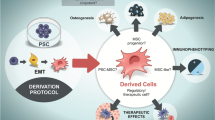
Abstract
The self-renewal and differentiation status of a stem cell is very important in the applications concerning regenerative medicine. Proliferation capacity, differentiation potentials and epigenetic properties of stem cells differ between sources. Studies have shown the high potentials of stem cells in iPS reprogramming. To examine this; we have compared the stem-ness and differential potential of four adult stem cells from common sources. We show a correlation between pluripotency and differentiation status of each stem cell with available data on the reprogramming efficiency. Four human adult stem cells including, adipose tissue-mesenchymal stem cells (AT-MSC), bone marrow mesenchymal stem cells (BM-MSCs), nasal septum derived multipotent progenitors (NSP) and umbilical cord blood stem cells (USSCs) were isolated and characterized. The self- renewal and differentiation potentials of each stem cell were assessed. Stem-ness transcription factors and the propagation potentials of all cells were analyzed. Furthermore the differentiation potentials were evaluated using treatment with induction factors and specific MicroRNA profile. Real-time PCR results showed that our stem cells express innate differentiation factors, miR145 and Let7g, which regulate the stem-ness and also the reprogramming potentials of each stem cell. To complete our view, we compared the propagation and differentiation potentials by correlating the stem-ness gene expression with differentiation MicroRNAs, also the direct effect of these factors on reprogramming. Our results suggest that the potentials of adipose tissue stem cells for GMP (Good Manufacturing Practice) compliant starting material are adequate for clinical applications. Our results indicate a low risk potential for AT-MSCs as starting material for iPS production. Although let7g and mir145 are well known for their differentiation promoting effects, but function more of a fine tuning system between self-renewal and differentiation status.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stem cells have opened a new era in cell based therapies. There are two types of stem cells: embryonic stem cells, which are isolated from the inner cell mass of blastocysts, and adult stem cells, which are found in various tissues. Three major accessible sources of autologous adult stem cells in the clinic are bone marrow, Adipose tissue and blood. stem cells are also isolated from umbilical cord blood just after birth. By definition, autologous cell therapy has been limited to the mentioned adult stem cells.
With the generation of induced pluripotent stem cells (iPS) a new revolution occurred in the field of regenerative medicine [1]. Resembling embryonic stem cells in their self-renewal potentials, iPS cells have the advantage of patient specificity, which brings the experimental limitations a step closer to the clinical level [2]. The initial protocol, better known as “Yamanaka method”, used a set of genes (Oct-3/4, Sox2, Klf4, cMyc)using a retroviral system to induce pluripotency [3, 4]. Likewise, The same results were achieved through exogenous expression of a different set of genes (Oct-3/4, Sox2, Nanog, Lin28)using a lentiviral system [5].
One of the main hurdles has been to develop efficient reprogramming techniques that are functional and safe [4–6]. To this extend, a wide range of cellular and molecular modulations have been applied to gain higher efficiency and efficacy for both research and clinical purposes [7].
iPS cells have been developed from a broad range of species [8–10], as well as different sources of somatic cells including fibroblasts [3, 4], adipose cells [11], keratinocytes [12], neural stem cells [13], hepatocytes [14] and astrocytes [15]. Epithelial cell types, such as keratinocytes, liver and stomach cells, can be converted to iPS cells with higher efficiency compared to skin fibroblast cells which suggests that different cell types may possess different degrees of plasticity [12, 16].
In addition, it seems there is a correlation between differentiation stage and reprogramming efficiency [17]. In recent studies it was found that early passage iPS cells still retain some degree of somatic cell memory. Although these remaining epigenetic memories appeared to attenuate after continuous in vitro culture but they can influence the differentiation preference of these cells [17–19]. Considering the remaining epigenetic memory on somatic cells, not only the efficiency but the requirements to induce such a state are also consequently affected.
It has been reported that using as many as two or just a single transcription factors can induce the same effect and gain pluripotency [13, 20]. It seems that the basic needs for induction are met according to the nature and potentials of each cell source.
MicroRNAs as part of the epigenetic signature of each cell, are approximately 22-nucleotide RNAs that regulate gene expression by binding to complementary sequences in the 3′ untranslated regions of protein coding mRNAs, which eventually induce their degradation or translation inhibition [21]. Two important and prevalent MicroRNAs that have been detected among somatic cells are mir145 and let7g [22–24].
It is found that the sequential expression of let7 RNAs regulate and synchronize specific stages of development [25, 26]. Let-7g levels have been demonstrated to be regulated by Lin28 through inhibition of Dicer-mediated processing of pre-let-7 to mature let-7 [27, 28]. Promoters of both let-7g and Lin28 are occupied by the embryonic transcription factors Oct4, Sox2, Nanog, and Tcf3 in mice, suggesting these factors promote the transcription of both primary let-7g and Lin28, which eventually blocks the maturation of let-7g [29].
It has been found that mirR-145 functions to regulate as well as modulate the differentiation progress in Human Embryonic Stem Cells (hESC) through Oct4/Sox2 pathway [23]. Identification and the unique conservation of mir145 in many species [30, 31]and organs [32, 33] shows the evolutionary importance and critical impact of this MicroRNAs in cell fate [34].It has been demonstrated that mir145 represses pluripotency and controls ESC differentiation through interaction with three other core pluripotency factors Oct4, Sox2, and Klf4 [23]. In addition, up-regulation of miR-145 expression caused a significant diminution of the self-renewal marker SSEA4 and an increase in multiple differentiation markers associated with all three germ layers [23].Also, the reverse has been observed, with down-regulation of both MicroRNAs which has gained an up-regulation in pluripotency-related genes [35].
Regarding the potential of stem cells in regenerative medicine and cell therapy, the source of stem cells is an essential factor. Many factors affect the final fate of a stem cell and commitment to certain functions [36]. Therefore, the initial step for optimization is choosing the best cell source, the availability and reliability followed by reprogramming efficiency and kinetics [37]. There has been continuous effort in generating a GMP-compliant system to produce and maintain iPS cells for clinical purposes. To date various types of adult stem cells have been separated [36].
In this study, considering the potentials of stem cells for iPS generation, we have compared four types of adult stem cells with mesenchymal origin. Stem cells were isolated from bone marrow, adipose tissue, umbilical cord blood and also nasal septum. We have compared their pluripotency-related genes with their differentiation potentials and their correlation to self-renewal and reprogramming. Features such as, ease of isolation (AT-MSCs), availability in cell banks (USSCs), high propagation level (NSPs) and high differentiation potential (BM-MSCs) can render stem cells as candidates for iPS generation. The aim of this study is to analyze the potentials of generating iPS cells under standard conditions, compared to the genetic make-up of MSCs derived from different ontogenic sources.
Material and method
Isolation and characterization of adult stem cells
Isolation and culture of unrestricted somatic stem cells from umbilical cord blood (USSCs)
All procedures were approved by the ethical committee of Stem Cell Technology Research Center (Tehran, Iran). Isolation of USSCs has been primarily described by Kogler et al. [38]. In brief, cord blood samples were collected from the umbilical cord vein of neonates with informed consent of their mother. In our work, USSCs were successfully isolated from 4 out of 11 cord blood samples and studied. Ficoll (Pharmacia-Amersham) gradient separation of the mononuclear cell fraction accompanied with lysis of RBCs by ammonium chloride. Cells were plated out in growth medium at an average 6 × 106 cells/ml in T25 culture flasks. Growth Medium consists of low glucose DMEM (GIBCO) with 30 % fetal calf serum (GIBCO), dexamethasone (107M; Sigma-Aldrich), penicillin (100 U/ml; GIBCO), streptomycin (0.1 mg/ml; GIBCO), and ultra-glutamine (2 mM; GIBCO). USSCs are expanded in the same medium with the dexamethasone omitted and lower concentration of FCS (10 %).
Isolation and culture of mesenchymal stem cells from human bone marrow (hBM-MSCs)
Bone marrow MSCs were isolated using BM aspirations collected from six healthy donors aged between 26 and 35 years old (both male and females),with informed consent of the patients (Bone Marrow Transplantation Center, Shariati Hospital, Tehran, Iran). As previously reported [38–40], BM aspirates were isolated over a Ficoll-Hypac gradient separation. The mononuclear cells were recovered and were seeded at 5 × 10 6cell density into 25 cm2 flasks containing DMEM supplemented with 2 mM GlutaMAX-I, 10 U/ml penicillin, 100 mg/ml streptomycin and 10 % FBS. Reaching 90 % confluency the MSCs were replated at 1.5 × 10 5cells in a 25 cm2 flask using 0.25 % trypsin–EDTA.
Isolation and culture of adipose tissue-mesenchymal stem cells (AT-MSC)
Human adipose tissues of four healthy donors, ages between 23 and 35 years, were obtained from elective liposuction procedures with informed consent of the patient. As described in previous literature, AT-MSC were isolated Using a two-step digest in Krebs–Ringer (pH 7.4) buffered with 25 mM HEPES containing 20 mg/ml bovine serum albumin(BSA) and 1.5 mg/ml collagenase (type I) [41, 42]. Following filtration through a 70 μm mesh filter cell suspensions were centrifuged, and decontaminated for erythrocytes using RBC lysis buffer at pH 7.3. After washing, filtered cells were cultivated in DMEM supplemented with 2 mM GlutaMAX-I, 10 U/ml penicillin, 100 mg/ml streptomycin and 10 % FBS as described for BM-MSC.
Isolation and culture of nasal septum derived multipotent progenitors (NSP)
As previously described [43], human nasal septal cartilage samples were obtained from patients undergoing septoplasty or septorhinoplasty operations with their informed consent. Cartilage samples were obtained from healthy male and female donors (25–40 years old). Briefly, the procedures involve incision, enzymatic digestion and filtration of cartilage specimens to separate the NSPs from their extra cellular matrix (ECM). After isolation of each individual specimen, the cells are re-suspended in low glucose-DMEM containing 15 % FBS (Gibco) 10 μg/ml ascorbic acid (Sigma), 1 % penicillin/streptomycin (Sigma) and 1.25 μg/ml amphotericin-B (Sigma).Plated at a density of 104 cell/cm2in a 75 cm2 culture flasks, the colonies were separated and sub cultured.
Flowcytometery analysis
Immunophenotyping profile of each stem cell was done using flowcytometery analysis for different cell surface antigens. All cell types including NSP, AT-MSC, BM-MSC and USSC were used to prepare a single cell suspension. To analyze intracellular markers, cells were permeabilized with 0.5 % Triton X-100. To block non-specific binding a solution of 3 % human serum in PBS was added to cell suspension for 30 min. Cell suspensions were stained with following mouse monoclonal antibodies against human CD105, CD106, CD90, CD166, CD45, HLA-ABC, HLA-DR (all from eBioscience), CD34, CD133 (Prominin-1) (Dako) and CD271,OCT4 (Santa Cruz). Relative to each marker the matching isotype was used as control to detect non-specific binding. Furthermore, the relevant PE-labeled secondary antibody was applied to cell suspensions, then fixed with 2 % paraformaldehyde in PBS and analyzed on a FACS Caliburcytometer (Becton–Dickinson) with WinMDI 2.8 software.
Stem cells were detached by trypsin/EDTA and incubated with the specific antibodies or corresponding isotype controls in 100 μl of PBS-BSA 3 % for 1 h at 4 °C. Approximately 105–106 cells were used for each analysis. The cells were then fixed with 1 % paraformaldehyde and analyzed with a periodic acid-Schiff (PAS) flowcytometer using FloMax software (Partec, Münster, Germany). Flowcytometry data are available in Table 1.
MTT assay
In order to determine the proliferation capacity of each stem cell we assessed the viability using MTT (Methylthiazolyldiphenyltetrazolium bromide) reduction assay. An average 1,500 cells were seeded in a 96 well plate Leaving 8 wells empty for blank controls. Stem cells were incubated (37 °C, 5 % CO2) for 1–5 days. MTT solution (5 mg/ml in PBS) was added and cells were incubated for further 3 h (37 °C, 5 % CO2). Formazancrystal (MTT metabolic product)was re-suspended in DMSO (Dimethyl sulfoxide) and the optical density was observed at 570 nm and subtracted from background at 670 nm.
MicroRNA isolation and real time PCR
MicroRNA extraction was carried out using our modified method of total RNA isolation with QIAzolLysis Reagent. Extended incubations and centrifugations at higher speeds were applied in this system.
The expression of MicroRNAs was analysed using 1st-Strand cDNA Synthesis Kit (Agilent Technologies, Stratagene Products Division). According to the instructions, first the MicroRNAs are elongate in a polyadenylation reaction (PAP product), which are then reverse transcribed using the universal reverse primer provided, into QPCR-ready cDNA. The product can then be amplified using a unique forward primer that is specific to MicroRNA target (Supplementary Table 1). cDNA Synthesis was followed by 40 cycles of real time PCR using the MicroRNA QPCR Master Mix (Agilent Technologies, Stratagene Products Division). MicroRNA levels were normalized against the snoRNA control (U6-snoRNA).
To screen for contamination and specific amplification, a no-PAP control cDNAwas prepared from a polyadenylation reaction in which the poly A polymerase is omitted. Quantitative real time pcr reactions were carried out according to manufacturer instructions. The Rotor gene 6000 detection system (Corbett) was used for MicroRNA transcript expressions.
Total RNA isolation and real-time polymerase chain reaction
Total RNA was extracted using QIAzol–reagent (Qiagen) from all stem cells under study. Synthesis of cDNA was carried out with MMuLV reverse transcriptase (RT) and random hexamer according to the manufacturer’s instructions (Fermentas). The Hypoxanthine–guanine phosphoribosyltransferase (HPRT) gene was used as the internal control. PCR amplification was performed using Maxima™SYBR Green/Fluorescein qPCR Master mix (Fermentas) with a two-step procedure of an initial denaturation at 95 °C for 10 min, followed by cycles circulating 15 s of 95 °C and 60 s of annealing/Extension at 60 °C. The sequence of primers and product lengths are listed in Supplementary Table 2. All reactions were performed in triplicates and normalized to HPRT gene. Changes in microRNA and mRNA expressions were normalized to the lowest expressing cell type which was subsequently calculated using the 2-ΔΔCt method. The Rotor gene 6000 detection system (Corbett) was used for quantitative mRNA transcript expressions.
In vitro differentiation potentials
Osteogenic differentiation
To assess the potential of osteogenic differentiation, 1–2 × 104 cells/cm2 were cultured in osteogenesis medium DMEM with 10 % FBS (Gibco) containing 10−7 M dexamethasone, 0.2 mM ascorbic acid 2-phosphate, and 10 mM β-glycerophosphate (Sigma products). Medium was changed every 3 days. After 3 weeks of induction, cells were stained with Alizarin Red solution (Sigma) to assess mineralization [44–46].
Adipogenic differentiation
For adipogenic differentiation potentials, cells were plated at 1–2 × 104 cells/cm2 density in DMEM with 10 % FBS (Gibco) medium supplemented with 0.5 mM Hydrocortisone, 0.5 mM Isobutylmethylxanthine and 60 mM Indomethacin (all from Sigma) was incubated for 3 weeks. Medium was changed every 3 days. After 3 weeks of induction, cells were stained with oil red O solution (Sigma) [39, 44].
Chondrogenic differentiation
Chondrogenenic potential of each stem cell was examined as follow; 2 × 105 cells were pelleted at 350 g for 4 min. After 24 h of incubation, media was changed with 500 μl of chondrogenic medium containing high glucose DMEM with 1 % ITS supplements (625 mg/ml insulin, 625 mg/ml transferrin, 625 ng/ml selenious acid) (Gibco), 50 μg/ml ascorbate 2-phosphate (Sigma), 100 nM dexamethasone (Sigma), 10 ng/ml fibroblast growth factor-2 (bFGF) (Peprotech). The pellets were maintained in this medium for 3 weeks with medium exchange of every 3 days. Then, the aggregates were harvested and sectioned using standard histology protocols and stained with Alcian Blue (Sigma) detecting glycoproteins in the extra cellular matrix.
Statistical analysis
The relative quantification of MicroRNA and gene expression was calculated using the comparative threshold cycle (Ct) method. The crossing point (Ct) for each transcript was determined using second derivative maximum method of Rotor gene software v3.5.3 (Rotorgen6).Data gained from MTT assay and the gene expression were analysed as mean–SD with one-way analysis of variance using SPSS v.16 software and a P value less than 0.05 was considered as statistical significance.
Results
Stem cell isolation and characterization
All Stem cells were expanded and cultivated up to two passages to gain the adequate, uniform and juvenile cells for further analysis. Stem cell isolation and characterizations were performed as described in pervious literature by flowcytometry analysis (Table 1) and also growth factor based induction of differentiation (Fig. 1) [38, 43, 44]. Regarding these data, it could be considered that the rate of accessibility of each stem cell could be ranked as NSP in the leading sector fallowed by USSC, AT-MSC and BM-MSC (data not shown).
The differentiation potential of isolated stem cells (to osteogenic, adipogenic and chondrogenic lineages) was assessed using induction media. All cultures were maintained under induction medium for a minimum of 3 weeks and stained with Alizarin Red (second column from left), Oil Red O (third column from the left) and Alician blue (forth column from left). The NSP cells did not differentiate to adipocytes after 3 weeks in differentiation medium. Bars: 200.m. 420 × 1 (300 × 300). (Color figure online)
Propagation and doubling time
MTT assay was performed at indicated time points (day 0, 1, 3 and 5) to measure the viability and proliferation rate of each stem cell. All tests were performed in triplicates and the statistical significance was assessed using one-way anova (p value < 0.05). As outlined in Fig. 2, there is an overall increasing trend in the proliferation rate of all examined cells [43].The increasing trend in cell proliferation from day 1 to day 3 was significant for all stem cells and the same for the next time point (day 5) where all the cells showed significant increase in proliferation. But in comparing stem cells with each other, the only significant increase could be dedicated to NSPs and on the transition from day 3 to day 5 none of the other stem cells (BM-MSCs, AT-MSCs and USSCs) showed significant data associated to each other.
MTT assay: The comparison of proliferation capacity between different cell types, i.e. USSC, BM-MSC, NSP and AT-MSC was done using MTT assay protocol; Asterisk (*): Significant difference in proliferation of NSPs was seen in comparison to other cell types. Statistical analysis was performed using one-way anova (p < 0.05). Error bars represent standard deviation (SD) of each data set, which is the result of triplicate samples
MicroRNA and gene expression analysis
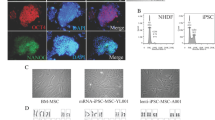
RNA samples were obtained from all undifferentiated stem cells and mRNA expression of pluripotency/reprogramming -specific genes, including Oct4, Nanog, Sox2, Klf4, cMyc, Lin28 and Rex-1 were analyzed by Real Time-PCR. Using HPRT as the internal control the gene expression was normalized and relatively compared in regards to the stem cell with the lowest expressing genes, i.e. USSCs (Fig. 3). As indicated in Fig. 3, AT-MSCs show the highest expression of Sox2, Klf4 and Lin28 but the lowest in Oct4 and cMyc genes. On the other hand, BM-MSCs express Nanog and cMyc more than any other line, but the lowest expression of Rex-1. And finally, USSC and NSP show the highest expression in Rex-1 and Oct4, respectively. Noteworthy, USSCs were the least expressing cells in almost all genes which make USSCs the base line of comparison and also with the most statistical significance with other gene expression data (Fig. 3).
Expression of specific reprogramming genes: The relative expression was calculated relative to USSCs using the 2-ΔΔCt method. Each data set is the result of triplicate samples. Statistical analysis was performed using one-way anova (p < 0.05). Asterisk (*) Statistically significant up-regulation in comparing four different stem cell groups (p < 0.05). (**): Statistically significant down-regulation of gene expression between examined stem cell groups. Each data set is the result of triplicate samples. Error bars represent standard deviation (SD) of each data set
Our results indicate an expression level of both differentiation MicroRNAs in all samples. MicroRNA expression was analyzed in regards to a no-PAP control which is the exact replicate of the test samples during cDNA synthesis without PolyA polymerase. This control will eliminate any non-specific expansions due to mispriming. The Expression of differentiation MicroRNAs, let7g and miR145, were normalized to U6 internal control. Relative quantification of miR145 and let7g were evaluated relevant to chondrogenic progenitor (NSP) and Adipose tissue MSCs, respectively (Fig. 4). Statistical analysis of the MicroRNAs was performed at two levels; first the lowest expressing cell;(i.e. AT-MSC for let7g and NSP for miR145). Secondly, to other stem cells. Data analysis of miR145, express that all groups are significantly up regulated relative to NSP, whereas let7g is only significantly higher in USSCs compared to AT-MSCs. Interesting to note that USSC’s expression of miR145 is also significantly higher compared to other stem cells (i.e. BM-MSCs and NSP).
Expression of differentiation MicroRNAs, let7g and miR145 normalized to U6 internal control in USSCs, BM-MSCs, NSP and AT-MSC. All PCR reactions were performed in triplicates. Relative quantification of miR145 and let7g were assessed relevant to chondrogenic precursors (NSP) and adipose tissue MSCs, respectively. The statistical significance (p < 0.05) was analyzed using one way-anova Test. Asterisk (*) Statistical significance relevant to the expression of the stem cell considered as base; (i.e. AT-MSC for let7g and NSP for miR145). (**): Statistical significance compared to other stem cell groups. Error bars represent standard deviation (SD) of each data set, which is the result of triplicate samples
Differentiation potential
Differentiation capacities of stem cells were assessed by in vitro differentiation to osteocytes, adipocytes and chondrocytes. Osteogenesis was detected with the creation of mineral aggregates as nodule like structures which was stained with Alizarin Red. All cells showed the potentials for osteogenesis (Fig. 1). No mineralization of calcium phosphate was observed in control cells maintained without the osteogenic medium (data not shown). Adipogenic induction after 21 days were assessed with oil red staining. Visualizing oily vesicles indicate the adipogenic differentiation. After 21 days of differentiation, all except NSP showed the potentials for adipogenesis. The Oil Red stained cultures of differentiated USSC, BM-MSC, NSP and AT-MSC are shown in Fig. 1. Chondrogenic differentiation was assessed using Alcian Blue staining. All stem cells showed the potential for this lineage.
Discussion
In our study, we have compared four available adult stem cells for reprogramming strategies. In this study we examined the effective factors for determining the best possible source for iPS generation for clinical purposes. Looking for special features of stem-ness in adult stem cells which should contribute to susceptibility in reprogramming, we examined the expression of known reprogramming factors including, Oct4, Nanog, Sox2, Rex-1, cMyc, Klf4 and Lin28.
Taking USSCs as the base line for observations, real time PCR data showed significant differences between each cell line. As indicated in Fig. 3, AT-MSCs show the highest expression of Sox2, Klf4 and Lin28 but the lowest in Oct4 and cMyc genes. On the other hand, BM-MSCs express Nanog and cMyc more than any other line, but the lowest expression of Rex-1. And finally, USSCs and NSP show the highest expression in Rex-1 and Oct4, respectively. Noteworthy, USSCs were the least expressing cells in almost all genes except Rex-1, which makes USSCs the base line of comparison.
For further assessment of the difference, we connected the reprogramming gene expression data with the expression of differentiation MicroRNAs. Interestingly, USSCs with the least expressing cells in almost all genes showed the highest expression in differentiating MicroRNAs. As previously demonstrated, mir145 regulates Oct4/Sox2 pathway as well as Klf4 [23], which could cause for the low expression of all three factors in USSCs (Fig 3). Similarly, it has been established that let7g regulates and is regulated by Lin28 directly [47], which again is highly expressed in USSCs. Consequently, Lin28 has the least expression in USSCs due to the high occurrence of let7g (Fig. 4). Logical as it may seem, the efficiency of reprogramming for USSCs is an average 0.13 %. Compared to conventional fibroblast generation of iPS which is about 0.2 %, the potential of USSCs for reprogramming are more than half as fibroblasts [3].
USSCs are a sub group of stem cells derived from umbilical cord blood [38]. Cord blood, like tooth germs of human third molar usually discarded as clinical wastes, are a valuable cell source for generation of iPS cells [48]. These pluripotent stem cells possess high propagation potentials and low immunogenicity. Initial generation of iPS cells using USSCs applied a retroviral vector encoding the human complementary DNAs of Oct4, Sox2, Klf4, and c-Myc [20]. Cord blood banks have made it possible for non-invasive access to juvenile stem cells aimed for clinical therapies. USSCs have been regarded as one of the best potential allografts due to their lack of HLA-DR surface antigens, making cross matching less of a hurdle [49].
Adipo tissue MSCs express the highest level of pluripotency genes compared to their counter parts. Interestingly, AT-MSCs possess very low expression of miR145, indicative of the direct effect of this MicroRNA on the pluripotency network [23]. The same coordination was observed with let7g, where the target gene Lin28, is at the highest level compared to other stem cells (Fig. 3). It is only logical that the regulatory MicroRNAs should control their targets. On contrary, the only conflict observed was the case with Oct4 where we expected a higher level of expression regarding the depletion of miR145. Although Oct4 is essential for pluripotency, but there are numerous parameters [50] that affect Oct4 expression, such as, long non coding RNAs [51], other MicroRNAs [52], transcription factors [53, 54] and even at post translational level by certain signaling pathways [55].
New approaching sources of stem cells such as adipose tissue MSCs provide alternative choices for the clinic. Considering that these stem cells exhibit high intrinsic expression of self-renewal supporting factors and can effectively serve as feeder layers of their own or become independent pluripotent cells [11].
Under specific conditions, reprogramming AT-MSCs has resulted in production of ES-like iPS cell colonies at comparable efficiencies (0.25 ± 0.11 % with feeders vs. 0.42 ± 0.17 % without feeders), indicating that Adipocytes do not require exogenous factors to support the growth of iPS cells [11].
Human adipose sources were also retro virally transduced with human Yamanaki’s factors and efficiently gave rise to hiPS colonies were 0.74 % of AT-MSCs cells formed iPS colonies, compared to 0.28 % for human keratinocytes, which is considered the most efficient human cells to give rise to hiPS cells to date [11, 12]. Interestingly, a recent report examined the effect of lineage markers (Lin) on generation of iPS cells from different sources of somatic stem cells [11]. According to their result, Lin+ cells such as erythrocytes (Ter119), endothelial cells (CD31), and hematopoietic (CD45) cells were not as efficient as the Lin− cells which consisted of preadipocytes and mADS cells [56]. The same was observed in other lineages such as the liver. It has been depicted that Liver progenitor cell (LPC) have intrinsic, cell proliferation–independent characteristics which results in 275-fold increase in reprogramming compared to differentiated liver cells [57]. AT-MSCs provide xeno- and feeder-free conditions for hiPS generation at similar efficiencies which are amenable for designing a GMP-compliant system.
The results here discussed are in accordance with previous reports which suggest notable intrinsic potentials for adult adipose-derived cells to support proliferation and preservation of self-renewal of autologous pluripotent cells [11, 58].
In contrast to adipose tissue-MSCs, NSPs are stem cells derived from an ecto-mesodermal source, but the gene and MicroRNA pattern is the opposite of adipose tissue derived MSCs. Although mir145 and let7g expression are also very low in NSPs, but in this case OCT4 expression is the highest among all cells examined, and a fare expression of the other two factors (i.e. Sox2 and Klf4 Fig. 3).
As shown in Fig. 2, NSPs shows great propagation potentials compared to its counterparts, which will make NSPs a good candidate for reprogramming. This feature could be dedicated to the high level of oct4 expression, although not entirely. Considering that stem cells have higher proliferation rate than somatic cells, further manipulations such as gene therapies could feasibly be acquired. The MTT assay shows the viability, which in this case correlates with proliferation of stem cells. It should be noted that in comparing the proliferation rate of stem cells, the significancy between cells shows a considerable difference. Therefore NSPs could possess high potentials for reprogramming, but further manifestations and experiments are required to prove the capacity of this progenitor.
As for bone marrow derived MSCs, gene and MicroRNA expression profile indicate a fairly high expression level for all genes and also their target MicroRNAs. It seems that BM-MSCs have been able to control the system in a mediocre level where all genes examined are maintained at medium. A reasonable coordination between genes and the regulatory MicroRNAs, is clearly prominent in our data.
According to previous reports, mouse BM-MSC have been used for iPS production [59]. For this, a retroviral system containing mouse Oct4, Sox2, Klf4, and c-Myc (OSKM package) were introduced into BM-MNCs (bone marrow-smono nuclear cells) and MEF (Mouse Embryonic Fibroblast) cells. Although this report refers to an animal model of stem cell derived iPS cells, our result suggest this stem cell to be of great potentials for human iPS generation. According to our results naive BM-MSCs express all reprogramming factors and also both differentiating MicroRNAs regulating the reprogramming circuit. BM-MSCs do not possess high self-renewal capacity, taking to account that chromosomal abnormalities have been reported with high passages, it is preferable to use low passage expands for clinical purposes [60].
Our results demonstrate a high capacity of stem-ness in all four isolates, with minor differences. Taken the differences observed, we looked for the cause; and among the regulators of the stem cell factors the regulatory role of MicroRNAs has been demonstrated. Mir145 and let7g are of the well-known regulators of differentiation. Although differentiating MicroRNAs are expressed consecutively with stem factors but it appears to fine tune the expression rather to down regulate it. This system allows the self-renewal to continue and under specific stimulus turn around to initiate differentiation.
As demonstrated in Fig. 4, USSCs have the highest level of mir145 and let7g expression. Consequently, USSCs should be regarded with the highest potential for differentiation, but according to our results it seems that BM-MSCs possess such characteristics. BM-MSCs with initial and intense differentiation properties compared to its counterparts alternate more easily [61] (data not shown).
Human iPSCs have offered exciting solutions to allay the ethical concerns with human Embryonic Stem Cells. Regarding the epigenetic memory left in iPS cells derived from fully differentiated cells, stem cells are considered better candidates as the starting material for reprogramming [62]. Providing that stem cells have high propagation potentials, such sources are amenable for further manipulations desired, such as gene therapy and tissue engineering purposes.
In a comprehensive study, it was demonstrated that the origin of the various mouse iPSCs influence the tumor-forming propensities in cell transplantation [63]. They established that mouse tail-tip fibroblast iPSCs (mesoderm origin) have the highest tumorigenic propensity, where gastric epithelial and hepatocyte derived iPSCs (endodermal origin) show significantly lower tumorigenic propensities [63].
This can show the importance of cell origin on the safety and functionality of human iPSCs. Human iPSCs have been derived mostly from the mesodermal (i.e. fibroblasts and blood cells) or the ectodermal origin cells (i.e. keratinocytes, and neural stem cells).To keep that majority, we have also chosen a group of stem cells with mesodermal (BM-MSC, USSC, AT-MSC) and ectomesodermal (NSP) origin. It is interesting to compare gene and MicroRNA expression patterns of iPS sources from different lineages.
Many factors affect the efficiency of iPS production including epigenetic factors such as chromosome remodeling, DNA methylation and histone acetylation [18, 64]. Another epigenetic factor considered a difference between induced and natural pluripotency is the MicroRNA profile [65]. The MicroRNA expression observed in iPS compared to embryonic stem cells show that the reprogramming occurred are relatively varied and dependent on many factors [66].
This study suggests that specific MicroRNAs are able to fine tune the reprogramming process and may be useful to reduce the heterogeneity in iPS cells. The fact that Klf4and cMYC can be replaced by NANOG and LIN28 in human fibroblast reprogramming experiments suggests that different molecular pathways can lead to reprogramming or, alternatively, that these factors perform highly similar functions during this process. In support of the latter, LIN28 was recently found to function as a negative regulator of MicroRNA processing in ES cells, specifically of members of the let-7 family [27]. cMYC represses the transcription of similar MicroRNAs, suggesting that LIN28 and cMYC could perturb the same regulatory mechanisms that contribute to reprogramming [67, 68]. For example, in another insightful study [69] it was shown that over-expression of Lin28 shortened the cell cycle in monoclonal B cells and speed up iPS cell generation.
Although the functional targets of let7g and mir145 have been identified [28, 33, 70–73], their role in reprogramming could be further examined through inhibition experiments. Altogether, we suggest that AT-MSCs may be the preferable source for iPS generation in the clinical field, considering the ease of isolation, high rate of reprogramming, feeder free conditions and the advantage of high expression of reprogramming genes and also, the low expression in differentiating markers miR145 and let7g. But further investigations are necessary to guarantee clinical safe protocols for generating therapeutic induced pluripotent stem cells.
References
Chen YC, Tsai KL, Hung CW, Ding DC, duChen LH, Chang YL et al (2011) Induced pluripotent stem cells and regenerative medicine. J Clin Gerontol Geriatrics 2(1):1–6
Caulfield T, Scott C, Hyun I, Lovell-Badge R, Kato K, Zarzeczny A (2009) Stem cell research policy and iPS cells. Nat Methods 7(1):28–33
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K et al (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131(5):861–872
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676
Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S et al (2007) Induced pluripotent stem cell lines derived from human somatic cells. Science 318(5858):1917
Narsinh KH, Wu JC (2010) Gene correction in human embryonic and induced pluripotent stem cells: promises and challenges ahead. Molecular therapy: the journal of the American Society of Gene Therapy 18(6):1061
Sun N, Longaker MT, Wu JC (2010) Human iPS cell-based therapy: considerations before clinical applications. Cell cycle (Georgetown, Tex) 9(5):880
Esteban MA, Xu J, Yang J, Peng M, Qin D, Li W et al (2009) Generation of induced pluripotent stem cell lines from Tibetan miniature pig. J Biol Chem 284(26):17634
Chang MY, Kim D, Kim CH, Kang HC, Yang E, Moon JI et al (2010) Direct reprogramming of rat neural precursor cells and fibroblasts into pluripotent stem cells. PLoS ONE 5:e9838
Wu Z, Chen J, Ren J, Bao L, Liao J, Cui C et al (2009) Generation of pig induced pluripotent stem cells with a drug-inducible system. J Mol Cell Biol 1(1):46
Sugii S, Kida Y, Kawamura T, Suzuki J, Vassena R, Yin YQ et al (2010) Human and mouse adipose-derived cells support feeder-independent induction of pluripotent stem cells. Proc Natl Acad Sci 107(8):3558
Trond Aasen AR, Maria JB, Elena Garreta AC, Federico Gonzalez RV (2008) Efficient and rapid generation of induced pluripotent stem cells from human keratinocytes. Nat Biotechnol 26(11):1276–1284
Kim JB, Greber B, Araúzo-Bravo MJ, Meyer J, Park KI, Zaehres H et al (2009) Direct reprogramming of human neural stem cells by OCT4. Nature 461(7264):649–653
Liu H, Ye Z, Kim Y, Sharkis S, Jang YY (2010) Generation of endoderm derived human induced pluripotent stem cells from primary hepatocytes. Hepatology 51(5):1810–1819
Ruiz S, Brennand K, Panopoulos AD, Herrerías A, Gage FH, Izpisua-Belmonte JC et al (2010) High-efficient generation of induced pluripotent stem cells from human astrocytes. PLoS ONE 5(12):e15526
Aoi T, Yae K, Nakagawa M, Ichisaka T, Okita K, Takahashi K et al (2008) Generation of pluripotent stem cells from adult mouse liver and stomach cells. Science 321(5889):699
Eminli S, Foudi A, Stadtfeld M, Maherali N, Ahfeldt T, Mostoslavsky G et al (2009) Differentiation stage determines potential of hematopoietic cells for reprogramming into induced pluripotent stem cells. Nat Genet 41(9):968–976
Kim K, Doi A, Wen B, Ng K, Zhao R, Cahan P et al (2010) Epigenetic memory in induced pluripotent stem cells. Nature 467(7313):285–290
Polo JM, Liu S, Figueroa ME, Kulalert W, Eminli S, Tan KY et al (2010) Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nat Biotechnol 28:848–855
Zaehres H, Kogler G, Arauzo-Bravo MJ, Bleidissel M, Santourlidis S, Weinhold S et al (2010) Induction of pluripotency in human cord blood unrestricted somatic stem cells. Exp Hematol 38(9):809–818
Stefani G, Slack FJ (2008) Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol 9(3):219–230
Peter M (2009) Let-7 and miR-200 microRNAs: guardians against pluripotency and cancer progression. Cell cycle (Georgetown, Tex) 8(6):843
Xu N, Papagiannakopoulos T, Pan G, Thomson J, Kosik K (2009) MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 137(4):647–658
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C et al (2007) Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131(6):1109–1123
Carrington J, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301(5631):336
Reinhart B, Slack F, Basson M, Pasquinelli A, Bettinger J, Rougvie A et al (2000) The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403(6772):901–906
Viswanathan SR, Daley GQ, Gregory RI (2008) Selective blockade of microRNA processing by Lin28. Science 320(5872):97
Newman MA, Hammond SM (2010) Lin-28: an early embryonic sentinel that blocks Let-7 biogenesis. Int j Biochem Cell Biol 42(8):1330–1333
Marson A, Levine S, Cole M, Frampton G, Brambrink T, Johnstone S et al (2008) Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134(3):521–533
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294(5543):853
Michael MZ, O’Connor SM, Van Holst Pellekaan NG, Young GP, James RJ (2003) Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1:882–891
Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A et al (2007) A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129(7):1401–1414
Cordes K, Sheehy N, White M, Berry E, Morton S, Muth A et al (2009) miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature 460(7256):705–710
Hatfield S, Ruohola-Baker H (2008) MicroRNA and stem cell function. Cell Tissue Res 331(1):57–66
Adegani FJ, Langroudi L, Arefian E, Soleimani M (2011) Differentiation microRNAs affect stemness status of USSCs. Iranian Red Crescent Medical Journal 13(10):726
Watt FM, Driskell RR (2010) The therapeutic potential of stem cells. Philosophical Transactions of the Royal Society B 365(1537):155
González F, Boué S, Belmonte JCI (2011) Methods for making induced pluripotent stem cells: reprogramming a la carte. Nat Rev Genet 12:231–242
Kögler G, Sensken S, Airey JA, Trapp T, Müschen M, Feldhahn N et al (2004) A new human somatic stem cell from placental cord blood with intrinsic pluripotent differentiation potential. J Exp Med 200(2):123–135
Pittenger M, Mackay A, Beck S, Jaiswal R, Douglas R, Mosca J et al (1999) Multilineage potential of adult human mesenchymal stem cells. Science 284:143–147
Majumdar MK, Thiede MA, Mosca JD, Moorman M, Gerson SL (1998) Phenotypic and functional comparison of cultures of marrow-derived mesenchymal stem cells (MSCs) and stromal cells. J Cell Physiol 176(1):57–66
Wagner W, Wein F, Seckinger A, Frankhauser M, Wirkner U, Krause U et al (2005) Comparative characteristics of mesenchymal stem cells from human bone marrow, adipose tissue, and umbilical cord blood. Exp Hematol 33(11):1402–1416
Hauner H, Entenmann G, Wabitsch M, Gaillard D, Ailhaud G, Negrel R et al (1989) Promoting effect of glucocorticoids on the differentiation of human adipocyte precursor cells cultured in a chemically defined medium. J Clin Investig 84(5):1663–1670
Shafiee A, Kabiri M, Ahmadbeigi N, Yazdani SO, Mojtahed M, Amanpour S et al (2011) Nasal septum-derived multipotent progenitors: a potent source for stem cell-based regenerative medicine. Stem Cells Dev 20(12):2077–2091
Hashemi SM, Soleimani M, Zargarian SS, Haddadi-Asl V, Ahmadbeigi N, Soudi S et al (2009) In vitro differentiation of human cord blood-derived unrestricted somatic stem cells into hepatocyte-like cells on poly(ε-caprolactone) nanofiber scaffolds. Cells Tissues Organs 190(3):135–149
Reyes M, Lund T, Lenvik T, Aguiar D, Koodie L, Verfaillie CM (2001) Purification and ex vivo expansion of postnatal human marrow mesodermal progenitor cells. Blood 98(9):2615–2625
Haynesworth SE, Baber MA, Caplan AI (1992) Cell surface antigens on human marrow-derived mesenchymal cells are detected by monoclonal antibodies. Bone 13(1):69–80
Rybak A, Fuchs H, Smirnova L, Brandt C, Pohl E, Nitsch R et al (2008) A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat Cell Biol 10(8):987–993
Oda Y, Yoshimura Y, Ohnishi H, Tadokoro M, Katsube Y, Sasao M et al (2010) Induction of pluripotent stem cells from human third molar mesenchymal stromal cells. J Biol Chem 285(38):29270–29278
Kögler G, Sensken S, Wernet P (2006) Comparative generation and characterization of pluripotent unrestricted somatic stem cells with mesenchymal stem cells from human cord blood. Exp Hematol 34(11):1589–1595
Kellner S, Kikyo N (2010) Transcriptional regulation of the Oct4 gene, a master gene for pluripotency. Histol Histopathol 25(3):405
Hawkins PG, Morris KV (2010) Transcriptional regulation of Oct4 by a long non-coding RNA antisense to Oct4-pseudogene 5. Transcription 1(3):165–175
Tay Y, Zhang J, Thomson AM, Lim B, Rigoutsos I (2008) MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455(7216):1124–1128
Vizlin-Hodzic D, Johansson H, Ryme J, Simonsson T, Simonsson S (2011) SAF-A has a role in transcriptional regulation of Oct4 in ES cells through promoter binding. Cellular Reprogramming(Formerly” Cloning and Stem Cells”) 13(1):13–27
Ding L, Paszkowski-Rogacz M, Nitzsche A, Slabicki MM, Heninger AK, de Vries I et al (2009) A genome-scale RNAi screen for Oct4 modulators defines a role of the Paf1 complex for embryonic stem cell identity. Cell Stem Cell 4(5):403
Saxe JP, Tomilin A, Schöler HR, Plath K, Huang J (2009) Post-translational regulation of Oct4 transcriptional activity. PLoS ONE 4(2):e4467
Rodeheffer MS, Birsoy KV, Friedman JM (2008) Identification of white adipocyte progenitor cells in vivo. Cell 135(2):240–249
Kleger A, Mahaddalkar P, Katz SF, Lechel A, Ju JY, Loya K et al (2012) Increased reprogramming capacity of mouse liver progenitor cells, compared with differentiated liver cells, requires the BAF complex. Gastroenterology 142(4):907–917
Sun N, Panetta NJ, Gupta DM, Wilson KD, Lee A, Jia F et al (2009) Feeder-free derivation of induced pluripotent stem cells from adult human adipose stem cells. Proc Natl Acad Sci 106(37):15720–15725
Kunisato A, Wakatsuki M, Kodama Y, Shinba H, Ishida I, Nagao K (2010) Generation of induced pluripotent stem cells by efficient reprogramming of adult bone marrow cells. Stem Cells and Development 19(2):229–238
Tarte K, Gaillard J, Lataillade JJ, Fouillard L, Becker M, Mossafa H et al (2010) Clinical-grade production of human mesenchymal stromal cells: occurrence of aneuploidy without transformation. Blood 115(8):1549
Shafiee A, Seyedjafari E, Soleimani M, Ahmadbeigi N, Dinarvand P, Ghaemi N (2011) A comparison between osteogenic differentiation of human unrestricted somatic stem cells and mesenchymal stem cells from bone marrow and adipose tissue. Biotechnol Lett 33(6):1257–1264
Niibe K, Kawamura Y, Araki D, Morikawa S, Miura K, Suzuki S et al (2011) Purified mesenchymal stem cells are an efficient source for iPS cell induction. PLoS ONE 6(3):e17610. doi:10.1371/journal.pone.0017610
Miura K, Okada Y, Aoi T, Okada A, Takahashi K, Okita K et al (2009) Variation in the safety of induced pluripotent stem cell lines. Nat Biotechnol 27(8):743–745
Doi A, Park I-H, Wen B, Murakami P, Aryee MJ, Irizarry R et al (2009) Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells embryonic stem cells and fibroblasts. Nat Genet 41(12):1350–1353. doi:10.1038/ng.471
Chin MH, Mason MJ, Xie W, Volinia S, Singer M, Peterson C et al (2009) Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell 5(1):111–123
Nelson TJ, Martinez-Fernandez A, Yamada S, Mael AA, Terzic A, Ikeda Y (2009) Induced pluripotent reprogramming from promiscuous human stemness-related factors. Clin Trans Sci 2(2):118–126
Chang HH, Hemberg M, Barahona M, Ingber DE, Huang S (2008) Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature 453(7194):544–547
Chang TC, Yu D, Lee YS, Wentzel EA, Arking DE, West KM et al (2007) Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet 40(1):43–50
Hanna J, Saha K, Pando B, Van Zon J, Lengner CJ, Creyghton MP et al (2009) Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462(7273):595–601
Chivukula R, Mendell J (2009) Abate and switch: miR-145 in stem cell differentiation. Cell 137(4):606–608
Sachdeva M, Zhu S, Wu F, Wu H, Walia V, Kumar S et al (2009) p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc Natl Acad Sci 106(9):3207
Arora H, Qureshi R, Jin S, Park AK, Park WY (2011) miR-9 and let-7g enhance the sensitivity to ionizing radiation by suppression of NFκB1. Exp Mol Med 43(5):298
Ji J, Zhao L, Budhu A, Forgues M, Jia HL, Qin LX et al (2010) Let-7g targets collagen type I [alpha] 2 and inhibits cell migration in hepatocellular carcinoma. J Hepatol 52(5):690–697
Acknowledgments
This work was supported by funding from Stem Cell Technology Research Center, Tehran, Iran.
Disclosure
Authors have nothing to declare.
Author information
Authors and Affiliations
Corresponding author
Additional information
Fatemeh Jamshidi Adegani, Lida Langroudi contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Adegani, F.J., Langroudi, L., Arefian, E. et al. A comparison of pluripotency and differentiation status of four mesenchymal adult stem cells. Mol Biol Rep 40, 3693–3703 (2013). https://doi.org/10.1007/s11033-012-2445-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2445-7