Abstract
The physiological, biochemical and molecular mechanisms regulating the initiation of a regenerative pathway remain partially unknown. Efforts to identify the biological features that confer transformation ability, or the tendency of some cells to induce transgene silencing, would help to improve plant genetic engineering. The objective of our study was to monitor the evolution of plant cell competencies in relation to both in vitro tissue culture regeneration and the genetic transformation properties. We used a simple wheat regeneration procedure as an experimental model for studying the regenerative capacity of plant cells and their receptivity to direct gene transfer over the successive steps of the regenerative pathway. Target gene profiling studies and biochemical assays were conducted to follow some of the mechanisms triggered during the somatic-to-embryogenic transition (i.e. dedifferentiation, cell division activation, redifferentiation) and affecting the accessibility of plant cells to receive and stably express the exogenous DNA introduced by bombardment. Our results seem to indicate that the control of cell-cycle (S-phase) and host defense strategies can be crucial determinants of genetic transformation efficiency. The results from studies conducted at macro-, micro- and molecular scales are then integrated into a holistic approach that addresses the question of tissue culture and transgenesis competencies more broadly. Through this multilevel analysis we try to establish functional links between both regenerative capacity and transformation receptiveness, and thereby to provide a more global and integrated vision of both processes, at the core of defense/adaptive mechanisms and survival, between undifferentiated cell proliferation and organization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For in vitro culture-based plant transformation methods, amenability to the culture stages required for gene transfer, selection and plant regeneration is a major determinant of transformation efficiency, which is controlled by intrinsic and external factors.
Although considerable progress has been made in the past decade, the ability to transform Triticeae species currently still lags behind that of other model plants, such as Arabidopsis thaliana or rice (Oryza sativa L.). Within Triticeae, while stable transformation is relatively efficient in barley (Hordeum vulgare L.), for the other crops, particularly wheat (Triticum aestivum L.), transformation efficiency continues to improve, but the establishment of a high throughput transformation platform and the strong genotype dependence of transformation methods, among others, remain technical hurdles [1–4].
The delivery of DNA into cells competent for in vitro culture regeneration is one of the crucial requirements for achieving genetic engineering of plant cells. Monocotyledonous species have long been described as recalcitrant in terms of in vitro regeneration. Compared with the two other major cereals, rice and maize (Zea mays L.), wheat appeared even more refractory [5]. Embryogenic cultures remain the reliable and main route in regenerating transgenic cereals, regardless of the DNA transfer strategy. Considered as the most responsive explants, immature zygotic embryos (and derived calli) are the usual target tissues for wheat genetic transformation, through either microprojectile or Agrobacterium-mediated DNA delivery [5, 6].
There is growing interest in developing transformation protocols and methods using mature seeds rather than the immature embryos as the starting explants in developing high throughput transformation pipelines, such as those developed for rice. The use of mature embryos as starting material provides a low-cost and time-saving alternative to explants from immature plants [4]. In rice, Agrobacterium-mediated transformation using mature embryo-derived calli is becoming the method of choice for most laboratories, allowing up to several thousand independent lines to be generated for large-scale applications [7, 8].
We had previously developed a simple and effective procedure for wheat, based on mature embryo in vitro culture and biolistic DNA delivery [9, 10]. This method provides an appropriate experimental model for studying the regenerative capacity of plant cells through somatic embryogenesis and their receptivity to exogenous DNA [11].
Only a subset of cells are competent for regeneration but those cells are not necessarily transformable and not every transformed cell can be regenerated into plants [12]. Our previous results demonstrated varying abilities in transient and stable marker gene expression between callus cells derived from an immature zygotic embryo and those derived from a mature one. Whether the recipient tissue variations mentioned in this report [9] could be attributed to their proliferative potential, physiological fate or the possible existence of different dynamics during tissue culture was an interesting issue to address.
In addition, in our model, among several culture periods tested the most suitable physiological state for achieving stable transformation was obtained with a 6 day-pre-culture period [9]. This pivotal period in terms of receptivity to exogenous DNA transfer coincides with a critical stage of the multi-step regenerative pathway. At this time the tissue cultures go through a transition phase between unorganized cell proliferation and differentiation [10]. The developmental switching of somatic cells towards the regenerative pathway involves a plethora of adjustments in response to the in vitro environmental conditions (i.e., a whole reorganization of their physiology, metabolism and morphology), dedifferentiation and the acquisition of a ‘stem cell’-like state that confers pluripotency, requiring remodeling of the gene expression program [13–15].
Our intention in this study was to improve the understanding of the functional links between the regenerative capacity and the transformation receptiveness. On the premise that cells are dynamic systems that go through a sequence of physiological stages in time and that these physiological events might interfere with their transformation receptiveness, we investigated molecular events underlying the responses to in vitro culture and bombardment transformation in tissues proliferating from mature wheat embryos.
Transcriptional profiling has shown that many genes are differentially expressed during somatic embryogenesis (SE) induction or differentiation in cereals. The genome-wide shift in the transcriptome involves multiple cellular pathways interconnecting within an intricate functional network; these pathways include signal transduction cascades, defense, anti-oxidation, programmed cell death/senescence, hormone response/metabolism and cell division [16, 17].
Choosing a candidate-gene strategy, semi-quantitative reverse transcription and polymerase chain reaction (RT-PCR) studies were conducted to follow the expression of genes whose functions were associated with key events or steps in both the expression of totipotency in vitro and the transformation mechanisms. We characterized the transcription profiles for genes for which expression data might provide spatio-temporal indicators relative to: (1) the establishment of plant defense/adaptive mechanisms and antioxidant apparatus dynamics; (2) cell adjustment to changing environmental conditions and reprogramming of genome expression; and (3) cell division activation and a proliferative state, competent when it comes to the expression of totipotency, as well as for gene transfer and transgene integration [11].
Genes investigated experimentally are associated with: (1) response to stress (glutathione S-transferase, GST), (2) transcriptional regulation (MADS-domain transcription factor, MADS-box), (3) cell proliferation and S phase-enriched cell populations (replication factor complex, RFC, known to be associated with S phase, i.e. DNA synthesis phase of the cell cycle) and (4) acquisition/maintenance of embryogenic competence (somatic embryogenesis receptor kinase, SERK). In addition, the involvement of germin oxalate oxidases (GerOXOs) in stress response, developmental reprogramming and differentiation (including through SE and microsporogenesis), reactive oxygen species (ROS) (H2O2) and calcium release, cell wall remodeling and reinforcement, and defense strategy in Poaceae makes this family particular attractive [18–22]. This gene family might provide an appropriate spatio-temporal molecular signature of the successive stages of both competencies in cereals.
The results from the transcript profilings and enzymatic oxalate oxidase (OXO) activity are discussed in relation to tissue culture response and transformation receptiveness. Finally, macroscopic, microscopic and molecular data were summarized from a holistic perspective and integrated with experimental data and theoretical aspects in the literature in order to address the question of tissue culture and transgenesis more broadly.
Any attempt to learn more about the events triggered during the somatic-to-embryogenic transition would help to increase understanding of totipotency and plant ontogenesis flexibility. Understanding the biological features that confer transformation ability, or the differences in the tendency of some cells to bring about transgene silencing, is desirable for a rational approach of plant genetic engineering. From a practical perspective in wheat transformation, molecular and biochemical signatures of the main steps in the expression of totipotency and receptivity to transgenesis would be useful tools, whatever the regenerative pathway or the gene transfer method.
Materials and methods
Plant material
The donor plants (T. aestivum L.) of the winter cultivar Dream were grown in the field. Harvested mature seeds were stored at room temperature.
Seed surface-sterilization and aseptic embryo isolation
Wheat caryopses were fungicide treated (Sibutol ® Bayer) against seed-borne pathogenic fungi at least 8 days before culture initiation, after which seeds were kept in a flask for several weeks for subsequent use. After a 16-h rehydration in sterile water at room temperature, the seeds were surface-sterilized with 70 % ethanol for 2 min, soaked in 8 % calcium hypochlorite containing 0.1 % Tween 80 (Merck) for 10 min and rinsed three times with sterile deionized water. The embryos were aseptically removed and protected from desiccation.
Tissue culture
Thin tissue fragments (≈500 fragments resulting from crushing 100 embryos) were re-suspended in 4 ml of liquid basal medium (i.e., MS medium, supplemented with 100 mg/l casein hydrolysate [Sigma] and 20 g/l sucrose [Merck] as the carbon source, pH adjusted to 5.8 with NaOH prior to autoclaving). The resulting suspension was distributed among five 9-cm disposable plastic Petri dishes containing the callus induction medium (i.e., the liquid basal medium containing 2 mg/l filter-sterilized 2,4-D (Sigma) as the growth regulator, semi-solidified with 7 % agar [Invitrogen]. Excess liquid medium was retrieved and discarded before sealing the dishes. Light and temperature conditions for callus culture and plant regeneration were as described earlier [9].
Molecular analysis
Sampling
Nine specimens were used for the RT-PCR experiments. The sampling consisted of five culture periods (2, 6, 8, 14 and 20 days) and two morphogenetic types (embryogenic and non-embryogenic that were distinguishable after a 6-day period of tissue culture).
Total RNA extraction
RNA was extracted in at least three independent replications. The tissues (T. aestivum L. Dream cv) (500 mg) were frozen in liquid nitrogen and disrupted using a mortar and pestle. Total RNA was extracted (‘Pure Script RNA isolation Kit®’ GENTRA SYSTEMS for GST and MADS-box analysis, ‘TRI REAGENT®’ FERMENTAS for RFc, SERK and GerOXO analysis) and treated with DNase I (‘Dnase I RNase free®’ ROCHE) to remove any residual genomic DNA, according to the manufacturer’s instructions. RNA yield was quantified by spectrophotometry and RNA integrity was controlled by separation on a 1.2 % agarose gel electrophoresis, each sample containing 0.5 mg/ml ethidium bromide. The RNA quantity, quality and integrity were assessed by agarose gel electrophoresis and by evaluating the 28S and 18S ribosomal RNA bands.
Semi-quantitative one-step RT-PCR
Sequence specific primers were designed for each gene as following. For 18SrRNA, GST and RFC, primers previously used in cited papers (Table 1) were selected for our study. For MADS-box, SERK and GerOXO, primers were designed from published sequences (Table 1) using Primer Premier Software (version 5.0). In all cases, a single amplicon of the expected size was detected.
Reverse transcription and cDNA amplification were performed in a single tube experiment using ‘Ready-To-Go™ RT-PCR Beads’ (GE HEALTHCARE, formerly Amersham Biosciences), according to the instruction manual protocol.
Briefly, first strand cDNAs was synthesized from 200 μg of total RNA using an oligo dT as the first-strand primer and the Moloney murine leukemia virus reverse transcriptase at 42 °C for 20 min. The enzyme and RNA:cDNA heteroduplex were denatured at 95 °C for 5 min. Sequence specific primers were then added to the reaction mixtures for the selective amplification of the cDNAs used as templates (Table 1). The PCR reactions were performed under the following conditions: 1 min denaturation at 95 °C, 1 min annealing, 2 min polymerization at 72 °C and a 10 min end-elongation step at 72 °C. Specific conditions (annealing temperature-cycle numbers) were 56 °C-30, 60 °C-35, 58 °C-35, 58 °C-35, 56 °C-35 and 56 °C-30, for 18SrRNA, GST, MADS-box, RFC, SERK and GerOXO analysis, respectively.
The amplified product sizes were confirmed by separation on a 1.2 % agarose gel electrophoresis. Ethidium bromide-stained signals were digitalized and quantified by the Quantity One® 4.2.1software package (BIORAD Laboratories) and expressed as total pixel values. The amount of transcripts was normalized to the amount of the housekeeping transcript, 18sRNA (i.e., normalized results were determined as ratios of the total pixel value of the band of interest to the total pixel value of the 18S rRNA band). For each transcript, the final relative abundance was expressed as a percentage of the highest value among the set of observations (i.e., nine samples: 2, 6 days embryogenic, 6 days non-embryogenic, 8 days embryogenic, 8 days non-embryogenic, 14 days embryogenic, 14 days non-embryogenic, 20 days embryogenic and 20 days non-embryogenic). All the molecular analyses were performed at least three times.
Statistical analysis
Calculated variables were normalized (arcsinrac[x] variable transformation) and the data were analyzed by analysis of variance (ANOVA). Mean differences were compared pair-wise with the Tukey Multiple comparison procedure, at a confidence interval of 95 % (Systat for Windows, Version 8, SYSTAT Software Inc.).
Histochemical detection of OXO activity
Oxalate oxidase activity in situ was detected as described by Caliskan and Cuming [23] using the procedure reported by Dumas et al. [24]. The tissues were incubated with an assay buffer (25 mM succinic acid, 3.5 mM EDTA, 2.5 mM oxalic acid, 0.6 mg/ml 4-chloro-1-naphthol, pH 4.0) at 25 °C in night darkness, and were fixed in 4 % paraformaldehyde in phosphate buffered saline (PBS). The control samples were incubated in assay buffer lacking oxalic acid.
Results
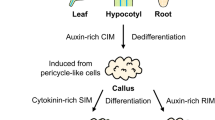
We had previously developed a simple, straightforward regeneration protocol based on mature zygotic embryos as starting material, rather than immature material [10]. This alternative protocol relies on 2,4-D-induced regenerable calli from mature embryos reduced in tissue fragments (Fig. 1). Histological studies were conducted in order to determine the morphogenetic pathway and its tissular origin. The identification of the major events along the SE pathway leading to the genesis of plantlets, the localization of the origin of the process and the identification of the most favorable tissue environment are discussed elsewhere (manuscript submitted). We used this simple and well-characterized wheat regeneration procedure as an experimental model for studying the tissue culture and transformation properties.
Wheat mature embryo culture, sampling for RNA extraction and relative quantification of target gene transcripts. Samples were taken at regular periods after calli induction (red arrows, pointing upwards) from tissues displaying distinct behavior (i.e., embryogenic vs. non-embryogenic, and lesser vs. optimal ability for long-term transformation). The embryogenic character becomes apparent after 6 days of culture; this period also coincides with the most suitable physiological state for bombardment DNA introduction and long-term transgene expression in calli [9, 10]. This 6 day-culture period corresponds to a pivotal developmental window. The two types of calli with distinct morphogenetic capacities are illustrated in this figure (on the right, 7-day-old calli). Calli termed ‘embryogenic’ (above) are white to greenish-white and compact, with smooth-looking structures formed by densely proliferating cells. These calli tend to engage in the SE differentiation pathway, subsequently confirmed by cell cluster greening (i.e., chloroplast differentiation) and followed by the emergence of shoots (i.e., somatic embryo germination). The calli considered to be ‘non-embryogenic’ (below) are white, limpid, watery, friable and formed by large highly vacuolated cells. Using semi-quantitative RT-PCR and gel analysis, the 18S ribosomal RNA gene was used as the control for normalization and relative quantification of target gene RNA abundance. 2, 6, 8, 14, and 20 number of days after 2,4-D culture initiation; the star marks the period corresponding to embryogenic character expression and optimal transgenesis ability. E embryogenic calli, NE non-embryogenic calli
Transcription profiling of target genes and enzymatic assays were conducted to gain better understanding of key events underlying the early processes involved in plant neoformation when wheat mature zygotic embryo fragments are 2.4-D induced in culture. We also sought to gain a better understanding of transformation receptiveness: by tracking the physiological evolution of somatic cells during the acquisition of embryogenic competence, we investigated jointly the reasons for the fluctuations in the transformation ability of those cells.
Competence in callogenesis (undifferentiated cellular proliferation, Fig. 1, ‘non-embryogenic callus’) and in embryogenesis (white to green-white, compact and smooth, plane or dome-shaped or nodular appearance calli formed with densely proliferating cells, Fig. 1, ‘embryogenic callus’) was identified. Those calli termed as ‘embryogenic’ tended to engage in differentiation, confirmed by cell cluster greening thereafter.
RNA was extracted from tissues displaying distinct behavior (i.e., embryogenic vs. non-embryogenic) and after varying culture periods (i.e., 2, 6, 8, 14 and 20 days), the latter producing different responses to genetic transformation, on the premise that the 6-day culture is a pivotal period (when the embryogenic character becomes apparent macroscopically, and this coincides with the most suitable physiological state for the particle bombardment-mediated DNA introduction and long-term transgene expression) [9] (Fig. 1).
Semi-quantitative RT-PCR (Fig. 2, 3, 4, 5, 6) was used to determine the relative levels of expression of genes involved in adaptive response to stress (GST), the transcriptional regulation of cellular processes and organ development program (MADS-box), cell proliferation (RFC large subunit), the transition of the somatic cells into an embryonic pattern of differentiation (SERK), the acquisition/maintenance of embryogenic competence and defense (GerOXO). The specific primers are listed in Table 1.
GST transcription profiles—relative abundance of transcripts and representative gels showing their RT-PCR amplification rate. Restricted temporal analysis of the relative transcription of individual GST genes (TtGSTU1, TtGSTU2 and TtGSTU3) (a), and complete temporal analysis of the TtGSTU1 (b, c, d); fluctuations in the relative level of transcripts over time and depending on the different tissue types (b), average values across the culture duration (c) and the morphogenic character displayed by tissues (d). Bars represent mean values ± SDM (standard deviation of the mean) from at least three independent experiments. Data were analyzed by ANOVA (SYSTAT 8.0) following arcsinrac(x) variable transformation. Means were compared using the Tukey test. Data with different letters are significantly different at p ≤ 0.05. 2, 6, 8, 14, and 20 number of days after 2,4-D culture initiation, E embryogenic tissues, N non-embryogenic tissues
MADS-domain transcription factor, MADS-box transcription profile—relative abundance of transcripts and representative gels showing their RT-PCR amplification rate. Fluctuations in the relative level of transcripts over time and depending on the tissue types (a), average values across the culture duration (b) and the morphogenic character displayed by tissues (c). Bars represent mean values ± SDM from at least three independent experiments. Data were analyzed by ANOVA (SYSTAT 8.0) following arcsinrac(x) variable transformation. Means were compared using the Tukey test. Data with different letters are significantly different at p ≤ 0.05. 2, 6, 8, 14, and 20 number of days after 2,4-D culture initiation, E embryogenic tissues, N non-embryogenic tissues
Replication factor C large subunit, Rfc-1 transcription profile—relative abundance of transcripts and representative gels showing their RT-PCR amplification rate. Fluctuations in the relative level of transcripts over time and depending on the tissue types (a), average values across the culture duration (b) and the morphogenic character displayed by tissues (c). Bars represent mean values ± SDM from at least three independent experiments. Data were analyzed by ANOVA (SYSTAT 8.0) following arcsinrac(x) variable transformation. Means were compared using the Tukey test. Data with different letters are significantly different at p ≤ 0.05. 2, 6, 8, 14, and 20 number of days after 2,4-D culture initiation, E embryogenic tissues, N non-embryogenic tissues
Somatic embryogenesis receptor kinase, TaSERK1 transcription profile—relative abundance of transcripts and representative gels showing their RT-PCR amplification rate. Fluctuations in the relative level of transcripts over time and depending on the tissue types (a), average values across the culture duration (b) and the morphogenic character displayed by tissues (c). Bars represent mean values ± SDM from at least three independent experiments. Data were analyzed by ANOVA (SYSTAT 8.0) following arcsinrac(x) variable transformation. Means were compared using the Tukey test. Data with different letters are significantly different at p ≤ 0.05. 2, 6, 8, 14, and 20 number of days after 2,4-D culture initiation, E embryogenic tissues, N non-embryogenic tissues
Germin oxalate oxidase, GerOXO transcription and enzymatic profiles—relative abundance of transcripts and representative gels, histochemical detection of OXO activity. Fluctuations in the relative level of transcripts over time and depending on the tissue types (a), average values across the culture duration (b) and the morphogenic character displayed by tissues (c). Fluctuation in the OXO activity over time and depending on the tissue types (d). Bars represent mean values ± SDM from at least three independent experiments. Data were analyzed by ANOVA (SYSTAT 8.0) following arcsinrac(x) variable transformation. Means were compared using the Tukey test. Data with different letters are significantly different at p ≤ 0.05. 2, 6, 8, 14, and 20 number of days after 2,4-D culture initiation, E embryogenic tissues, N non-embryogenic tissues
Glutathione S-transferases (GSTs)
The differential transcription of three members of the GST tau wheat family [25] was considered in this study: the two tandemly repeated GST genes, TtGSTU2 and TtGSTU1 [GenBank: AY013753] and the related GST-like sequence, TtGSTU3 [GenBank: AY013754].
As confirmed by RT-PCR analysis, each specific primer pair allowed each T. aestivum individual GST sequence to be amplified, and the amplified fragments were of the expected sizes. A higher inducible expression of the TtGSTU1 gene was observed (Fig. 2a).
An early transcriptional response of the cells submitted to 2,4-D in vitro induction was illustrated by the TtGSTU1 gene (Fig. 2). In the detailed analysis (temporal and tissue specific measures of mRNA abundance), the level was significantly higher early in the culture (i.e., day 2, p = 0.03) (Fig. 2b). In the temporal analysis (values averaged, whatever the morphogenetic character of the tissues), this level was more than 1.9 times greater at 2 days than during the rest of the investigated period (p = 0.03). The term of the temporal analysis was marked by a clear decrease (Fig. 2c). In the embryogenic versus non-embryogenic analysis (on average, covering the whole investigated period), the mRNA level of this member of the wheat GST family did not differ significantly (p ≤ 0.05) (Fig. 2d).
MADS-box transcription factor (MADS-box)
PCR with degenerate primers was used for cloning a wheat MADS-box cDNA (T. aestivum L.) putatively homologous to the maize ZmMADS1 transcription factor, using RNA extracted from embryogenic cultures as the template (paper in preparation). The partial wheat cDNA (113 bp, see Additional file 1 in Electronic Supplementary Material) showed 100 % homology to the T. aestivum TaMADS#12 mRNA for MADS box transcription factor [GenBank: AB007505] [26] and encompassed the highly conserved sequence motif MADS (DNA binding domain), typically found in this transcription factor family.
This MADS-box gene produced a transcription profile similar to that of the TtGSTU1 gene (Figs. 2, 3). In the detailed analysis, the mRNA level was significantly higher 2 days after 2,4-D culture initiation (p ≤ 0.05) (Fig. 3a). In the temporal analysis, this level was 1.7 times higher, on average, in the 2 day-old culture than during the rest of the investigated period (p = 0.03). A progressive decrease was observed temporally (Fig. 3b). The embryogenic versus non-embryogenic analysis revealed no difference between the studied tissue types (Fig. 3c).
Replication factor C (Rfc-1)
In rice the five subunits that make up the RFC (OsRFC) were expressed strongly in proliferating tissues [27].
A search in the public database of the National Centre for Biotechnology Information (NCBI, http://www.ncbi.nim.nih.gov), with the five subunits of the OsRFC ([GenBank: AB109200, GenBank: AB045677, GenBank: AB038319, GenBank: AB066661, GenBank: AB109201] corresponding to OsRFC1, 2, 3, 4, 5, respectively) as the query entries, produced only one homologous sequence in wheat, encoding the large subunit replication factor C (Rfc-1 gene, [GenBank: AJ318783]), which shares 92 and 89 % sequence identities (at the nucleic acid and amino acid level, respectively) with OsRFc1. This wheat Rfc-1 gene has been shown to be governed by cell-cycle regulation, with mRNA accumulating in cell populations enriched for early S-phase [28].
In the detailed analysis, using primers specific for Rfc-1 sequence, a significantly higher mRNA level than for all other samples was observed on day 6 in the samples identified as embryogenic (p ≤ 0.05) (Fig. 4a). In the temporal analysis, a peak was observed in samples collected on day 6, followed by a progressive reduction (Fig. 4b). Lastly, in the embryogenic versus non-embryogenic analysis, a significantly higher level marked the tissues that were embryogenic (p = 0.03) (Fig. 4c).
Somatic embryogenesis receptor kinase (SERK)
At the time that this study was initiated, no gene had been found in searches for sequences annotated as ‘SERK’ or ‘SERK homologous’ in wheat (T. aestivum) using the database of the NCBI. The functional characterization of homologous SERKs in Poaceae had been reported in maize [29].
A search in the public database using the coding sequences of Z. mays SERK1, SERK2 and SERK3 as query entries [GenBank: AJ277702, GenBank: AJ277703, GenBank: AJ400870] retrieved two wheat clones, full mRNA sequences [GenBank: BT009426, GenBank: BT009223], with the former sharing 83 % sequence identity with ZmSERK1 and ZmSERK2, 74 % with DcSERK [GenBank: U93048] and 73 % with AtSERK1 [GenBank: A67827], at the nucleotide level.
This clone [GenBank: BT009426] has since been assigned as T. aestivum TaSERK1. Its expression has been showed to be auxin inducible and to correlate with the initiation of SE in wheat leaf base cultures [30].
In the detailed analysis, a noticeably higher abundance of TaSERK1 mRNA was observed on day 8 in tissues identified as embryogenic (p = 0.02) (Fig. 5a). In the temporal analysis, on average this level peaked significantly in samples collected 8 days after the 2,4-D in vitro induction (p = 0.02) (Fig. 5b). In the embryogenic versus non-embryogenic analysis, the tissues involved in an embryogenic differentiation process showed a greater tendency to express the TaSERK1 gene (20 % more), although this difference did not appear statistically significant at p ≤ 0.05 (Fig. 5c).
Germin oxalate oxidase (GerOXO)
In the detailed analysis, the level of GerOXO messengers appeared to fluctuate, reaching a maximum value for embryogenic tissues after 6 days (Fig. 6a). In the temporal analysis, their abundance increased over time, marking a significant peak day 6 (p ≤ 0.05), after which the mRNA content gradually decreased (Fig. 6b). In the embryogenic versus non-embryogenic analysis, despite a high rate of expression of GerOXO in embryogenic tissues, the difference (30 % more) did not appear to be significant at p ≤ 0.05 (Fig. 6c).
Oxalate oxidase (OXO) enzymatic activity
The fundamental nature of the germin family, which is involved in both adaptability and developmental processes, including through SE [21, 31, 32], motivated us to conduct a deeper analysis. Within the wheat germin family, the member considered in this study has received considerable attention and was the object of numerous characterization studies, in both homologous and heterologous expression systems. A clear correlation has been demonstrated between the level of transcription, the synthesis of the protein and the level of its enzymatic activity when this gene was expressed either in wheat [19, 23, 33] or in other plant species (e.g., [34–38]). Taking advantage of the ease of identifying the presence of the corresponding proteins via analysis of their OXO enzymatic function (i.e., readily detectable through an oxalate-dependent histochemical detection of its reaction product (H2O2) fuelling a peroxidase-linked colorimetric reaction [24, 39]), OXO activity was temporally monitored in 2,4-D induced calli from mature embryo fragments, in tissues growing after 2, 6, 8, 10 and 14 days of culture, either embryogenic or non-embryogenic (Fig. 6d).
Temporally, significant activity observed in the 2-day old calli (the dark color detected) was followed by a transient decrease on day 6 and a peak on day 8 in the tissues engaged in the embryogenic differentiation process. Subsequently, there was a gradual decrease (Fig. 6d, upper series of pictures). These temporal changes tracked through the enzymatic assays accorded with the patterns in the gene transcript levels, but with the kinetics of OXO activity following that of the accumulation of messengers (Fig. 6b). The important enzymatic activity visible in the most responsive 2-day old calli suggests a substantial induction of transcript synthesis during the previous period.
When comparing the embryogenic versus non-embryogenic tissues (Fig. 6d, the upper vs. the bottom series of pictures), specific changes tracked through enzymatic assays confirmed the differential mRNA levels previously recorded (i.e., transcripts were most abundant in embryogenic tissues, Fig. 6c). Embryogenic tissues showed a biphasic colorimetric reaction over time (Fig. 6d, upper series of pictures), but this was always clearly paler in non-embryogenic tissues (Fig. 6d, bottom series of pictures).
In order to complete our analysis of OXO activity, an enzymatic assay was conducted during the mature zygotic embryo culture by applying the same protocol to whole embryos rather than fragmented tissues (Fig. 7). Imbibing seed water alone triggered OXO activity in some embryo tissues, but a synergic effect was observed when these embryos were 2,4-D induced after a 16-h rehydration (i.e., as described in our in vitro culture procedure [9]). After a 2-day 2,4-D treatment, the visible dark color that indicated important enzymatic activity had extended to the remaining tissues.
Wheat caryopsis imbibition and entire embryo culture—histochemical detection of OXO activity. Enzymatic activity after increasing the duration of seed imbibition (a), and when mature embryos were 2,4-D induced after a 16 h-rehydration (b). Upper alignment of pictures, embryo scutellum side up; lower alignment, scutellum side down. h hour, d day
Discussion
To gain a better understanding of the molecular events involved in the acquisition of competence for gene transfer and tissue culture, target gene expression patterns were profiled.
The GST, MADS-box, RFC, SERK and GerOXO gene families were considered promising for learning more about the chronology of the critical cellular processes, such as the adaptation to the in vitro context, regulation of the cellular fate, induction of cell division, and their commitment in the embryogenic pattern of development.
Tissue samples were randomly collected at each of five time points after the 2,4-D induction (day 0): (i) one time point during cell proliferation initiation and callus induction (day 2); (ii) two time points during callus development and around the acquisition of the embryogenic character and transformation ability (day 6 and day 8); and (iii) two time points during further differentiation and morphogenetic development (day 14 and day 20). Where possible, the tissues visually engaged in the process of embryogenesis were analyzed separately from those that were not (after day 6, a crucial period in terms of both the macroscopic manifestation of the embryogenic character and the physiological receptivity to the introduction of transgenes) (Fig. 1).
In vitro culture as stress signal (GSTs)
A profusion of physiological and metabolic adjustments occur in response to a multitude of stress factors of a physical and chemical nature when plant tissues are subjected to in vitro culture conditions [13].
“Somatic embryogenesis is the process by which somatic cells, under induction conditions, generate embryogenic cells, which go through a series of morphological and biochemical changes that result in the formation of a somatic embryo” [40]. Starting with de-differentiation, SE is a multi-step, complex and highly regulated process that occurs as part of natural plant development in vivo (e.g., apomixis, a form of asexual reproduction, [41]) or is achievable in plant tissue culture [14]. The ability of several somatic cell types to engage in embryogenic development illustrates their adaptive response to the in vitro environmental constraints. This response includes the re-programming of gene expression in parallel with stress signaling cascade activation and defense response induction [13].
Gene expression analyses at both the proteome and transcriptome levels have led to the identification and characterization of many stress-related genes and proteins associated with the early stage of SE [42, 43]. Belonging to the antioxidant enzyme system, GSTs (EC 2.5.1.18) form a superfamily of multifunctional proteins, playing an important role in the overall natural defense mechanisms in all living organisms, whose expression is induced after exposure to a multitude of stresses (for review, see [44]).
Plant GSTs respond to a wide range of agents and conditions in which increased level of ROS production is a common factor (reviewed by [45]). Biotic and abiotic stresses (pathogens, natural and synthetic toxins, heavy metals, heat shock), ROS themselves and phytohormones activate the transcription of individual GST genes differentially [46].
Beyond their functions associated with detoxification and other stress responses in plants, recent studies show that several GSTs also play a role in normal plant growth and development in vivo, and in regeneration in vitro [22, 47, 48]. Lastly, in the interacting molecular network triggered very early (the first 24 h) during the auxin-induction phase in wheat leaf base segments, the GST protein family emerged as one of the three major nodes [16].
The GSTs of the Phi and Tau classes are plant-specific and are the most abundant subclasses in plant tissues. Besides their well-known conjugating activity to exogenous compounds both classes possess GSH-dependent peroxidase activity (GPOX) which uses GSH to reduce organic hydroperoxides. Also observed for several wheat tau GSTs [49] the latter activity is of particular significance for the prevention of hydroperoxide-induced damages (reviewed by [45, 50, 51]). The primers we used were designed to selectively amplify three members of the GST Tau wheat family (i.e., the two highly homologous genes, TtGSTU1 and TtGSTU2, and the GST-like sequence named TtGSTU3) for expression studies in Triticum species in terms of their biochemical function in herbicide metabolism [25].
In Xu et al. [25] gene-specific semi-quantitative RT-PCR analyses, the tandemly duplicated TtGSTU1 and TtGSTU2 genes (but not the TtGSTU3 sequence) were induced in root and shoot tissues by the synthetic auxin 2,4-D. TtGSTU1 expression was always higher than that of TtGSTU2. Both TtGSTU1 and TtGSTU2 proteins were thought to have important roles in xenobiotic metabolism in wheat, and might also have significant yet undefined roles in response to plant stresses [25].
In a pilot assay, we initially assessed the selective amplification of each individual transcript for two-sample tests (i.e., two time points, days 6 and 20) (Fig. 2a). In our results we recorded the amplification of the TtGSTU3 sequence in embryo tissue cultures. The expression of this gene had not previously been detected by Xu et al. [25] whose analyses were performed on shoot and root tissues. The TtGSTU1 transcripts were more abundant, as observed earlier [25], and we followed the transcription profile of the latter more closely (Fig. 2b–d).
Unique combinations of often multiple interactive signaling pathways (i.e. various phytohormones, ROS or antioxidants) produce the distinct transcriptional activation patterns of individual GSTs during stress (reviewed by [46]). An early response of wheat cells submitted to 2,4-D in vitro induction was illustrated by a quick transcriptional activation of the TtGSTU1 (Fig. 2b, c), reflecting the role of this gene superfamily in the immediate response of plants to environmental alterations and constraints.
TtGSTU1 expression could be triggered either through an overall response to stressful in vitro culture conditions, associated with a global deterioration of the environment (i.e., via the central role of GSTs in antioxidant functions and defense mechanisms), or through a 2,4-D specific induction as a growth regulatory substance (putative ABA-, ethylene- and, in particular, auxin-responsive regulatory elements were identified in the promoters of those GST genes [25]).
TtGSTU1 messengers were the most abundant when cell proliferation was initiated (2 days, Fig. 8). Cell proliferation and active aerobic metabolism occurring upon auxin SE induction is associated with a burst of reactive oxygen species, including hydrogen peroxide [52, 53].
Events surrounding the acquisition of competence in tissue culture and genetic transformation: an integrated approach of data and concepts at the tissular, cellular and molecular levels. With wheat as the observed biological system, this model summarizes data and concepts aimed at providing an analytic-to-holistic view of the regenerative property of plant cells and their receptivity to direct gene transfer. Cells are complex and dynamic systems that go through a sequence of physiological stages along the regenerative pathway. The molecular signatures of the responses to stress/auxin and the release of embryogenic development are integrated with the series of microscopic/macroscopic observations, together with the cellular processes that might interfere with transformation ability over time (time scale in days, −1 to 12, relative to embryo culture initiation). In particular, the germin oxalate oxidase gene family provides an appropriate spatio-temporal molecular signature of the successive stages of both competencies in cereals. (1) Both the higher level of intracellular H2O2 and the activity of cellular antioxidant machinery have been shown to be crucial for cell division and the expression of totipotency [161–163, 196, 197]. (2) An initial oxidative signaling (a rapid and transient production of huge amounts of ROS) induces antioxidant defenses. Notably, belonging to the antioxidant apparatus, plant GSTs, with GSH, play a central protective role in counteracting oxidative stress [198], sensing and maintaining redox homeostasis [199]. A growing number of studies recognize the importance of the redox environment for growth and development, and of the GSTs link with the early developmental phases of SE (reviewed in [200]), making GST transcript abundance an early signal for the identification of embryogenic cultures. (3) An acclimation response is complex and occurs in different phases. Immediately after the cellular changes that occur as a result of stress exposure, defense processes are triggered; the concomitant activation of a set of specific genes is indicative of the adaptive behavior of cells adapt [201]. Reprogramming the gene expression pattern is achieved by regulation at the transcriptional level, as could be reflected by the activation of the MADS-box TF, probably involved in the integration of environmental and developmental cues. (4) Re-entry into the cell cycle plays a crucial role in the expression of cellular totipotency and in the transformation competent state [11, 92]. Strongly expressed in proliferating tissues, an increased Rfc transcript synthesis is on average correlated with the highest response to embryogenesis, as expected. But this critical component of the DNA replication machinery is cell-cycle regulated and was also used as a marker of S phase-enriched cell populations in our study [28]: the Rfc transcript level marks a sharp increase in the 6-day-old proliferating cells that are the most responsive to receiving and stably expressing transgenes [9]. Histological observations show that the physiological state reached at day 6 corresponds to a pivotal developmental window between undifferentiated cell proliferation and organization, constituting a transition phase between two different dynamic regimes. (5) First discovered as a marker of the transition of the somatic cells into an embryonic pattern of differentiation, SERKs were shown to confer embryogenic competence. As previously observed [104], SERK maximal transcription occurred at the globular stage of SE differentiation. The SERK homologue considered in this study is auxin inducible and calcium dependent. It is expressed rapidly and long before the first appearance of somatic embryos [30]. (6) All along the steps of the regenerative process, among the stress responses, the accumulation of germin OXOs appears to be of particular interest. Involved in SE, these defense-related proteins are H2O2 inducible, and they themselves produce Ca2+ and H2O2 from oxalate under acidic conditions. The release of Ca2+ and H2O2 has long been considered as the biochemically important result of their OXO activity. Both these molecules play key roles in cell elongation and division, in relaying a stress signal, in cell wall reinforcement and in establishing a second-wave long-lasting defense (see “Discussion” section). After the transient first burst that occurs immediately after an abiotic stimulus, a second oxidative burst of H2O2 resulting from OXO activity would be involved in long-lasting adaptive behavior and constitutive defense, as observed earlier in an another member of the Poaceae family [149, 152]. H2O2 and Ca2+ are known to be interconnected signaling molecules that are essential in the multiple pathways of plant innate immunity [192]. During the Agrobacterium transformation process, successful DNA transfer is possible due to the ability of Agrobacterium to hijack fundamental cellular processes and to interfere with plant immunity pathways in order to counteract host defense [181, 182]. According to the temporal–spatial concept of ROS wave signaling [178], OXOs appear as attractive candidates in propagating the ROS wave throughout the cells, carrying the signal in time and in feeding the cascade of cell-to-cell communication events and over time
This result could also be interpreted in terms of the role of the synthetic auxin 2,4-D itself, which is still a matter of investigation: it is a key substance for efficiently inducing embryogenic competence (i.e., as a growth regulator that mimics auxins, with a concentration-dependent mode of action, triggering a specific developmental adaptive response), but is also known for its herbicide and therefore phytotoxic activity, and possibly acts as an individual stress factor (i.e., the physical or chemical stress factors by themselves are known to be SE inductors) (reviewed by [13, 54]).
Lastly, in contrast to Singla et al. [16], the transcripts were identically induced in tissues, whether embryogenic or not (Fig. 2b, d); this suggests that TtGSTU1 was a protein acting at the very early phase during the auxin induction in this study, with wheat mature embryo fragments as a culture system.
Transcriptional regulatory network: release of the embryogenic program (MADS-box)
During the somatic-to-embryogenic transition period, chromatin remodeling is implicated in the coordinated reorganization of the cellular state [13, 55]. Dynamic changes in chromatin structure are central for the reversion of the differentiation process, leading to a resetting of the gene expression program and activation of silent genes, and the release of the embryogenic program otherwise repressed in vegetative plant cells [55–57].
Signal transduction cascades are still largely unexplored, but the presumed increased activity of the various signaling pathways is assumed to be associated with a differential expression of many families of transcription factors (TFs).
TFs are master regulators of developmental patterns, in both animals and plants [58–60], notably in regulating different aspects of embryonic processes in higher plants [61–64].
The MADS group members include many examples where the change in activity of a single TF has been shown to have a profound effect on an important aspect of plant biology [65–71].
Among them, MIKC-type proteins are plant specific and the most characterized MADS group in plants [69, 72]. In dicot species, the AGAMOUS-like 15 (AGL15) gene is highly expressed only in cells that take on an embryonic identity, whatever the embryo origin (i.e., zygotic, somatic, apomictic) [63, 73]; and its constitutive expression promotes somatic embryo development [74, 75].
There is little information about the implication of the members of this very large MADS-box gene family in the in vitro embryogenic process in Poaceae. Nevertheless, within the molecular network of the genes shortly expressed (24 h) in 2,4-D-induced wheat leaf bases, genes involved in the transcription machinery formed one of the largest groups, and included the MADS-box protein family [16]. In maize, a member of MADS-box family of type II (MIKC) has been functionally characterized. ZmMADS1 [GenBank: AF112148] was shown to be specifically induced in cells able to form somatic embryos (i.e., during early SE between the embryogenic competent cell stage and the globular phase), but not to be expressed in non-embryogenic suspension cultures neither in immature or mature zygotic embryos [76].
Based on sequence information from ZmMADS1 and given the high degree of synteny among species sharing a common evolutionary history in the cereal lineage, we designed degenerate primers surrounding the conserved MADS-box region and isolated a partial cDNA from wheat, representing a putative ZmMADS1 orthologous.
The MADS-box genes constitute a large multigene family dispersed throughout the genome in wheat [26, 77–79]. This is the first report of a spatio-temporal analysis of a MADS-box gene during SE in wheat. A differential expression level of this sequence was detected temporally (Fig. 3b), but not according to the type of tissue analyzed (Fig. 3c): the transcripts appeared at the early stage of the culture induction, and then gradually decreased (Fig. 3a, b).
It is worth mentioning that MADS-domain TFs characterized in wheat have been shown not only to be involved in plant developmental control [77–79], but also to be differentially expressed in stress-induced tissues [80–82] and to act at the confluence of the integration of environmental and developmental signals [83]. Stimulation of transcript accumulation at the time of the culture initiation (Fig. 3b) suggests that the cloned cDNA could be involved in early embryogenesis. It could also be involved in stress response associated with the conditions of in vitro survival, or both. An overlap between functions of genes involved in development and stress response has been documented in rice [70, 84] and wheat [85].
One could also explore a possible evolutionary functional variation between the cloned sequence and that of maize used as a reference. This important and complex MADS family has greatly expanded during the evolution of land plants [86, 87]. In particular, the MIKC-type proteins display an unusually efficient and high degree of evolutionary plasticity and functional diversification [88]. This could be especially true for wheat, as the evolutionary history of its genome is considerably richer in multiple events of large-scale duplication and polyploidization, stigmata of the evolution of Poaceae and the allohexaploid bread wheat, in particular.
Finally, high TF transcriptional activity at the beginning of tissue culture could simply mark the competent chromatin state associated with the reprogramming of the gene expression pattern [89–91].
Somatic embryogenesis and transgenesis: re-entry in cell cycle as a crucial step (RFC)
The production of nuclear transgenic plants requires the delivery of foreign DNA into the nucleus and its stable insertion into the host genome, followed by the multiplication and differentiation of transformed cells into organs or embryos, culminating in the subsequent regeneration of whole plants [92]. Transgenic plants can be obtained only from those cells that are competent for both transformation and regeneration, with efficiency depending on the degree of overlap between both types of cells [12].
The regeneration and transformation ability of cells are closely linked to their proliferation potential.
Regenerative ability depends largely on the ability of somatic cells to undergo dedifferentiation, a process preceding re-entry into the cell cycle, trans- or redifferentiation [56, 57]. In plants, the remarkable cellular plasticity is exemplified by the reversion of certain mature cells to a stem cell state and the regeneration of the whole plant [93]. This extraordinary feature requires cell-cycle reactivation at the G1-S transition phase, with the G1 phase that is central to the integration of signals that regulate the reactivation of cell proliferation or the exit from the cell division cycle to differentiation [94], and with the S-phase (DNA synthesis) culminating in full genome replication.
Actively dividing cells are the most responsive targets for transgene nuclear insertion [95, 96], including via particle bombardment [96–98]. A strong positive correlation was also observed between the cell cycle phase at the time of DNA transfer and Agrobacterium transformation efficiency. This pivotal importance of S-phase control functions for the Agrobacterium-mediated stable integration mechanism has been independently demonstrated previously [99–102]. To our knowledge, no such clear developmental control (cell cycle control and progression) has been established for direct DNA transfer strategies, notably via biolistic DNA delivery.
The large subunit of the wheat RFC considered in this study (Rfc-1, [GenBank: AJ318783]) is one of the essential components of the cell DNA replication machinery, its expression has been shown to be subjected to cell-cycle regulation, with a peak in early S-phase [28].
Rfc-1 messengers were most abundant in the 6 day-old calli that were embryogenic (Fig. 4). As in the case of the rice RFC homologue [27], Rfc-1 messengers were more abundant in the proliferating meristematic cells (day 6, Fig. 8). This is consistent with the fact that actively dividing cells are essential for establishing a new developmental program, organ morphogenesis and, in this case, somatic embryo completion [13]. Rfc-1 messengers were also more abundant (Fig. 4b) in tissues shown to have the best stable transformation ability (6-day-old tissues, [9]), which accords with the fact that dividing cells are the most effective targets for transgene insertion [95–97]. These findings therefore support the view mentioned earlier that the re-entry into the cell cycle has a crucial role in the expression of cellular totipotency [94] and also in gene transfer, nuclear integration and stable expression [92, 99–102].
However, the Rfc-1 transcripts were less abundant after 6 days (Fig. 4a, b). With the embryogenic developmental program being a continuous process, a proliferation state is still needed to some extent for its achievement at this time (see successive stages of development since day 8, Fig. 8). Given that the wheat Rfc-1 gene is cell-cycle regulated, this result suggests that a higher proportion of cells would be S phase-synchronized on day 6 of culture (i.e., during the pivotal period that coincides with the most suitable physiological state for DNA bombardment and long-term transgene expression) [9]. A previous study of the influence of the cell cycle on the efficiency of direct DNA transfer in the synchronized cultured cells of tobacco (Bright Yellow-2) reported that cells bombarded at the M and G2 phases gave higher transformation efficiency than those bombarded at the S and G1 phases, but in the absence of evidence in discriminating between transient and stable expression of the marker gene [103]. Therefore, as demonstrated for Agrobacterium-mediated transformation [99–102], in our study there appeared to be a positive correlation between the cell cycle S-phase at the time of biolistic DNA transfer and the stable transformation efficiency.
SERK: marker for SE differentiation and transformability
Among the genes whose expression pattern is altered during early SE induction, the SERK is considered to play an essential role in the vegetative-to-embryogenic transition. First characterized in Daucus carrota (carrot) from auxin-induced embryogenic cell cultures, DcSERK encoded a leucine-rich repeat (LRR) transmembrane receptor-like kinase (RLK) and was found to mark unequivocally the transition to embryogenesis [104]. Its expression was specifically and transiently expressed during the initiation of embryogenic development up to the globular stage, in suspension cell cultures, initially in the single cells that were able to form somatic embryos, and then during the initial phases of their development. In addition, during zygotic embryogenesis, the expression was detectable transiently in young zygotic embryos of up to 100 cells [104, 105].
SERK homologues were identified in others species, both dicotyledonous and monocotyledonous plants. As in carrot, homologous genes were shown to be expressed in the embryogenic cell cultures and somatic embryos [29, 106–110].
Arabidopsis AtSERK1 ectopic expression has been reported to confer embryogenic competence in culture [106]. Its expression was characteristic of those cells capable of rapid response to hormonal signals and able to form somatic embryos or embryogenic cell cultures [106, 111, 112].
The corresponding protein would be a component of an embryogenesis-signaling pathway. Competent cells might contain an inactive receptor, which is activated by the presence of the proper ligand to switch on the embryogenic program. SERK1 is a component of plasma membrane LRR-RLKs receptors organized into heterooligomeric protein complexes and involved in the brassinolide signaling pathway [113, 114]. SERK1 appears as an active member in the extracellular signal perception, followed by intracellular signal transduction via phosphorylation of specific targets that results in the altered expression of hundreds of genes [114–117].
Three SERK genes have been isolated from wheat and characterized. The signaling of 2,4-D-mediated induction of SE in leaf tissues was mediated by the SERK pathway, of which TaSERK1 is one component [16, 30].
In this study, there was a significantly higher relative abundance of TaSERK1 transcripts in tissues identified as embryogenic on day 8 of culture (Fig. 5a).
On average throughout the investigated period (Fig. 5c), a measurable higher mRNA level was observed in calli annotated as embryogenic, although transcripts were also detectable in the other samples. A broader expression pattern than that of DcSERK and AtSERK had previously been observed in other species [29, 118]. Actually, SERK homologue expression not only marks cells able to form somatic embryos, and is involved in conferring this embryogenic competence, but they also may have a wider developmental role (i.e., including zygotic embryogenesis, apomixes, in vitro organogenesis) [106, 108, 110, 119–121]. SERK proteins are involved in the general mechanisms of biotic and abiotic stress perception. They might partially mediate defense signal transduction for disease resistance response, in addition to their basic role in SE [107, 122–124].
Based on the spatio-temporal analyses, a higher transcript level was measured in the 8 day-old embryogenic structures (Fig. 5a, b), coinciding with the globular stage of embryo development (see histological picture day 8, Fig. 8). After that, the transcript level gradually decreased (Fig. 5b). This finding accords with results observed for carrot, where DcSERK expression is characteristic of embryogenic development up to the globular stage and stopped thereafter [104].
It is worth noting that this globular developmental stage coincides with the appearance of the first signs of differentiation (i.e., histogenesis begins with the demarcation of a protoderm covering the globular embryo, Fig. 8, day 8). On the premise that, along with differentiation, cells exit the mitotic cycle [125], this finding is congruent with the lesser transformation ability of wheat calli for culture periods exceeding 6 days [9]. In addition, after 6 days, clusters of dividing cells are enclosed within a structure bounded by cells that differentiate (the protoderm) in terms of their future basic protective function (the epidermis). Knowing that most DNA-coated particles penetrate only the epidermal and first few sub-epidermal layers [126], this configuration is a further constraint to the success of the physical DNA transfer.
Cell wall GerOXO: central role in development and defense
The cell wall is a highly dynamic entity and a fundamental structural element at the heart of the life strategy of plants, their development and defense [127–131].
The processes of wall assembly and remodeling, along with apoplastic metabolism, would be expected to play an important role during both somatic embryogenesis and direct DNA transfer strategies.
First, during in vitro culture, the auxin-induced structural changes of the plant cell wall associated with cell division cycle have long been known. Those changes prelude to cell enlargement preparing for mitosis and later to cytokinesis, involving the deposition of a new, diffusely fibrillar primary wall [132].
Second, cell wall formation in higher plants is central to cellular differentiation and morphogenesis. Its architecture and remodeling reflect functional differentiation of the cells [127], including through SE [133]. As part of a supracellular structure, called the cytoskeleton–plasma membrane–cell wall continuum [128], the cell wall (ECM: extracellular matrix) plays a crucial regulatory role during embryogenesis induced in vitro (for review, see [133]). Detailed temporal reports of ultrastructural changes also show that during the transition from the meristematic to the embryogenic state, ECM remodeling progressively leads to the physical and physiological individualization of single embryogenic cells from their surroundings (closure of plasmodemata, which breaks the symplasmic continuum, callose deposition and local cell wall thickening) and, later, of the globular unit formed through successive divisions of the initial embryogenic cell [134, 135].
Third, during genetic transformation experiments, plasma membranes and the walls of some cell types are the first biological barriers to be pierced or to pass, depending on the biological system being targeted and the cell compartment one wants to attain (nuclear, mitochondria or chloroplast organelles in eukaryotic cells) [95, 98, 136–142]. Plant cell differentiation leads to the increased mechanical strength of the plant wall [130]. Depending on the mode of particle acceleration, the momentum of particles can be more finely controlled, however, the degree of penetration will depend on the thickness of the cell wall [98], which is an effective penetration barrier [143].
In all these contexts, among the proteins associated with enzymatic cell wall metabolism, the apoplastic oxalate oxidases (OXOs) are particularly remarkable given the diversity of information in the literature supporting their implication in biosynthesis, remodeling and reinforcement of plant cell wall structures [144], but also in developmental reprogramming and differentiation [23, 145–148], including via SE [19, 22, 146], and in stress and defense responses triggered by biotic and abiotic aggressions (for review, see [21]).
OXOs (the so-called germin OXOs or simply germins) found in the ‘true cereals’ (barley, maize, oat, rice, rye, wheat) and ryegrass are able to break down oxalate liberated from calcium oxalate crystals dissolution [149–151]. The OXO oxygen-dependent degradation of calcium oxalate leads to the release of H2O2 and Ca2+. They are defense-related and H2O2 inducible [32, 152, 153]. They have been considered as a main provider of H2O2 in oxalate-producing plants [146, 152] and would also be an important provider of Ca2+ [144, 145, 154]. Both these small and diffusible molecules behave as signals or ‘second messengers’ at lower concentrations [155, 156] and are implicated in the biochemistry of the ECM at higher concentrations [144]. They are known to be key elements for cross-linking plant cell wall components; they are required to synthesize new walls during meristematic growth and for wall-stiffening in differentiated or stressed cells [144, 145, 151, 157, 158].
The OXO gene considered in this work responded to a wide range of biotic and abiotic factors in wheat, and its heterologous expression conferred resistance to bacteria, fungi or herbivorous insects [33–38].
In our study, a particular feature of this OXO enzymatic function was its biphasic nature apparent during the observation period (Fig. 6d), with the kinetics of enzymatic activity following that of the accumulation of messengers (Fig. 6b). A similar delay in the kinetics of accumulation of OXO proteins following that of the accumulation of messengers in response to auxin was observed by Caliskan et al. [19] in wheat immature embryo cultures, the time-lag there being about 20 h.
The first peak of OXO activity observed within 2 days of culture initiation (Fig. 6d) coincided with the first signs of auxin-induced proliferation and growing observed macroscopically [10] and confirmed by histological observations (Fig. 8).
This result accords with the analysis of the promoter structure of the wheat GerOXO gene: found to contain several auxin-response elements [159], this promoter was auxin-inducible in transgenic plants [160].
This result also accords with the previously established implication of H2O2 and Ca2+ in ECM during meristematic growth: in totipotent protoplasts [161–163], in the actively dividing and expanding cell [164], and during auxin-induced SE in gymnosperm [165, 166].
With regard to the appearance of this first peak, it is worth recalling that the OXO gene product considered in this study had long been known to be a protein marker of the onset of growth in germinating wheat (‘germin’, [167]) following rehydration. The exposure of immature embryos to auxin is also known to trigger OXO protein accumulation [19, 148]. The embryos treated according to our protocol had been subjected to water imbibition before being exposed to 2,4-D induction. The conjunction of these two treatments led to an intensified OXO activity after 2 days of 2,4-D exposure (Fig. 7). This result coincides with the burst of ROS (H2O2) associated with cell proliferation and active metabolism, during germination [52, 53, 168] and auxin SE induction [52, 53].
The second peak of OXO activity, evident at 8 days (Fig. 6d exclusively in the responsive embryogenic cultures), was displayed by tissues that were embarking on the embryogenic differentiation process at that time, in accord with the first signs of SE differentiation identified histologically that appear at the globular stage (the demarcation of a protoderm on day 8, Fig. 8).
This result accords with several studies showing specific transcription profiles of germins/germin-like proteins (GLPs) during the study of SE in wheat and gymnosperms [19, 165, 169, 170]. In wheat immature embryo culture, there is a clear correspondence between the bulk of the OXO activity found within the regenerating calli of nodular appearance (i.e., in small, densely cytoplasmic cells forming organ primordial) occurring after 7 days and the subsequent organogenic development [19]. In gymnosperms, specifically expressed at the early globular stage [165], the requirement of GLP expression for the maturation and normal development of somatic embryos was established by gene silencing: in silenced lines, the efficiency of maturation is reduced and most of them were blocked at the pre-globular stage [171].
This result also agrees with the presumed structural changes of the plant cell wall associated with differentiation and with the biological function of H2O2 and Ca2+ in mature walls during growth arrest and rigidification [151, 158].
Our results illustrate once again the apparent paradox associated with germin OXO proteins coupled with the increase in cell wall extensibility in cereals [144, 147, 172], but also with terminal growth arrest in differentiating structures [23, 24, 144, 148] and with wall reinforcement triggered in response to biotic and abiotic aggressions (reviewed by [21, 173]).
Regarding receptivity to gene transfer, the cell structure remodeling, derived from OXO enzymatic activity, could be expected to affect the success of DNA delivery through particle bombardment. The changes of cellular ultrastructure, particularly in terms of thickness, composition (callose and pectin) and hence the rigidity properties of the wall, reported to occur during somatic-to-embryogenic transition, are likely to account for much in cell receptiveness to the physical DNA introduction into the nucleus.
In addition, OXO proteins are also apparently deeply anchored in the defense strategy of monocot plants. Some OXOs and GLPs have now been recognized as members of pathogenesis-related protein families typical of monocots [174, 175]. More importantly, convergent literature data suggest that, together with some members of GLPs, the OXOs seem to form the foundation of a basal defense mechanism in Poaceae, as both immediate and long-lasting responses [33, 149, 152, 153, 176, 177].
According to the temporal–spatial concept of ROS wave signaling [178], OXOs could be attractive candidates in terms of relaying and feeding the cascade of cell-to-cell communication events over time. The study of the response to wounding [152] supports the early hypothesis put forward by Berna and Bernier [33] that OXOs are involved in stress-induced signaling. According to Le Deunff et al. [152], in line with a biphasic model, OXO proteins might represent a second wave and an alternative enzymatic pathway for relaying protection mechanisms and for establishing long-lasting defense and adaptive behavior.
It is worth noting that the establishment of plant defense mechanisms has been recognized as an efficient obstacle against an effective Agrobacterium transformation. During the transformation process, differential gene expression analysis demonstrated a reverse correlation of the efficiency of transformation with the expression level of the host defense genes [179–182]. Further, the fate of the transgene depends on the plant’s diverse adaptive defense mechanisms that act ordinarily on natural foreign, parasitic or ‘invasive’ nucleic acids (such as transposable elements, viroids, RNA and DNA viruses, and bacterial DNA) with a number of parallels with the immune system of mammals [183–185]. We suggest that the establishment of defense mechanisms might exert a prominent influence on the success of any genetic transformation process.
Lastly, our results lend credence to the idea that germin OXOs are ideally suited to playing a part in both initiation and termination of wall expansion [144], as well in SE development [19], in relaying the culture-induced oxidative burst [22, 152] and in the developmental adaptive response ensuring survival by producing H2O2 and Ca2+ [149, 186]. Taken together, this makes these OXO proteins critical enzymes in cereal embryogenic cultures with a potentially significant role in the receptivity of cells to genetic transformation mediated by biolistic delivery.
Integrated discussion
In the attempt to produce a synopsis (Fig. 8), we propose to integrate the elements and the data gathered in this study at the tissular, cellular and molecular levels together with the experimental data and theoretical aspects from the literature in order to address the question of tissue culture and transgenesis more broadly.
Stress in plants could be defined as any change in growth conditions that alters or disrupts metabolic homeostasis. This requires an adjustment of metabolic pathways, aimed at achieving a new state of homeostasis, in a process that is usually referred to as acclimation [187–189]. Regardless of the nature of biotic and abiotic stimuli, one of the consequences of stress is an increase in the cellular concentration of ROS that are subsequently converted to H2O2 [190].
A close connection, or an overlapping, between in vitro embryogenesis and stress response pathways has often been highlighted [13–15, 40]. Induction of SE is usually achieved by a stress and/or hormone treatment of somatic cells [14, 15, 43]. Activation of key regulators of embryogenesis is preceded by the stress-induced reprogramming of cellular metabolism [191]. H2O2 scavenging inhibits not only the activation of cellular defense reactions, but also cell division [163].
Figure 8 summarizes the successive treatments applied to plant tissues, as well as their effects (i.e., the renewed growth when wheat embryos are germinated in water, followed by the abrupt suppression of the sprouting of the embryonic axis, its wounding and torn tissues exposed to the physico-chemical/hormonal in vitro environmental context), which collectively disrupt metabolic homeostasis and cause stress, triggering a first oxidative burst (Fig. 8, auxin and culture induced oxidative stresses, first H2O2 burst). ‘Oxidative stress’, formerly a pejorative term denoting a harmful process to be avoided, actually represents a positive process known as ‘oxidative signaling’ [192], an essential component of the repertoire of signals that plant cells use to perceive the environment and make appropriate metabolic/physiological adjustments. ROS participate in signal transduction cascades in processes as diverse as mitosis, growth, development and adaptation. Intimately interconnected with calcium signaling, ROS/redox signaling/antioxidant interaction is seen as a switching node in plant cell physiology [192–195]. Through crosstalk with phytohormones (mainly auxin and ethylene) and sugars, this convergence hub between stress signaling and development allows specific responses to environmental in vitro challenges to be orchestrated [11] (Fig. 8, cellular adjustment to environmental stresses). Dynamic changes in chromatin structure are central to the reversion of the differentiation process and the developmental transition. The developmental switching in somatic cells towards the embryogenic pathway is a release from suppression, controlled at the chromatin level, through a global and dynamic reorganization of the chromatin structure, giving the somatic cells the ability to manifest the embryogenic pathway [14, 15] (Fig. 8, nuclear reorganization based on chromatin remodeling and resetting of gene expression program). During the ongoing SE process, after dedifferentiation, cell division reactivation is the second requirement for pluripotentiality, before the coordinated proliferation process progressively culminates in the development of organized structures (Fig. 8, cell cycle e-entry, followed by cell proliferation and embryogenic pattern differentiation). Meanwhile, a dynamic process of ROS signaling is thought to occur within and between cells, relaying defense- and stress-adaptive processes [178]. It is worth recalling that, during the Agrobacterium transformation process, repressing the host defense response is prerequisite to successful transformation. Differential gene expression analysis demonstrated a reverse correlation of transformation efficiency with the expression level of the host defense genes, notably the GSTs [179–182].
Figure 8 provides an integrated view of the functional links between regenerative and gene transfer properties, indicating the possible significance of gene profiling results in parallel with the regenerative pathway events and their impact on plant cell receptivity to gene transfer.
Conclusions
This paper presents the results of studies conducted at three observation levels (i.e., macroscopic, histological and molecular).
Surprisingly, to the best of our knowledge there has been no report to date that establishes the molecular interconnections between tissue culture and genetic transformation competencies in plant cells. Cells undergo an extensive change in gene regulation during the successive steps along a regenerative pathway. In this study, GST, MADS-box TF, SERK and RFc gene expression data provide spatio-temporal indicators relative to cell adjustment, chromatin remodeling, reprogramming of gene expression during SE induction and the proliferation state required for transgene integration, respectively.
Suitability for stable transformation is correlated with intense cell proliferation, as would be expected in any transformation procedure, but it also appears to be associated with indicators of a higher proportion of S phase-synchronized cells and a temporarily weakened defense system, as previously reported in the Agrobacterium transformation strategy.
We hypothesize that, in the course of the multi-step defense adaptation process that takes place during our in vitro culture protocol, the ideal moment for particle bombardment is probably an optimal period that coincides with (i) the chance to reach the nuclei of dividing cells (S-phase), (ii) the ease of passing through the physical barriers (before mature thickened walls are established in differentiating cells) and (iii) just before the establishment of durable biological defense mechanisms, one of whose functions would be to preserve genome integrity (between the first line of defense and the second wave of long-lasting defense processes).
As far as both tissue culture and direct gene transfer abilities are concerned, however, OXO activity appears to occur in parallel with the overall process of auxin-induced dedifferentiation, proliferation and differentiation, as well gene transfer competency over time. In the latter case, the involvement of OXO proteins, locally in cell wall structural reinforcement and globally in consolidating plant innate immunity, might be critical.
The idea that GerOXOs might be crucial during critical events in the life of plants was put forward some time ago [144, 145] and again more recently [149, 152, 186]. Such a broad and vital role in relaying stress response, adaptive behavior and cell survival might originate from the evolutionary history of this family of genes within the larger group of the multifunctional Cupin superfamily, of which the cereal OXOs are the archetypal members (for review, see [21, 31, 32]).
References
Chen H, Lin Y, Zhang Q (2010) Rice. Genetic modification of plants, vol 64. Springer, Berlin
Hensel G, Himmelbach A, Chen W, Douchkov DK, Kumlehn J (2011) Transgene expression systems in the Triticeae cereals. J Plant Physiol 168(1):30–44. doi:10.1016/j.jplph.2010.07.007
Kumlehn J, Hensel G (2009) Genetic transformation technology in the Triticeae. Breeding Sci 59:553–560
Xia L, Ma Y, He Y, Jones HD (2012) GM wheat development in China: current status and challenges to commercialization. J Exp Bot 63(5):1785–1790. doi:10.1093/jxb/err342
Vasil IK (2005) The story of transgenic cereals: the challenge, the debate, and the solution—a historical perspective. In Vitro Cell Dev Biol Plant 41:577–583
Shewry PR, Jones HD (2005) Transgenic wheat: where do we stand after the first 12 years? Ann Appl Biol 147:1–14
Nakamura H, Hakata M, Amano K, Miyao A, Toki N, Kajikawa M, Pang J, Higashi N, Ando S, Toki S, Fujita M, Enju A, Seki M, Nakazawa M, Ichikawa T, Shinozaki K, Matsui M, Nagamura Y, Hirochika H, Ichikawa H (2007) A genome-wide gain-of-function analysis of rice genes using the FOX-hunting system. Plant Mol Biol 65:357–371
Shrawat AK, Good AG (2010) A high-throughput Agrobacterium tumefaciens-mediated transformation system for molecular breeding and functional genomics of rice (Oryza sativa L.). Plant Biotechnol 27(1):47–58
Delporte F, Li S, Jacquemin JM (2005) Calluses initiated from thin mature embryo fragments are suitable targets for wheat transformation as assessed by long-term GUS expression studies. Plant Cell Tissue Organ Cult 80:139–149
Delporte F, Mostade O, Jacquemin JM (2001) Plant regeneration through callus initiation from thin mature embryo fragments of wheat. Plant Cell Tissue Organ Cult 67:73–80
Delporte F, Jacquemin J-M, Masson P, Watillon B (2012) Insights into the regenerative property of plant cells and their receptivity to transgenesis: wheat as a research case study. Plant Signal Behav 7(12):1608–1620
Potrykus I (1991) Gene transfer to plants: assessment of published approaches and results. Annu Rev Plant Physiol Plant Mol Biol 42:205–225
Fehér A, Pasternak TP, Dudits D (2003) Transition of somatic plant cells to an embryogenic state. Plant Cell Tissue Organ Cult 74:201–228
Fehér A (2006) Why somatic plant cells start to form embryos? In: Mujib A, Šamaj J (eds) Somatic embryogenesis. Plant cell monographs, vol 2. Springer, Berlin, pp 85–101
Fehér A (2008) The initiation phase of somatic embryogenesis: what we know and what we don’t. Acta Biol Szegediensis 52(1):53–56
Singla B, Tyagi AK, Khurana JP, Khurana P (2007) Analysis of expression profile of selected genes expressed during auxin-induced somatic embryogenesis in leaf base system of wheat (Triticum aestivum) and their possible interactions. Plant Mol Biol 65(5):677–692
Chakrabarty D, Trivedi PK, Shri M, Misra P, Asif MH, Dubey S, Kumar S, Rai A, Tiwari M, Shukla D, Pandey A, Nigam D, Tripathi RD, Tuli R (2010) Differential transcriptional expression following thidiazuron-induced callus differentiation developmental shifts in rice. Plant Biol 12(1):46–59
Caliskan M (2011) The roles of germin gene products in plants under salt stress. In: AK Shanker, B Venkateswarlu (eds) Abiotic stress response in plants—physiological, biochemical and genetic perspectives. InTech, Rijeka Croatia, pp 297–320. doi:10.5772/23313. Availble from: http://www.intechopen.com/books/abiotic-stress-response-in-plants-physiological-biochemical-and-genetic-perspectives/the-roles-of-germin-gene-products-in-plants-under-salt-stress1, ISBN:978-953-307-672-0
Caliskan M, Turet M, Cuming AC (2004) Formation of wheat (Triticum aestivum L.) embryogenic callus involves peroxide-generating germin-like oxalate oxidase. Planta 219:132–140
Davidson RM, Reeves PA, Manosalva PM, Leach JE (2009) Germins: a diverse protein family important for crop improvement. Plant Sci 177:499–510
Dunwell JM, Gibbins JG, Mahmood T, Naqvi SMS (2008) Germin and germin-like proteins: evolution, structure, and function. Crit Rev Plant Sci 27:342–375
Jacquard C, Mazeyrat-Gourbeyre F, Devaux P, Boutilier K, Baillieul F, Clément C (2009) Microspore embryogenesis in barley: anther pre-treatment stimulates plant defence gene expression. Planta 229(2):393–402
Caliskan M, Cuming AC (1998) Spatial specificity of H2O2 generating oxalate oxidase gene expression during wheat embryo germination. Plant J 15:165–171
Dumas B, Freyssinet G, Pallett KE (1995) Tissue-specific expression of germin-like oxalate oxidase during development and fungal infection of barley seedlings. Plant Physiol 107(4):1091–1096
Xu F, Lagudah ES, Moose SP, Riechers DE (2002) Tandemly duplicated safener-induced glutathione S-transferase genes from Triticum tauschii contribute to genome- and organ-specific expression in hexaploid wheat. Plant Physiol 130:362–373
Murai K, Murai R, Ogihara Y (1997) Wheat MADS box genes, a multigene family dispersed throughout the genome. Genes Genet Syst 72(5):317–321
Furukawa T, Ishibashi T, Kimura S, Tanaka H, Hashimoto J, Sakaguchi K (2003) Characterization of all the subunits of replication factor C from a higher plant, rice (Oryza sativa L.), and their relation to development. Plant Mol Biol 53(1–2):15–25
Luque A, Sanz-Burgos AP, Ramirez-Parra E, Castellano MM, Gutierrez C (2002) Interaction of geminivirus Rep protein with replication factor C and its potential role during geminivirus DNA replication. Virology 302(1):83–94
Baudino S, Hansen S, Brettschneider R, Hecht VFG, Dresselhaus T, Lorz H, Dumas C, Rogowsky PM (2001) Molecular characterization of two novel maize LRR receptor-like kinases, which belong to the SERK gene family. Planta 213:1–10
Singla B, Khurana JP, Khurana P (2008) Characterization of three somatic embryogenesis receptor kinase genes from wheat, Triticum aestivum. Plant Cell Rep 27:833–843
Dunwell JM, Khuri S, Gane PJ (2000) Microbial relatives of the seed storage proteins of higher plants: conservation of structure and diversification of function during evolution of the cupin superfamily. Microbiol Mol Biol Rev 64:153–179
Dunwell JM, Purvis A, Khuri S (2004) Cupins: the most functionally diverse protein superfamily? Phytochemistry 65:7–17
Berna A, Bernier F (1999) Regulation by biotic and abiotic stress of a wheat germin gene encoding oxalate oxidase, a H2O2-producing enzyme. Plant Mol Biol 39(3):539–549
Schneider M, Droz E, Malnoë P, Chatot C, Bonnel E, Métraux J (2002) Transgenic potato plants expressing oxalate oxidase have increased resistance to oomycete and bacterial pathogens. Potato Res 45(2):177–185
Liang H, Maynard CA, Allen RD, Powell WA (2001) Increased Septoria musiva resistance in transgenic hybrid poplar leaves expressing a wheat oxalate oxidase gene. Plant Mol Biol 45(6):619–629
Donaldson PA, Anderson T, Lane BG, Davidson AL, Simmonds DH (2001) Soybean plants expressing an active oligomeric oxalate oxidase from the wheat gf-2.8 (germin) gene are resistant to the oxalate-secreting pathogen Sclerotina sclerotiorum. Physiol Mol Plant Pathol 59(6):297–307
Ramputh AI, Arnason JT, Cass L, Simmonds JA (2002) Reduced herbivory of the European corn borer (Ostrinia nubilalis) on corn transformed with germin, a wheat oxalate oxidase gene. Plant Sci 162(3):431–440
Hu X, Bidney DL, Yalpani N, Duvick JP, Crasta O, Folkerts O, Lu G (2003) Overexpression of a gene encoding hydrogen peroxide-generating oxalate oxidase evokes defense responses in sunflower. Plant Physiol 133(1):170–181
Simmonds J, Cass L, Routly E, Hubbard K, Donaldson P, Bancroft B, Davidson A, Hubbard S, Simmonds D (2004) Oxalate oxidase: a novel reporter gene for monocot and dicot transformations. Mol Breeding 13(1):79–91
Quiroz-Figueroa FR, Rojas-Herrera R, Galaz-Avalos RM, Loyola-Vargas VM (2006) Embryo production through somatic embryogenesis can be used to study cell differentiation in plants. Plant Cell Tissue Organ Cult 86:285–301. doi:10.1007/s11240-006-9139-6
Raghavan V (2006) Can carrot and Arabidopsis serve as model systems to study the molecular biology of somatic embryogenesis? Curr Sci 90:1336–1343
Karami O, Saidi A (2010) The molecular basis for stress-induced acquisition of somatic embryogenesis. Mol Biol Rep 37(5):2493–2507. doi:10.1007/s11033-009-9764-3
Zavattieri MA, Frederico AM, Lima M, Rui S, Arnholdt-Schmitt B (2010) Induction of somatic embryogenesis as an example of stress-related plant reactions. Electron J Biotechnol 13(1). doi:10.2225/vol2213-issue2221-fulltext-2224
Frova C (2006) Glutathione transferases in the genomics era: new insights and perspectives. Biomol Eng 23(4):149–169
Riechers DE, Kreuz K, Zhang Q (2010) Detoxification without intoxication: herbicide safeners activate plant defense gene expression. Plant Physiol 153(1):3–13
Moons A (2005) Regulatory and functional interactions of plant growth regulators and plant glutathione S-transferases (GSTs). Plant Horm 72:155–202
Gong H, Jiao Y, Hu W-W, Pua E-C (2005) Expression of glutathione-S-transferase and its role in plant growth and development in vivo and shoot morphogenesis in vitro. Plant Mol Biol 57(1):53–66
Kim SG, Kim ST, Kang SY, Wang Y, Kim W, Kang KY (2008) Proteomic analysis of reactive oxygen species (ROS)-related proteins in rice roots. Plant Cell Rep 27(2):363–375
Gallé Á, Csiszár J, Secenji M, Guóth A, Cseuz L, Tari I, Györgyey J, Erdei L (2009) Glutathione transferase activity and expression patterns during grain filling in flag leaves of wheat genotypes differing in drought tolerance: response to water deficit. J Plant Physiol 166(17):1878–1891
Edwards R, Dixon D (2005) Plant glutathione transferases. Methods Enzymol 401:169–186
Dixon DP, Skipsey M, Edwards R (2010) Roles for glutathione transferases in plant secondary metabolism. Phytochemistry 71(4):338–350
Pfeiffer W, Höftberger M (2001) Oxidative burst in Chenopodium rubrum suspension cells. Physiol Plant 111(2):144–150
Stasolla C, Bozhkov PV, Chu T-M, van Zyl L, Egertsdotter U, Suarez MF, Craig D, Wolfinger RD, Von Arnold S, Sederoff RR (2004) Variation in transcript abundance during somatic embryogenesis in gymnosperms. Tree Physiol 24(10):1073–1085
Gaj MD (2004) Factors influencing somatic embryogenesis induction and plant regeneration with particular reference to Arabidopsis thaliana (L.) Heynh. Plant Growth Regul 43:27–47
Fehér A (2006) Why somatic plant cells start to form embryos? In: Mujib A, Šamaj J (eds) Somatic embryogenesis. Plant cell monographs, vol 2. Springer, Berlin, pp 85–101. doi:10.1007/7089_019
Grafi G (2004) How cells dedifferentiate: a lesson from plants. Dev Biol 268(1):1–6
Williams L, Zhao J, Morozova N, Li Y, Avivi Y, Grafi G (2003) Chromatin reorganization accompanying cellular dedifferentiation is associated with modifications of histone H3, redistribution of HP1, and activation of E2F-target genes. Dev Dyn 228:113–120
Benfey PN, Weigel D (2001) Transcriptional networks controlling plant development. Plant Physiol 125:109–111
Meyerowitz EM (2002) Plants compared to animals: the broadest comparative study of development. Science 295:1482–1485
Levine M, Davidson EH (2005) Gene regulatory networks for development. Proc Natl Acad Sci USA 102(14):4936–4942
Boutilier K, Offringa R, Sharma VK, Kieft H, Ouellet T, Zhang LM, Hattori J, Liu CM, van Lammeren AAM, Miki BLA, Custers JBM, van Lookeren Campagne MM (2002) Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell 14:1737–1749
Lotan T, Ohto M, Yee KM, West MAL, Lo R, Kwong RW, Yamagishi K, Fischer RL, Goldberg RB, Harada JJ (1998) Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93:1195–1205
Perry SE, Lehti MD, Fernandez DE (1999) The MADS-domain protein AGAMOUS-like 15 accumulates in embryonic tissues with diverse origins. Plant Physiol 120:121–129
Zuo J, Niu QW, Frugis G, Chua NH (2002) The WUSCHEL gene promotes vegetative-to-embryonic transition in Arabidopsis. Plant J 30:349–359
Vasil IK (1988) Progress in the regeneration and genetic manipulation of cereal crops. Biotechnology 6:397–402
Alvarez-Buylla ER, Pelaz S, Liljegren SJ, Gold SE, Burgeff C, Ditta GS, Ribas de Pouplana L, Martínez-Castilla L, Yanofsky MF (2000) An ancestral MADS-box gene duplication occurred before the divergence of plants and animals. Proc Natl Acad Sci USA 97(10):5328–5333
Theissen G, Becker A, Di Rosa A, Kanno A, Kim JT, Munster T, Winter KU, Saedler H (2000) A short history of MADS-box genes in plants. Plant Mol Biol 42:115–149
Henschel K, Kofuji R, Hasebe M, Saedler H, Münster T, Theissen G (2002) Two ancient classes of MIKC-type MADS-box genes are present in the moss Physcomitrella patens. Mol Biol Evol 19:801–814
de Folter S, Angenent GC (2006) Trans meets cis in MADS science. Trends Plant Sci 11:224–231
Arora R, Agarwal P, Ray S, Singh AK, Singh VP, Tyagi AK, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8:242
Century K, Reuber TL, Ratcliffe OJ (2008) Regulating the regulators: the future prospects for transcription-factor-based agricultural biotechnology products. Plant Physiol 147:20–29
Veron AS, Kaufmann K, Bornberg-Bauer E (2007) Evidence of interaction network evolution by whole-genome duplications: a case study in MADS-box proteins. Mol Biol Evol 24(3):670–678
Perry SE, Nichols KW, Fernandez DE (1996) The MADS domain protein AGL15 localizes to the nucleus during early stages of seed development. Plant Cell l 8:1977–1989
Harding EW, Tang W, Nichols KW, Fernandez DE, Perry SE (2003) Expression and maintenance of embryogenic potential is enhanced through constitutive expression of AGAMOUS-like15. Plant Physiol 133:653–663
Thakare D, Tang W, Hill K, Perry SE (2008) The MADS-domain transcriptional regulator AGAMOUS-LIKE15 promotes somatic embryo development in Arabidopsis and soybean. Plant Physiol 146:1663–1672
Heuer S, Hansen S, Bantin J, Brettschneider R, Kranz E, Lörz H, Dresselhaus T (2001) The maize MADS box gene ZmMADS3 affects node number and spikelet development and is co-expressed with ZmMADS1 during flower development, in egg cells, and early embryogenesis. Plant Physiol 127:33–45
Ciaffi M, Paolacci AR, D’Aloisio E, Tanzarella OA, Porceddu E (2005) Identification and characterization of gene sequences expressed in wheat spikelets at the heading stage. Gene 346:221–230
Zhao T, Ni Z, Dai Y, Yao Y, Nie X, Sun Q (2006) Characterization and expression of 42 MADS-box genes in wheat (Triticum aestivum L.). Mol Genet Genomics 276(4):334–350
Paolacci AR, Tanzarella OA, Porceddu E, Varotto S, Ciaffi M (2007) Molecular and phylogenetic analysis of MADS-box genes of MIKC type and chromosome location of SEP-like genes in wheat (Triticum aestivum L.). Mol Genet Genomics 278(6):689–708
Houde M, Belcaid M, Ouellet F, Danyluk J, Monroy AF, Dryanova A, Gulick P, Bergeron A, Laroche A, Links MG, MacCarthy L, Crosby WL, Sarhan F (2006) Wheat EST resources for functional genomics of abiotic stress. BMC Genomics 7:149
Mochida K, Kawaura K, Shimosaka E, Kawakami N, Shin-I T, Kohara Y, Yamazaki Y, Ogihara Y (2006) Tissue expression map of a large number of expressed sequence tags and its application to in silico screening of stress response genes in common wheat. Mol Genet Genomics 276(3):304–312
Kawaura K, Mochida K, Ogihara Y (2008) Genome-wide analysis for identification of salt-responsive genes in common wheat. Funct Integr Genomics 8(3):277–286
Trevaskis B, Hemming MN, Dennis ES, Peacock WJ (2007) The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci 12(8):352–357
Cooper B, Clarke JD, Budworth P, Kreps J, Hutchison D, Park S, Guimil S, Dunn M, Luginbühl P, Ellero C, Goff SA, Glazebrook J (2003) A network of rice genes associated with stress response and seed development. Proc Natl Acad Sci USA 100(8):4945–4950
Tardif G, Kane NA, Adam H, Labrie L, Major G, Gulick P, Sarhan F, Laliberte JF (2007) Interaction network of proteins associated with abiotic stress response and development in wheat. Plant Mol Biol 63:703–718
De Bodt S, Raes J, Van de Peer Y, Theißen G (2003) And then there were many: MADS goes genomic. Trends Plant Sci 8(10):475–483
Nam J, Kaufmann K, Theißen G, Nei M (2005) A simple method for predicting the functional differentiation of duplicate genes and its application to MIKC-type MADS-box genes. Nucl Acids Res 33(2):e12
Kaufmann K, Melzer R, Theißen G (2005) MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants. Gene 347(2):183–198
Efroni S, Duttagupta R, Cheng J, Dehghani H, Hoeppner DJ, Dash C, Bazett-Jones DP, Le Grice S, McKay RDG, Buetow KH, Gingeras TR, Misteli T, Meshorer E (2008) Global transcription in pluripotent embryonic stem cells. Cell Stem Cell 2(5):437–447
Grafi G, Chalifa-Caspi V, Nagar T, Plaschkes I, Barak S, Ransbotyn V (2011) Plant response to stress meets dedifferentiation. Planta 233:433–438
Fransz P, de Jong H (2011) From nucleosome to chromosome: a dynamic organization of genetic information. Plant J 66(1):4–17
Arias RS, Filichkin SA, Strauss SH (2006) Divide and conquer: development and cell cycle genes in plant transformation. Trends Biotechnol 24:267–273
Gutierrez C (2005) Coupling cell proliferation and development in plants. Nat Cell Biol 7(6):535–541
Gutierrez C, Ramirez-Parra E, Castellano MM, Del Pozo JC (2002) G(1) to S transition: more than a cell cycle engine switch. Curr Opin Plant Biol 5:480–486
Sharma KK, Bhatnagar-Mathur P, Thorpe TA (2005) Genetic transformation technology: status and problems. In Vitro Cell Dev Biol Plant 41:102–112
Altpeter F, Baisakh N, Beachy R, Bock R, Capell T, Christou P, Daniell H, Datta K, Datta S, Dix PJ, Fauquet C, Huang N, Kohli A, Mooibroek H, Nicholson L, Nguyen TT, Nugent G, Raemakers K, Romano A, Somers DA, Stoger E, Taylor N, Visser R (2005) Particle bombardment and the genetic enhancement of crops: myths and realities. Mol Breeding 15:305–327
Sanford JC, Smith FD, Russel JA (1993) Optimising the biolistic process for different biological applications. Methods Enzymol 217:483–509
Taylor NJ, Fauquet CM (2002) Microparticle bombardment as a tool in plant science and agricultural biotechnology. DNA Cell Biol 21(12):963–977
Villemont E, Dubois F, Sangwan RS, Vasseur G, Bourgeois Y, Sangwan-Norreel BS (1997) Role of the host cell cycle in the Agrobacterium-mediated genetic transformation of Petunia: evidence of an S-phase control mechanism for T-DNA transfer. Planta 201:160–172
Chen LFO, Hwang JY, Charng YY, Sun CW, Yang SF (2001) Transformation of broccoli (Brassica oleracea var. italica) with isopentenyltransferase gene via Agrobacterium tumefaciens for post-harvest yellowing retardation. Mol Breeding 7(3):243–257
Gordon-Kamm W, Dilkes BP, Lowe K, Hoerster G, Sun X, Ross M, Church L, Bunde C, Farrell J, Hill P, Maddock S, Snyder J, Sykes L, Li Z, Woo YM, Bidney D, Larkins BA (2002) Stimulation of the cell cycle and maize transformation by disruption of the plant retinoblastoma pathway. Proc Natl Acad Sci USA 99(18):11975–11980
Peña LA, Pérez RM, Cervera M, Juárez JA, Navarro L (2004) Early events in Agrobacterium-mediated genetic transformation of citrus explants. Ann Bot 94:67–74
lida A, Yamashita T, Yamada Y, Morikawa H (1991) Efficiency of particle-bombardment-mediated transformation is influenced by cell cycle stage in synchronized cultured cells of tobacco. Plant Physiol 97:1585–1587
Schmidt ED, Guzzo F, Toonen MA, de Vries SC (1997) A leucine-rich repeat containing receptor-like kinase marks somatic plant cells competent to form embryos. Development 124(10):2049–2062
de Vries SC (2000) Signals and their transduction in early plant embryogenesis. In: de Vries, GE, Metzlaff K (eds) Highlights in European plant biotechnology research and technology transfer. Proceedings of the second European conference on plant biotechnology, Rome. Elsevier, Amsterdam, pp 141–148
Hecht V, Vielle-Calzada J-P, Hartog MV, Schmidt ED, Boutilier K, Grossniklaus U, de Vries SC (2001) The Arabidopsis somatic embryogenesis receptor kinase 1 gene is expressed in developing ovules and embryos and enhances embryogenic competence in cultures. Plant Physiol 127:803–816
Huang X, Lu X-Y, Zhao J-T, Chen J-K, Dai X-M, Xiao W, Chen Y-P, Chen Y-F, Huang X-L (2010) MaSERK1 gene expression associated with somatic embryogenic competence and disease resistance response in banana (Musa spp.). Plant Mol Biol Rep 28(2):309–316
Nolan KE, Irwanto RR, Rose RJ (2003) Auxin up-regulates MtSERK1 expression in both Medicago truncatula root-forming and embryogenic cultures. Plant Physiol 133(1):218–230
Somleva MN, Schmidt EDL, de Vries SC (2000) Embryogenic cells in Dactylis glomerata L. (Poaceae) explants identified by cell tracking and by SERK expression. Plant Cell Rep 19(7):718–726
Zakizadeh H, Stummann BM, Lütken H, Müller R (2010) Isolation and characterization of four somatic embryogenesis receptor-like kinase (RhSERK) genes from miniature potted rose (Rosa hybrida cv. Linda). Plant Cell Tissue Organ Cult 101(3):331–338
Kwaaitaal MACJ, de Vries SC (2007) The SERK1 gene is expressed in procambium and immature vascular cells. J Exp Bot 58(11):2887–2896
Salaj J, von Recklinghausen IR, Hecht V, de Vries SC, Schel JHN, van Lammeren AAM (2008) AtSERK1 expression precedes and coincides with early somatic embryogenesis in Arabidopsis thaliana. Plant Physiol Biochem 46(7):709–714
Karlova R, Boeren S, Russinova E, Aker J, Vervoort J, de Vries SC (2006) Arabidopsis somatic embryogenesis receptor-like kinase 1 protein complex includes brassinosteroid-insensitive. Plant Cell 18:626–638
Aker J, de Vries SC (2008) Plasma membrane receptor complexes. Plant Physiol 147(4):1560–1564
Belkhadir Y, Chory J (2006) Brassinosteroid signaling: a paradigm for steroid hormone signaling from the cell surface. Science 314:1410–1411
Karlova R, de Vries SC (2006) Advances in understanding brassinosteroid signaling. Sci STKE pe36
Clouse SD (2008) Brassinosteroid signaling. In: Bögre L, Beemster G (eds) Plant growth signaling. Plant cell monograph, vol 10. Springer, Berlin, pp 179–197
Ben Mahmoud K, Delporte F, Muhovski Y, Elloumi N, Jemmali A, Druart P (2013) Expression of PiABP19, Picdc2 and PiSERK3 during induction of somatic embryogenesis in leaflets of Prunus incisa (Thunb.). Mol Biol Rep 40(2):1569–1577. doi:10.1007/s11033-012-2205-8
Albertini E, Marconi G, Reale L, Barcaccia G, Porceddes A, Ferranti F, Falcinelli M (2005) SERK and APOSTART. Candidate genes for apomixis in Poa pratensis. Plant Physiol 138:2185–2194
Sharma SK, Millam S, Hein I, Bryan GJ (2008) Cloning and molecular characterisation of a potato SERK gene transcriptionally induced during initiation of somatic embryogenesis. Planta 228:319–330
Thomas C, Meyer D, Himber C, Steinmetz A (2004) Spatial expression of a sunflower SERK gene during induction of somatic embryogenesis and shoot organogenesis. Plant Physiol Biochem 42(1):35–42
Hu H, Xiong L, Yang Y (2005) Rice SERK1 gene positively regulates somatic embryogenesis of cultured cell and host defense response against fungal infection. Planta 222:107–117
Song D, Li G, Song F, Zheng Z (2008) Molecular characterization and expression analysis of OsBISERK1, a gene encoding a leucine-rich repeat receptor-like kinase, during disease resistance responses in rice. Mol Biol Rep 35:275–283
Santos MO, Romano E, Vieira LS, Baldoni AB, Aragão FJL (2009) Suppression of SERK gene expression affects fungus tolerance and somatic embryogenesis in transgenic lettuce. Plant Biol 11:83–89
John P, Sek F, Carmichael J, McCurdy D (1990) p34cdc2 homologue level, cell division, phytohormone responsiveness and cell differentiation in wheat leaves. J Cell Sci 97(4):627–630
Taylor LP, Vasil IK (1991) Histology of, and physical factors affecting, transient GUS expression in plant millet (Pennisetum glaucum (Li) R. Br.) embryos following microprojectile bombardment. Plant Cell Rep 10:120–125
Wojtaszek P (2000) Genes and plant cell walls: a difficult relationship. Biol Rev Camb Philos Soc 75:437–475
Baluska F, Samaj J, Wojtaszek P, Volkmann D, Menzel D (2003) Cytoskeleton-plasma membrane-cell wall continuum in plants. Emerging links revisited. Plant Physiol 133:482–491
Cosgrove DJ (2000) Loosening of plant cell walls by expansins. Nature 407:321–326
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861
Humphrey TV, Bonetta DT, Goring DR (2007) Sentinels at the wall: cell wall receptors and sensors. New Phytol 176:7–21
Bowes BG, Torrey JG (1976) Ultrastructural changes in cells of pea root cortical explants cultured In vitro. Protoplasma 90(1–2):99–118
Šamaj J, Bobák M, Blehová A, Pretová A (2006) Importance of cytoskeleton and cell wall in somatic embryogenesis. In: Mujib A, Samaj J (eds) Somatic embryogenesis, plant cell monographs (2). Springer, Berlin, Heidelberg, pp 35–50. doi:10.1007/7089_024
Verdeil JL, Hocher V, Huet C, Grosdemange F, Escoute J, Ferriere N, Nicole M (2001) Ultrastructural changes in coconut calli associated with the acquisition of embryogenic competence. Ann Bot 88(1):9–18
Konieczny R, Bohdanowicz J, Czaplicki AZ, Przywara L (2005) Extracellular matrix surface network during plant regeneration in wheat anther culture. Plant Cell Tissue Organ Cult 83(2):201–208
Daniell H, Cohill PR, Kumar S, Dufourmantel N (2004) Chloroplast genetic engineering. In: Daniell H, Chase C (eds) Molecular biology and biotechnology of plant organelles. Springer, Dordrecht
Maliga P (2004) Plastid transformation in higher plants. Annu Rev Plant Biol 55(1):289–313
Remacle C, Cardol P, Coosemans N, Gaisne M, Bonnefoy N (2006) High-efficiency biolistic transformation of Chlamydomonas mitochondria can be used to insert mutations in complex I genes. Proc Natl Acad Sci USA 103(12):4771–4776
Koop H-U, Herz S, Golds T, Nickelsen J (2007) The genetic transformation of plastids. Cell Mol Biol Plastids 19:457–510
Villemejane J, Mir LM (2009) Physical methods of nucleic acid transfer: general concepts and applications. Br J Pharmacol 157(2):207–219
Cadoret J-P, Bardor M, Lerouge P, Cabigliera M, Henriquez V, Carlier A (2008) Les microalgues: usines cellulaires productrices de molécules commerciales recombinantes. Méd Sci 24(4):375–382
Riveline D, Nurse P (2009) ‘Injecting’ yeast. Nat Methods 6(7):513–514
Kao C-Y, Huang S-H, Lin C-M (2008) A low-pressure gene gun for genetic transformation of maize (Zea mays L.). Plant Biotechnol Rep 2(4):267–270
Lane BG (1994) Oxalate, germin, and the extracellular matrix of higher plants. FASEB J 8:294–301
Lane BG, Dunwell JM, Ray JA, Schmitt MR, Cuming AC (1993) Germin, a protein marker of early plant growth, is an oxalate oxidase. J Biol Chem 268:12239–12242
Çalişkan M (2000) Germin, an oxalate oxidase, has a function in many aspects of plant life. Turk J Biol 24:717–724
Caliskan M, Cuming AC (2000) Temporal and spatial determination of germin biosynthesis in wheat tissues. Turk J Biol 24:775–782
Lane BG (2000) Oxalate oxidases and differentiating surface structure in wheat: germins. Biochem J 349:309–321
Davoine C, Le Deunff E, Ledger N, Avice J-C, Billard J-P, Dumas B, Huault C (2001) Specific and constitutive expression of oxalate oxidase during the ageing of leaf sheaths of ryegrass stubble. Plant Cell Environ 24(10):1033–1043
Lane BG (2002) Oxalate, germins, and higher-plant pathogens. IUBMB Life 53(2):67–75
Vreeburg RA, Fry SC (2005) Reactive oxygen species in cell walls. In: Smirnoff N (ed) Antioxidants and reactive oxygen species in plants. Blackwell, Oxford, pp 215–249
Le Deunff E, Davoine C, Le Dantec C, Billard J-P, Huault C (2004) Oxidative burst and expression of germin/oxo genes during wounding of ryegrass leaf blades: comparison with senescence of leaf sheaths. Plant J 38(3):421–431
Zimmermann G, Baumlein H, Mock H-P, Himmelbach A, Schweizer P (2006) The multigene family encoding germin-like proteins of barley. Regulation and function in basal host resistance. Plant Physiol 142(1):181–192
Çalişkan M (2000) The metabolism of oxalic acid. Turk J Zool 24:103–106
Luttrell BM (1993) The biological relevance of the binding of calcium ions by inositol phosphates. J Biol Chem 268(3):1521–1524
Apostol I, Heinstein PF, Low PS (1989) Rapid stimulation of an oxidative burst during elicitation of cultured plant cells: role in defense and signal transduction. Plant Physiol 90(1):109–116
Demarty M, Morvan C, Thellier M (1984) Calcium and the cell wall. Plant Cell Environ 7(6):441–448
Hepler PK (2005) Calcium: a central regulator of plant growth and development. Plant Cell 17(8):2142–2155
Lane BG, Bernier F, Dratewka-Kos E, Shafai R, Kennedy TD, Pyne C, Munro JR, Vaughan T, Walters D, Altomare F (1991) Homologies between members of the germin gene family in hexaploid wheat and similarities between these wheat germins and certain Physarum spherulins. J Biol Chem 266(16):10461–10469
Berna A, Bernier F (1997) Regulated expression of a wheat germin gene in tobacco: oxalate oxidase activity and apoplastic localization of the heterologous protein. Plant Mol Biol 33(3):417–429
de Marco A, Roubelakis-Angelakis K (1996) Hydrogen peroxide plays a bivalent role in the regeneration of protoplasts. J Plant Physiol 149(1–2):109–114
de Marco A, Roubelakis-Angelakis KA (1996) The complexity of enzymatic control of hydrogen peroxide concentration may affect the regeneration potential of plant protoplasts. Plant Physiol 110(1):137–145
Pasternak T, Ötvös K, Domoki M, Fehér A (2007) Linked activation of cell division and oxidative stress defense in alfalfa leaf protoplast-derived cells is dependent on exogenous auxin. Plant Growth Regul 51(2):109–117
Fry SC (2004) Primary cell wall metabolism: tracking the careers of wall polymers in living plant cells. New Phytol 161:641–675
Neutelings G, Domon JM, Membré N, Bernier F, Meyer Y, David A, David H (1998) Characterization of a germin-like protein gene expressed in somatic and zygotic embryos of pine (Pinus caribaea Morelet). Plant Mol Biol 38(6):1179–1190
Mathieu M, Neutelings G, Hawkins S, Grenier E, David H (2003) Cloning of a pine germin-like protein (GLP) gene promoter and analysis of its activity in transgenic tobacco Bright Yellow 2 cells. Physiol Plant 117(3):425–434
Thompson EW, Lane BG (1980) Relation of protein synthesis in imbibing wheat embryos to the cell-free translational capacities of bulk mRNA from dry and imbibing embryos. J Biol Chem 255(12):5965–5970
Liu Y, Ye N, Liu R, Chen M, Zhang J (2010) H2O2 mediates the regulation of ABA catabolism and GA biosynthesis in Arabidopsis seed dormancy and germination. J Exp Bot 61:2979–2990
Bishop-Hurley SL, Gardner RC, Walter C (2003) Isolation and molecular characterization of genes expressed during somatic embryo development in Pinus radiata. Plant Cell Tissue Organ Cult 74(3):267–281
Domon JM, Dumas B, Laine E, Meyer Y, David A, David H (1995) Three glycosylated polypeptides secreted by several embryogenic cell cultures of pine show highly specific serological affinity to antibodies directed against the wheat germin apoprotein monomer. Plant Physiol 108(1):141–148
Mathieu M, Lelu-Walter M, Blervacq A, David H, Hawkins S, Neutelings G (2006) Germin-like genes are expressed during somatic embryogenesis and early development of conifers. Plant Mol Biol 61(4):615–627
Caliskan M, Ozcan B, Turan C, Cuming AC (2004) Localization of germin genes and their products in developing wheat coleoptiles. J Biochem Mol Biol 37(3):339–342
Patnaik D, Khurana P (2001) Germins and germin like proteins: an overview. Indian J Exp Biol 39(3):191–200
Muthukrishnan S, Liang GH, Trick HN, Gill BS (2001) Pathogenesis-related proteins and their genes in cereals. Plant Cell Tiss Organ Cult 64(2):93–114
van Loon LC, Rep M, Pieterse CMJ (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44(1):135–162
Christensen AB, Thordal-Christensen H, Zimmermann G, Gjetting T, Lyngkjaer MF, Dudler R, Schweizer P (2004) The germinlike protein GLP4 exhibits superoxide dismutase activity and is an important component of quantitative resistance in wheat and barley. Mol Plant Microbe Interact 17(1):109–117
Manosalva PM, Davidson RM, Liu B, Zhu X, Hulbert SH, Leung H, Leach JE (2009) A germin-like protein gene family functions as a complex quantitative trait locus conferring broad-spectrum disease resistance in rice. Plant Physiol 149:286–296
Mittler R, Vanderauwera S, Suzuki N, Miller G, Tognetti VB, Vandepoele K, Gollery M, Shulaev V, Van Breusegem F (2011) ROS signaling: the new wave? Trends Plant Sci 16(6):300–309. doi:10.1016/j.tplants.2011.03.007
Veena HJ, Doerge RW, Gelvin SB (2003) Transfer of T-DNA and Vir proteins to plant cells by Agrobacterium tumefaciens induces expression of host genes involved in mediating transformation and suppresses host defense gene expression. Plant J 35(2):219–236
Ditt RF, Nester E, Comai L (2005) The plant cell defense and Agrobacterium tumefaciens. FEMS Microbiol Lett 247(2):207–213
Citovsky V, Kozlovsky SV, Lacroix B, Zaltsman A, Dafny-Yelin M, Vyas S, Tovkach A, Tzfira T (2007) Biological systems of the host cell involved in Agrobacterium infection. Cell Microbiol 9(1):9–20
Djamei A, Pitzschke A, Nakagami H, Rajh I, Hirt H (2007) Trojan horse strategy in Agrobacterium transformation: abusing MAPK defense signaling. Science 318(5849):453–456
Matzke MA, Mette MF, Matzke AJM (2000) Transgene silencing by the host genome defense: implications for the evolution of epigenetic control mechanisms in plants and vertebrates. Plant Mol Biol 43(2–3):405–415
Waterhouse PM, Wang MB, Lough T (2001) Gene silencing as an adaptive defence against viruses. Nature 411:834–842
Matzke MA, Matzke AJM (2004) Planting the seeds of a new paradigm. PLoS Biol 2(5):e133
Davoine C (2003) Rôle de l’oxalate oxydase dans la production de peroxyde d’hydrogène au cours de la sénescence des gaines foliaires de ray-grass (Lolium perenne L.) et dans la réponseà la blessure des limbes. PhD thesis, Université de Caen
Mittler R (2006) Abiotic stress, the field environment and stress combination. Trends Plant Sci 11:15–19
Suzuki N, Mittler R (2006) Reactive oxygen species and temperature stresses: a delicate balance between signaling and destruction. Physiol Plant 126:45–51
Shulaev V, Cortes D, Miller G, Mittler R (2008) Metabolomics for plant stress response. Physiol Plant 132:199–208
Quan LJ, Zhang B, Shi WW, Li HY (2008) Hydrogen peroxide in plants: a versatile molecule of the reactive oxygen species network. J Integr Plant Biol 50:2–18
Maraschin SF, de Priester W, Spaink HP, Wang M (2005) Androgenic switch: an example of plant embryogenesis from the male gametophyte perspective. J Exp Bot 56(417):1711–1726
Foyer C, Noctor G (2009) Redox regulation in photosynthetic organisms: signaling, acclimation, and practical implications. Antioxid Redox Signal 11(4):861–905. doi:10.1089/ars.2008.2177
Gadjev I, Stone JM, Gechev TS (2008) Programmed cell death in plants: new insights into redox regulation and the role of hydrogen peroxide. Int Rev Cell Mol Biol 270:87–144
Gechev TS, Van Breusegem F, Stone JM, Denev I, Laloi C (2006) Reactive oxygen species as signals that modulate plant stress responses and programmed cell death. BioEssays 28:1091–1101
Mazars C, Thuleau P, Lamotte O, Bourque S (2010) Cross-talk between ROS and calcium in regulation of nuclear activities. Mol Plant 3(4):706–718. doi:10.1093/mp/ssq024
Kairong C, Gengsheng X, Xinmin L, Gengmei X, Yafu W (1999) Effect of hydrogen peroxide on somatic embryogenesis of Lycium barbarum L. Plant Sci 146:9–16
Papadakis AK, Siminis CI, Roubelakis-Angelakis KA (2001) Reduced activity of antioxidant machinery is correlated with suppression of totipotency in plant protoplasts. Plant Physiol 126(1):434–444
Yeung EC, Belmonte MF, Tu LTT, Stasolla C (2005) Glutathione modulation of in vitro development. In Vitro Cell Dev Biol Plant 41:584–590
Noctor G (2006) Metabolic signalling in defence and stress: the central roles of soluble redox couples. Plant Cell Environ 29:409–425
Stasolla C (2010) Glutathione redox regulation of in vitro embryogenesis. Plant Physiol Biochem 48(5):319–327. doi:10.1016/j.plaphy.2009.10.007
Wu G, Shao HB, Chu LY, Cai JW (2007) Insights into molecular mechanisms of mutual effect between plants and the environment. A review. Agron Sustain Dev 27:69–78
Kim B-R, Nam H-Y, Kim S-U, Kim S-I, Chang Y-J (2003) Normalization of reverse transcription quantitative-PCR with housekeeping genes in rice. Biotechnol Lett 25:1869–1872
Acknowledgments
We wish to thank Dr P. Druart and Dr J-M. Jacquemin for their helpful insight. Many thanks, too, to C. Leroy for his excellent technical assistance throughout this work and to M-R. Bollen for her technical support in the histological study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Delporte, F., Muhovski, Y., Pretova, A. et al. Analysis of expression profiles of selected genes associated with the regenerative property and the receptivity to gene transfer during somatic embryogenesis in Triticum aestivum L.. Mol Biol Rep 40, 5883–5906 (2013). https://doi.org/10.1007/s11033-013-2696-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-013-2696-y