Abstract
Despite the association of several miRNAs with bladder cancer, little is known about the miRNAs’ regulatory networks. In this study, we aimed to construct potential networks of bladder-cancer-related miRNAs and their known target genes using miRNA expression profiling and bioinformatics tools and to investigate potential key molecules that might play roles in bladder cancer regulatory networks. Global miRNA expression profiles were obtained using microarray followed by RT-qPCR validation using two randomly selected miRNAs. Known targets of deregulated miRNAs were utilized using DIANA-TarBase database v6.0. The incorporation of deregulated miRNAs and target genes into KEGG pathways were utilized using DIANA-mirPath software. To construct potential miRNA regulatory networks, the overlapping parts of three selected KEGG pathways were visualized by Cytoscape software. We finally gained 19 deregulated miRNAs, including 5 ups- and 14 down regulated in 27 bladder-cancer tissue samples and 8 normal urothelial tissue samples. The enrichment results of deregulated miRNAs and known target genes showed that most pathways were related to cancer or cell signaling pathways. We determined the hub CDK6, BCL2, E2F3, PTEN, MYC, RB, and ERBB3 target genes and hub hsa-let-7c, hsa-miR-195-5p, hsa-miR-141-3p, hsa-miR-26a-5p, hsa-miR-23b-3p, and hsa-miR-125b-5p miRNAs of the constructed networks. These findings provide new insights into the bladder cancer regulatory networks and give us a hypothesis that hsa-let-7c, hsa-miR-195-5p, and hsa-miR-125b-5p, along with CDK4 and CDK6 genes might exist in the same bladder cancer pathway. Particularly, hub miRNAs and genes might be potential biomarkers for bladder cancer clinics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bladder cancer is the 9th most common cancer in the world with 430,000 new cases diagnosed in 2012 and Belgium had the highest rate of bladder cancer in both sexes (17.5 per 100,000), followed by Lebanon (16.6 per 100,000), Malta (15.8 per 100,000) and Turkey (15.2 per 100,000) [1]. In Turkey 10,757 new cases were diagnosed and 4,690 patients died from bladder cancer complications in both sexes in 2012 [1]. Both environmental and genetic factors contribute to carcinogenesis of bladder cancer; and tobacco smoking, occupational exposure to aromatic hydrocarbons, family history of cancer, chemotherapy and radiotherapy are major etiological factors [2, 3].
Micro ribonucleic acids (miRNAs) are short (19–22 nucleotides) non-protein coding RNAs and are found in all eukaryotic cells. They regulate gene expression epigenetically through complementarily binding to the 3′ UTR region of a target mRNA transcript, and resulting in translational repression and gene silencing. More than 60 % of mammalian mRNAs contain conserved regions that serve as targets for miRNAs. In addition, one mRNA can be the target of multiple miRNAs, and each individual miRNA has the capacity to target hundreds of genes, with an average of 500 targets per miRNA [4]. As of its most recent release in June of 2013, over 2,578 mature miRNAs were identified in humans and cataloged in the miRBase 20.0 databases [5].
Accumulating evidence suggests aberrant miRNA expression patterns in most human malignancies, and some highly expressed miRNAs might function as oncogenes by repressing tumor suppressors; conversely, miRNAs expressed at low levels might function as tumor suppressors by negatively regulating oncogenes [6, 7]. Based on reports describing miRNA signatures, several down regulated and up regulated miRNAs have been discovered in bladder cancer [8], [9] and [10]. Some of those miRNAs are thought to be potential biomarkers for bladder cancer in diagnosis and prognosis prediction, as well as a treatment target [11, 12]. Especially current studies mainly focus on miRNAs for providing prognostic information of related cancer [12], [13], and [14].
However, the exact roles of these miRNAs and their target genes in oncogenic pathways have not been clearly elucidated in bladder cancer. Insights into these molecular mechanisms of bladder cancer initiation and progression can provide targets for preventative and therapeutic approaches while providing reliable biomarkers.
In this study, we aimed to predict potential mechanisms of bladder-cancer related miRNAs and target genes by miRNA profiling and bioinformatics tools. By combining miRNA profiling and bioinformatics tools in our study, we sought not only to provide insights into the pathogenesis of bladder cancer but also to discover potential biomarkers for diagnosis, prognosis and treatment.
Matherials and methods
Tissue preparation and RNA isolation
The collection of samples and research protocols were reviewed and approved by the Ethics Committee of Eskisehir Osmangazi University, and the investigation was performed with written informed consent from all the patients. Tissue samples were taken from 30 bladder cancer patients from the central Anatolian part of Turkey who had undergone prospective cystectomy or transurethral resection of BCs, and 9 normal urothelial tissue samples were derived from patients who underwent control cystoscopy. All patients were males ranging from 35 to 65 years old (50.63 + 9.87). Hematoxylin eosin stained sections were examined for tumor cell percentage, and tumor tissues containing more than 85 % tumor cells were selected for microarray analysis.
The tissue samples were deposited in RNAlater solution (Qiagen, USA) and subsequently stored at −20 °C until the RNA extraction. Total RNA including miRNAs was extracted from frozen bladder cancer and normal urothelial tissues using Trizol reagent (Invitrogen, USA) according to the manufacture’s protocol. Total RNA concentrations were measured using a spectrophotometer (NanoDrop Technologies, Wilmington, USA) and quantification analysis of total RNA was performed by a microfluidics-based platform (2100 Bioanalyzer, Agilent, Santa Clara, CA). Three tumor samples and one normal urothelial tissue sample were excluded from conducting miRNA array because of poor RNA quality.
MiRNA expression profiling
After quality assessment remaining 27 tumor samples (12 high grade and 15 low grade; 6 muscle invasive and 21 non-invasive) and 8 normal urothelial tissue samples were loaded onto Agilent’s miRNA arrays with 723 human and 76 viral miRNA represented probes based on Sanger 10.1 (Agilent V 2.0). The input for the miRNA labeling system was 100 ng of total RNA. Following dephosphorylation and denaturation, total RNA was labeled with cyanine 3-pCp (Cy-3) and then hybridized on Human miRNA Microarray (Agilent V 2.0) using the miRNA Complete Labeling and Hyb Kit (Agilent). Each sample was hybridized at 55 °C for 20 h in an agitated hybridization oven at 20 rpm. Then, the slides were washed using the Gene Expression Wash Buffer Kit (Agilent) and scanned by an Agilent Scanner (G2505C). The images were processed and analyzed with Feature Extraction Software (Agilent Technology Ver. 10.1). For genomic profiling, spots were background-corrected using the median foreground minus the median background signal intensities for dye, and log2 ratios were calculated. Following the microarray signal data transformation (linear to log values), a filtration process based on the flag value of spots was performed. Raw microarray data were normalized and analyzed using GeneSpring GX 11.5 (Agilent Technologies, San Francisco, USA). We performed quantile normalization for signals and used the median (50th percentile) method,which normalizes each chip on its median for “per chip” normalization.
Quantitative reverse transcriptase-polymerase chain reaction (qRT-PCR) analysis
Independent RNA samples prepared in parallel with the samples for the miRNA array were used to validate the miRNA array data. In the validation of microarray data, quantitative RT-PCR reactions were performed on randomly selected deregulated miRNAs including hsa-miR-143 and hsa-miR-145. The reverse-transcription reactions were carried out according to the manufacturer’s protocol by using a MiRCURY LNA Universal Reverse Transcriptase kit (Exiqon, Denmark). Initially, reverse transcription was performed with 5 ng/µl total RNA in a 20-µl volume. The assays were heated to 42 °C for 60 min and then 95 °C for 5 min.Before real-time assays, total RNAs were diluted 80 fold in nuclease-free water. Real-time PCR was carried out on a Roche Lifecycler 480 by using SYBR Green master mix and LNA primers of hsa-miR-143, hsa-miR-145, and housekeeping 5S RNA according to the manufacturer’s protocol. The transcription level of 5SRNA was used as an internal control. Then, melting curve analysis was performed to evaluate the specificity of assays. Each sample was analyzed in triplicate.
Bioinformatic analysis
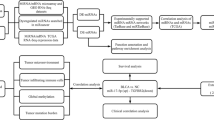
Mature sequences of miRNAs were loaded into the DIANA-Tarbase v 6.0 databases for target analysis [15]. We checked the mature sequences of our deregulated miRNAs from the Mirbase 20.0 databasebefore target analysis. Target genes of deregulated miRNAs were listed using DIANA-TarBase database v 6.0, which include experimentally validated miRNA targets in the literature. To explore the potential function of the whole miRNA and target gene signature, DIANA-mirPath, a web-based computational tool that was developed to identify molecular pathways that are potentially altered by the expression of multiple miRNAs, was used to incorporate miRNAs into KEGG molecular pathways [16]. The pathways were obtained with a p value < 0.05 and gene count >2. The overlapping parts of the “Pathways in cancer (hsa05200)”, “Bladder cancer (hsa05219)”, and “ErbB signaling pathway (hsa04012)” categories of enriched KEGG pathways and related miRNAs were visualized by Cytoscape software [17].
Statistical analysis
All miRNAs in the bladder cancer and normal samples were compared using a t-test to define differentially expressed miRNAs. Multiple testing corrections were done using the Benjamini-Hochberg method [18]. A false discovery rate (FDR) less than 0.05 and absolute log fold change (logFC) greater than 2 were set as the significant cut-offs. The expression levels of the transcripts were evaluated using the comparative CT method (2−∆∆Ct) in qRT-PCR.
Results
Identification of differentially expressed genes
According to the cut-off criteria of |logFC| > 2 and p value < 0.05, we finally gained 19 deregulated miRNAs, including 5 up- and 14 downregulated (Table 1).
Validation of differentially expressed miRNAs using qRT-PCR
Independent RNA samples that were prepared in parallel with the samples for the miRNA array were used to validate the miRNA array data. The expression of randomly selected deregulated miRNAs including hsa-miR-143 and hsa-miR-145 was validated using qRT-PCR (Table 4).
Target genes of miRNAs and their roles in the KEGG pathways
The experimentally validated target genes of deregulated miRNAs were obtained from DIANA-Tarbase v 6.0 databases. The results showthat hsa-miR-30a-5p had the highest number of targets (411), whereas hsa-miR-370 had only 4 targets. Hsa-miR-195-3p, hsa-miR-574-3p, hsa-miR-638, and hsa-miR-193b-3p had no validated targets in the database (Table 2). To gain further insights into the function of deregulated miRNAs and their targets, DIANA-mirPath was applied to identify the significant KEGG pathways. Enrichment results showed that most pathways were related to cancer or cell signaling pathways (Table 3). Most target genes of miRNAs (35) were enriched in the “Pathways in cancer (hsa05200)” category (Table 5). Significantly,13 cancer-associated target genes and 7 related miRNAs were enriched in the “ErbB signaling pathway (hsa04012)” category (Table 5). E2F1, ERBB2, NRAS, RAF1, CDKN2A, KRAS, TP53, CCND1, MMP1, E2F3, MYC, RB1, HRAS, CDKN1A, and VEGFA target genes (Fig. 1) as well as the related hsa-let-7c, hsa-miR-125b-5p, hsa-miR-143-3p, hsa-miR-145-5p, hsa-miR-195-5p, hsa-miR-572, hsa-miR-26a-5p, hsa-miR-23b-3p,and hsa-miR-210 were enriched in the “Bladder cancer (hsa05219)” category (Table 5).
MiRNA regulated gene networks associated with bladder cancer
The “Pathways in cancer (hsa05200)”, “Bladder cancer (hsa05219)”, and “ErbB signaling pathway (hsa04012)” categories from enriched KEGG pathways were selected for further investigation of the miRNA regulatory networks in bladder cancer. All of the interactions of enriched target genes and miRNAs of three pathways are listed in Table 6. Based on these data, the overlapping parts of three pathways were visualized by Cytoscape software (Fig. 2). We determined hub CDK6, BCL2, E2F3, PTEN, MYC, RB, and ERBB3 target genes along with hub hsa-let-7c, hsa-miR-195-5p, hsa-miR-141-3p, hsa-miR-26a-5p, hsa-miR-23b-3p, and hsa-miR-125b-5p miRNAs for these interaction networks.
Cytoscape representation of overlapping parts of three selected KEGG pathways and related miRNAs. Pink circles represent target genes of miRNAs, and green circles represent hub genes. Pink diamonds represent miRNAs, and yellow diamonds represent hub miRNAs. Colored lines represent target genes and miRNA interactions. We used “C”, “B”, and “E” abbreviations for “Pathways in cancer (hsa05200)”, “Bladder cancer (hsa05219)”, “ErbB signaling pathway (hsa04012)”, respectively. Blue lines represent interactions in only C. Black lines represent interactions in only B. Green lines represent interactions in only E. Pink lines represent interactions of overlapping parts of C + B + E. Yellow lines represent interactions of overlapping parts of C + B. Light blue lines represent interactions of overlapping parts of C + E. Red lines represent interactions of overlapping parts of B + E
Discussion
In this study, we analyzed the expression profile in 27 tissue samples of histologically confirmed bladder cancer samples (12 high grade and 15 low grade; 6 muscle invasive and 21 non-invasive) and 8 non-malignant tissue samples of the bladder. A total of 19 miRNAs, including 5 ups- and 14 down regulated, displayed significant differential expression in cancerous tissue compared to non-cancerous tissue.To date, differential expressions of miRNAs in bladder cancer have been published in several studies [19, 20]. Although the results of these studies are not the same, the down regulation of hsa-miR-143, hsa-miR-145, hsa-miR-125b, and hsa-miR-26a and the up regulation of hsa-miR-141 are in agreement with most studies, including our study [19, 20].
Known targets of 19 deregulated miRNAs and their roles in biological processes were demonstrated via KEGG pathway analysis. The KEGG enrichment analysis results in this study confirmed the reliability of our findings, and many of them have been implicated in various cancers and related cell signaling pathways. We considered that the “Pathways in cancer (hsa05200)”, “Bladder cancer (hsa05219)”, and “ErbB signaling pathway (hsa04012)” categories from KEGG pathways were highly related to bladder cancer initiation and progression [21].Studies suggest that more centralized genes in the network are more prone to be key deliverers to proper cellular function than peripheral genes [22, 23]. Therefore; we constructed miRNA regulatory networks based on overlapping parts of these three pathways. Our results showed that CDK6, BCL2, E2F3, PTEN, MYC, RB, and ERBB3 hub genes along with hsa-let-7c, hsa-miR-195-5p, hsa-miR-141-3p, hsa-miR-26a-5p, hsa-miR-23b-3p, and hsa-miR-125b-5p hub miRNAs might play key roles in miRNA regulatory networks in bladder cancer.
We also reviewed some of the literature and reconfirmed the roles of some hub genes in bladder cancer.Bladder cancer has been reported to be associated with loss of function, affecting tumor-suppressor genes such as Tp53, RB, and PTEN [24, 25]. Oncogenes, especially E2F genes (e.g.E2F1and E2F3), which affect the RB pathways, have also been reported to be involved in the progression of this neoplasm [26]. Huang et al. [27] previously reported on the association between hsa-miR-125b and E2F3. Similar to previous studies, we demonstrated that PTEN, RB1, and E2F3 were hub genes in our network, and their regulatory hub miRNAs, including hsa-miR-141-3p, hsa-miR-26a-5p, hsa-miR-23b-3p, and hsa-miR-125b-5p, might play a central role in bladder cancer development in our regulatory networks (Fig. 2). Accumulating evidence suggests that aggressive bladder tumors are associated with the overexpression of some oncogenes [28]. The overexpression of MYC, BCL2, andERBB3 hub oncogenes might be related to the downregulation of hsa-let-7c, hsa-miR-195-5p, hsa-miR-26a-5p, hsa-miR-23b-3p, and hsa-miR-125b-5p in our study. These gene network associations have to be confirmed with further comprehensive functional studies.
The CDK6 gene is a member of the cyclin-dependent protein kinase (CDK) family.This kinase is a catalytic subunit ofthe protein kinase complex that is important for cell cycle G1 phase progression and G1/S transition.This kinase, along with CDK4, has been shown tophosphorylate and thus regulate the activity of tumor-suppressor protein RB. Although increased expression of this oncogene has rarely been reported in bladder cancer, we speculate that CDK6 might have a significant effect on bladder cancer development [29].
Lin et al. [30] reported that CDK4 is a novel target in has-miR-195-mediated cell cycle arrest in bladder cancer cells. As mentioned, CDK4 and CDK6 had similar functions in the cell cycle. In our networks, CDK6 was related to hsa-let-7c, hsa-miR-195-5p, and hsa-miR-125b-5p (Fig. 2). From this point, we speculated that these miRNAs, CDK4, and CDK6 genes might exist in the same bladder cancer pathway. However, the exact regulatory networks remain elusive, and it is hard to estimate the actual false-positive rate of bioinformatic tools. Therefore, further experimental studies should be carried out in order to confirm our results.
Conclusion
Overall, deregulated miRNAs and known target genes might play key roles in the pathogenesis of bladder cancer according to the pathway enrichment analysis. Particularly, hub genes and miRNAs of our constructed network might be central actors of molecular alterations in bladder cancer and candidate biomarkers for diagnostic, prognostic and therapeutic purposes. Of course, further research is needed to confirm their exact roles.
References
Ferlay J, Soerjomataram I, Ervik M, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray, F (2013) GLOBOCAN 2012 v1.0, Cancer incidence and mortality worldwide: IARC cancerbase No. 11 Lyon, France: international agency for research on cancer. Available from: http://globocan.iarc.fr, accessed on 04/06/2014.
Taioli E, Raimondi S (2005) Genetic susceptibility to bladder cancer. Lancet 366:610–612
Clapp RW, Jacobs MM, Loechler EL (2008) Environmental and occupational causes of cancer: new evidence 2005-2007. Rev Environ Health 23:1–37
Garzon R, Marcucci G, Croce CM (2010) Targeting microRNAs in cancer: rationale, strategies and challenges. Nat Rev Drug Discov 9:775–789
Visone R, Croce CM (2009) MiRNAs and cancer. Am J Pathol 174:1131
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Chen YH et al (2013) Characterization of microRNAs expression profiling in one group of Chinese urothelial cell carcinoma identified by Solexa sequencing. Urol Oncol 31:219–227
Dyrskjøt L et al (2009) Genomic profiling of microRNAs in bladder cancer: miR-129 is associated with poor outcome and promotes cell death in vitro. Cancer Res 69:4851–4860
Han Y et al (2011) MicroRNA expression signatures of bladder cancer revealed by deep sequencing. PLoS ONE 6:e18286
Puerta-Gil P, Garcia-Baquero R, Jia AY, Ocana S, Alvarez-Mugica M, Alvarez-Ossorio JL, Cordon-Cardo C, Cava F, Sanchez-Carbayo M (2012) miR-143, miR-222, and miR-452 are useful as tumor stratification and noninvasive diagnostic biomarkers for bladder cancer. Am J Pathol 180:1808–1815
Rosenberg E, Baniel J, Spector Y, Faerman A, Meiri E, Aharonov R, Margel D, Goren Y, Nativ O (2013) Predicting progression of bladder urothelial carcinoma using microRNA expression. BJU Int 112(7):1027–1034
Wang S, Hu J, Zhang D, Li J, Fei Q, Sun Y (2014) Prognostic role of microRNA-31 in various cancers: a meta-analysis.Tumour Biol. [Epub ahead of print]
Fu W, Pang L, Chen Y, Yang L, Zhu J, Wei Y (2014) The MicroRNAs as prognostic biomarkers for survival in esophageal cancer: a meta-analysis. Sci World J. 2014: 523979. Epub 2014 Jul 6. Review.
Vergoulis T, Vlachos IS, Alexiou P, , Georgakilas G, , Maragkakis M, Reczko M, Gerangelos S, Koziris N, Dalamagas T, Hatzigeorgiou AG (2012) Tarbase 6.0: capturing the exponential growth of miRNA targets with experimental support. Nucl Acids Res 40(D1):D222–D229. doi:10.1093/nar/gkr1161
Vlachos IS, Kostoulas N, Vergoulis T, Georgakilas G, Reczko M, Maragkakis M, Paraskevopoulou MD, Prionidis K, Dalamagas T, Hatzigeorgiou AG (2012) DIANA miRPath v.2.0: investigating the combinatorial effect of microRNAs in pathways nucleic acids research (Web server issue)
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B Stat Methodol 57:289–300
Yoshino H, Seki N, Itesako T, Chiyomaru T, Nakagawa M (2013) Enokida H (2013) Aberrant expression of microRNAs in bladder cancer. Nat Rev Urol 10(7):396–404. doi:10.1038/nrurol.2013.113.Epub.Review
Guancial EA, Bellmunt J, Yeh S, Rosenberg JE, Berman DM (2014) The evolving understanding of microRNA in bladder cancer. UrolOncol 32(1):e31–e40. doi:10.1016/j.urolonc.2013.04.014 Epub 2013 Aug 2
http://www.kegg.jp/kegg-bin/search_pathway_text?map=map&keyword=cancer&mode=1&viewImage=true
Abdollahi A, Schwager C, Kleeff J, Esposito I, Domhan S, Peschke P, Hauser K, Hahnfeldt P, Hlatky L, Debus J, Peters JM, Friess H, Folkman J, Huber PE (2007) Transcriptional network governing the angiogenic switch in human pancreatic cancer. Proc Natl Acad Sci USA 104(31):12890–12895 Epub 2007 Jul 24
Horvath S, Zhang B, Carlson M, Lu KV, Zhu S, Felciano RM, Laurance MF, Zhao W, Qi S, Chen Z, Lee Y, Scheck AC, Liau LM, Wu H, Geschwind DH, Febbo PG, Kornblum HI, Cloughesy TF, Nelson SF, Mischel PS (2006) Analysis of oncogenic signaling networks in glioblastoma identifies ASPM as a molecular target. Proc Natl Acad Sci USA 103(46):17402–17407 Epub 2006 Nov 7
Castillo-Martin M, Domingo-Domenech J, Karni-Schmidt O, Matos T, Cordon-Cardo C (2010) Molecular pathways of urothelial development and bladder tumorigenesis. UrolOncol 28(4): 401–8. doi: 10.1016/j.urolonc.2009.04.019. Review.
Sfakianos JP, Lin Gellert L, Maschino A, Gotto GT, Kim PH, Al-Ahmadie H, Bochner BH (2014) The role of PTEN tumor suppressor pathway staining in carcinoma in situ of the bladder. UrolOncol S1078–1439(14):00034–00039. doi:10.1016/j.urolonc.2014.02.003 Epub ahead of print
Olsson AY, Feber A, Edwards S, TePoele R, Giddings I, Merson S, Cooper CS (2007) Role of E2F3 expression in modulating cellular proliferation rate in human bladder and prostate cancer cells. Oncogene 26(7):1028–1037 Epub 2006 Aug 14
Huang L, Luo J, Cai Q, Pan Q, Zeng H, Guo Z, Dong W, Huang J, Lin T (2011) MicroRNA-125b suppresses the development of bladder cancer by targeting E2F3. Int J Cancer 128(8):1758–1769. doi:10.1002/ijc.25509
Wu XR (2005) Urothelial tumorigenesis: a tale of divergent pathways. Nat Rev Cancer 5(9):713–725
Wang G, Zheng L, Yu Z, Liao G, Lu L, Xu R, Zhao Z, Chen G (2012) Increased cyclin-dependent kinase 6 expression in bladder cancer. Oncol Lett 4(1):43–46 Epub 2012 Apr 25
Lin Y, Wu J, Chen H, Mao Y, Liu Y, Mao Q, Yang K, Zheng X, Xie L (2012) Cyclin-dependent kinase 4 is a novel target in micoRNA-195-mediated cell cycle arrest in bladder cancer cells. FEBS Lett 586(4):442–447. doi:10.1016/j.febslet.2012.01.027 Epub
Acknowledgments
This research was supported by the Eskisehir Osmangazi University Research Fund with Grant no. 200811031. We are also thankful to Ece TURKMEN, the Product Manager at SEM Laboratuar Cihazlari Paz. San. Tic. A. S, (Istanbul-Turkey), who provided facilities to carry out this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Canturk, K.M., Ozdemir, M., Can, C. et al. Investigation of key miRNAs and target genes in bladder cancer using miRNA profiling and bioinformatic tools. Mol Biol Rep 41, 8127–8135 (2014). https://doi.org/10.1007/s11033-014-3713-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3713-5