Abstract
The researches attempting to associate the PPARγ C161T polymorphism with coronary artery disease (CAD) yielded complicated and contradictory results. We aimed for more precise estimate of the relationship and conducted a comprehensive meta-analysis. Publications written in English or Chinese were screened in MEDLINE, Embase, CNKI, Wanfang and CBM. Data on 11 studies including 3,020 cases and 2,853 controls were extracted. A random-effects model was available to synthesize the inconsistent outcomes of the individual studies, while addressing between-study heterogeneity and publication bias. The PPARγ C161T polymorphism followed Hard-Weinberg Equilibrium for all studies (P > 0.05).Overall, there was no evidence for a significant association under all genetic models but with distinct heterogeneity (T vs. C: P = 0.29, OR = 0.91, 95 %CI 0.77–1.08, P heterogeneity = 0.004, I 2 = 61.2 %). However, in the subgroup analysis by ethnicity, the T allele carriers showed a prominent 26 % risk reduction of CAD among Chinese (dominant genetic model: P = 0.03, 95 %CI 0.57–0.97, P heterogeneity = 0.03, I 2 = 56.1 %). After dividing into population source, the significance of CAD risk reduction was strengthened in hospital-based studies (allele comparison: P = 0.04, OR = 0.82, 95 %CI 0.67–1.00, P heterogeneity = 0.04, I 2 = 52.5 %; dominant model: P = 0.01, OR = 0.73, 95 %CI 0.57–0.92, P heterogeneity = 0.05, I 2 = 50.8 %). There was no obvious publication bias verified in the method of funnel plot and Egger’s linear regression test (t = −0.11, P = 0.913). Taken together, our results revealed the PPARγ C161T polymorphism might play a moderate protective effect on developing CAD among Chinese, but not among Caucasians.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent decades, great progress has been made in diagnosis and treatment of coronary artery disease (CAD) and myocardial infarction (MI), resulting in favorable improvement in prognosis. However, CAD and MI are still the leading causes of morbidity and death in western societies. In 2008, the CAD morbidity was 405309 in Unite States and ~1 of every 6 deaths was caused by CAD [1]. Except for some modifiable environment risks [2], the genetic factors may contribute to the majority of variation in susceptible to CAD [3]. Numerous researches have investigated the genetic background of CAD, but little crucial genetic variants that determined the pathogenicity of CAD were verified.
The peroxisome proliferator-activated receptor γ (PPARγ) is a member of the nuclear receptor superfamily. As a ligand activated transcription factor, PPARγ masters the regulation of fatty acid and carbohydrate homeostasis [4, 5] and is thought to play a key role in metabolic diseases such as diabetes [6], hyperlipidemia [7], obesity [6] and atherosclerosis [8]. The gene PPARγ is located on chromosome 3p25 and one of the most common polymorphism of PPARγ gene is C161T (also known as C1431T [9]), a C to T substitution at position 161 in exon 6. The silent C161T polymorphism is a synonymous polymorphism that encodes histidine with either allele and has been found to connect with insulin resistance [10] and metabolic syndrome [11]. Recently, evidences showed that this polymorphism might be associated with CAD, but the observed results were inconclusive. The diversity of these outcomes was probably owing to the small sample bias, inadequate statistical power and various racial descents of single researches. Large meta-analysis is acknowledged as a feasible method to evaluate the reliability of the predetermined candidates in genetic association researches and to clarify the discrepancies. For the sake of a more accurate estimation of the association between the PPARγ C161T polymorphism and the risk of CAD, we conducted a meta-analysis of published studies to date and investigated the potential origin of between-study heterogeneity as well as whether publication bias was present.
Materials and methods
Search strategy
We did a systematic computerized literature search from PubMed/MEDLINE, Embase, China Nation Knowledge Infrastructure Platform (CNKI), Wanfang and China Biological Medicine Database (CBM) electronic databases until September 2012. A combination of the following text search string was used: ‘C161T’ or ‘C1431T’, ‘Peroxisome proliferator-activated receptor gamma’ or ‘PPARγ’ and ‘coronary’ or ‘ischemic heart disease’ or ‘myocardial infarction’ or ‘atherosclerosis’. We also retrieved additional studies through the MEDLINE option ‘related articles’ and the results of manual search were also added into the database at the same time. References from the retrieved articles, reviews, and previous meta-analysis were also screened to improve the databank. We connected with the corresponding authors to obtain the supplementary information of the studies with insufficient genotyping data. The following standards were applied to the search: (1) Published articles (full texts or abstracts) in English or Chinese journals or their supplements, (2) Studies only in human subjects without country restrictions and if articles containing more than one geographic or ethnic heterogeneous group, each subgroup was considered separately, (3) When multiple studies from the same study cohort, only the study with largest population was included to avoid data duplication, (4) The genotype frequency of control population must be in line with Hardy–Weinberg equilibrium (HWE).
CAD endpoints were defined as previous MI, angina pectoris, coronary artery bypass grafting or significant angiographic stenosis of 50 % in at least one or more major coronary artery [12]. ACS included unstable angina pectoris, fatal and non-fatal MI [13] and MI was diagnosed according to the clinical symptoms of angina, the presence of typical electrocardiographic changes, elevation in the levels of cardiac enzymes [14].
Extracted information
A standard procedure and data-collection scenario was used according to the inclusion criteria described above. Two authors (Z. W and Y. L) extracted the relevant characteristics of each eligible study respectively and the data were entered into separate databases in duplicate. Moreover, any encountered discrepancies were checked for by a discussion in order to reach a consensus. The following information was collected on the genotype of the C161T polymorphism according to different cohort: First author’s name, publication year, geographic location and population ethnicity, study design, population source, endpoints, baseline characteristics of the study population [such as age, gender, body mass index (BMI) and total-cholesterol (TC)], the percentage of diabetes and smoking status, the frequency of the C161T genotype both in patients and controls, genotyping methods and consistency of genotype frequencies with HWE. Continuous variables were expressed as mean ± standard deviation (SD) or median (5th and 95th percentiles).
Statistical analysis
The combined studies’ odd ratio (OR) corresponding to 95 % confidence interval (CI) were calculated to evaluate the association of the PPARγ C161T polymorphism with CAD risk under four genetic models, involving allele comparison (T vs. C), dominant genetic model (CT + TT vs. CC), recessive genetic model (TT vs. CT + CC) and homozygote comparison (TT vs. CC). A random-effects model using the DerSimonian and Laired method was applied to evaluate individual effect size together and adjust the study weights according to the in-study variance. We checked on the possibility of heterogeneity in the method of the Mantel–Haenszel model [15]. The between-study heterogeneity were assessed in approach to a Chi square-based Q statistic test [16] and P > 0.1 was considered homogenetic among the studies. In addition, the heterogeneous degree was estimated via inconsistency index I 2 statistic, which ranged from 0 to 100 % and determined whether the between-study variation was resulted from heterogeneity. High values of I 2 strongly suggested the presence of between-study heterogeneity [17, 18]. The significance of the pooled OR was estimated by the Z test and P < 0.05 was considered to be significant. Then we categorized the data into subgroups according to the study characteristics with homogeneous effects, such as racial descent (Chinese and Caucasian), study design (prospective and retrospective) and population source (hospital-based [H–B] and population-based [P–B]). In addition, we undertook a meta-regression, as a supplement of the random-effects meta-analysis, to estimate the effect of covariates on genetic heterogeneity.
Sensitivity analysis was performed by sequential ruling out an individual study each time in order to identify the predominant influential studies which had a possibility of biasing the overall estimation. Furthermore, we used the visual funnel plot and Egger’s test to estimate the probability of publication bias. The standard error of log (OR) of each study was plotted against its OR. An asymmetric plot may suggest publication bias probably and be verified by Egger’s linear regression test [19]. We also performed a T test to determine whether the intercept was significant. P < 0.05 of I 2 statistic and Egger’s test was regarded significant.
We evaluated the expected power of each individual study as determined by the probability to reject a null hypothesis at the 0.05 level of significance, under the assumption that OR equals 1.5 and 2.0 for differences in allele frequency, and the minor allele frequency is of 0.2. The power was estimated on the basis of the method described earlier [20].The statistical power of 80 % is used to avoid false negative rates and to determine whether the sample size is effective.
We also performed a cumulative meta-analysis to clarify whether the first published study influenced the subsequent publications and the evolution of the combined effects over time in line with the ascending date of publications. Conformity of the C161T polymorphism to HWE was tested via the χ2 test or Fisher’s exact test based on a Web program (http://ihg2.helmholtz-muenchen.de/cgi-bin/hw/hwa1.pl). Review Manager software release 5.0 (Oxford, England) and Stata 11.0 (Stata Corporation, College Station, TX, USA) were used to manage data and statistically analysis in all studies, and all P values were 2-sided.
Result
Description of studies search result
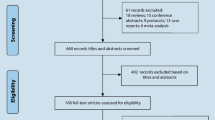
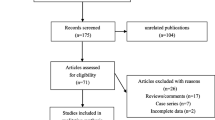
The flowchart schematizing the process of study search and selection was presented in Fig. 1 and the specific reasons of excluded articles were also listed. According to our search strategy and inclusion criteria, the primary literature search in PubMed, EMBASE, CNKI, Wangfang and CBM yielded 341 relevant articles altogether. As a result of the subsequent selection, 18 studies describing the association between the PPARγ C161T polymorphism and CAD risk were provisionally included. Among these studies, 2 Chinese articles were further excluded because of the overlapped data by their latter studies [21, 22] with larger cohort written in English. However, four studies with the genotyping data deviated from HWE in control population were also excluded, including Wang et al. [23] (P HWE = 0.013), Liu et al. [24] (P HWE = 0.009), Bluher et al. [25] (P HWE = 0.004) and Zhou et al. [26] (P HWE = 0.019). The data of Doney et al. [27] was inadequate. Although we have tried to contact with the corresponding authors via E-mail to obtain the relevant data, they did not reply and provide us the raw data. The data from two independent studies [Prospective Study of Myocardial Infarction (PRIME) and Atherosclerotic Disease, Vascular Function, and Genetic Epidemiology (ADVANCE)] were provided by Dallongeville et al. [28], so we considered them as 2 studies individually in our meta-analysis.
10 articles including 11 studies (6 English articles [21, 22, 28–31] and 4 Chinese articles) with sufficient information were identified on the basis of the inclusion criteria. All the eligible studies were published between 2003 and 2011, with 7 in Chinese [21, 22, 29] and 4 in Caucasian population [28, 30, 31]. In addition, 3 studies were P–B [28, 31] and the other 8 study were H–B [21, 22, 29, 30]. Except the studies by Dallongeville et al. [28] were prospective, the rest 9 qualified studies [21, 22, 29–31] were retrospective.
Overall analysis
Eleven studies comprising 3,020 cases and 2,853 controls were pooled into the meta-analysis. The baseline characteristics of the qualified studies were summarized in Table 1. The distribution of the PPARγ C161T genotypes and alleles in the individual studies was listed in Table 2. The genotype distribution of the C161T polymorphism among the control population was consistent with HWE for all the eligible studies (P > 0.05). The pooled overall frequency of the T allele was 17.2 % in cases and 18.4 % in controls. The T allele had a higher proportion among Chinese (22.5 % cases vs. 27.6 % controls) than that among Caucasians (13.2 % cases vs. 12.0 % controls). In addition, the expected power of each individual study and subgroup analysis to identify the association of the PPARγ C161T polymorphism with the CAD risk was shown in Table 3. However, while an allele with an OR of 1.5 and 2, 2 studies and 9 studies achieved 80 % statistical power respectively.
Then we evaluated the association of the PPARγ C161T polymorphism with CAD risk under four genetic models for each study. The main results of the meta-analysis and the heterogeneity test were presented in Table 4. The overall comparison of T allele with C allele yielded an non-significant risk for CAD (allele comparison: P = 0.28, OR = 0.91, 95 % CI: 0.77–1.08; dominant model: P = 0.07, OR = 0.83, 95 %CI: 0.68–1.02; recessive model: P = 0.20, OR = 1.34, 95 %CI: 0.85–2.12, homozygote comparison: P = 0.36, OR = 1.25, 95 %CI: 0.78–2.00) (Fig. 2). A significant between-study heterogeneity (allele comparison: I 2 = 61.2 %, P heterogeneity = 0.004; dominant model: I 2 = 59.7 %, P heterogeneity = 0.01; recessive model: I 2 = 47.5 %, P heterogeneity = 0.04; homozygote comparison: I 2 = 48.6 %, P heterogeneity = 0.04) was observed for all the subjects. The expected power of all the subgroups attained 100 % assuming OR equals 1.5.
Sensitivity analysis
Considering there was a striking heterogeneity in the total meta-analysis, we performed a sensitivity analysis to ascertain the origin of heterogeneity among all the studies. We aimed to find out the potential influential studies which could bias the overall outcome. As a result, the study by Chao et al. [29] was verified to account for the majority of the heterogeneity by means of the sensitivity analysis. After ruling out this most influential study, the magnitude of CAD risk reduction in T allele carriers was progressively reinforced with OR tending to decrease (allele comparison: P = 0.12, OR = 0.87, 95 %CI: 0.74–1.04; dominant model: P = 0.04, OR = 0.80, 95 %CI: 0.65–0.99; recessive model: P = 0.45, OR = 1.19, 95 %CI: 0.76–1.88; homozygote comparison: P = 0.73, OR = 1.09, 95 %CI: 0.68–1.73). Meanwhile, between-study heterogeneity was substantially attenuated (allele comparison: P heterogeneity = 0.02, I 2 = 56.3 %; dominant model: P heterogeneity = 0.01, I 2 = 59.4 %; recessive model: P heterogeneity = 0.10, I 2 = 39.2 %; homozygote comparison: P heterogeneity = 0.10, I 2 = 38.8 %).
Cumulative analysis
There was no remarkable evidence suggesting that the first published study had potential impact on the subsequent publication replication by the cumulative meta-analysis (data not shown).
Publication bias
The funnel plot showed there was not obvious asymmetry and directly interpreted the results as showing evidence against the presence of publication bias. Egger’s regression asymmetry statistic further verified that there was low possibility of publication bias (t = −0.10, P = 0.92 for allele comparison). Even when the most influential study was excluded, there was not any material change in the evaluation of publication bias (t = −0.12, P = 0.91) (Fig. 3).
Subgroup analysis
Given the conspicuous heterogeneity was presented in total analysis, subgroup analysis is an appropriate method to explore the potential origin of the heterogeneity. All the genotypes and allele frequencies of the characteristic-homogeneous groups were listed in Table 4. Data were categorized according to the 2 racial descent groups (Chinese and Caucasian) to assess the possible effect of ethnicity on the variability of overall estimates. The association of the C161T polymorphism with CAD risk among Chinese was at odds with that among Caucasians. In Chinese population, the ORs of the PPARγ C161T polymorphism approached to decrease (allele comparison: P = 0.10, OR = 0.83, 95 %CI 0.66–1.04, P heterogeneity = 0.03, I 2 = 58.4 %) and the T allele carriers showed a significant 26 % risk reduction of CAD under the dominant model (P = 0.03, 95 %CI 0.57–0.97, P heterogeneity = 0.03, I 2 = 56.1 %) (Fig. 4). In contrast, the ORs of CAD approached to unity without significance among Caucasians (allele comparison: P = 0.71, OR = 1.05, 95 %CI: 0.81–1.37, P heterogeneity = 0.05, I 2 = 61 %; dominant model: P = 0.88, OR = 0.98, 95 %CI 0.75–1.29, P heterogeneity = 0.09, I 2 = 54.3 %).
Meta-analysis for the association between the PPARγ C161T polymorphism and CAD among Chinese. The T allele carriers shows a significant reduced risk of CAD under the dominant model (CT + TT vs. CC). ‘Events’ indicates the total number of CT + TT genotype. ‘Total’ indicates the total number of CT + TT genotype plus CC genotype
Further subgroup analysis by population source suggested the significance of the risk reduction of CAD in T allele carriers was remarkably strengthened in H–B group (allele comparison: P = 0.04, OR = 0.82, 95 %CI 0.67–0.99, P heterogeneity = 0.04, I 2 = 52.5 %, dominant model: P = 0.01, OR = 0.73, 95 %CI 0.57–0.92, P heterogeneity = 0.05, I 2 = 50.8 %), accompanying with high between-study homogeneity (Fig. 5). In contrast, the lack of dramatic association was observed of the PPARγ C161T polymorphism with CAD risk in P–B studies under any genetic model (allele comparison: P = 0.31, OR = 1.15, 95 %CI 0.88–1.52, P heterogeneity = 0.10, I 2 = 56.6 %). Similarly, there was not any distinct association of the PPARγ C161T polymorphism with CAD risk was detected under the all genetic models when data were categorized according to study design. The expected power of all the subgroups achieved 100 % assuming OR equals 1.5.
Meta-analysis for the association between the PPARγ C161T polymorphism and CAD in hospital-based studies. The C161T polymorphism shows a significant reduced risk of CAD under the allele comparison (T vs. C, a) and under the dominant model (CT + TT vs. CC b). ‘Events’ indicates the total number of T allele (a) and CT + TT genotype (b), ‘Total’ indicates the total number of T allele plus C allele (a) and CT + TT genotype plus CC genotype (b) respectively
Meta-regression analysis
Meta-regression is in common practice to clarify the potential reasons for between-study heterogeneity in consideration of various environmental factors. Multiple study-level covariates, including the average levels of age, BMI and TC as well as the percent of male, smoke and diabetes were incorporated in our meta-regression. However, we failed to detect the evidence which suggested these above factors influenced the between-study heterogeneity.
Discussion
The definite association of the PPARγ C161T polymorphism with CAD has not been studied extensively. Although some studies have been performed to connect the PPARγ C161T polymorphism with CAD, the results remained conflicting. This lack of uniformity stems from multiple causes, such as research design, sample size, statistic power issues and real variability across different populations [32]. Meta-analysis affords an opportunity to benefit the clarification of genuine relationship by addressing some of these obstacles [33], combining the results and overcoming the limitations of statistical power found in each moderate size studies [34]. To our best knowledge, this is the first systematic review assessing the association between the PPARγ C161T polymorphism and CAD by means of a meta-analysis. We embarked on a meta-analysis with 11 studies totaling 3020 CAD patients and 2853 controls to reconcile the inconsistent results. As a result, we found out a nonsignificant association of C161T polymorphism with CAD risk under different genetic models overall, accompanying with distinct heterogeneity. Sensitive analysis implied that the study by Chao et al. [29] was the major source of heterogeneity. After excluding this most influential article, a marginal association of the C161T polymorphism with the CAD risk reduction was presented and the magnitude of heterogeneity was attenuated.
Genetic heterogeneity is inevitable in any disease identification strategy [35], we conducted the subgroup analysis to explore the diverse role of the PPARγ C161T polymorphism on different ethnic population. Indeed, the T allele carriers were at striking risk reduction of CAD among Chinese, however the protective effect was absent among Caucasians. The diverse results of ethnic groups may be due to different genetic ancestral background. In addition, some individual studies with constrained sample size [30] and insufficient statistical power may be responsible for the discrepancy, as reflected from our power analysis. So the data from large-scale and well-designed researches with adequate statistical power are required for the confirmation of our result.
Besides the interference of ethnicity, another interpretation of the inconsistency between different studies is that the single locus effects contribute to the minority of CAD risk prediction. The genetics of disease aetiology and drug dependance is mostly based on interactions among multiple genes and environmental factors. We centered on the single polymorphism on the PPARγ gene in spite of the probability that other candidate genes or polymorphisms might cooperatively exert effect on the development of CAD. Optimized models to characterize the polygenic or haplotype-based effects are needed to fully understand disease causation.
It should be noticed that most common diseases cannot be explained by single genetic risk factors in isolation. The elucidation of between-study heterogeneity is often confounded by the environment exposures including lifestyle, socioeconomic factors, prescription policies, endocrine disruptors, maternal behavior and so on. Environmental agents can independently alter gene expression and modulate the disease risk via the process called epigenetics [36]. Genetic factors modify the epigenetic response to the environment [37].Epigenetic marks, such as DNA methylation and histone modifications, are dynamically changing throughout the life course. Measuring epigenetic influences derived from environmental risk factors and including them in models of the assessments of genetic risk criteria will become pivotal in future investigations. Nevertheless, we did not find out any environment exposures as primary causes of between-study heterogeneity in the way of meta-regression, implying interaction of the C161T genotype with the environment factors barely influenced our result.
In addition, another estimate should be cautiously treated when data were stratified by population source. The association of C161T polymorphism with the CAD risk reduction in H–B group was obviously at odds with that in P–B group. The CAD risk reduction dramatically and progressively reduced in H–B group. Interestingly, the magnitude of association was reversed in P–B studies although this relationship was not significant. Population classified remained problematic [38], especially in H–B studies. Despite high participation and less information bias may favor H–B studies, they still have some drawbacks. Subjects are recruited from a given hospital in spite of the population from which they arise. Even in some studies, controls are selected from an ill-defined study base. Thus there is probably a biased case–control comparison between cases and controls because subjects could not reflect the true exposure experience of the source population, resulting in an overestimation of the OR. By contrast, the controls sampled from community or general population are acknowledged as preferable for reasons of representativeness and reliability. Considering a wide range of confidence intervals of in the H–B subgroup analysis, further studies are called for to ascertain the reliability of effect size.
Despite our meta-analysis included relatively large sample size consistent of HWE and offered an analysis with the potential for high power [39], there are some weakness of methodology for characterizing meta-analyses of observational studies [40]. Small negative studies are prone to be refused and the unpublished studies as well as the “grey” literature (articles in languages other than English and Chinese) were likely not to be involved, literature bias might not be ruled out absolutely, although the Egger’s test and funnel plots did not document the presentation of publication bias in our meta-analysis. In addition, some studies free of nested or matched case–control design also accounted for heterogeneity because the different distribution of these study exposures is poorly comparable. Additional studies estimating this kind of interaction are required.
Experimental studies have identified the anti-inflammatory effects of PPARγ. Although the function significance of silent mutation can be explained in some ways [41], there were scarce evidences determining the role of the C161T polymorphism on the expression of PPARγ. Meirhaeghe et al. [42] detected a dramatic interaction presented between the C161T polymorphism and BMI for plasma leptin levels, which implied that the C161T polymorphism possibly participated in modifying the activation of PPARγ, resulting in altered transcription levels of target genes in the metabolic pathway [43]. Our research complemented the previous studies and provided clues that the C161T polymorphism might exert pleiotropic and positive effects in preventing atherosclerosis.
Although the statistical bias could not be avoided completely, our meta-analysis, comprising 5,873 people, suggested that the T allele might have a potential protective effect on CAD. The effect is heterogeneous, being distinct among Chinese and lack of significance among Caucasians. Our meta-analysis also emphasizes the necessity of great caution when trying to interpret and combine data observed in different ethnic population. However, it is confused whether the T allele carriers have more advantages in aspect of anti-atherosclerosis because of the low frequency of T allele. More large-scale cohort studies with well study design may facilitate to explore the true relationship.
References
Roger VL, Go AS, Lloyd-Jones DM, Benjamin EJ, Berry JD et al (2012) Heart disease and stroke statistics–2012 update: a report from the American Heart Association. Circulation 125:e2–e220
Expert Panel on Detection E, Treatment of High Blood Cholesterol in A (2001) Executive summary of the third report of the National Cholesterol Education Program (NCEP) expert panel on detection, evaluation, and treatment of high blood cholesterol in adults (Adult Treatment Panel III). JAMA 285:2486–2497
Zdravkovic S, Wienke A, Pedersen NL, Marenberg ME, Yashin AI et al (2002) Heritability of death from coronary heart disease: a 36-year follow-up of 20 966 Swedish twins. J Intern Med 252:247–254
Duval C, Chinetti G, Trottein F, Fruchart JC, Staels B (2002) The role of PPARs in atherosclerosis. Trends Mol Med 8:422–430
Fisman EZ, Tenenbaum A (2009) A cardiologic approach to non-insulin antidiabetic pharmacotherapy in patients with heart disease. Cardiovasc Diabetol 8:38
Kota BP, Huang TH, Roufogalis BD (2005) An overview on biological mechanisms of PPARs. Pharmacol Res 51:85–94
Wahli W, Michalik L (2012) PPARs at the crossroads of lipid signaling and inflammation. Trends Endocrinol Metab 23:351–363
Huang JV, Greyson CR, Schwartz GG (2012) PPAR-gamma as a therapeutic target in cardiovascular disease: Evidence and uncertainty. J Lipid Res 53(9):1738–1754
Fahmi H, Martel-Pelletier J, Pelletier JP, Kapoor M (2011) Peroxisome proliferator-activated receptor gamma in osteoarthritis. Mod Rheumatol 21:1–9
Moffett SP, Feingold E, Barmada MM, Damcott CM, Marshall JA et al (2005) The C161– > T polymorphism in peroxisome proliferator-activated receptor gamma, but not P12A, is associated with insulin resistance in Hispanic and non-Hispanic white women: evidence for another functional variant in peroxisome proliferator-activated receptor gamma. Metabolism 54:1552–1556
Dongxia L, Qi H, Lisong L, Jincheng G (2008) Association of peroxisome proliferator-activated receptorgamma gene Pro12Ala and C161T polymorphisms with metabolic syndrome. Circ J 72:551–557
Wright RS, Anderson JL, Adams CD, Bridges CR, Casey DE Jr et al (2011) 2011 ACCF/AHA focused update incorporated into the ACC/AHA 2007 Guidelines for the Management of Patients with Unstable Angina/Non-ST-Elevation Myocardial Infarction: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines developed in collaboration with the American Academy of Family Physicians, Society for Cardiovascular Angiography and Interventions, and the Society of Thoracic Surgeons. J Am Coll Cardiol 57:e215–e367
Joensen AM, Jensen MK, Overvad K, Dethlefsen C, Schmidt E et al (2009) Predictive values of acute coronary syndrome discharge diagnoses differed in the Danish National Patient Registry. J Clin Epidemiol 62:188–194
Anderson JL, Adams CD, Antman EM, Bridges CR, Califf RM et al (2011) 2011 ACCF/AHA Focused Update Incorporated Into the ACC/AHA 2007 Guidelines for the Management of Patients With Unstable Angina/Non-ST-Elevation Myocardial Infarction: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. Circulation 123:e426–e579
Cohn LD, Becker BJ (2003) How meta-analysis increases statistical power. Psychol Methods 8:243–253
Lau J, Ioannidis JP, Schmid CH (1997) Quantitative synthesis in systematic reviews. Ann Intern Med 127:820–826
Higgins JP, Thompson SG (2002) Quantifying heterogeneity in a meta-analysis. Stat Med 21:1539–1558
Higgins JP, Thompson SG, Deeks JJ, Altman DG (2003) Measuring inconsistency in meta-analyses. Br Med J 327:557–560
Egger M, Davey Smith G, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. Br Med J 315:629–634
Ramsey DM, Futschik A (2012) DNA pooling and statistical tests for the detection of single nucleotide polymorphisms. Stat Appl Genet Mol Biol 11: Article 1
Peng DQ, Zhao SP, Nie S, Li J (2003) Gene-gene interaction of PPARgamma and ApoE affects coronary heart disease risk. Int J Cardiol 92:257–263
Wan J, Xiong S, Chao S, Xiao J, Ma Y et al (2010) PPARgamma gene C161T substitution alters lipid profile in Chinese patients with coronary artery disease and type 2 diabetes mellitus. Cardiovasc Diabetol 9:13
Wang XL, Oosterhof J, Duarte N (1999) Peroxisome proliferator-activated receptor gamma C161– >T polymorphism and coronary artery disease. Cardiovasc Res 44:588–594
Liu Y, Yuan Z, Zhang J, Yin P, Wang D et al (2007) PPARgamma gene C161T substitution is associated with reduced risk of coronary artery disease and decreased proinflammatory cytokine expression. Am Heart J 154:718–724
Bluher M, Klemm T, Gerike T, Krankenberg H, Schuler G et al (2002) Lack of association between peroxisome proliferator-activated receptor-gamma-2 gene variants and the occurrence of coronary heart disease in patients with diabetes mellitus. Eur J Endocrinol 146:545–551
Zhou X, Chen J, Xu W (2012) Association between C1431T polymorphism in peroxisome proliferator-activated receptor-gamma gene and coronary artery disease in Chinese Han population. Mol Biol Rep 39:1863–1868
Doney AS, Fischer B, Leese G, Morris AD, Palmer CN (2004) Cardiovascular risk in type 2 diabetes is associated with variation at the PPARG locus: a Go-DARTS study. Arterioscler Thromb Vasc Biol 24:2403–2407
Dallongeville J, Iribarren C, Ferrieres J, Lyon L, Evans A et al (2009) Peroxisome proliferator-activated receptor gamma polymorphisms and coronary heart disease. PPAR Res 2009:543746
Chao TH, Li YH, Chen JH, Wu HL, Shi GY et al (2004) The 161TT genotype in the exon 6 of the peroxisome-proliferator-activated receptor gamma gene is associated with premature acute myocardial infarction and increased lipid peroxidation in habitual heavy smokers. Clin Sci (Lond) 107:461–466
Yilmaz-Aydogan H, Kurnaz O, Kurt O, Akadam-Teker B, Kucukhuseyin O et al (2011) Effects of the PPARG P12A and C161T gene variants on serum lipids in coronary heart disease patients with and without Type 2 diabetes. Mol Cell Biochem 358:355–363
Evangelisti L, Attanasio M, Lucarini L, Sofi F, Marcucci R et al (2009) PPARgamma promoter polymorphisms and acute coronary syndrome. Atherosclerosis 205:186–191
Colhoun HM, McKeigue PM, Davey Smith G (2003) Problems of reporting genetic associations with complex outcomes. Lancet 361:865–872
Whitehead A (2003) Meta-analysis of controlled clinical trials. Wiley, Chichester
Attia J, Thakkinstian A, D’Este C (2003) Meta-analyses of molecular association studies: methodologic lessons for genetic epidemiology. J Clin Epidemiol 56:297–303
Hemminki K, Lorenzo Bermejo J, Forsti A (2006) The balance between heritable and environmental aetiology of human disease. Nat Rev Genet 7:958–965
Su J, Qi Y, Liu S, Wu X, Lv J et al (2012) Revealing epigenetic patterns in gene regulation through integrative analysis of epigenetic interaction network. Mol Biol Rep 39:1701–1712
Bjornsson HT, Fallin MD, Feinberg AP (2004) An integrated epigenetic and genetic approach to common human disease. Trends Genet 20:350–358
Salanti G, Sanderson S, Higgins JP (2005) Obstacles and opportunities in meta-analysis of genetic association studies. Genet Med 7:13–20
Munafo MR, Flint J (2004) Meta-analysis of genetic association studies. Trends Genet 20:439–444
Stroup DF, Berlin JA, Morton SC, Olkin I, Williamson GD et al (2000) Meta-analysis of observational studies in epidemiology: a proposal for reporting. Meta-analysis Of Observational Studies in Epidemiology (MOOSE) group. J Am Med Assoc 283:2008–2012
Kimchi-Sarfaty C, Oh JM, Kim IW, Sauna ZE, Calcagno AM et al (2007) A “silent” polymorphism in the MDR1 gene changes substrate specificity. Science 315:525–528
Meirhaeghe A, Fajas L, Helbecque N, Cottel D, Lebel P et al (1998) A genetic polymorphism of the peroxisome proliferator-activated receptor gamma gene influences plasma leptin levels in obese humans. Hum Mol Genet 7:435–440
Dragojević JMJMB (2008) Association between Pro12Ala and His477His polymorphisms in PPARG gene and insulin resistance in patients with polycystic ovary syndrome. Biochemia Medica 18:342–350
Acknowledgments
This work was supported by the Shanghai excellent young teachers training fund of colleges and universities (jdy09097) and the science and technology fund of Shanghai Jiao Tong University School of Medicine (11XJ21001).
Conflict of interest
The authors have declared that no competing interests exist.
Author information
Authors and Affiliations
Corresponding author
Additional information
Zhijun Wu, Yuqing Lou contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wu, Z., Lou, Y., Jin, W. et al. The C161T polymorphism in the peroxisome proliferator-activated receptor gamma gene (PPARγ) is associated with risk of coronary artery disease: a meta-analysis. Mol Biol Rep 40, 3101–3112 (2013). https://doi.org/10.1007/s11033-012-2384-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2384-3