Abstract
Litchi (Litchi chinensis Sonn., L. chinensis), a type of tree growing in most areas of southern China, produces an edible fruit that is also a source of traditional medicine. Genetic identification of litchi species or cultivars using molecular markers is very important. In this study, a total of six litchi samples from Fujian, Hainan, Guangdong, Guangxi and Sichuan province, as well as one wild Dimocarpus confinis (D. confinis) sample from Guangxi province were collected for genetic analysis. The cluster dendrograms were constructed for genetic analysis on the basis of DNA amplification results by RAPD and ISSR. The improved RAPD amplified DNA with consistent and clear banding patterns. A total of 176 bands were found, indicating a 72.7 % polymorphism in L. chinensis DNA samples. Significant genetic distances were found among the different species or cultivars, with an index of similarity coefficient ranging from 0.59 to 0.87. Similar to RAPD results, ISSR analysis of the L. chinensis DNA samples showed a range of 0.70–0.93 similarity coefficients. The genetic distance between Hainan sample and Sichuan samples was the farthest, which is consistent with their geographic distance. Furthermore, the index of similarity coefficient between D. confinis and L. chinensis was 0.35–0.41 by RAPD and 0.38–0.48 by ISSR, indicating that these two species have significant genetic difference. This study reveals the high level of genetic differences between different litchi species or cultivars, and confirms the significance of the improved RAPD method in genetic characterization of organisms. Taken together, the improved RAPD combined with ISSR analysis can be used frequently for the genetic diversity, germplasm resources preservation, molecular-assisted breeding, and genetic characterization of various organisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Litchi chinensis Sonn. (litchi or lychee), belonging to the family of Sapindaceae, is a tropical and subtropical evergreen tree, native to the Hainan, Guangdong and Fujian provinces of China. It is also cultivated in many other parts of the world, particularly in Southeast Asia. Litchi has been cultivated in China since 2000 BC. Litchi is an edible fruit and source of traditional medicine. As a traditional medicine, litchi fruit and its secondary metabolic products, have been reported to have anticancer [1, 2], anti-inflammatory [3], antifungal [4], antiviral [5], antioxidant [6–11], antiplatelet and anticoagulant [12], and antidiabetic activities [13–15].
Random amplified polymorphic DNA (RAPD) analysis was developed in 1990 [16]. Since then, RAPD analysis, combined with other molecular techniques such as internal transcribed spacer (ITS), simple sequence repeat (SSR), inter-simple sequence repeat (ISSR), inter-retrotransposon amplified polymorphism (IRAP), amplified fragment length polymorphism (AFLP) and restriction fragment length polymorphism (RFLP) [17–22], has been widely used in the various unicellular and multi-cellular organisms across various fields for assessments of genetic diversity and characterization of germplasm, identification and fingerprinting of genotypes, and molecular-assisted breeding in ecological, evolutionary, taxonomical, phylogenic and genetic studies. Although RAPD has gained popularity because of its advantages, it also has some disadvantages, including low production and poor reproducibility. Interestingly, the resolution and production can be greatly increased by an improved RAPD technique (also called RAMP-PCR), in which ramp time from the stage of annealing to extension is elongated [23, 24].
There are numerous litchi species or cultivars, and considerable confusion has been noticed regarding their naming and identification. The same cultivar grown in different climates may produce very different fruit. Cultivars can also have different synonyms in various parts of the world. Therefore, the nutritional or medicinal values of litchi may vary. Heretofore there are very limited genetic studies of this edible and medicinal species exist. Ding et al. [25] established the segregation patterns by RAPD markers in an F1 population. Liu et al. [26] investigated the genetic relationship between superior individual late-maturity Yurongwanli and other cultivars from Fuqing city of Fujian province by POD isoenzyme and RAPD analysis. Zhou et al. [27] constructed two high density genetic linkage maps by RAPD, AFLP and SRAP techniques based on the mapping population derived from a cross of ‘Maguili’ and ‘Jiaohesanyuehong’. However, no study has been conducted for the measurement of genetic distance and diversity between species of litchi (including D. confinis) from different geographic origins using the improved RAPD together with ISSR.
In this study, we collected the DNA materials from different geographic localities, characterized them genetically by using our developed RAMP-PCR technique, and validated the RAPD results by ISSR technique.
Materials and methods
Experimental reagents
The RAPD primers (SBS Genetech Corporation) (Table 1), ISSR primers (UBC Primer Set #9) (Table 2), 2 × PCR Taq Mastermix (TianGen Biotech Co. Ltd, Beijing, China) and DNA Marker (Takara Biotechnology Co. Ltd, Dalian, China) were used in this study. Other reagents were analytical grade reagents that we described in our previous studies [23, 24, 28–30].
Collection of plant samples
A total of six litchi samples were collected in southern China: 1 from Quanzhou of Fujian, 1 from Wanning of Hainan, 1 from Dongguan of Guangdong, 1 from Yulin of Guangxi, and 2 from Luzhou of Sichuan including 1 from the Hejiang county and 1 from Zhangba Longan Park (Fig. 1); (Table 1). One wild D. confinis sample was provided from Agricultural College of Guangxi University, Guangxi province and was described previously [31].
The localities of samples of L. chinensis or D. confinis species from different regions of southern China. The spots in dark blue indicate the cities and the line in light blue indicates the Yangtze River. The detailed information for each sample is shown in Table 1. (Color figure online)
DNA extraction
Total genomic DNA was extracted from fresh leaves using a modified Cetyl trimethylammonium bromide (CTAB) method, described previously [24, 32]. The leaves were first fixed in fixing solutions containing chloroform, PVP, 2-Hydroxy-1-ethanethiol (without liquid nitrogen), and then ground into tiny pieces by silica (SiO2) for the extraction of DNA using the CTAB method. DNA quality was checked by 0.8 % agarose gel electrophoresis and spectrophotometry [33]. The final concentration of all DNA samples was adjusted to10 ng/μl for PCR, and stored at −20 °C until use.
RAPD-PCR
Twenty three different SBS primers were initially evaluated for the polymorphism detection by improved RAPD analysis, among which nineteen primers amplified DNA with polymorphic profiles for data analysis (Table 2). The contents of PCR system (10 μl) were as follow: 1 μl of primers (2.5 μmol/L), 1 μl (10 ng) of L. chinensis or D. confinis DNA templates, 5 μl of 2 × PCR Taq Mastermix and 3 μl of ddH2O. The PCR condition was as follows: initial denaturation at 95 °C for 90 s, followed by 40 cycles of 40 s at 94 °C, 60 s at 36 °C, 90 s at 72 °C, and final extension of 5 min at 72 °C. PCR was performed in Applied Biosystems Veriti® 96-Well Thermal Cycler (Life Technology, USA). The RAMP rate from annealing to extension was adjusted from 2.5 °C/s (100 % ramp rate) to 0.125 °C/s (5 % ramp rate) for L. chinensis and D. confinis species using our previously established ramp PCR conditions [24] to compare the resolution and production of the two methods in the present study. PCRs were repeated three times for all 7 samples.
ISSR amplification
ISSR amplifications were performed in 10 μl reactions volumes consisting 1 μl of 2.5 μmol/L primers, 1 μl of DNA template of L. chinensis or D. confinis species samples, 5 μl of 2 × PCR Taq Mastermix, and 3 μl of ddH2O. The PCR condition was as follows: initial denaturation at 95 °C for 90 s, followed by 35 cycles of 40 s at 94 °C, 30 s at 50 °C, 90 s at 72 °C, and final extension of 5 min at 72 °C [24]. PCR was executed in an above mentioned “Applied Biosystems Veriti® 96-Well Thermal Cycler”. Totally twenty-two ISSR primers (from UBC Primer Set #9), were tested initially. Twelve primers (Table 3) amplified DNA well with polymorphic bands and were selected for further use.
Agarose gel electrophoresis
The amplified PCR products were separated by electrophoresis on a 1.8 % agarose gel in 1 × TAE buffer. Gels were visualized by 0.5 μg/mL ethidium bromide staining and the images were documented using the ChemiDoc XR (Bio-Rad, USA) under UV light. Unambiguous and reproducible bands in successive amplifications were selected for scoring.
Data analysis
The presence of each selected clearly DNA band in the amplified gel profiles was recorded as “1”, and absence of this corresponding band in other sample(s) was recorded as “0”. The similarity matrix (SM) and the similarity index (SI) were calculated by using SM coefficient. The dendrograms based on unweighted pair group method with arithmetic mean algorithm (UPGMA) were generated using the SAHN module in a NTSYS pc 2.1 package [34].
Results
Comparison between regular RAPD amplification and improved RAPD technique for litchi DNA analysis
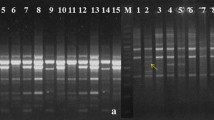
To increase the RAPD amplification efficiency and get more specific bands, PCR was first used to amplify DNA by adjusting RAMP time from annealing to extension with a ramp rate for 5 % (0.125 °C/s) and 100 % (2.5 °C/s) from 7 of L. chinensis or D. confinis species samples (listed in Table 1) by RAPD primers SBS-Q2 and SBS-Q10. Specifically, with primer SBS-Q2, the range of amplified bands was 0-6 by regular PCR (Fig. 2a, left panel), whereas the amounts of PCR products and the numbers of DNA bands (8–11) were clearly increased in RAMP PCR (in which RAMP rate was adjusted from 100 to 5 %) (Fig. 2a, right panel); In the case of primer SBS-Q10, there were 2–5 amplified bands in regular PCR (Fig. 2b, left panel), whereas the amounts of PCR products and the numbers of DNA bands (6–12) were clearly increased by RAMP PCR with 5 % RAMP rate (Fig. 2b, right panel). Particularly, the signals of some PCR bands in the right panels of Fig. 2 were much stronger than the corresponding bands in the left panels, which demonstrates that the yield was higher by improved RAPD PCR than by regular RAPD PCR. Hence, our modified RAPD condition with prolonging ramp time from annealing to extension, would be useful for genetic analysis of L. chinensis or D. confinis species as the yield, resolution, and reproducibility were significantly improved. This improved condition was then used to complete amplification of 7 samples of L. chinensis or D. confinis species by other RAPD primers.
Comparison between regular RAPD amplification with 100 % RAMP rate and improved RAPD amplification with 5 % RAMP rate. Lanes 1–7 represent different samples of L. chinensis or D. confinis species listed in Table 1. a PCR results by primer SBS-Q2. b PCR results by primer SBS-Q10. Left panels indicate the results of regular RAPD amplification (regular PCR); right panels indicate the results of improved RAPD amplification (RAMP PCR). Lane “M” represents DL2000 DNA marker with indicated molecular size (bp)
Amplification of DNAs of L. chinensis and D. confinis species by RAMP RAPD
A total of twenty three primers were used in improved RAPD analysis for the evaluation of DNA polymorphism, and nineteen primers (Table 1) obtained reproducible polymorphic amplification bands. Figure 3a, b showed that three representative primers (SBS-I4, SBS-Q1 and SBS-Q19) generated reproducible polymorphic amplification bands by RAMP PCR in these 7 samples. From these polymorphic amplification primers, a total of 218 bands were obtained, where each primer had amplified 2–11 bands with an average of 5.99 bands per primer. The approximate range of band size was 200–2200 bp, and 93 % of the bands were found polymorphic in these 7 samples. In six L. chinensis cultivars, a total of 176 RAPD bands were found with an average of band number of 6.06 per primer, and 72.7 % of the bands were polymorphic.
The representative results of banding profiles obtained by improved RAPD amplification (5 % RAMP rate) by primers SBS-I4, SBS-Q1 and SBS-Q19. Lanes 1–7 represent different samples listed in Table 1. Lane “M” represents DL2000 DNA marker
Genetic distance and cluster analysis of RAMP RAPD
Based on the RAMP-PCR amplification profiles, cluster dendrogram was obtained (Fig. 4a, b). The dendrogram showed that the index of similarity coefficients ranged from 0.35 to 0.87. The index of similarity coefficient between sample 1 and 6 or 7 (D. confinis in Guangxi and L. chinensis in Sichuan) was lowest (0.35), while that between sample 6 and 7 (L.chinensis from Zhangba and Hejiang in Sichuan province), was the highest (0.93) (Fig. 4b). In six L. chinensis cultivars, the dendrogram showed that the index of similarity coefficients ranged from 0.59 to 0.87. The index of similarity coefficient between sample 3 and 6 (Hainan sample and Zhangba of Luzhou in Sichuan) was the lowest (0.59), while that between sample 6 and 7 (L. chinensis from Zhangba and Hejiang in Sichuan) was still the highest (0.87) (Fig. 4b).
Dendrogram of L. chinensis and D. confinis species samples by improved RAPD. a Dendrogram of L. chinensis and D. confinis species based on improved RAPD PCR amplification files. Bars on the bottom indicate similarity index based on S.M coefficients. b Genetic distance dendrogram for L. chinensis and D. confinis species using improved RAPD markers
Amplification of D. confinis and L. chinensis species DNAs by ISSR PCR
Twelve of ISSR primers amplified well, producing 99 clear and reproducible fragments from 7 samples. Each primer had amplified 0–14 bands with an average number of 4.43 bands per primer. The approximate range of band size was 200–1900 bp, and 85.9 % of the bands were found polymorphic in these samples. The representative results by ISSR primers UBC825, UBC873 and UBC879 are showed in Fig. 5. These results provide a clear detection of polymorphism in DNA samples of L. chinensis species. In six L. chinensis cultivars, there was a total of 73 ISSR bands with an average number of 4.26 bands per primer, while 61.64 % of the bands were found polymorphic.
The representative results of different ISSR maker patterns in L. chinensis and D. confinis species obtained by primers for UBC825, UBC873 and UBC879. Lanes 1–7 represent different samples listed in Table 1. Lane “M” represents DL2000 DNA marker
Genetic distance and cluster analysis of ISSR PCR
Cluster dendrogram was also obtained based on the ISSR banding profiles, showing similar results to those obtained by improved RAPD of the similarity coefficients among seven varieties of D. confisis and L. chinensis species (Fig. 6a, b). The dendrogram showed that the index of similarity coefficients among the seven samples ranged from 0.38 to 0.93. The index of similarity coefficient between sample 1 (D. confinis) and 2 (L. chinensis in Fujian) was the lowest (0.38), and that between sample 6 and 7 (L. chinensis from Zhangba and Hejiang in Sichuan), was maximum (0.93) (Fig. 6b). In the six L. chinensis cultivars, the dendrogram showed that the index of similarity coefficients ranged from 0.70 to 0.93. The index of similarity coefficient between sample 3 and 6 (Hainan sample and Zhangba sample of Luzhou in Sichuan, respectively) was the lowest (0.70), and that between sample 6 and 7 (L. chinensis from Zhangba and Hejiang in Sichuan, respectively) was the highest (0.93), which is consistent with the RAPD results (Fig. 6b).
Dendrogram of L. chinensis and D. confinis species by ISSR technique. a Dendrogram of L. chinensis or D. confinis species based on with amplification files by ISSR primers. Bars on the bottom indicate similarity index based on S.M coefficients. b Genetic distance dendrogram for L. chinensis and D. confinis species using ISSR markers
Discussion
To increase the RAPD amplification efficiency and get more specific bands, RAPD PCR was first used to amplify in a machine of “Applied Biosystems Veriti® 96-Well Thermal Cycler” by adjusting RAMP time from annealing to extension with a ramp rate for 5 % (0.125 °C/s) and 100 % (2.5 °C/s). We found that the yield, resolution, and reproducibility of RAPD were significantly improved by this modified RAMP rate (Fig. 2). These improvements are likely due to the appropriately prolonged ramp time (from 0.125 to 2.5 °C/s) that increases the capacity for the binding of the 10-bp primer with DNA template, and the stability between primer and DNA, and prevents falling off primers from the DNA templates [23]. This improved condition was used to amplify 7 DNA samples in our study. Of twenty three primers tested, nineteen primers generated unambiguous and bright bands.
Using improved RAPD and ISSR marker techniques, we successfully authenticated and characterized L. chinensis from different geographic localities. In six L. chinensis cultivars, the dendrogram generated by RAPD showed that the index of similarity coefficients ranged from 0.59 to 0.87. The index of similarity coefficient between sample 3 and 6 was the lowest (0.59), while that between sample 6 and 7 was the highest (0.87) (Fig. 4b). The dendrogram generated by ISSR showed that the index of similarity coefficients ranged from 0.70 to 0.93 in the six L. chinensis cultivars. The index of similarity coefficient between sample 3 and 6 was the lowest (0.70), and that between sample 6 and 7 was the highest (0.93) (Fig. 6b), which is consistent with the RAPD results. These findings demonstrate that not only increases the resolution and yield by the improved RAPD technique, but also it is a reliable molecular tool for the genetic characterization of various organisms, which was reported in previous studies [23, 24]. To our knowledge, this is the first report of genetic characterization of L. chinensis by combining improved RAPD and ISSR analysis, and this characterization could be useful for the preservation of genetic diversity and litchi population. Importantly, our RAPD results in Fig. 4 and ISSR results in Fig. 6, demonstrate that the genetic distance between Hainan sample (sample 3 in Table 1) and Sichuan samples (sample 6 and 7 in Table 1) is longest, which is consistent with the geographic distance. The different geographic location and host’s nature may have significant influence on this variation. This information might be helpful for the analysis of genetic diversity, germplasm resources preservation, molecular-assisted breeding and identification of litchi species or cultivars.
D. confinis, also known as “longli” in Chinese, has similarity in the morphology of trees and fruits with L. chinensis and D. longan. D. confinis is a very ancient species in Guangxi province and was first described by Dacheng Fan in his book “Guihai Yuheng zhi” at the year of 1162. One wild D. confinis sample from Guangxi province, a species of plant in the genus Dimocarpus family, was described previously [31]. However, it is hard to distinguish D. confinis from L. chinensis only by morphology. In this study, genetic distance and cluster analysis showed that the index of similarity coefficient between D. confinis and L. chinensis is 0.35–0.41 by RAPD, and 0.38–0.48 by ISSR, respectively, indicating that the two species have significant difference at molecular level. Our improved RAPD and ISSR analysis thus showed potentiality to distinguish L. chinensis from related genus or species.
In summary, our study indicates that the improved RAPD combined with ISSR techniques would be used frequently for the genetic diversity, germplasm resources preservation, molecular-assisted breeding and genetic characterization of various organisms.
References
Wang X, Yuan S, Wang J, Lin P, Liu G, Lu Y, Zhang J, Wang W, Wei Y (2006) Anticancer activity of litchi fruit pericarp extract against human breast cancer in vitro and in vivo. Toxicol Appl Pharmacol 215:168–178
Hsu CP, Lin CC, Huang CC, Lin YH, Chou JC, Tsia YT, Su JR, Chung YC (2012) Induction of apoptosis and cell cycle arrest in human colorectal carcinoma by Litchi seed extract. J Biomed Biotechnol 2012:341479
Huang F, Zhang R, Yi Y, Tang X, Zhang M, Su D, Deng Y, Wei Z (2014) Comparison of physicochemical properties and immunomodulatory activity of polysaccharides from fresh and dried litchi pulp. Molecules 19:3909–3925
Xu L, Xue J, Wu P, Wang D, Lin L, Jiang Y, Duan X, Wei X (2013) Antifungal activity of hypothemycin against Peronophythora litchii in vitro and in vivo. J Agric Food Chem 61:10091–10095
Ichinose T, Musyoka TM, Watanabe K, Kobayashi N (2013) Evaluation of antiviral activity of Oligonol, an extract of Litchi chinensis, against betanodavirus. Drug Discov Ther 7:254–260
Yang B, Wang J, Zhao M, Liu Y, Wang W, Jiang Y (2006) Identification of polysaccharides from pericarp tissues of litchi (Litchi chinensis Sonn.) fruit in relation to their antioxidant activities. Carbohydr Res 341:634–638
Duan X, Wu G, Jiang Y (2007) Evaluation of the antioxidant properties of litchi fruit phenolics in relation to pericarp browning prevention. Molecules 12:759–771
Kong F, Zhang M, Liao S, Yu S, Chi J, Wei Z (2010) Antioxidant activity of polysaccharide-enriched fractions extracted from pulp tissue of Litchi Chinensis Sonn. Molecules 15:2152–2165
Yang DJ, Chang YZ, Chen YC, Liu SC, Hsu CH, Lin JT (2012) Antioxidant effect and active components of litchi (Litchi chinensis Sonn.) flower. Food Chem Toxicol 50:3056–3061
Zhang R, Zeng Q, Deng Y, Zhang M, Wei Z, Zhang Y, Tang X (2013) Phenolic profiles and antioxidant activity of litchi pulp of different cultivars cultivated in Southern China. Food Chem 136:1169–1176
Su D, Zhang R, Hou F, Zhang M, Guo J, Huang F, Deng Y, Wei Z (2014) Comparison of the free and bound phenolic profiles and cellular antioxidant activities of litchi pulp extracts from different solvents. BMC Complement Altern Med 14:9
Sung YY, Yang WK, Kim HK (2012) Antiplatelet, anticoagulant and fibrinolytic effects of Litchi chinensis Sonn. extract. Mol Med Rep 5:721–724
Guo J, Li L, Pan J, Qiu G, Li A, Huang G, Xu L (2004) Pharmacological mechanism of Semen Litchi on antagonizing insulin resistance in rats with type 2 diabetes. Zhong Yao Cai 27:435–438
Noh JS, Kim HY, Park CH, Fujii H, Yokozawa T (2010) Hypolipidaemic and antioxidative effects of oligonol, a low-molecular-weight polyphenol derived from lychee fruit, on renal damage in type 2 diabetic mice. Br J Nutr 104:1120–1128
Noh JS, Park CH, Yokozawa T (2011) Treatment with oligonol, a low-molecular polyphenol derived from lychee fruit, attenuates diabetes-induced hepatic damage through regulation of oxidative stress and lipid metabolism. Br J Nutr 106:1013–1022
Williams JG, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Hess J, Kadereit JW, Vargas P (2000) The colonization history of Olea europaea L. in Macaronesia based on internal transcribed spacer 1 (ITS-1) sequences, randomly amplified polymorphic DNAs (RAPD), and intersimple sequence repeats (ISSR). Mol Ecol 9:857–868
Agarwal M, Shrivastava N, Padh H (2008) Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Rep 27:617–631
Varela ES, Lima JP, Galdino AS, Pinto Lda S, Bezerra WM, Nunes EP, Alves MA, Grangeiro TB (2004) Relationships in subtribe Diocleinae (Leguminosae; Papilionoideae) inferred from internal transcribed spacer sequences from nuclear ribosomal DNA. Phytochemistry 65:59–69
Noormohammadi Z, Hasheminejad-Ahangarani Farahani Y, Sheidai M, Ghasemzadeh-Baraki Alishah S, Alishah O (2013) Genetic diversity analysis in Opal cotton hybrids based on SSR, ISSR, and RAPD markers. Genet Mol Res 12:256–269
Pachuau L, Atom AD, Thangjam R (2014) Genome classification of Musa cultivars from Northeast India as revealed by ITS and IRAP markers. Appl Biochem Biotechnol 172:3939–3948
Khadivi-Khub A, Soorni A (2014) Comprehensive genetic discrimination of Leonurus cardiaca populations by AFLP, ISSR, RAPD and IRAP molecular markers. Mol Biol Rep 41:4007–4016
Fu J, Li L, Xu X, Wang Z, Tang G, Yin C, Lu G (2000) An improved method for increasing the efficiency of the technique of random amplified polymorphic DNA (RAPD). Hereditas 22:251–252
Fu J, Yang L, Khan MA, Mei Z (2013) Genetic characterization and authentication of Lonicera japonica Thunb. by using improved RAPD analysis. Mol Biol Rep 40:5993–5999
Ding XD, Lu LX, Chen XJ (2001) Segregation patterns of RAPD markers in an F1 population of Litchi chinensis Sonn. Acta Hort 558:167–172
Liu LS, Pan DM, Zhong FL, Wang JB (2010) POD isoenzyme and RAPD analysis on litchi in Fuqing. The Third international conference symposium on Longan, Lychee and other fruit trees in Sapindaceae family. Acta Hort 863:189–194
Zhou J, Fu JX, Wu ZX, Huang SS, Zhao YH, Zhang B, Hu YL, Hu GB, Liu CM (2012) Construction of two high density genetic linkage maps in litchi. Acta Hort 929:207–213
Mei ZQ, Fu SY, Yu HQ, Yang LQ, Duan CG, Liu XY, Gong S, Fu JJ (2014) Genetic characterization and authentication of Dimocarpus longan Lour. using an improved RAPD technique. Genet Mol Res 13:1447–1455
Mei Z, Yang L, Khan MA, Yang M, Wei C, Yang W, Peng X, Tania M, Zhan Hg, Li X, Fu J (2014) Genotyping of ganoderma species by improved random amplified polymorphic DNA (RAPD) and inter-simple sequence repeat (ISSR) analysis. Biochem Syst Ecol 56:40–48
Yang L, Khan MA, Mei Z, Yang M, Zhang T, Wei C, Yang W, Zhu L, Long Y, Fu J (2014) Development of RAPD-SCAR markers for Lonicera japonica Thunb. (Caprifolicaceae) variety authentication by improved RAPD and DNA cloning. Rev Biol Trop 62:(in press)
Yang L, Fu S, Khan MA, Zeng W, Fu J (2013) Molecular cloning and development of RAPD-SCAR markers for Dimocarpus longan variety authentication. SpringerPlus 2:501
Sharma P, Joshi N, Sharma A (2010) Isolation of genomic DNA from medicinal plants without liquid nitrogen. Indian J Exp Biol 48:610–614
Fu JJ (2012) Short protocols in medical molecular biology. China Medical Science Press, Beijing
Rohlf FJ (2002) NTSYS-pc: numerical taxonomy system ver.2.1. Exeter Pub Ltd, Setauket, New York. http://www.exetersoftware.com/cat/ntsyspc/ntsyspc.html. Accessed 6 June 2014
Acknowledgments
This work was supported in part by the National Natural Science Foundation of China (81172049), Science and Technology Innovation Team of Colleges and Universities of Sichuan Province (13TD0032), Applied Basic Research Program of Science and Technology Department of Sichuan Province (14JC0797) and Luzhou City Special Foundation (2013LZLY-J10). The authors thank all individuals who provided plant leaves or DNAs.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Yan Long and Jingliang Cheng are co-first authors.
Rights and permissions
About this article
Cite this article
Long, Y., Cheng, J., Mei, Z. et al. Genetic analysis of litchi (Litchi chinensis Sonn.) in southern China by improved random amplified polymorphic DNA (RAPD) and inter-simple sequence repeat (ISSR). Mol Biol Rep 42, 159–166 (2015). https://doi.org/10.1007/s11033-014-3755-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3755-8