Abstract
Many studies have been performed to identify regulatory circuit underlying plant stress tolerance. However, the reliability of some findings has been criticized because of exclusive use of stress sensitive plant species such as Arabidopsis thaliana. Sensitive plant species often harbor narrow defensive mechanisms and have relatively low capacity for adaptive responses. Therefore, it is useful to employ tolerant model plants, such as Eutrema salsugineum, to provide comprehensive insights into various mechanisms involved in response to abiotic stresses. In this study, comparative transcriptome and regulatory network analysis of stress-sensitive (A. thaliana) and -tolerant (E. salsugineum) model plants uncovered regulatory hierarchies underlying response to abiotic stresses and suggested the transcription factor genes, MYB44 and VIP1 as the candidate hub genes to perform molecular analyses on their Brassica napus homologs, BnMYB44 and BnVIP1. The full-length coding sequence of BnMYB44 and BnVIP1 with 891 and 969 bp long were cloned and sequenced. They shared high similarity with their counterparts in other plants at nucleotide and amino acid levels. The expression patterns of BnMYB44 and BnVIP1 genes of the two B. napus cultivars under drought and salt stress conditions coupled with the data obtained from the physiological measurements as well as analysis of the BnMYB44 and BnVIP1 promoters suggested that BnMYB44 and BnVIP1 genes may contribute to responses to drought and salt stresses in B. napus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Crop plants frequently encounter adverse environmental factors that limit their growth and productivity. Drought and salinity stresses are known as the most crucial environmental constraints for agricultural production worldwide [1, 2]. To cope with such stresses, plants reprogram their genome activities which essentially cause changes in expression and function of genes [3,4,5]. Protein families of transcription factors (TFs), protein kinases (PKs) and protein phosphatases (PPs) are the core regulatory components underlying downstream functions involved in plant stress responses [5,6,7,8]. For example, expression change of only one regulatory gene, namely histidine kinase 1 (ATHK1) may lead to alter activity of more than six-hundred downstream genes in A. thaliana [9]. Thus, understanding the roles of regulatory components will be of great benefit to improve crop stress tolerance.
Several studies have been performed to identify important regulatory genes involved in plant stress tolerance. However, the reliability of some findings has been criticized because of exclusive use of stress sensitive plant species such as A. thaliana [10]. In contrast to stress tolerant plant species, sensitive plant species often harbor narrow defensive mechanisms and have relatively low capacity for adaptive responses [10, 2]. Therefore, it is useful to employ model plants that can provide comprehensive insights into various mechanisms conferring tolerance to abiotic stresses.
Eutrema salsugineum (also known as Thellungiella halophila) is widely used as a model plant to study abiotic stress responses and tolerance [11,12,13,14,15]. This stress tolerant species is a close relative of Arabidopsis, and its genes show high homology to those of Arabidopsis at the coding sequence level [16, 17]. Interestingly, E. salsugineum does not recruit complex adaptation mechanisms such as dramatic morphological changes, and its stress tolerance is simply derived from enhancement of mechanisms involved in controlling water status, metabolic rates, ion homeostasis and so on, which are also present in stress sensitive species [11, 16, 18]. A large number of E. salsugineum homologs of Arabidopsis stress-inducible genes are highly expressed under normal growth conditions [19, 20]. The availability of whole-genome sequence and transcriptome data of E. salsugineum [21] along with various Arabidopsis genomic resources has opened a new vista for a better understanding of genome function under abiotic stress conditions in Brassicaceae family, including the agronomically important Brassica genus.
Rapeseed (B. napus L.) is a globally important oilseed crop. Rapeseed production is often limited by abiotic stresses in arid and semi-arid regions. Identification of regulatory genes involved in response mechanisms to abiotic stresses will facilitate development of rapeseed cultivars with improved stress tolerance [22,23,24]. B. napus is an obvious target crop to exploit comparative genomics data between Arabidopsis and E. salsugineum. Several abiotic stress responsive genes have been identified in B. napus based on homology with Arabidopsis [25,26,27,28,29]; however, their significance in relation to other co-expressed genes in the context of a network system have remained unclear. Comparative transcriptome analysis of Arabidopsis and E. salsugineum has the potential to uncover regulatory hierarchies underlying response and tolerance to abiotic stresses and suggests candidate hub genes to be characterized in B. napus.
This study aimed to use comparative transcriptomics and experimental approaches to identify and characterize key regulatory genes that respond to drought and salt tolerance in B. napus. A gene regulatory network was constructed based on comparative analysis of the microarray data from Arabidopsis thaliana and E. salsugineum. The transcription factor genes namely MYB44 and VIP1 were selected from the regulatory network to perform molecular analyses on their B. napus homologs, BnMYB44 and BnVIP1. The coding sequence of BnMYB44 and BnVIP1 were isolated, cloned and sequenced. Afterwards, the expressions of BnMYB44 and BnVIP1 genes in response to drought and salt stresses were analyzed in two B. napus cultivars with different levels of tolerance to drought and salt. Finally, the expression data coupled with the measurements of physiological characteristics as well as the results of promoter analysis of the genes of interest provided an insight into the role and significance of BnMYB44 and BnVIP1 in response to drought and salt stresses in B. napus.
Materials and methods
Comparative analysis of microarray data from Arabidopsis thaliana and Eutrema salsugineum
The microarray raw data from A. thaliana and E. salsugineum were retrieved from European Bioinformatics Institute (EBI) array expressed sub-division (http://www.ebi.ac.uk/arrayexpress/experiments/E-GEOD-19740/). Two independent leaf samples per species collected at the growth stage 6.10 (i.e. about 35.9 days after sowing in which, 10% flowers to be produced are open) according to Boyes et al. [30] were used to perform two color microarray assay.
The microarray expression data were normalized by Robust Multi-array Averaging (RMA) algorithm [31] using FlexArray software 1.6.1 [32]. In order to identify Differentially Expressed Genes (DEGs) between leaves of A. thaliana and E. salsugineum, a Bayesian t test was applied followed by the false discovery rate (FDR, q-value) method [33] to correct p-values for multiple testing. All the DEGs at q-value ≤ 0.05 were subjected to further analysis to specify those genes encoding regulatory proteins. To do so, first, a local database was made comprising Arabidopsis transcription factors collected from PlantTFact (http://plantgrn.noble.org/PlantTFcat/) [34] as well as a complete set of Arabidopsis protein kinases and protein phosphatases obtained from Plant Protein Phosphorylation Database (P3DB) (http://www.p3db.org/) [35]. The DEGs were then searched against the local database to specify those genes encoding TFs, PKs and PPs.
The regulatory genes were subjected to Gene Set Enrichment (GSE) analysis to extract significant sub-networks based on Fisher exact test (p-values ≤ 0.05), using Pathway Studio webtool [36, 37], as previously described [38,39,40]. After that, all the significant sub-networks were merged by “union selected pathways” of Pathway studio software 7 to construct the hierarchical regulatory network. In order to delineate significance of the DEGs in the regulatory network, topological features including betweenness and closeness centrality as well as edge betweenness centrality [41], were determined for the network using Cytoscape software 3.1.1 [42, 43].
Cloning and sequencing of B. napus MYB44 and VIP1 genes
Total RNA was extracted from leaf samples of B. napus cv. SLM046 using column RNA isolation kit (DENA Zist Asia, Iran) according to the manufacturer’s protocol. RNA quantification and integrity were double-checked by agarose gel electrophoresis and spectrophotometry. DNA contamination was removed from RNA using Fermantas DNaseІ treatment (Thermo scientific, USA). First strand cDNA was synthesized from total RNA by Viva 2-steps RT-PCR kit (Vivantis, Malaysia). Since the genome sequence of Brassica napus was not publicly available at the time of primer design, the primers were designed based on Brassica rapa orthologs of MYB44 and VIP1 identified by searching B. rapa genome sequence (http://www.phytozome.net) against Arabidopsis MYB44 and VIP1 (Table 1). The two pairs of primers were used to clone two fragments with overlapping region to cover up the full-length coding sequence of BnVIP1.
PCR reaction was carried out as follows: initial denaturation at 94 °C for 5 min followed by 35 cycles of 1 min at 94 °C, 1 min at 56.5 °C and 1 min at 72 °C, and a final extension at 72 °C for 10 min. The PCR products were analyzed by agarose gel electrophoresis (1% w/v). Subsequently, the PCR products were purified and cloned into pTZ57R/T vector (InsTAclone PCR Cloning kit, Fermentas, USA). Transformation of competent cells of E. coli strain DH5α was performed by electroporation. The cloned amplicons were finally sequenced.
The coding sequences were subjected to similarity searches against non-redundant nucleotide and protein database using BLASTN and BLASTX (E-value ≤ 10−5), respectively, using CLC Genomics workbench 3.6 software. Also, the structure and synteny of BnMYB44 and BnVIP1 were assigned using Blat tool of GenoScope (http://www.genoscope.cns.fr/spip/), the B. napus genome sequence database. Moreover, protein features of the deduced amino acid sequences of BnMYB44 and BnVIP1 genes were specified using CLC Genomics Workbench software 3.6.
Plant materials, growth conditions and stress treatments
The B. napus cultivars, SLM046 and Zarfam, with different responses to abiotic stresses [25, 44,45,46], were used to perform drought and salt stress experiments. The drought experiment was conducted in factorial completely randomized design with three replicates. Plants were grown in plastic pots filled with sterilized sandy-clay soil under growth room conditions (16 h light/8 h dark; 24 °C day/8 °C night). Plants with five true leaves were subjected to drought stress by withholding water to reach soil moisture to 40% of field capacity (FC). Sampling was carried out under non-stress along with three drought stress treatments including when soil moisture reached to 40% of FC as well as maintaining that for 6 and 24 h. The non-stress condition was performed by regularly irrigating plants to keep soil moisture around FC during the experiment.
The salt stress experiment was also carried out in factorial completely randomized design with three replicates. The surface-sterilized seeds of the B. napus cultivars, SLM046 and Zarfam, were sown in sandy soil under a long-day photoperiod (16 h light/8 h dark) with 24 and 8 °C day and night temperature, respectively. 10-day-old seedlings were then transferred to a hydroponic culture system supplemented with half-strength Hoagland solution (pH ≈ 6.0) under growth room conditions. Plants with five true leaves were exposed to salt stress by adding 150 mM NaCl solution to the medium. Sampling was carried out under non-stress as well as three salt stress treatments including 3 h, 24 h and 7 days after adding salt solution.
Physiological measurements
In order to evaluate drought tolerance of the B. napus cultivars, a number of physiological characteristics were measured under all the drought stress conditions. Relative Water Content (RWC) was determined according to the method of Slatyer and Shmueli [47]. Proline content was measured using the protocol described by Bates et al. [48]. Cell membrane damage caused by drought stress was estimated by measuring Electrolyte leakage (EL) following the method described by Sairam and Srivastava [49]. In addition, the concentration of Na+ and K+ ions in leaf and root tissues of the B. napus cultivars were determined under all the salt stress conditions according to Munns et al. [50] protocol in order to obtain K+/Na+ ratio as an indicator for salt tolerance. The data were subjected to analysis of variance followed by comparison of means using Least Significant Difference (LSD) test at p-value ≤ 0.05.
Expression profiling of BnMYB44 and BnVIP1 under stress conditions
Real time quantitative RT-PCR (qRT-PCR) was used to detect expression levels of BnMYB44 and BnVIP1 in leaf and root tissues of the cultivars during drought and salt stresses. Total RNA extraction and cDNA synthesis were done as described earlier for isolating the genes. The specific primers, listed in Table 1, were designed for each gene. qRT-PCR was performed on a Bioer thermal cycler (Bioer Technology Co., Ltd., China) by using RealQ PCR Master Mix kit (Ampliqon, Denmark) according to the manufacturer’s instructions. qRT-PCR reactions were also performed for the reference gene, B. napus Elongation Factor 1α (Bnelf1α; GenBank Accession number: FJ529181) for normalization of expression data. Three independent biological samples were used to do qRT-PCR. Each reaction was also repeated two times as technical replicates. The relative expression levels of the target genes were calculated using the 2−ΔΔCT method [51].
Analysis of BnMYB44 and BnVIP1 promoters
1500 bp upstream of BnMYB44 and BnVIP1 genes (including 5′ UTR), as putative promoter regions were obtained from GenoScope (http://www.genoscope.cns.fr/spip/). The promoter sequences were searched for cis regulatory elements (CREs) using PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/; [52]). Then, a statistical test for pairwise comparison of the promoters of BnMYB44 and BnVIP1 genes was performed according to the method of Shamloo-Dashtpagerdi et al. [53] to identify CREs with significantly different number of occurrences between the promoters, which may contribute to different expression patterns of the genes.
Results
Comparative analysis of microarray data
The microarray data obtained from leaf samples of Arabidopsis and E. salsugineum were analyzed to identify differentially expressed regulatory genes. Considering q-value ≤ 0.05, we found 3706 genes that their expressions significantly differed between leaves of Arabidopsis and E. salsugineum. Of those, 564 genes fell into the three major groups of regulatory genes encoding transcription factors, protein kinases and protein phosphatases (Supplementary 1). 307 transcription factors belonging to 50 different families were found with different magnitudes and directions of expression changes indicating that there is a great difference in the regulatory functions of Arabidopsis and E. salsugineum transcriptomes. bZip family with 31 members was the largest group of transcription factors (Supplementary 1). In addition, 226 protein kinase and 31 protein phosphatase genes belonging to 52 and 22 families, respectively, showed significantly different expressions between Arabidopsis and E. salsugineum. The receptor like cytoplasmic kinase VII family with 18 members was the largest protein kinase family. Among protein phosphatase families, the protein phosphatase 2C family was the largest group containing eight members (Supplementary 1).
Regulatory network analysis
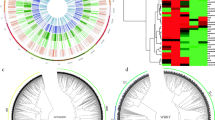
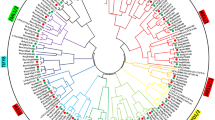
The hierarchal gene regulatory network (Fig. 1a) was predicted using the differentially expressed regulatory genes in order to identify major stress-responsive regulatory genes. The network clearly identified the regulatory genes involved in the drought and osmotic stress pathways. The network represented that signaling pathways of ABA and auxin hormones as well as Ca2+ ion govern most of the abiotic stress responses. The protein kinase gene, RPK1, and the transcription factor gene, VIP1 were positioned at the forefront of the regulatory genes.
Regulatory network analysis. a The hierarchical regulatory network based on the differentially expressed regulatory genes between Arabidopsis thaliana and Eutrema salsugineum. b The centrality analysis of the gene network: the nodes with bigger size indicate higher betweenness centrality. Red color represents the highest closeness centrality, while green color indicates the lowest value of closeness centrality. The edges with more thickness represent high values of edge betweenness centrality. The positions of MYB44 and VIP1 have also been shown. (Color figure online)
Topology analysis revealed that the network contained a total number of 81 nodes and 261 edges (supplementary 2). Among different components of the network, ABA, auxins, osmotic stress and drought showed the largest number of connectivity (more than 20 connections). Also, transcription factor MYB44 had the largest number of connectivity (19 connections) among the genes. A summary of topology analysis of the regulatory network was illustrated in Fig. 2b. The nodes with bigger size indicated higher betweenness centrality and had greater influence. Betweenness is a measure of the relevance of a given node obtained by counting the number of shortest paths going through it to quantify the importance of a node in a network [41, 42]. In addition, the closeness centrality is scaled using gradient color-coding system in which, red color represents the highest closeness centrality, while green color indicates the lowest value of closeness centrality. The closeness of a node is defined as the inverse of the average distance from all other nodes [41, 54]. The edges with more thickness represent high values of edge betweenness centrality. Edge betweenness is defined as the number of shortest paths between pairs of nodes that run through that edge and is known as a standard measure of the impact of a node or a linkage in a network [41, 54]. Accordingly, topology analysis uncovered that ABA, auxin and NaCl nodes obtained high values for the centrality indices. Of those, ABA with the highest values was presumed as the main node in the network (Fig. 1b). Moreover, among the functional classes within the network; root development, stomatal movement, cell death and root morphogenesis had the maximum values for the centrality indices, respectively, illustrating the main strategies to cope with abiotic stresses based on the network’s output. The results also revealed that the transcription factor MYB44 achieved the highest centrality values implying its pivotal role in the regulatory network. As a result, MYB44 and its direct regulator, the transcription factor gene VIP1, were selected to perform molecular cloning and characterization of their B. napus homologs.
Cloning and sequence analysis of BnMYB44 and BnVIP1 genes
BnMYB44 (GenBank Accession Number: KT071672) was 891 bp in length and encoded a deduced protein of 296 amino acid residues with a predicted molecular weight of 32.457 kDa and isoelectric point of 9.02. BnMYB44 protein contained two conserved MYB-like DNA binding domains, identified at positions 6th -52nd and 58th -103rd amino acid residues, indicating that BnMYB44 protein is a member of MYB R2R3 subfamily whose members have two conserved DNA-binding domains [55]. The amino acid sequence and some structural features of BnMYB44 protein were shown in Fig. 2a.
Blast search showed that the BnMYB44 gene had high homology (more than 80%) with other plant MYB44 genes at nucleotide and amino acid levels. The structural analysis revealed that the BnMYB44 gene was located on chromosome 7 of B. napus A genome and consisted of one exon, which resembled the genomic structure of AtMYB44 gene [56]. Synteny analysis showed that orthologs of the BnMYB44 in B. napus progenitor species, B. rapa and B. oleracea, were located on the chromosome 7 of their corresponding genomes.
The two isolated fragments of BnVIP1 gene were assembled resulting in an open reading frame of 969 bp (GenBank Accession Number: KR537434). The deduced protein consisted of 322 amino acids with molecular weight of 35.988 kDa and isoelectric point of 7.89. One bZIP domain was detected within the protein of BnVIP1 (from 187th to 235th amino acid residue) implying that BnVIP1 is a member of the bZIP transcription factor family (Fig. 2b). BnVIP1 gene and its homologs from other members of Brassicaseae family including E. salsugineum, A. lyrata, Camelina sativa and (A) thaliana shared more than 80% nucleotide sequence homology in the coding regions. At the amino acid level, BnVIP1 protein showed 88, 83, 75 and 73% identity with its homologs in (B) rapa, E. salsugineum, A. lyrata and (A) thaliana, respectively. The results also revealed that BnVIP1 gene was located on the chromosome 8 of (B) napus C genome, originated from its progenitor, B. oleracea. The genomic structure of BnVIP1, similar to its Arabidopsis counterpart [57], contained four exons with different lengths. Moreover, orthologs of the BnVIP1 in B. rapa and B. oleracea were located on the chromosome 8 of their corresponding genomes.
Physiological responses of B. napus to drought stress
A number of physiological characteristics were measured in the B. napus cultivars, SLM046 and Zarfam, under non-stress and drought stress conditions. The results showed that drought stress significantly affected all the physiological characteristics (Table 2). While no variation was observed between the cultivars for the physiological characteristics under non-stress conditions, the cultivars showed significantly different physiological responses to drought stress treatments (Table 2). RWC significantly decreased during drought stress conditions, however the cultivars exhibited different rates of reduction of RWC under more severe drought stress treatments (D2 and D3). SLM046 (stress-tolerant cultivar) had significantly higher amount of RWC than Zarfam, under such conditions. High accumulation of Proline content occurred during drought stress conditions, although there were different levels of proline accumulation between the cultivars. SLM046 showed significantly higher proline contents compared to Zarfam under more severe drought stress treatments. Cell membrane damage, as measured by EL, significantly increased under drought stress conditions. Zarfam showed much higher amount of EL than SLM046 indicating higher degree of cell membrane injury during drought stress (Table 2). Altogether, evaluation of the physiological characteristics revealed distinct responses of the B. napus cultivars to drought stress, as SLM046 was more efficiently capable of coping with drought stress effects and consequently showed more tolerance to drought compared to Zarfam. This finding provided physiological evidence to explain possible variation between the cultivars in the expression patterns of BnMYB44 and BnVIP1 genes in response to drought stress.
Expression patterns of BnMYB44 and BnVIP1 under drought stress
The expression patterns of BnMYB44 and BnVIP1 in response to drought stress were analyzed by qRT-PCR in the time course studies. Different expression profiles were observed for the genes not only between the cultivars but also between leaf and root tissues during drought stress period. In leaves of SLM046, BnMYB44 expression was more quickly induced in response to drought stress and also showed significantly higher level of increase compared to that of the other B. napus cultivar (Fig. 3a). In roots of SLM046, BnMYB44 expression was significantly upregulated 6 h after maintaining soil moisture at 40% of field capacity (D2) and then declined to the level with no significant difference with its transcript level under non-stress conditions (Fig. 3b). On the other hand, drought stress caused a significant downregulation in the expression of BnMYB44 in root tissue of Zarfam.
The expression of BnMYB44 and BnVIP1 in response to drought stresses. a BnMYB44 in leaves. b BnMYB44 in roots. c BnVIP1 in leaves. d BnVIP1 in roots. The expression was determined by Real time quantitative RT-PCR. Three drought stress treatments include the time that soil moisture reached to 40% of field capacity (D1) as well as maintaining that for 6 h (D2) and 24 h (D3). Different letters indicate significant differences according to Duncan’s test (P < 0.01)
The expression of BnVIP1 gene in leaves of SLM046, rapidly and significantly responded to drought stress and showed the maximum level of transcript accumulation at the first drought stress treatment (D1) (Fig. 3c). On the contrary, no induction was detected for the expression of BnVIP1 during the drought stress period in leaves of Zarfam. The highest level of BnVIP1 expression in roots of both cultivars was observed 6 h after maintaining soil moisture at 40% of field capacity (D2), however, Zarfam had a lower level of transcript accumulation (Fig. 3d).
Ion homeostasis status under salt stress
Salt stress had a significant effect on sodium and potassium contents of leaves and roots of the B. napus cultivars, SLM046 and Zarfam. Consequently, K+/Na+ ratio, as a reliable indicator for salt tolerance in Brassica species [58], significantly decreased due to salt stress (Table 3). Significant variation was also observed between the cultivars for K+/Na+ ratio indicating their different levels of salt tolerance (Table 3). SLM046 showed higher leaf K+/Na+ ratios than those of Zarfam under salt stress treatments. While Zarfam showed a higher level of root K+/Na+ ratio than that of SLM046 under non-stress conditions, salt stress caused a dramatic fall in root K+/Na+ ratios of Zarfam which eventually resulted in slightly higher root K+/Na+ ratios for SLM046 under 3 and 6 h after salt stress induction. Overall, SLM046 exhibited more effective ion homeostasis, possibly associated with more tolerance to salt stress compared to Zarfam. This result helped to find out whether the expression patterns of BnMYB44 and BnVIP1 genes under salt stress may be associated with salt tolerance of the cultivars.
Expression patterns of BnMYB44 and BnVIP1 under salt stress
The expression patterns of BnMYB44 and BnVIP1 in response to salt stress were examined by qRT-PCR in leaves and roots of the B. napus cultivars. The results revealed significant differences for the expression trends of the genes between the cultivars and also between leaf and root tissues (Fig. 4). In leaves of SLM046 (the salt tolerant cultivar), BnMYB44 expression rapidly and sharply increased and reached its maximum level at 3 h following salt stress (S1) in which the transcript accumulation of BnMYB44 was almost seven times higher than that of non-stress conditions (Fig. 4a). BnMYB44 expression was then significantly downregulated and showed the minimum transcript level at 24 h (S2) and 7 days (S3) after salt stress application. Similar trend was detected for BnMYB44 expression in roots of SLM046, as BnMYB44 expression exhibited a tenfold increase at S1 compared to non-stress conditions, and then significantly reduced during S2 and S3 treatments (Fig. 4b). On the other hand, BnMYB44 expression in leaves of Zarfam showed no induction during salt stress period. Moreover, salt stress caused a significant reduction in BnMYB44 expression in roots of Zarfam, although no significant variation was observed for its expression across the salt stress treatments (Fig. 4b).
The expression of BnMYB44 and BnVIP1 in response to salt stresses. a BnMYB44 in leaves. b BnMYB44 in roots. c BnVIP1 in leaves. d BnVIP1 in roots. The expression was determined by Real time quantitative RT-PCR. Three salt stress treatments include 3 h (S1), 24 h (S2) and 7 days (S3) after adding salt solution. Different letters indicate significant differences according to Duncan’s test (P < 0.01)
BnVIP1 expression was significantly upregulated in leaves of SLM046 at 3 h post salt stress and then declined to finally reach the level with no significant difference of that under non-stress conditions at 7 days post salt stress (Fig. 4c). On the contrary, BnVIP1 expression was not induced in leaves of Zarfam by salt stress and exhibited no significant changes during salt stress (Fig. 4c). BnVIP1 expression generally showed a declining trend in roots of both cultivars during salt stress period (Fig. 4d). No significant difference was detected for BnVIP1 expressions between roots of the cultivars in each of the salt stress treatments (Fig. 4d).
Analysis of BnMYB44 and BnVIP1 promoters
The promoter regions of BnMYB44 and BnVIP1 genes were analyzed to eventually identify CREs relevant to stress responsiveness. The total number of 136 and 94 CREs from 37 to 26 different types were identified within BnMYB44 and BnVIP1 promoter sequences, respectively (Supplementary 3). The results showed that BnMYB44 promoter contained several hormone-responsive CREs associated with activity and regulation of ABA, gibberellin, salicylic acid and auxin suggesting that BnMYB44 may be involved in hormone crosstalk. In addition, several biotic and abiotic stress responsive CREs involved in anaerobic, fungal, heat, low temperature and wounding stresses were found in BnMYB44 promoter indicating its possible contribution in response to environmental changes (Supplementary 3). Interestingly, one VIP1 response element (VRE) motif (TTGACC) was also detected within the BnMYB44 promoter.
In contrast to BnMYB44, BnVIP1 promoter contained few hormone related CREs (Supplementary 3). Also, there were a number of stress responsive CREs involved in fungal and abiotic stresses within BnVIP1 promoter, however they were fewer than the ones within BnMYB44 promoter (Supplementary 3). This is in accordance with the level of responsiveness of BnMYB44 and BnVIP1 to drought and salt stresses. The transcript accumulation of BnMYB44 and BnVIP1 genes under drought and salt stress conditions revealed that BnMYB44 gene was significantly more responsive to these stresses than BnVIP1 gene.
According to the statistical pairwise test [53], eleven CREs with significantly different numbers between the two promoters of BnMYB44 and BnVIP1 genes were identified (Table 4). The six significant CREs, involved in various biological processes, namely BOXII, G-BOX, AS-2-BOX, E-box, GT1-MOTIF and ABRE were more abundant in the promoter of BnMYB44. On the other hand, the CREs including CAAT-BOX, HSE, GAG-MOTIF, CGTCA-MOTIF and TGACG-MOTIF had higher number of occurrences in the BnVIP1 promoter. The result of the pairwise comparative promoter analysis coupled with the expression data of BnMYB44 and BnVIP1 genes will be further discussed in order to explain biological significance of some of the differentially significant CREs.
Discussion
Transcriptomics based networks have been widely used to elucidate plant responses to abiotic stresses [8, 59,60,61,62]. In this study, the regulatory network derived from the microarray data from Arabidopsis and E. salsugineum, shed light on the significance of some regulatory components involved in metabolisms and signaling of ABA and auxin hormones, root development and morphogenesis, stomatal movements, oxidative stress, cell death and Ca2+ signaling. Based on the topological features of the network, MYB44 transcription factor was known as the network bottleneck and the network central protein. MYB44 and its upstream regulator gene, VIP1 had significantly higher levels of expressions in E. salsugineum compared to Arabidopsis. Previous findings have indicated that E. salsugineum homologs of Arabidopsis stress-responsive genes are highly expressed under normal growth conditions [19, 20]. Arabidopsis MYB44 is a R2R3-MYB transcription factor gene expressed in different plant tissues and after various treatments [63]. Arabidopsis plants overexpressing MYB44 gene showed tolerance to abiotic stresses [64]. Moreover, transgenic rice plants harboring AtMYB44 gene exhibited significantly enhanced tolerance to drought stress [65]. As shown in the regulatory network, VIP1 transcription factor was located upstream of MYB44 (Fig. 1a). Arabidopsis VIP1 is a bZIP protein known for its role in nuclear import of T-DNA during Agrobacterium-mediated plant transformation [57]. Tsugama et al. [66] suggested that VIP1 functions as a regulator of osmosensory signaling in Arabidopsis through up-regulating of osmotic stress responsive genes. The phosphorylated form of VIP1 translocates from cytoplasm to nucleus in a stress dependent manner and regulates target genes such as MYB44 [67, 68].
This study has provided evidence indicating that BnMYB44 and BnVIP1 genes may be involved in adaptive responses to abiotic stresses in B. napus, however it is essential to perform overexpression and knockout experiments to resolve the direct involvement of these genes in tolerance mechanisms to abiotic stresses. The expression analysis showed that BnVIP1 was quickly induced under drought stress in the drought tolerant B. napus cultivar (SLM046). Accordingly, a strong and quick upregulation of BnMYB44 expression was observed in SLM046 compared to the drought sensitive B. napus cultivar (Zarfam). Arabidopsis Plants overexpressing VIP1 also accumulate high levels of MYB44 transcripts [68]. VRE is known as the specific binding site of Arabidopsis VIP1 protein and exists within the promoters of VIP1 target genes such as Trxh8 and MYB44 [68]. The existence of one VRE motif within the BnMYB44 promoter suggested that BnMYB44 transcription may be influenced by BnVIP1.
The different expression patterns of BnVIP1 and BnMYB44 genes between the cultivars during drought stress were somewhat associated with the physiological responses of the cultivars. The results indicated that SLM046 exhibited higher cell membrane stability and leaf relative water content under drought stress conditions in comparison with Zarfam. High relative water content and proline over-accumulation have been considered as the beneficial drought and salinity tolerance indicators in B. napus [69, 70]. This was evidenced by higher level of proline concentration in leaves of the drought tolerant cultivar during drought stress compared to the drought sensitive cultivar. Osmotic stress-induced proline accumulation is partially modulated by an ABA signaling pathway [71]. Furthermore, a meaningful relationship has been found between leaf relative water content and stomatal closure in various plant species [72]. stomatal closure in response to abiotic stress conditions is a powerful strategy to reduce water loss and to adjust water status in plants [73]. Stomatal closure is mainly under control of an ABA-positive signaling gene, SnRK2 (SNF1-related protein kinase 2) which is kept inactive by the type 2C protein phosphatase genes (PP2Cs) known as negative regulators of ABA signaling [74, 75]. It has been previously reported that MYB44 transcription factor plays a role in the ABA-mediated signaling pathway in Arabidopsis via reducing the expression of PP2Cs genes [64]. Arabidopsis plants overexpressing MYB44 exhibited enhanced stomatal closure [63] and reduced level of superoxide [76].
BnMYB44 and BnVIP1 genes showed significantly different expression profiles between roots of the drought tolerant and sensitive B. napus cultivars. Interestingly, it has already been unveiled that an ABA receptor, PYL8, recruits MYB44, MYB77 and MYB73 transcription factors to reinforce auxin signaling and thereby promoting lateral root growth under water limited conditions in Arabidopsis [77]. In addition, Arabidopsis MYB44 is involved in regulation of root architecture by modulating antagonistic interaction between salicylic acid and jasmonic acid signaling [78]. It is required to measure root morphological characteristics of the cultivars as well as performing in-depth analyses about possible functions of BnVIP1 and BnMYB44 in relation to root architecture of B. napus.
Homeostasis of sodium and potassium ions is of particular importance for plant salt tolerance [79]. The results demonstrated that the B. napus cultivars had different levels of salt tolerance evidenced by K/Na ratio. The significant differences observed in the salt-induced expression levels of BnMYB44 and BnVIP1 between the B. napus cultivars may be related to salt stress response of the cultivars. The cultivar with higher salt tolerance (SLM046) generally showed higher levels of BnMYB44 and BnVIP1 transcripts during salt stress period. Further investigations are required to determine the regulatory roles of BnVIP1 and BnMYB44 under salt stress conditions. The previous reports discovered that AtMYB44 gene was activated under salinity [63] and Arabidopsis plants overexpressing AtMYB44 showed enhanced salt tolerance [76]. It has recently been reported that salt stress induces the eviction of H2A.Z containing nucleosomes from the promoter region of AtMYB44 leading to transcript accumulation of this gene [80].
Computational-based promoter analysis can provide valuable knowledge for understanding and prediction of gene expression pattern [81,82,83]. The expression profiles of BnVIP1 and BnMYB44 under drought and salt stress conditions revealed that BnMYB44 was more responsive to such stresses. In accordance with this finding, the stress related CREs were generally more over-represented within the BnMYB44 promoter. In particular, some of the CREs with significantly different number of occurrences between the BnVIP1 and BnMYB44 promoters may contribute to the level of responsiveness of these genes. The number of occurrences of b-ZIP transcription factor binding sites including G-BOX, BOXII and ABRE within the BnMYB44 promoter was significantly greater than those within the promoter of BnVIP1. G-BOX and ABRE are the ABA responsive CREs that have been found in promoter regions of several ABA signaling genes such as MYB2 and MYC2 [84, 85]. These CREs are involved in early response to abiotic stresses to activate a broad range of adaptation mechanisms [86]. The greater number of occurrences of these CREs within the BnMYB44 promoter, compared to BnVIP1 promoter, may be connected with its faster induction under stress conditions. The greater numbers of CAAT-Box element, which is a common enhancer [87], within the BnVIP1 promoter may likely be involved in transcribing BnVIP1 at sufficient level to provide cytoplasmic concentration of VIP1 as a protein at the forefront of the signaling pathways.
Conclusion
In conclusion, the comparative transcriptome analysis of stress-sensitive (Arabidopsis) and -tolerant (E. salsugineum) model plants resulted in the regulatory network as a reliable source of the regulatory genes for further investigations in the agronomically important species such as B. napus. This study is the first report on cloning and expression analysis of B. napus transcription factor genes, BnMYB44 and BnVIP1, aimed at elucidating their functions in response to drought and salt stresses. The expression patterns of BnMYB44 and BnVIP1 genes of the two B. napus cultivars under drought and salt stress conditions along with the data obtained from the measurements of the stress-related physiological characteristics as well as analysis of the BnMYB44 and BnVIP1 promoters suggested that BnMYB44 and BnVIP1 genes may contribute to responses to drought and salt stresses in B. napus. Further detailed analyses are required to fully understand the roles and importance of BnMYB44 and BnVIP1 genes in stress regulatory pathways.
References
Farooq M, Hussain M, Wahid A, Siddique KHM (2012) Drought stress in plants: an overview. In: Aroca R (ed) Plant responses to drought stress. Springer, Berlin, pp 1–33
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Atkinson NJ, Urwin PE (2012) The interaction of plant biotic and abiotic stresses: from genes to the field. J Exp Bot 63:3523–3543
Farrant JM, Ruelland E (2015) Plant signalling mechanisms in response to the environment. Environ Exp Bot 114:1–3
Golldack D, Lüking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Cristina MS, Petersen M, Mundy J (2010) Mitogen-activated protein kinase signaling in plants. Annu Rev Plant Biol 61:621–649
Duque AS, Farinha AP, da Silva AB, de Almeida AM, Santos D, da Silva JM, Fevereiro P, de Sousa Araújo S (2013) Abiotic stress responses in plants: unraveling the complexity of genes and networks to survive. INTECH Open Access Publisher, Rijeka
Gehan MA, Greenham K, Mockler TC, McClung CR (2015) Transcriptional networks—crops, clocks, and abiotic stress. Curr Opin Plant Biol 24:39–46
Wohlbach DJ, Quirino BF, Sussman MR (2008) Analysis of the Arabidopsis histidine kinase ATHK1 reveals a connection between vegetative osmotic stress sensing and seed maturation. The Plant Cell Online 20:1101–1117
Deyholos MK (2010) Making the most of drought and salinity transcriptomics. Plant Cell Environ 33:648–654
Amtmann A (2009) Learning from evolution: Thellungiella generates new knowledge on essential and critical components of abiotic stress tolerance in plants. Mol Plant 2:3–12
Gong Q, Li P, Ma S, Indu Rupassara S, Bohnert HJ (2005) Salinity stress adaptation competence in the extremophile Thellungiella halophila in comparison with its relative Arabidopsis thaliana. Plant J 44:826–839
Mucha S, Walther D, Müller TM, Hincha DK, Glawischnig E (2015) Substantial reprogramming of the Eutrema salsugineum (Thellungiella salsuginea) transcriptome in response to UV and silver nitrate challenge. BMC Plant Biol 15:1
Wang X, Chang L, Wang B, Wang D, Li P, Wang L, Yi X, Huang Q, Peng M, Guo A (2013) Comparative proteomics of Thellungiella halophila leaves from plants subjected to salinity reveals the importance of chloroplastic starch and soluble sugars in halophyte salt tolerance. Mol Cell Proteom 12:2174–2195
Wong C, Li Y, Whitty B, Diaz-Camino C, Akhter S, Brandle J, Golding G, Weretilnyk E, Moffatt B, Griffith M (2005) Expressed sequence tags from the Yukon ecotype of Thellungiella reveal that gene expression in response to cold, drought and salinity shows little overlap. Plant Mol Biol 58:561–574
Bressan RA, Zhang C, Zhang H, Hasegawa PM, Bohnert HJ, Zhu J-K (2001) Learning from the Arabidopsis experience. The next gene search paradigm. Plant Physiol 127:1354–1360
Zhu J-K (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Xu X, Feng J, Lü S, Lohrey GT, An H, Zhou Y, Jenks MA (2014) Leaf cuticular lipids on the Shandong and Yukon ecotypes of saltwater cress, Eutrema salsugineum, and their response to water deficiency and impact on cuticle permeability. Physiol Plant 151:446–458
Taji T, Sakurai T, Mochida K, Ishiwata A, Kurotani A, Totoki Y, Toyoda A, Sakaki Y, Seki M, Ono H (2008) Large-scale collection and annotation of full-length enriched cDNAs from a model halophyte, Thellungiella halophila. BMC Plant Biol 8:1
Taji T, Seki M, Satou M, Sakurai T, Kobayashi M, Ishiyama K, Narusaka Y, Narusaka M, Zhu J-K, Shinozaki K (2004) Comparative genomics in salt tolerance between Arabidopsis and Arabidopsis-related halophyte salt cress using Arabidopsis microarray. Plant Physiol 135:1697–1709
Yang R, Jarvis DE, Chen H, Beilstein MA, Grimwood J, Jenkins J, Shu S, Prochnik S, Xin M, Ma C (2013) The reference genome of the halophytic plant Eutrema salsugineum. Front Plant Sci 4:14
Diepenbrock W (2000) Yield analysis of winter oilseed rape (Brassica napus L.): a review. Field Crops Res 67, 35–49.
Raymer PL (2002) Canola: an emerging oilseed crop. Trends New Crops New Uses 1:122–126
Xie, F.L., Huang, S.Q., Guo, K., Xiang, A.L., Zhu, Y.Y., Nie, L. and Yang, Z.M., 2007. Computational identification of novel microRNAs and targets in Brassica napus. FEBS Lett 581:1464–1474.
Aliakbari M, Razi H (2013) Isolation of Brassica napus MYC2 gene and analysis of its expression in response to water deficit stress. Mol Biol Res Commun 2:63–71
Bakhtari B, Razi H (2014) Differential expression of BnSRK2D gene in two Brassica napus cultivars under water deficit stress. Mol Biol Res Commun 3:241–251
Dalal M, Tayal D, Chinnusamy V, Bansal KC (2009) Abiotic stress and ABA-inducible Group 4 LEA from Brassica napus plays a key role in salt and drought tolerance. J Biotechnol 139:137–145
Orr W, Iu B, White TC, Robert LS, Singh J (1992) Complementary DNA sequence of a low temperature-induced Brassica napus gene with homology to the Arabidopsis thaliana kin1 gene. Plant Physiol 98:1532
Zhuang J, Sun C-C, Zhou X-R, Xiong A-S, Zhang J (2011) Isolation and characterization of an AP2/ERF-RAV transcription factor BnaRAV-1-HY15 in Brassica napus L. HuYou15. Mol Biol Rep 38:3921–3928
Boyes DC, Zayed AM, Ascenzi R, McCaskill AJ, Hoffman NE, Davis KR, Görlach J (2001) Growth stage—based phenotypic analysis of Arabidopsis: a model for high throughput functional genomics in plants. Plant Cell 13:1499–1510
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Blazejczyk M, Miron M, Nadon R (2007) FlexArray: a statistical data analysis software for gene expression microarrays. Genome Quebec, Montreal
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (Methodol) 57:289–300
Dai X, Sinharoy S, Udvardi M, Zhao P (2013) PlantTFcat: an online plant transcription factor and transcriptional regulator categorization and analysis tool. BMC Bioinform 14:321
Yao Q, Ge H, Wu S, Zhang N, Chen W, Xu C, Gao J, Thelen JJ, Xu D (2014) P3DB 3.0: from plant phosphorylation sites to protein networks. Nucleic Acids Res 42:D1206–D1213
Nikitin A, Egorov S, Daraselia N, Mazo I (2003) Pathway studio—the analysis and navigation of molecular networks. Bioinformatics 19:2155–2157
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Panahi B, Mohammadi SA, Khaksefidi RE, Mehrabadi JF, Ebrahimie E (2015) Genome-wide analysis of alternative splicing events in Hordeum vulgare: highlighting retention of intron-based splicing and its possible function through network analysis. FEBS Lett 589:3564–3575
Ebrahimi Khaksefidi R, Mirlohi S, Khalaji F, Fakhari Z, Shiran B, Fallahi H, Rafiei F, Budak H, Ebrahimie E (2015) Differential expression of seven conserved microRNAs in response to abiotic stress and their regulatory network in Helianthus annuus. Front Plant Sci 6:741
Panahi B, Abbaszadeh B, Taghizadeghan M, Ebrahimie E (2014) Genome-wide survey of alternative splicing in Sorghum bicolor. Physiol Mol Biol Plants 20:323–329
Boccaletti S, Latora V, Moreno Y, Chavez M, Hwang D-U (2006) Complex networks: structure and dynamics. Phys Rep 424:175–308
Scardoni G, Petterlini M, Laudanna C (2009) Analyzing biological network parameters with CentiScaPe. Bioinformatics 25:2857–2859
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Darjani A, Rad AHS, Gholipour S, Haghighat A (2013) Investigation the effects of water stress on yield and yield components of canola winter varieties. Int J Agron Plant Prod 4:370–374
Mirzaee M, Moieni A, Ghanati F (2013) Effects of drought stress on the lipid peroxidation and antioxidant enzyme activities in two canola (Brassica napus L.) cultivars. J Agric Sci Technol 15:593–602
Rameeh V, Cherati A, Abbaszadeh F (2012) Salinity effects on yield, yield components and nutrient ions in rapeseed genotypes. J Agric Sci Belgrade 57:19–29
Slatyer R, Shmueli E (1967) Measurements of internal water status and transpiration. In: Hagan RM, Haise HR, Edminster TW (eds.) Irrigation of Agricultural Lands. American Society of Agronomy. pp 337–353
Bates L, Waldren R, Teare I (1973) Rapid determination of free proline for water-stress studies. Plant soil 39:205–207
Sairam R, Srivastava G (2002) Changes in antioxidant activity in sub-cellular fractions of tolerant and susceptible wheat genotypes in response to long term salt stress. Plant Sci 162:897–904
Munns R, Wallace PA, Teakle NL, Colmer TD (2010) Measuring soluble ion concentrations (Na+, K+, Cl–) in salt-treated plants, plant stress tolerance. Springer, New York, pp 371–382
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 25:402–408
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Shamloo-Dashtpagerdi R, Razi H, Aliakbari M, Lindlöf A, Ebrahimi M, Ebrahimie E (2015) A novel pairwise comparison method for in silico discovery of statistically significant cis-regulatory elements in eukaryotic promoter regions: application to Arabidopsis. J Theor Biol 364:364–376
Koschützki D, Schreiber F (2008) Centrality analysis methods for biological networks and their application to gene regulatory networks. Gene Regul Syst Biol 2:193
Du H, Zhang L, Liu L, Tang X-F, Yang W-J, Wu Y-M, Huang Y-B, Tang Y-X (2009) Biochemical and molecular characterization of plant MYB transcription factor family. Biochemistry 74:1–11
Kirik V, Kölle K, Miséra S, Bäumlein H (1998) Two novel MYB homologues with changed expression in late embryogenesis-defective Arabidopsis mutants. Plant Mol Biol 37:819–827
Tzfira T, Vaidya M, Citovsky V (2001) VIP1, an Arabidopsis protein that interacts with Agrobacterium VirE2, is involved in VirE2 nuclear import and Agrobacterium infectivity. EMBO J 20:3596–3607
Chakraborty K, Bose J, Shabala L, Shabala S (2016) Difference in root K+ retention ability and reduced sensitivity of K+-permeable channels to reactive oxygen species confer differential salt tolerance in three Brassica species. J Exp Bot 67:4611–4625
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149:88–95
Shamloo-Dashtpagerdi R, Razi H, Ebrahimie E (2015) Mining expressed sequence tags of rapeseed (Brassica napus L.) to predict the drought responsive regulatory network. Physiol Mol Biol Plants 21:329–340
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581
Jung C, Seo JS, Han SW, Koo YJ, Kim CH, Song SI, Nahm BH, Choi D, Y. and Cheong J-J (2008) Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis. Plant Physiol 146:623–635
Joo J, Oh N-I, Nguyen NH, Lee YH, Kim Y-K, Song SI, Cheong J-J (2017) Intergenic transformation of AtMYB44 confers drought stress tolerance in rice seedlings. Appl Biol Chem 60:447–455
Tsugama D, Liu S, Takano T (2014) Analysis of functions of VIP1 and its close homologs in osmosensory responses of Arabidopsis thaliana. PLoS ONE 9:e103930
Djamei A, Pitzschke A, Nakagami H, Rajh I, Hirt H (2007) Trojan horse strategy in Agrobacterium transformation: abusing MAPK defense signaling. Science 318:453–456
Pitzschke A, Djamei A, Teige M, Hirt H (2009) VIP1 response elements mediate mitogen-activated protein kinase 3-induced stress gene expression. Proc Natl Acad Sci 106, 18414–18419
Norouzi M, Toorchi M, Salekdeh GH, Mohammadi S, Neyshabouri M, Aharizad S (2008) Effect of water deficit on growth, grain yield and osmotic adjustment in rapeseed. J Food Agric Environ 6:312
Szabados L, Savoure A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97
Rejeb KB, Abdelly C, Savouré A (2014) How reactive oxygen species and proline face stress together. Plant Physiol Biochem 80:278–284
Brodribb TJ, Holbrook NM (2003) Stomatal closure during leaf dehydration, correlation with other leaf physiological traits. Plant Physiol 132:2166–2173
Geiger D, Maierhofer T, AL-Rasheid KA, Scherzer S, Mumm P, Liese A, Ache P, Wellmann C, Marten I, Grill E (2011) Stomatal closure by fast abscisic acid signaling is mediated by the guard cell anion channel SLAH3 and the receptor RCAR1. Sci Signal 4:ra32
Acharya BR, Jeon BW, Zhang W, Assmann SM (2013) Open Stomata 1 (OST1) is limiting in abscisic acid responses of Arabidopsis guard cells. New Phytol 200:1049–1063
Yoshida R, Umezawa T, Mizoguchi T, Takahashi S, Takahashi F, Shinozaki K (2006) The regulatory domain of SRK2E/OST1/SnRK2. 6 interacts with ABI1 and integrates abscisic acid (ABA) and osmotic stress signals controlling stomatal closure in Arabidopsis. J Biol Chem 281:5310–5318
Persak H, Pitzschke A (2014) Dominant repression by Arabidopsis transcription factor MYB44 causes oxidative damage and hypersensitivity to abiotic stress. Int J Mol Sci 15:2517–2537
Zhao Y, Xing L, Wang X, Hou Y-J, Gao J, Wang P, Duan C-G, Zhu X, Zhu J-K (2014) The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci Signal 7:ra53
Walker L, Boddington C, Jenkins D, Wang Y, Grønlund JT, Hulsmans J, Kumar S, Patel D, Moore JD, Carter A (2017) Root architecture shaping by the environment is orchestrated by dynamic gene expression in space and time. Plant Cell. https://doi.org/10.1105/tpc.16.00961
Benito B, Haro R, Amtmann A, Cuin TA, Dreyer I (2014) The twins K+ and Na+ in plants. J Plant Physiol 171:723–731
Nguyen NH, Cheong J-J (2018) H2A. Z-containing nucleosomes are evicted to activate AtMYB44 transcription in response to salt stress. Biochem Biophys Res Commun 499:1039–1043
Hosseinpour B, Bakhtiarizadeh MR, Khosravi P, Ebrahimie E (2013) Predicting distinct organization of transcription factor binding sites on the promoter regions: a new genome-based approach to expand human embryonic stem cell regulatory network. Gene 531:212–219
Bakhtiarizadeh MR, Moradi-Shahrbabak M, Ebrahimie E (2013) Underlying functional genomics of fat deposition in adipose tissue. Gene 521:122–128
Mahdi LK, Deihimi T, Zamansani F, Fruzangohar M, Adelson DL, Paton JC, Ogunniyi AD, Ebrahimie E (2014) A functional genomics catalogue of activated transcription factors during pathogenesis of pneumococcal disease. BMC Genom 15:769
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell Online 15:63–78
Deppmann CD, Alvania RS, Taparowsky EJ (2006) Cross-species annotation of basic leucine zipper factor interactions: insight into the evolution of closed interaction networks. Mol Biol Evol 23:1480–1492
Corrêa LGG, Riaño-Pachón DM, Schrago CG, Dos Santos RV, Mueller-Roeber B, Vincentz M (2008) The role of bZIP transcription factors in green plant evolution: adaptive features emerging from four founder genes. PLoS ONE 3:e2944
Ramji D, Foka P (2002) CCAAT/enhancer-binding proteins: structure, function and regulation. Biochem J 365:561–575
Acknowledgements
This research was supported by Department of Crop Production and Plant Breeding, College of Agriculture, Shiraz University.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shamloo-Dashtpagerdi, R., Razi, H., Ebrahimie, E. et al. Molecular characterization of Brassica napus stress related transcription factors, BnMYB44 and BnVIP1, selected based on comparative analysis of Arabidopsis thaliana and Eutrema salsugineum transcriptomes. Mol Biol Rep 45, 1111–1124 (2018). https://doi.org/10.1007/s11033-018-4262-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-018-4262-0