
Among 73 P. aeruginosa isolates, LasA and LasB proteases were detected in 24 (32.87) isolates, and 22 (30.13) isolates, respectively (Table 2).
Distribution of virulence factor genes
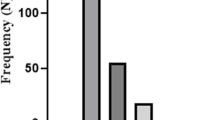
Out of 73 P. aeruginosa isolates, 24 (36.87%) isolates carried lasA gene, 20 (27.39%) isolates carried lasB gene. Also, the frequency of apr and plcH were 26.02% (19 isolates) and 21.91% (16 isolates), respectively. Moreover, phzI and phzM were detected in 21 (28.76%) and 23 (31.50%) isolates, respectively (Table2) (Fig. 1).
Distribution of antibiotic resistance genes
Among 73 P. aeruginosa isolates, 12 (16.43%) isolates carried the KPC gene, 10 (13.69%) isolates carried the IMP gene. VIM, FOX, and MOX genes were detected in 10 (13.69%) isolates, 55 (75.34%), and 48 (65.75%) isolates, respectively. 11 (15.06%) isolates carried the SIM gene, 9 (12.32%) isolates carried the GIM gene. Moreover, 29 (39.72%) and 27 (36.98%) isolates were identified to carry TEM and SHV plasmid genes, respectively (Table1) (Fig. 1).
Gene sequencing
All PCR products were assigned to the microbial genome using the Basic Local Alignment Search Tool (BLAST) and showed the same DNA sequences, which confirmed all PCR assay results.
Resistance and Virulence genes functionality evaluation:
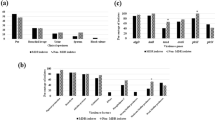
According to Fig. 2a, the activity of antibiotic resistance and virulence factors genes in XDR strains is more than MDR strains. In non-MDR-XDR strains, the activity of virulence factor genes was increased, and the activity of antibiotic resistance genes was decreased. In Fig. 2b, the expression of the virulence factor genes was higher in the biofilm producing strains. However, the expression of antibiotic resistance genes in weak/non-biofilm producer strains was reduced.
Virulence and antibiotic resistance gene expression average’s in MDR and XDR strains. a Gene expression of virulence and antibiotic resistance genes in MDR and XDR strains. b Gene expression of virulence and antibiotic resistance genes in biofilm positive and biofilm negative strains. c Gene expression of virulence and antibiotic resistance genes in strains with/without lasA and lasB enzymes. d Gene expression of virulence and antibiotic resistance genes in strains with/without phenazine and pyocyanin enzymes. Bars represent means ± SD of the results of three independent experiments. Asterisks indicate significant differences in gene expression levels between (*P < 0.05; **P < 0.01; ***P < 0.001)
Strains with strong biofilm showed increased expression of virulence factor and antibiotic resistance genes. Also, KPC and MBL genes were associated with reduced expression in non-biofilm forming strains. Due to the distribution of LasA and LasB protease enzymes in different strains (Tables 1 and 2), the increased frequency of these enzymes was associated with increased expression of the virulence factor and antibiotic resistance genes.
The frequency of phenazine and pyocyanin in MDR and XDR strains was higher than sensitive strains. Moreover, the expression of virulence factor and antibiotic resistance genes in XDR and MDR strains comparing to sensitive strains was extremely high. Of course, the expression of the MBL and ampC genes differed from other genes and did not follow the distribution of these enzymes.
Phylogenetic analysis of the MLST sequences
MLST was performed on all 73 isolates with reference strains. MLST analysis identified 16 known sequence types (ST108, ST260, ST217) and found 3 sequence types (ST2452, ST2427, and ST2542) to be novel (Fig. 3). The phylogenetic tree shows that ST108, ST217, ST260, ST274, ST501, ST1075, and ST3335 were highly resistant and pathogenic isolates (Fig. 3). Several close clusters were identified. Typing by MLST further illustrated the large diversity found within the strains, as isolates with similar β-lactamase genes had different STs.
Statistical analysis
The frequency of antibiotic resistance genes and virulence factor genes had a significant correlation with the antibiotic resistance pattern (p ≤ 0.05). Also, the frequency of antibiotic resistance genes and virulence factors were significantly correlated with biofilm expression (p ≤ 0.05) (Table 3).
Data analysis by Wilcoxon signed-rank test showed that the activity of QS genes had an inductive effect on each other. Also, the results clearly showed that there was a significant relationship between ampC and lasR gene expressions. On the other hand, a weak correlation was observed between the activity of KPC and lasR genes in the isolates (Fig. 2) (Table 3).
Discussion
Pseudomonas aeruginosa, as an opportunistic pathogen, is capable of causing a wide range of infections, particularly in patients with a compromised immune system due to long term hospitalization or other damages [18]. Prevalence of P. aeruginosa resistant to at least one of the studied antibiotics was 70%, higher than reported by Dou et al. [26], in their study investigating antibiotic resistance among P. aeruginosa, also during the seven-year period, resistant strains of this bacterium have increased. In our study, the most frequent resistance was detected against cefoxitin (79.45%), cefotaxime (97.26%), and ciprofloxacin (75.34%) (approximately 70% of isolates). This prevalence rate was comparatively higher than the reports of Acar et al. [27], Liews et al. [28], and less than Choudhary et al. [29] studies.
During a study in Turkey, it was found that 55% of bacteria isolated from the hospital during 2007–2017 were resistant to many beta-lactamase antibiotics [27]. In this case, the prevalence of ESBL, MBL, AmpC, and KPC-producing P. aeruginosa was around 38.35%, 13.69%, 70.54%, and 16.43%, respectively, relatively lower than the figures reported by Ghasemian et al. [30] and Choudhary et al. [29]. One of the most important reasons for these differences is the type of weather in the regions, the pattern of antibiotic use, various mutations in bacteria, and the health of the communities. Although some issues, including the emergence of new antibiotics and the inter-relationship of bacteria, have contributed to the appearance of such a resistance.
There are many reports that show an association between antimicrobial resistance patterns and resistance genes and virulence factor genes in P. aeruginosa. Our results showed that there is a significant relationship between antibiotic resistance and pathogenesis. Nearly all MDR and XDR strains strongly formed biofilm (more than 95%), and about 80% of the MDR and XDR strains of P. aeruginosa produced phenazine, pyocyanin and protease enzymes. Hwang et al. [31] and Tahmasebi et al. [32] showed a significant correlation between antibiotic resistance and pathogenicity in different strains of P. aeruginosa.
Our findings also indicated a direct relationship between phenazine production and lasI/R expression. lasI/R was expressed in 11 isolates, which produced phenazine, protease, and siderophores. According to Abd El-Aziz et al. [33], lasI/R system plays a key role in the regulation and production of phenazine. There was a converse relationship in one isolate. It might be due to unidentified mutations in lasI/R. Proteases including lasA, lasB, and aprA are positively regulated by lasI/R QS system [5, 34], and as our result showed so, the proteases were detected in 10 isolates out of 11 MDR-XDR isolates. Furthermore, There was not an established association between MBL and lasI/R expression in our study except for MDR-XDR isolates, which were consistent with Goncalves et al., study [35]. In contrast to Al Dawodeyah et al., and Lee et al. studies, there was a high expression level of KPC in 10 isolates in accordance with lasI/R expression. These results may indicate that the KPC enzymes are responsible for carbapenem resistance in MBLs [36, 37].
Our results confirm a relationship between mexR expression and MDR-XDR strains and virulence enzyme. In addition, the expression of antibiotic resistance and virulence factors in XDR strains is higher than MDR strains. In non-MDR and non-XDR strains, the activity of virulence factor genes was increased, and the activity of antibiotic resistance genes was decreased. Although, it seems that more factors might influence the association, such as environmental elements. Efflux pump families as functional and metabolic pumps significantly act in drug resistance of P. aeruginosa. Resistance- nodulation- division (RND) family is commonly regulated via mexR, nalC, or nalD.
The initial regulator of RND efflux pumps-mexR can determine efflux function as a negative regulator, and Cabot et al. reported that the mexR mutant indicates a significant reduction in susceptibility to β-lactams and fluoroquinolones [11, 38]. As our findings demonstrated, the mexR expression level was increased in 11 isolates in accordance with lasR/I and, as a consequence, kpc expression raised in the XDR and MDR strains.
Therefore, QS systems could impressively interconnect virulence and efflux pump functions [39, 40]. Poole [41] and Al Dawodeyah et al. [36] reported that there was a significant relationship between the presence of beta-lactamase enzymes and pathogenicity of bacteria so that the MBL and AmpC possessing strains of P. aeruginosa were more pathogenic than antibiotic susceptible strains. In line with previously mentioned studies, there was a direct relationship between kpc and lasI/R, and also the expression of efflux pump genes was decreased.
In the present study, we have done MLST for 73 P. aeruginosa and found predominantly high-risk clones of ST108, ST217, ST260, ST274, ST501, ST1075, and ST3335 of MDR-XDR positive P. aeruginosa among different isolates (Fig. 3). We have also confirmed a high frequency of ST108 and ST3335 in virulent strains. Similar findings have been reported from Estonia more than 50% of beta-lactamase-producing P. aeruginosa belong to ST108 and ST260 [42]. Furthermore, the study also testified ST108, ST147, ST250, and ST260 in ESBL, NDM, and KPC-producing P. aeruginosa isolated from Poland and Germany [43, 44]. However, Vatansever et al. [45] demonstrated that ST235 is a high-risk P. aeruginosa producing clone, but to our knowledge, ST235 has not been identified in NDM positive P. aeruginosa so far.
The presence of sequence types belonging to ST260, ST274, and ST3335 beta-lactamase-producing isolates is also worrisome as their expansion has been reported previously from other countries [46, 47]. Identification of high-risk clone ST260, ST274, and ST3335 isolates in association with ESBL, MBL, and ampC genes is significant. ST260, ST274, and ST3335 have been associated with the successful dissemination of biofilm formation and virulence factors with high prevalence worldwide. P. aeruginosa ST274 encoding biofilm forming genes, which have been reported to cause serious infections in patients from Spain [47].
In order to define a comprehensive and accurate relationship between antibiotic and virulence, further studies are necessary to investigate a considerable number of isolates from different sources, various antibiotic resistance patterns, and virulence factors. In this regard, host-related factors should be taken into account. The low number of isolates and less diverse patterns of antibiotic and virulence factors could be acknowledged as some limitations of this research.
Conclusion
The correlation of virulence factors with antibiotic resistance is a complex topic which has yet to be explained. However, based on our results, there is a clear association between QS-regulated virulence and antibiotic resistance. Furthermore, in P. aeruginosa, QS tends to be able to regulate antibiotic tolerance and pathogenicity. In addition, the expression of genes involved in the pathogenesis of P. aeruginosa showed that MDR and XDR strains are more virulent than antibiotic-sensitive strains. Also, according to phylogenetic tree, dangerous strains were identified in ST108, ST260, and ST274, which indicates the high distribution of dangerous STs in Iran.
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- MDR:
-
Multidrug-resistant
- XDR:
-
Extensively drug-resistant
- ESBL:
-
Extended-spectrum β-lactamase
- MBL:
-
Metallo β-lactamase
- QS:
-
Quorum sensing
- CF:
-
Cystic fibrosis
- MEGA5:
-
Molecular evolutionary genetics analysis version 5
References
Ding F et al (2018) The Pseudomonas aeruginosa orphan quorum sensing signal receptor QscR regulates global quorum sensing gene expression by activating a single linked operon. mBio 9(4):e01274-18
Rutherford ST, Bassler BL (2012) Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harbor perspectives in medicine 2(11):a012427
Venturi V (2006) Regulation of quorum sensing in Pseudomonas. FEMS Microbiol Rev 30(2):274–291
Steindler L et al (2009) LasI/R and RhlI/R quorum sensing in a strain of Pseudomonas aeruginosa beneficial to plants. Appl Environ Microbiol 75(15):5131–5140
Lee J, Zhang L (2015) The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6(1):26–41
Lau GW et al (2004) The role of pyocyanin in Pseudomonas aeruginosa infection. Trends Mol Med 10(12):599–606
Hall S et al (2016) Cellular effects of pyocyanin, a secreted virulence factor of Pseudomonas aeruginosa. Toxins (Basel) 8(8):e236
Furiga A et al (2016) Impairment of Pseudomonas aeruginosa biofilm resistance to antibiotics by combining the drugs with a new quorum-sensing inhibitor. Antimicrob Agents Chemother 60(3):1676–1686
Bonomo RA et al (2017) Carbapenemase-producing organisms: a global scourge! Clin Infect Dis 66(8):1290–1297
Maseda H et al (2004) Enhancement of the mexAB-oprM efflux pump expression by a quorum-sensing autoinducer and its cancellation by a regulator, MexT, of the mexEF-oprN efflux pump operon in Pseudomonas aeruginosa. Antimicrob Agents Chemother 48(4):1320–1328
Cabot G et al (2016) Evolution of Pseudomonas aeruginosa antimicrobial resistance and fitness under low and high mutation rates. Antimicrob Agents Chemother 60(3):1767–1778
Bogiel T et al (2017) The prevalence of exoenzyme S gene in multidrug-sensitive and multidrug-resistant Pseudomonas aeruginosa clinical strains. Pol J Microbiol 66(4):427–431
Jorth P et al (2017) Evolved aztreonam resistance is multifactorial and can produce hypervirulence in Pseudomonas aeruginosa. MBio 8(5):e00517–e617
Mahon CR, Lehman DC, Manuselis G (2007) Textbook of Diagnostic microbiology. Saunders Elsevier, Amsterdam
El-Fouly MZ et al (2015) Biosynthesis of pyocyanin pigment by Pseudomonas aeruginosa. J Radiat Res Appl Sci 8(1):36–48
O'Toole GA (2011) Microtiter dish biofilm formation assay. J Vis Exp 47:2437
Ołdak E, Trafny EA (2005) Secretion of proteases by Pseudomonas aeruginosa biofilms exposed to ciprofloxacin. Antimicrob Agents Chemother 49(8):3281–3288
Fazeli N, Momtaz H (2014) Virulence gene profiles of multidrug-resistant Pseudomonas aeruginosa isolated from Iranian hospital infections. Iran Red Crescent Med J 16(10):e15722
Fazeli H et al (2014) Molecular epidemiology and mechanisms of antimicrobial resistance in Pseudomonas aeruginosa isolates causing burn wound infection in Iran. J Chemother 26(4):222–228
Lima JLDC et al (2018) Biofilm production by clinical isolates of Pseudomonas aeruginosa and structural changes in LasR protein of isolates non biofilm-producing. Braz J Infec Dis 22(2):129–136
Quale J et al (2006) Interplay of efflux system, ampC, and oprD expression in carbapenem resistance of Pseudomonas aeruginosa clinical isolates. Antimicrob Agents Chemother 50(5):1633–1641
Uz Zaman T et al (2014) Multi-drug carbapenem-resistant Klebsiella pneumoniae infection carrying the OXA-48 gene and showing variations in outer membrane protein 36 causing an outbreak in a tertiary care hospital in Riyadh Saudi, Arabia. Int J Infect Dis 28:186–192
Geyer CN, Hanson ND (2014) Multiplex high-resolution melting analysis as a diagnostic tool for detection of plasmid-mediated AmpC β-lactamase genes. J Clin Microbiol 52(4):1262–1265
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9):e45
Vernez I et al (2005) Population genetic analysis of Pseudomonas aeruginosa using multilocus sequence typing. FEMS Immunol Med Microbiol 43(1):29–35
Dou Y et al (2017) Pseudomonas aeruginosa prevalence, antibiotic resistance and antimicrobial use in Chinese burn wards from 2007 to 2014. J Int Med Res 45(3):1124–1137
Acar A et al (2019) Pooled prevalence and trends of antimicrobial resistance in Pseudomonas aeruginosa clinical isolates over the past 10-years in Turkey: A meta-analysis. J Glob Antimicrob Resist 18:64–70
Liew SM et al (2019) Antimicrobial susceptibility and virulence genes of clinical and environmental isolates of Pseudomonas aeruginosa. PeerJ 7:e6217–e6217
Choudhary V, Pal N, Hooja S (2019) Prevalence and antibiotic resistance pattern of Metallo-β-lactamase-producing Pseudomonas aeruginosa isolates from clinical specimens in a tertiary care hospital. J Mahatma Gandhi Inst Med Sci 24(1):19–22
Ghasemian A et al (2018) Prevalence of clinically isolated metallo-beta-lactamase-producing Pseudomonas aeruginosa, coding genes, and possible risk factors in Iran. Iran J Pathol 13(1):1–9
Hwang W, Yoon SS (2019) Virulence characteristics and an action mode of antibiotic resistance in multidrug-resistant Pseudomonas aeruginosa. Sci Rep 9(1):487–487
Tahmasebi H et al (2019) Role and function of KPC and MBL enzymes in increasing the pathogenicity of Pseudomonas aeruginosa isolated from burn wounds. J Babol Univ Med Sci 21(1):127–134
El-Aziz NKA, El-Hamid MIA, El-Naenaeey E-SY (2018) A complex hierarchical quorum-sensing circuitry modulates phenazine gene expression in Pseudomonas aeruginosa. J Infect Dev Ctries 11(12):919–925
Hwang S et al (2016) Network-assisted investigation of virulence and antibiotic-resistance systems in Pseudomonas aeruginosa. Sci Rep 6:26223
Rossi Gonçalves I et al (2017) Carbapenem-resistant Pseudomonas aeruginosa: association with virulence genes and biofilm formation. Braz J Microbiol 48(2):211–217
Al Dawodeyah HY et al (2018) Antimicrobial resistance and putative virulence genes of Pseudomonas aeruginosa isolates from patients with respiratory tract infection. Germs 8(1):31–40
Lee J-Y, Peck KR, Ko KS (2013) Selective advantages of two major clones of carbapenem-resistant Pseudomonas aeruginosa isolates (CC235 and CC641) from Korea: antimicrobial resistance, virulence and biofilm-forming activity. J Med Microbiol 62(7):1015–1024
Beceiro A, Tomás M, Bou G (2013) Antimicrobial resistance and virulence: a successful or deleterious association in the bacterial world? Clin Microbiol Rev 26(2):185–230
Rampioni G et al (2017) Effect of efflux pump inhibition on Pseudomonas aeruginosa transcriptome and virulence. Sci Rep 7(1):11392–11392
Alcalde-Rico M et al (2016) Multidrug efflux pumps at the crossroad between antibiotic resistance and bacterial virulence. Front Microbiol 7:1483
Poole K (2011) Pseudomonas aeruginosa: resistance to the max. Front Microbiol 2:65
Telling K et al (2018) Multidrug resistant Pseudomonas aeruginosa in Estonian hospitals. BMC Infect Dis 18(1):513
Guzvinec M et al (2014) Sequence types 235, 111, and 132 predominate among multidrug-resistant Pseudomonas aeruginosa clinical isolates in Croatia. Antimicrob Agents Chemother 58(10):6277–6283
Boehmer T et al (2018) Phenotypic characterization and whole genome analysis of extended-spectrum beta-lactamase-producing bacteria isolated from dogs in Germany. PLoS ONE 13(10):e0206252
Vatansever C et al (2020) Co-existence of OXA-48 and NDM-1 in colistin resistant Pseudomonas aeruginosa ST235. Emerg Microbes Infect 9(1):152–154
Chen SH et al (2014) Multilocus sequencing typing of Pseudomonas aeruginosa isolates and analysis of potential pathogenicity of typical genotype strains from occupational oxyhelium saturation divers. Undersea Hyperb Med 41(2):135–141
Ocampo-Sosa AA et al (2015) Draft genome sequence of the quorum-sensing and biofilm-producing Pseudomonas aeruginosa strain Pae221, belonging to the epidemic high-risk clone sequence type 274. Genome Announc 3(1):e01343–e1414
Acknowledgments
The authors of this article are grateful to Hamadan University of Medical Sciences for their financial support.
Funding
This work was supported by the Research Centre of Hamadan University of Medical Sciences on the Grant Number 9510075755. This funding's devoted just to purchasing materials used in our study.
Author information
Authors and Affiliations
Contributions
SD performed microbiological and molecular tests and wrote the manuscript. MRA supervised all of the stages of designing the study, conducting the research, and writing the manuscript. MRP, MYA and SSA play a role in Project Administration.
Corresponding author
Ethics declarations
Conflicts of interest
All authors declare that they have no conflict of interest.
Ethics approval
This study was approved by the Ethics Committee of Hamadan University of Medical Sciences (Code No: IR.UMSHA.REC.1395402.) about the consent to participate is not applicable.
Consent for publication
Not Applicable.
Research involving Human Participants and/or Animals
In this study, there isn’t any research involving Human participant or animals.
Informed consent
Not Applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dehbashi, S., Pourmand, M.R., Alikhani, M.Y. et al. Coordination of las regulated virulence factors with Multidrug-Resistant and extensively drug-resistant in superbug strains of P. aeruginosa. Mol Biol Rep 47, 4131–4143 (2020). https://doi.org/10.1007/s11033-020-05559-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05559-4