Abstract
Breast cancer is the most common malignancy in women and a significant cause of morbidity and mortality. Sub-types of breast cancer defined by the expression of steroid hormones and Her2/Neu oncogene have distinct prognosis and undergo different therapies. Besides differing in their phenotype, sub-types of breast cancer display various molecular lesions that participate in their pathogenesis. BRCA1 is one of the common hereditary cancer predisposition genes and encodes for an ubiquitin ligase. Ubiquitin ligases or E3 enzymes participate together with ubiquitin activating enzyme and ubiquitin conjugating enzymes in the attachment of ubiquitin (ubiquitination) in target proteins. Ubiquitination is a post-translational modification regulating multiple cell functions. It also plays important roles in carcinogenesis in general and in breast carcinogenesis in particular. Ubiquitin conjugating enzymes are a central component of the ubiquitination machinery and are often perturbed in breast cancer. This paper will discuss ubiquitin and ubiquitin-like proteins conjugating enzymes participating in breast cancer pathogenesis, their relationships with other proteins of the ubiquitination machinery and their role in phenotype of breast cancer sub-types.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the most common malignancy in women and about one in eight women will be attained by the disease during her lifetime. Increasing understanding of its molecular pathogenesis has led to the definition of several sub-types with corresponding progress in treatments for some of these sub-groups [1]. Genomically-defined sub-types are not used in clinical practice due to practical issues. Instead, an approximate equivalent of genomic sub-groups used in clinical practice is based on immunohistochemical (and/or in situ hybridization) evaluation of a few proteins including estrogen receptor alpha (ERα), progesterone receptor (PR), Her2/Neu (a member of the epidermal growth factor receptor family) and the MIB-1 antigen (also named Ki-67) as a measurement of proliferation [2]. These evaluations are important for prognostication and therapeutic decisions as the sub-types defined in this manner have significantly different prognosis and may receive different treatments.
Ubiquitination (also referred to as ubiquitylation) is one of several post-translational protein modifications which regulates virtually every function of the cell and involves the covalent attachment of one or several molecules of the protein ubiquitinin to a target protein. It is carried out by a stepwise process executed with the help of three enzymes, a ubiquitin-activating enzyme (also called E1), a ubiquitin-conjugating enzyme (or E2) and a ubiquitin ligase (or E3) [3, 4]. There also exist several ubiquitin-like proteins (UBLs) that can similarly be attached to target proteins with the help of an analogous to ubiquitin enzymatic cascade. These include small ubiquitin-like modulator (SUMO), neural precursor cells expressed and developmentally down-regulated 8 (NEDD8), interferon-stimulated gene 15 (ISG15) and HLA-F adjacent transcript 10 (FAT10) [5].
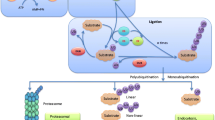
Ubiquitination is a versatile post-translational modification. Ubiquitin is a 76 amino-acids molecule and has seven lysine residues at positions 6, 11 27, 29, 33, 48 and 63. Attachment through each of these lysine residues as well as through the amino-terminal methionine residue has been confirmed to possess signaling potential [6, 7]. The number of ubiquitin molecules attached encodes also for different outcomes [8]. A target protein may become mono-ubiquitinated (a single ubiquitin molecule attached), multi-ubiquitinated (one ubiquitin molecule attached in several different lysine residues of the target protein) or poly-ubiquitinated (a chain of ubiquitins attached through the same lysine residue).
The importance of ubiquitination in breast cancer is exemplified by the fact that protein BRCA1, one of the proteins mutated in hereditary breast cancer syndromes, is a ubiquitin ligase. Moreover several key proteins in breast cancer and actual or potential targets of therapy such as ERα, PR and transcription factor NF-κB are regulated by ubiquitination and UBLs modifications. Ubiquitin conjugating enzymes that co-operate with BRCA1 as well as with other E3 ligases to perform ubiquitination or UBLs conjugation have also a role in breast cancer and will be discussed in this paper.
E2 enzymes
Ubiquitin-conjugating enzymes are situated in the middle of ubiquitin and UBLs enzymatic conjugation cascade between the ubiquitin-activating enzymes and the ubiquitin ligases. The number of E2s in the human genome is more limited than E3 ligases. There are about 35 E2s scattered across almost all human chromosomes, while there are more than 600 E3s in human genome belonging to two main types, the RING type and HECT type [9]. There are two E1 enzymes performing ubiquitination in humans UBA1 and UBA6, the latter serving also the FATylation (conjugation of FAT10) cascade. Other UBLs have dedicated E1 s, for example the heterodimer APPBP1/UBA3 is the E1 for NEDD8 and UBE1L is the E1 for ISG15. E2 enzymes interact with E1 enzyme that carries the activated ubiquitin and accept ubiquitin forming a thiolester bond using its active site cysteine. A conformational change in E1 occurring only after ubiquitin loading allows this interaction. After the transfer to E2, E1 retakes its prior conformation which promotes dissociation of the ubiquitin molecule-loaded E2 [10]. This is followed by an interaction of the ubiquitin-loaded E2 with an ubiquitin ligase and transfer of ubiquitin directly on a substrate concomitantly attached to E3, in the case of RING type E3s, or to a cysteine residue of the E3, in the case of HECT type E3s, which then transfers it to the substrate. A similar enzymatic cascade takes place for the attachment of ubiquitin-like molecules such as SUMO, NEDD8 and ISG15 but it uses specific enzymes some of which may also function in ubiquitination but others such as the SUMO E2 and E3s function uniquely in this ubiquitin-like molecule attachment. E2s are structurally characterized by a conserved domain of about 150–200 amino-acids called ubiquitin-conjugating (UBC) fold which serves as a platform for E1, E3 and ubiquitin binding (Fig. 1a). The catalytic cysteine is embedded in this domain. About 10 residues amino-terminal to the catalytic cysteine there usually exist an histidine, proline, asparagine (HPN) motif which plays roles in the active site formation of the isopeptide bond between ubiquitin and the ε amine group of a lysine in the substrate protein [11]. The general structure of the UBC fold comprises four α helices, a four-stranded β-sheet, and a small 310 helix (a helical structure with hydrogen bonds between the amino-acids separated by two other amino-acids, while in α helices these bonds link amino-acids separated by three amino-acids in between). Two loops between the β-sheet and the active site and between the second and third α helix are also important in the formation of the shallow groove that surrounds the active cysteine. The active cysteine accepting ubiquitin or a UBL is found after the fourth β-sheet strand and immediately amino-terminal to the small 310 helix in a shallow groove constructed, in addition to these two structures, by residues from the third and fourth α helices [12] (Fig. 1b). The first α helix and two loops between the third and fourth β strand and between the small 310 helix and the second α helix constitute the interacting surface with E3s. Except from the UBC fold several E2s have additional domains amino-terminal or carboxy-terminal to UBC or both. A classification based on the presence of these additional domains has been introduced in which E2s without additional domains besides UBC fold are considered class I, E2s with amino-terminal to UBC fold extensions are class II, E2s with carboxy-terminal extensions are considered in class III and those with both amino- and carboxy-terminal extensions are considered as belonging in class IV [13] (Fig. 1c).
a. Linear representation of the conserved UBC fold of E2 enzymes. The first α helix is followed by the four-stranded β-sheet which is connected with the 310 helix by a loop containing the active cysteine and the HPN motif. Three additional α helices are more carboxy-terminally in the UBC fold. H α helices, S strands of the β-sheet, L loops between α helices and strands of the β-sheet, C active cysteine, HPN histidine–proline–asparagine motif. b 3D representation of the UBE2C homolog from clam, E2-C. α helices are represented in red and β-sheet in blue. From Jiang and Basavappa [131]. c Classification of E2s and examples of each class playing a role in breast cancer and discussed in the paper. Class I E2s have only the UBC fold, class II have amino-terminal additional domains, class III have carboxy-terminal extensions and class IV both amino-terminal and carboxy-terminal extensions
The recognition of the target protein to be modified is mainly a task of the ubiquitin ligase involved. In contrast, in many occasions, the E2 enzyme plays a significant role in the type of ubiquitin link or chain to be attached, at least when co-operating with RING type E3s. This is very important because the type of attachment defines the ultimate outcome. A lysine 48 linked chain of at least four ubiquitin molecules and lysine 11 chains result for example in recognition of the target protein by the proteasome for degradation, while lysine 63-linked ubiquitination leads to lysosome-mediated degradation or non-degradative outcomes [14]. Another task in which E2 enzymes play an important role in collaboration with the specific E1 enzyme is the recognition of the E1 and the fidelity of transferring only their cognate UBL molecule and of concomitantly excluding all others [15]. In addition, E2s are significant in the processivity of the assembled ubiquitin chains formation. This refers to the number of ubiquitins that an E2 may add in a single interaction with a given E3 to a given target protein and depends on the affinity of the substrate for the E3 and on the rate of E2 catalysis [16].
An E2 enzyme may co-operate with several E3s and participate to the ubiquitination of several different substrate proteins and conversely an E3 ligase may use different E2s for performing attachment of diverse ubiquitin modifications. For example E2 UBE2L3 (UbcH7) may co-operate with both RING type casitas B cell lymphoma (E3 c-Cbl) [17] and HECT type E3 E6-associated protein (E6AP) [18] and conversely E3 ligase APC/C co-operates with two E2s UBE2C and UBE2S to perform substrate mono-ubiquitination and lysine11 poly-ubiquitination respectively.
Regarding their nomenclature, E2s were initially given various names that did not denote their function. In an effort to systematically categorize E2s in a homogeneous manner, a name consisting of the designation ubiquitin enzyme E2 (UBE2) followed by a serial capital letter according to their consecutive discovery has been proposed and will be used in the subsequent discussion [13]. Alternative names commonly used in the literature will be given in the first encounter of each enzyme in the discussion.
Specific E2 enzymes with a role in breast cancer
Several E2 enzymes taking part in ubiquitination or UBLs conjugation have been studied and found to play a role or be altered in breast cancer (Table 1). Depending on the substrate protein conjugated, important processes in carcinogenesis such as transcription, DNA repair, chromosome segregation during mitosis and apoptosis may be affected.
UBE2I (Ubc9), the SUMO conjugating enzyme
UBE2I is the only known E2 enzyme for the SUMOylation cascade. This cascade uses the heterodimeric AOS1/UBA2 as the E1 enzyme and several E3s. UBE2I being the sole E2 in SUMOylation, perturbations of its expression or function may theoretically influence all the processes in which SUMOylation plays a regulatory role. These include DNA damage response, DNA transcription and an increasing number of signaling cascades. Specific examples pertaining to breast cancer include ERα signaling and transcription regulation, BRCA1-dependent DNA repair and transcription and NF-κB signaling. Other proteins that are targets of SUMOylation may play a role in breast cancer. SUMO tags function basically through two mechanisms [19]: In some cases they promote protein–protein interactions through binding of a protein containing a SUMO interactive motif (SIM) to a SUMO-tagged protein (Fig. 2, upper panel). In other cases SUMO prevents protein–protein interactions, for example by binding to the same lysine residue of a protein that also could bind ubiquitin [20] (Fig. 2, lower panel). In this manner a SUMO-tagged protein is prevented from interacting with a protein possessing an ubiquitin-binding motif. SUMOylation plays a role in the transcription function of several transcription factors including ERα [21]. ERα signaling is implicated in the physiologic development of mammary gland but also in breast cancer [22]. The majority of breast cancers express ERα and activation of this receptor by estrogens promotes cell proliferation and survival. Endocrine therapies used clinically in ERα expressing breast cancers work by blocking ERα signaling through interruption of ligands binding or through inhibition of estrogen production or down-regulation of the receptor itself. After estradiol ligation ERα binds to DNA either directly through ER specific elements or indirectly through tethering to other transcription factors. Nuclear receptor binding triggers recruitment of co-factors leading to chromatin remodeling and further recruitment of basal transcription machinery for transcription to start. SUMOylation ranks among the modifications that take part in ERα transcription regulation despite the fact that ERα protein possesses no consensus SUMOylation sequence [23]. It takes place only after ligand binding to the receptor through conserved residues in the hinge region of ERα molecule and with the co-operation of UBE2I with SUMO E3 ligases protein inhibitor of activated STAT 1 (PIAS1) and PIAS3 [23]. SUMOylation positively modulates transcriptional activity of ERα. UBE2I may also have a role in ERα transcription regulation independently from its SUMO-conjugating activity based on the fact that mutant UBE2I disabled for conjugation is still able to modulate ERα transcription. In addition to ERα itself the transcriptional co-activator of ERα steroid receptor co-activator 3 (SRC3, also known as AIB1, amplified in breast cancer 1 or NCOA3, nuclear receptor co-activator 3) is a target of SUMOylation [24]. SRC3 SUMOylation is facilitated by PIAS1 and inhibits the interaction of SRC3 with ERα, suppressing ERα transcriptional activity and prolonging the stability of SRC3 in breast cancer cells [24].
Mechanisms of SUMOylation function. In the upper panel SUMOylation promotes the interaction of the SUMO-decorated protein with another protein that bears a SUMO interacting motif (SIM). In the lower panel SUMO competes with ubiquitin for the same lysine of a substrate protein preventing ubiquitination. Ubiquitin interacting motif (UIM)—containing protein is prevented from interacting with the SUMOylated protein
BRCA1 is a protein playing roles in DNA repair and is mutated in hereditary cases of breast cancer. In addition, it is dysfunctional in sporadic breast cancers that exhibit features of hereditary breast cancer biology such as negativity for the expression of ERα and PR receptors and absence of amplification of Her2/Neu receptor (the so-called triple negative phenotype). These similarities have led to coining the term ‘BRCAness’ to include breast cancers with defective BRCA1 activity either due to germline mutations or somatic dysfunction [25]. BRCA1 is a RING finger E3 ubiquitin ligase which participates in the repair of double strand DNA breaks in the homologous recombination pathway. It is recruited at sites of double strand DNA damage together with its partner BRCA1 associated RING domain (BARD1), a protein with a RING domain but without innate ligase activity. This recruitment depends on phosphorylation of histone H2AX by damage sensing kinases ATM and ATR and previous recruitment of E3 ligases RNF8 and RNF168 which ubiquitinate histones H2A and H2AX around the damaged site. Ubiquitinated histones help directly or indirectly in the accrual of protein receptor associated protein 80 (RAP80) which then recruits BRCA1 [26]. Both RAP80 and BRCA1 are targets of SUMOylation and this modification is required for BRCA1 localization to sites of DNA damage and augments its ubiquitin ligase activity [27, 28]. RAP80 SUMOylation takes place in its amino-terminal part with the help of UBE2I [29] and an unknown ligase.
Besides double strand DNA repair by homologous recombination, BRCA1 participates in cell cycle checkpoint signaling and transcription regulation including transcription of ERα gene. BRCA1 cell cycle checkpoint control complexes have many participants in common with DNA damage repair [30] and thus SUMOylation is expected to play roles in this process. BRCA1 SUMOylation in an amino-terminal consensus sequence is required for repression of ERα activity by BRCA1-dependent ubiquitination [31, 32]. UBE2I participates in BRCA1 SUMOylation as well as its nuclear localization and retention [33]. It is worth mentioning here that SUMOylation is a nuclear localization signal for several proteins [20, 34, 35]. In breast cancer cells, for example, SUMOylation and UBE2I over-expression promote nuclear accumulation of receptor kinase IGF-1R after ligand binding where it acts as a transcription factor [36, 37]. In an additional twist of the tale SUMO suppresses BRCA1-regulated transcription independently of SUMOylation [38]. This effect depends on the recruitment of deacetylases on the promoter of BRCA1-regulated genes and takes place even in cells expressing a dominant negative UBE2I impairing SUMOylation [38]. How this effect is mechanistically mediated remains unknown although one could speculate that free SUMO competes with bound SUMO for protein interactions.
NF-κB represents a family of transcription factors with a role in inflammation and carcinogenesis. The role of ubiquitination in NF-κB regulation is well-established and indeed NF-κB inhibition was one of the first mechanisms proposed for the explanation of the anti-tumor activity of proteasome inhibitors, although it is now clear that additional targets of this inhibition is clinically relevant [39]. Ubiquitin modifications leading to proteasome degradation and non-degradational outcomes involve many NF-κB pathway components. SUMOylation is an additional player in NF-κB regulation. The inhibitor of NF-κB IκBα is an example of regulation by SUMOylation. SUMO binding takes place through the same lysine residue of IκBα that also binds ubiquitin. SUMOylation of this lysine in response to adenosine signaling prevents ubiquitin binding and stabilizes IκBα inhibiting NF-κB [40]. Given that IκBα ubiquitination is regulated by both previous phosphorylation and by NEDDylation (attachment of the ubiquitin-like protein NEDD8) which activates the E3 ligase βTRCP, degradation of this key inhibitor of NF-κB is co-ordinated by at least four post-translational modifications securing a tight control of the pathway and allowing activation only if several prerequisite conditions are fulfilled (Fig. 3). SUMOylation also regulates NF-κB a step above IκBα at the level of the IKK complex by modulation of IKKγ (also named NEMO, NF-κB essential modulator) [41]. This modification appears to take place specifically following DNA damage and not after other NF-κB-activating signals and thus defines distinct modes for activation of the same transcription factor possibly helping to fine-tune this activation. Two other E2 enzymes, UBE2D2 and UBE2D3, participating in the regulation of NF-κB signaling by ubiquitination will be discussed in a later section.
NF-κB activation has been linked with endocrine treatment resistance of ERα positive breast cancer [42] and with sensitivity or resistance to neo-adjuvant chemotherapy [43–45]. These contradictory results regarding the role of NF-κB in chemotherapy resistance probably relate to different sub-types of breast cancer studied, different sub-units of NF-κB examined that may play differing roles [46] and the fact that the canonical and non-canonical NF-κB pathways have distinct roles in the epithelial breast cell hierarchy [47]. An additional experimental factor to consider when evaluating those results relates to immunohistochemistry which measures the expression or the nuclear presence of the transcription factor but is not able to inform functionality, a major limitation when studying transcription factors solely with this method.
UBE2I has been found to be up-regulated in breast cancer initiating cells due to down-regulation of microRNA miR-30 in these cells [48]. miR-30 reduction promotes the ability of breast cancer initiating cells to self-renew and resist apoptosis and anoikis. Breast cancer xenografts expressing miR-30 through a lentiviral vector system displayed reduced growth in mice compared with control xenografts. Xenografts transduced with UBE2I targeting shRNA displayed reduced growth but in a lesser degree suggesting that part of the miR-30 effect is due to UBE2I but other targets of the microRNA play a role. A proposed such target is integrin β3 which is a structural and signaling component of focal adhesions with a role in cell motility [48]. UBE2I may also have a role in breast cancer that is dependent on transcription regulation of another miR, miR-224 but independent of its SUMO conjugating activity [49]. This probably relates to its function as transcription co-factor [50]. Acting independently of protein SUMOylation, even a dominant negative mutant UBE2I without SUMO-conjugating activity is able to down-regulate expression of miR-224 which, in turn, up-regulated CDC42 and CXCR4, two invasion-promoting proteins [49]. Putative transcription factors co-operating with UBE2I for miR-224 suppression have not been reported.
UBE2I shows an increased expression in invasive ductal carcinoma compared with normal breast and is associated with larger and less differentiated tumors, nodal metastases, resistance to neo-adjuvant FEC chemotherapy and decreased disease free survival [51]. A higher percentage of triple negative and triple positive (ER/PR positive and Her2 positive) patients had high UBE2I expression as opposed to patients with ER/PR positive, Her2 negative and ER/PR negative, Her2 positive tumors in whom the percentage of high UBE2I expression was lower [51]. These data pinpoint to a role of high expression of UBE2I in associating the expression of ERα and PR with Her2/Neu while lower expression leads more often to dissociation of these expressions. A hypothesis on how these molecular associations might take place is given in a subsequent section.
UBE2C (UbcH10) and UBE2S (E2-EPF)
Two other E2 enzymes with a role in breast cancer are UBE2C (also called UbcH10) and UBE2S (also called E2-EPF) which are both working with E3 ligase anaphase promoting complex/cyclosome (APC/C) in the regulation of cell cycle. UBE2C co-operates with APC/C for the initial ubiquitin attachment to substrates while UBE2S performs with APC/C subsequent ubiquitin attachments producing lysine 11 chains [52]. These enzymes are the only known enzymes constructing ubiquitin chains through lysine 11. In the end of the metaphase stage of mitosis chromosomes are aligned at the equator of the cell and develop connections through the centromere with both poles of the mitotic spindle [6]. The completion of the attachment of all chromosomes to both poles gives the signal for each sister chromatid to detach from its pair and be pulled towards a spindle pole. Till this signal is given sister chromatids are kept attached at the centromere with the action of proteins cohesins. When all chromosomes are attached, APC/C ubiquitinates the protein securin, an inhibitor of the enzyme separase, and promotes its degradation by the proteasome [53]. After securin destruction, separase is activated and cleaves cohesins allowing sister chromatids to be pulled to the two poles at anaphase. In parallel APC/C promotes the destruction of Cyclin B allowing dephosphorylation and inactivation of CDK1, another prerequisite for progression from anaphase to telophase and completion of mitosis [54]. In the end of mitosis APC/C auto-regulates its action by ubiquitinating its own activating factors and E2 enzymes [55].
Lysine 11 chains constructed by APC/C and E2s UBE2C and UBE2S have a close configuration with additional non-covalent attachments between the neighbouring ubiquitin molecules similarly to lysine 48 chains and are also recognized by the proteasome leading to substrate degradation. Nevertheless they differ from lysine 48 in the kinetics of the degradation [56]. Thus the recognition of different ubiquitin chains by the proteasome may serve the purpose of conferring different priorities in proteasome destruction giving for example priority in specific proteins in order for the cell cycle to proceed. The timing of the destruction of different proteins is of paramount importance during the physiologic cell cycle progression and perturbations of the machinery that serves this destruction may lead to carcinogenesis. Given the importance of cell cycle and mitosis regulation which includes an error free and even distribution of chromosomes to daughter cells, eukaryotic cells have dedicated a specific ubiquitin linkage through lysine 11 and specific machinery to this process possibly in order to avoid interferences with other types of ubiquitin chains [57].
UBE2C is over-expressed in diverse tumor cell lines and primary tumor tissues compared with normal tissues [58]. UBE2C over-expressing transgenic mice display whole chromosome instability and are prone to spontaneous tumor formation [59]. Breast cancer is among the carcinomas that over-express UBE2C gene and protein [60] and there is a correlation of this over-expression with higher grade of carcinomas [61], proliferation index as measured by the Ki-67 antigen staining and Her2/Neu positivity [62]. High expression of UBE2C mRNA by RT-PCR was found to be associated with poor disease free and overall survival in patients with breast cancer treated with adjuvant chemotherapy [63] and protein evaluation by IHC was a prognostic marker for metastasis free and overall survival in another series of node positive breast cancer patients [64]. In this last study of 92 patients, a positive UBE2C staining (defined as more than 11 % positive cells) added prognostic information to the well-validated Nottingham Prognostic Index [64]. The E2 enzyme UBE2S that co-operates with APC/C ligase for the elongation of lysine 11 chains is also over-expressed in the protein level in many breast tumors and often in correlation with other cell cycle regulators [65]. An implication of UBE2S in topoisomerase II inhibitors etoposide and doxorubicin sensitivity has been suggested based on the fact that knock-down of the conjugating enzyme in vitro sensitized cells to the cytotoxicity of these inhibitors [65].
UBE2D2 and UBE2D3
UBE2D3 (UbcH5c) is one of the E2 partners of BRCA1/BARD1 ubiquitin ligase heterodimer co-operating with this ligase to build lysine 6 ubiquitin chains [66] in DNA damage repair when, as discussed, BRCA1 is recruited to double strand DNA damaged sites to participate to repair by homologous recombination [67]. UBE2D3 is directly regulated by the transcription regulator SLUG (also termed SNAIL2) which recruits co-repressor CtBP1 and histone deacetylase HDAC1 to the UBE2D3 promoter and represses its expression [68]. UBE2D3 participates in the ubiquitination and degradation of cyclin D1 which is stabilized after UBE2D3 suppression and promotes proliferation of breast cancer cells in vitro [68]. In addition cyclin D acts as an ERα transcription co-activator, competing with BRCA1 for receptor binding [69]. In this manner SLUG which is a core transcription regulator of epithelial to mesenchymal transition program, links proliferation with invasion and metastasis (Fig. 4). Nevertheless a study of 25 mostly non-metastatic breast cancers found increased expression of UBE2D in cancer specimens compared with adjacent normal tissue [70]. This increase was specific for breast cancer because colorectal cancer samples did not show an increased UBE2D expression. The rabbit polyclonal antibody used in this study recognizes all three isoforms of UBE2D and thus it is not possible to ascertain if this over-expression concerns one or more of the three isoforms UBE2D1-3 of the conjugating enzyme. Thus, it remains plausible that increased expression observed in breast cancer concerns also UBE2D2 isoform and confers a benefit for cancer cells by inhibiting p53 and activating NF-κB through IκBα degradation (see next).
Schematic representation of the roles of ubiquitin conjugating enzyme UBE2D3 in breast cancer. UBE2D3 co-operates with RING type ligases BRCA1, Mdm2 and βTrCP to down-regulate ERα, p53 and IκB respectively. It also interferes with ERα transcription function by antagonizing cyclin D. Epithelial to mesenchymal transition regulator slug is a repressor of UBE2D3 in co-operation with factor CtBP1 and deacetylase HDAC1
UBE2D2 (UbcH5b) and UBE2D3 are E2 enzymes that co-operate with ligase Mdm2 in the ubiquitination of p53 [71]. Their down-regulation in breast cancer cells in vitro leads to accumulation of p53, but it is not sufficient to induce p53 transcriptional activity [71]. This is possibly due to the fact that Mdm2 is able to partially suppress p53 activity independently from its ligase activity. UBE2D2 and UBE2D3 participate also in NF-κB regulation by co-operating with ligase β-TrCP in the degradation of NF-κB inhibitor IκBα [72]. This is one of the most important points of NF-κB regulation and is also regulated by additional post-translational modifications as previously described (Fig. 3). These effects of UBE2D in p53 and NF-κB regulation may contribute to cancer promotion in early breast cancer with over-expression of UBE2D [70]. It is tempting to speculate that, in more advanced cases with SLUG over-expression and features of EMT, UBE2D becomes down-regulated, an event that would also promote ERα transcription suppression and genomic instability through BRCA1 dysfunction.
Ube2b
UBE2B (hHR6B or Rad6B) is an E2 enzyme that together with the highly homologous UBE2A (hHR6A or Rad6A) contribute to DNA repair [73]. UBE2B is up-regulated following adriamycin-induced DNA damage in breast cancer cells and is localized to sites of damage [74]. UBE2B plays also a role in stabilization of β-catenin [75]. This effect is mediated by building of Lysine 63 ubiquitin chains in contrast to Lysine 48 chains which are built with the aid of ligase βTrCP and lead to proteasome degradation [75]. The Wnt/β-catenin signalling pathway is important in development but also in carcinogenesis. Its activation promotes EMT in epithelial cancers and metastasis [76]. In addition of being a positive regulator of β-catenin, UBE2B is a target gene of the Wnt/β-catenin pathway completing a positive feed-forward loop [77, 78].
UBE2B forms a complex with ligase Mdm2 that leads to p53 degradation [79]. This complex is dissociated in stress conditions when the conjugating enzyme is freed and accumulates on chromatin, participating in histone methylation and induction of p53 transcriptional activity [79]. It is evident that Mdm2 co-operates with different E2 enzymes for the regulation of p53 stability [71, 79] but the functional status of p53 differs after E2-E3 dissociation depending on the E2 participating in the initial complex. UBE2B supports p53 transcriptional activity whereas if UBE2D2 or UBE2D3 were the initial partners, p53 is stabilized without transcriptional activation. The choice of different E2 that participate in p53 regulation may depend on their expression in different cell types or other post-translational modifications. For example phosphorylation of UBE2B by CDK kinases has been reported to affect its functional capability in breast cancer cells [80]. As a result of the base-line availability and functionality of the E2s, a cell that possesses a functional p53 may respond differently to stress by activating p53-dependent transcription or stabilization of p53 without initializing transcription. In an integrative scenario p53 associated with UBE2D3 could represent an inactive and non-degradable pool of p53 protected from UBE2B-Mdm2 degradation and in constant equilibrium with it. In stress conditions the equilibrium is flipped towards the UBE2B-Mdm2 complex because there is a signal for dissociation that creates free UBE2B molecules. These molecules can either promote p53 transcription activity or participate with newly synthesised Mdm2 to new p53-Mdm2-UBE2B complexes in a negative feed-back loop (Fig. 5). Additional regulators such as the Mdm2 partner Mdm4 (also known as Mdmx) fine-tune p53 stability and function [81].
Model of regulation of tumor suppressor p53 by E2 enzymes. Association with conjugating enzyme UBE2D3 prevents binding to Mdm2-UBE2B and stabilizes p53 in an inactive form. Binding to Mdm2-UBE2B promotes ubiquitination and proteasome degradation of p53 but also allows in stressful conditions the activation of the transcription factor after dissociation of Mdm2 and UBE2B. In these conditions UBE2B associates with chromatin and acts as a p53 co-activator for the transcription of Mdm2 and other target genes
In a study of 20 patients with breast cancer of various molecular subtypes receiving neo-adjuvant doxorubicin-based chemotherapy, patients with higher UBE2B nuclear staining had a higher clinical response rate than patients with lower expression of the E2 enzyme [82]. Nevertheless, there was no statistically significant difference in pathologic response between higher and lower nuclear expressors possibly due to the low number of patients in the study [82]. As a result the confirmation of a possible predictive role of nuclear UBE2B in neo-adjuvant chemotherapy of breast cancer awaits further studies. Mechanistically a nuclear localization of UBE2B would be expected to have a role in therapeutic response to chemotherapeutic drugs. This response could follow a dissociation of p53 from the UBE2B-Mdm2 complex after drug-induced stress. Of interest there was no association of Rad18, a known factor localizing UBE2B to damaged sites, with nuclear localization of UBE2B and thus other factors are probably mediating this recruitment following chemotherapy [82].
Other E2s with a role in breast cancer
Additional E2s have been reported to have a role in breast cancer cell proliferation. E2 UBE2L3 (UbcH7) regulates the abundance of phosphatase PTEN in breast cancer cells. Immuno-precipitation experiments showed that UBE2L3 as well as the SUMO E2 UBE2I associate with PTEN [83]. Signalling by the bone morphogenetic protein (BMP) pathway decreases these associations and may stabilize PTEN (Fig. 6). UBE2L3 also acts as a co-activator of nuclear receptors including ERα and PR [84]. Co-activation requires the enzymatic activity of UBE2L3 and the presence of co-activator steroid receptor co-activator 1 (SRC1) on the promoter site occupied by the hormone-bound steroid receptor. Knock-down of UBE2L3 in breast cancer cells reduced the transcriptional activity from the PR promoter. In addition, exogenous administration of UBE2L3 was able to reduce squelching that is observed between ERα and PR-dependent transcription due to competition for limiting amounts of co-factors, implying that the conjugating enzyme is critical for both ERα and PR transcription [84]. UBE2L3 has been found to bind the BRCA1/BARD1 dimer but is unable to co-operate with it in ubiquitination [85]. It has been suggested that this interaction interferes with ability of BRCA1/BARD1 dimers to trigger an intra-S phase checkpoint of the cell cycle [86]. It would also be interesting to examine if this interaction interferes with the BRCA1 repressing activity on ERα transcription.
A conjugating enzyme network regulating steroid receptors in breast cancer. E2 UBE2T inhibits BRCA1, indirectly promoting ERα transcription. NEDDylation E2 UBE2M participates in the turn-over of ERα. UBE2L3 promotes ERα and PR transcription both directly and in two indirect ways; by interfering with stability of BRCA1 and of PTEN. This last event promotes kinase Akt signaling which is involved in ERα activation independently from estrogens and is also involved in hormonotherapy resistance. PR is a target of ERα. A functional ERα leads to PR production which becomes available for functioning as transcription factor
The conjugating enzyme UBE2L6 (UbcH8) is the E2 for the ubiquitin-like protein ISG15 also called, ubiquitin cross-reacting protein (UCRP). ISG15 was initially identified as one of the proteins up-regulated following treatment of cells with interferon-α and -β and has two domains homologous to ubiquitin. As a result the whole molecule has a structure similar to an ubiquitin linear dimer [87]. ISGylation, the conjugation of ISG15 to target proteins, antagonizes in many occasions ubiquitination of these proteins and results in their stabilization by inhibiting proteasome degradation in a manner analogous to SUMOylation [88]. ISG15 expression is elevated in breast cancer cells [89] and both ISG15 and UBE2L6 contribute to cell motility and invasion by interfering with the ubiquitin–proteasome system and by modifying the function of proteins with roles in signal transduction such as the JAK/STAT and MAPK pathways [90–92]. In contrast, stable knock-down of UBE2L6 by a small hairpin RNA in breast cancer cell lines increased focal adhesions and decreased cell motility in a wound healing assay in vitro [90]. Elevated ISG15 in cancer cells may confer sensitivity to topoisomerase I inhibitors camptothecins (used clinically as anti-neoplastic drugs) by blocking the ubiquitin-dependent repair of camptothecin-induced DNA/topoisomerase I complexes [93]. Interestingly, knock-down of the tumor suppressor BRCA2 associated with hereditary breast cancer results in a decrease of both UBE2L6 and ISG15 in breast cancer cells [94].These results suggest an explanation for the association of BRCA2-mutated breast cancers with resistance to certain drugs.
The role of the voluminous E2 enzyme Apollon (also named BIRC6 or BRUCE but has no UBE2 designation) has been investigated in breast cancer cells [95]. Apollon is a class IV E2 that has an amino-terminal BIR [Baculoviral inhibitor of apoptosis (IAP) repeat] domain and a carboxy-terminal conjugating enzyme domain. It antagonizes apoptosis by suppressing p53 activity [96] and by participating in the degradation of protein second mitochondria-derived activator of caspases/direct IAP binding protein with low isoelectric point (SMAC/DIABLO) [97]. It also plays a role in abscission, the very late event of mitosis when the final bridge between the two daughter cells is cut [98]. In breast cancer cells knock-down of Apollon leads to decreased proliferation and enhanced apoptosis associated with p53 stabilization and caspase 3 activation [95]. Nevertheless, it remains to be investigated if Apollon has a role in human breast cancers in vivo.
UBE2T (HSPC150) is an E2 enzyme that regulates proteasome degradation of BRCA1 [99]. It is up-regulated in breast cancer cell lines and clinical samples. Its knock-down by RNA interference caused BRCA1 up-regulation and cell growth inhibition [99]. Thus UBE2T may be one of the proteins whose up-regulation contributes to dysfunction of BRCA1-related processes in breast cancers that have wild type BRCA1 (Fig. 6).
The NEDDylation pathway is intimately involved in ubiquitination reactions mediated by RING E3 ligases of the cullin sub-family because these ligases require NEDDylation for proper function [100]. The role of NEDDylation in the regulation of NF-κB signalling has been mentioned above. Additionally, NEDDylation regulates ERα turn-over in breast cancer (Fig. 6). The NEDDylation conjugating enzyme UBE2M (also called Ubc12) participates in this process and the presence of mutated UBE2M in breast cancer cells stabilizes the receptor [101].
Two homologous putative E2 ubiquitin conjugating enzymes UBE2Q1 and UBE2Q2 have been shown to be over-expressed in breast cancer tissues compared to adjacent normal breast in a subset of breast cancer cases [102–104]. These enzymes have an amino-terminal RWD (RING, WD repeat and DEAD-like) domain in addition to their carboxy-terminal conjugating domain and may be involved in the regulation of the cytoskeleton [103]. Thus over-expression of these E2 enzymes in breast cancer would lead to deregulation of processes involving the cytoskeleton such as cytokinesis, adhesion and metastasis.
Ubiquitin conjugating enzymes and breast cancer sub-types
The above discussion illustrates the role of ubiquitination and other ubiquitin-like protein modifications in the pathogenesis of breast cancer. These modifications regulate many important molecular events in normal breast physiology and breast carcinogenesis. UBC enzymes have, as a result, a central role in these events. Important proteins in breast cancer pathogenesis such as ERα, BRCA1 and NF-κB are regulated by ubiquitin and UBLs attachment. Thus, the machinery serving these attachments and E2 enzymes in particular may have a defining role in breast cancer phenotypes, natural history and treatment response. BRCA1 is a positive regulator of ERα expression and this fact may explain the negativity of BRCA1 mutated breast cancers for ERα [105]. Moreover it represses basal markers such as cytokeratins 5 and 17 and P cadherin [106]. ERα negativity makes these cancers concomitantly negative for the PR, given that PR is an ERα transcriptional target [107]. In addition, BRCA1 directly down-regulates PR activity by inhibiting recruitment of additional PR molecules to promoters and favouring recruitment of co-repressors instead of co-activators [108]. PR serves as a transcriptional inducer of receptor activator of NF-κB ligand (RANKL) and thus may stimulate NF-κB signalling in a paracrine manner. This has implications for understanding the regulation of ERα negative breast cancers by estrogens. In these cancers estrogens may stimulate adjacent normal breast cells expressing the receptor which then up-regulate PR and produce RANKL which, in its turn, binds to RANK receptor on adjacent neoplastic cells and stimulates them [109, 110]. The role of BRCA1 in ERα expression regulation is independent of its role in DNA damage repair. This also explains the fact that cancers mutant for BRCA2, the other common culprit in hereditary breast cancers, are not of a restricted phenotype. Instead these cancers follow the distribution of sporadic breast cancers, given that BRCA2 plays a role in DNA damage repair but not in ERα expression. ERα down-regulation is associated with the aggressive phenotype of epithelial to mesenchymal transition in breast cancer cells [111]. As a result, despite being derived from a process different from DNA damage repair in which BRCA1 participates, i.e. ERα expression regulation, ERα negativity leads also to aggressiveness and treatment resistance which are both features of epithelial to mesenchymal transition. Transcription factor NF-κB is a target of ERα transcriptional repression and is thus up-regulated in ERα negative breast cancer [112]. The small percentage of BRCA1 mutant cancers with ERα positivity may be produced by a dissociation of the two functions of BRCA1. In these cases BRCA1 has a defective DNA repair function while it retains the ability to positively regulate ERα. On the other hand, the significant percentage of breast cancers with BRCA1 wild type but ERα negativity may be produced by dysregulation of BRCA1 function despite lack of mutations, for example from defective post-translation modifications including ubiquitination and SUMOylation. As mentioned in the previous section, up-regulation of E2 UBE2T may lead to BRCA1 down-regulation [99]. Alternatively ERα expression loss may be due to dysfunction of other ERα expression regulators.
Recent data show that the phenotype of BRCA1 mutant breast cancers depends on other concomitant mutations that precede the loss of heterozygosity (LOH) in the BRCA1 locus [113]. If PTEN loss of function is present before this LOH, cancers of a basal triple negative phenotype arise. In the case that PTEN is functional, p53 mutations often precede BRCA1 LOH and luminal type cancers develop [113]. These data add another dimension in the regulation of breast cancer phenotypes by E2s given that E2s UBE2L3 and UBE2I participate in PTEN regulation [83]. PTEN loss of function may also explain the third element of triple negativity, Her2/Neu lack of amplification or hyper-expression. If PTEN is not functional the PI3K/Akt pathway down-stream is up-regulated without the need of Her2/Neu amplification. In other words Her2/Neu amplification would not offer additional benefit for the survival of cancer cells. It would also explain the activation of NF-κB in triple negative cancers, as this transcription factor is activated by kinase Akt [114]. In addition the repressive role of ERα on NF-κB transcription [112] is absent in triple negative cancers. SUMO E2 UBE2I is over-expressed in many cases of breast cancer and especially of the triple positive or triple negative phenotype [51]. The association with triple negativity may be related to down-regulation of BRCA1 function which suppresses ERα expression and promotes at the same time “BRCAness” leading to PTEN loss of function which disfavours Her2/Neu amplification, as argued above. In contrast the triple positivity may be associated with UBE2I over-expression when, due to other concomitant regulations, “BRCAness” is absent, ERα remains expressed and promotes PR expression leading to RANKL expression. Nevertheless, despite the expression of the two hormone receptors, their function remains sub-optimal due to SUMOylation deregulation and PR target gene RANKL is not optimally produced. In addition, due to IκB SUMOylation, this inhibitor becomes more resistant to ubiquitination and NF-κB becomes more refractory to activation. In this scenario NF-κB becomes more dependent on additional pathways for activation and thus amplification or hyper-expression of Her2/Neu may confer additional advantage for the cancer cell to obtain this activation. In breast cancers which do not over-express UBE2I, ERα and PR function normally and RANKL is produced to activate NF-κB without the need for additional input. In addition, if hormone receptors function well, cells may depend less on NF-κB for survival [115]. This may explain the observed dissociation of the expressions of hormone receptors from Her2/Neu amplification [51].
Therapeutic perspectives
The UPS has been validated as a target of anti-cancer treatment with the approval and use of the proteasome inhibitor bortezomib in multiple myeloma. Other proteasome inhibitors are in clinical development [116]. Nevertheless, in other malignancies bortezomib had not the same success [117]. The exquisite sensitivity of myeloma to proteasome inhibition may derive from the specialized function of plasma cells which have the machinery for the production of great amounts of immunoglobulins and perturbations of this process may easily promote apoptosis in these cells [118]. In contrast other malignant cells may be less sensitive to such inhibition, being able to recompense by various mechanisms. The proteasome possesses three enzymatic activities. Bortezomib is a specific inhibitor of the chymotrypsin activity of the proteasome and, although this is the most important enzymatic activity of the three, the two other activities, trypsin and caspase-like, may suffice to functionally substitute for this inhibition in other than myeloma cell types. Indeed, in breast cancer cell lines, acute exposure to bortezomib, even in concentrations 2 orders of magnitude higher than those that inhibit chymotrypsin activity by 95 % failed to stabilize ERα [119]. In contrast chronic exposure to bortezomib in the same cells down-regulated ERα by interfering with its transcription [119]. Another reason for the failure of bortezomib in solid tumors may relate to the lack of specificity of proteasome inhibition. Dozens of proteins are proteasome substrates and the net effect of up-regulations and down-regulations following proteasome inhibition in each of them as part of complex networks may lead to various outcomes in cancer cells depending on the context. Two alternative strategies could overcome this lack of selectivity. The first could be interference with parts of the UPS besides the proteasome per se that work with a more restricted set of partners. A second strategy could target sub-sets of cancers with specific characteristics or molecular lesions that would make them sensitive to proteasome inhibition or sensitive to modulation of other steps of the UPS. Inhibition of specific ubiquitination enzymes or enzymes participating in other UBLs modifications is an example of the first strategy. NEDDylation is a post-translational modification that often takes place in the Cullin component of the SCF type RING ligases and helps the E2 enzyme binding to the ligase complex in the ubiquitination cascade. MLN4924 is a small molecule inhibitor of the NEDD8 activating enzyme (NAE, the E1 for NEDD8) and is in early clinical development [120]. Nevertheless the selectivity of this intervention is only partial, given that RING ligases of the SCF sub-family include several members and in addition other proteins such as p53 and caspases are targets of NEDDylation [121, 122]. As a result, it remains to be seen if a sub-set of cancers with a particular sensitivity to this drug can be defined. In breast cancer, interference with the NEDDylation pathway could be a way to promote expression of ERα in ERα negative cancers and promote sensitivity to hormonal therapies.
Alternatively inhibition of specific E3 ligases or E2 ubiquitin conjugating enzymes could offer greater specificity. For example, inhibition of Mdm2 is a strategy in development but it would be expected to be more effective in cancers with wild type p53 [123]. Concerning E2 inhibitors, a specific conjugating enzyme inhibitor inhibiting the E2 UBE2R1 has been developed and shown to inhibit ubiquitination of target protein CDK inhibitor p27 [124]. In another example of interference with specific E2 enzymes, it was shown that expression of a variant UBE2I protein representing the amino-terminal part of the enzyme interferes with the function of p53 and PTEN and synergizes with chemotherapy to kill cells in vitro [125]. Clinical development of E2 specific inhibitors has not started yet.
The second strategy would involve defining malignancies with specific molecular lesions that could be particularly sensitive to ubiquitin–proteasome system function inhibition as exemplified by multiple myeloma. This task requires development of molecular markers that would be easily and reproducibly performed in a clinical setting. Protein p130Cas, an adaptor protein of cellular adhesions, has been proposed as a marker of sensitivity to bortezomib [126]. Cells that lack p130Cas are resistant to apoptosis after bortezomib treatment and trigger autophagy while, in contrast, cells expressing this protein are sensitive to proteasome inhibition [126]. Nevertheless, p130Cas deficient cells are concomitantly resistant to doxorubicin. Thus, it remains possible that this resistance is generalized and related to adhesion destabilization and may not represent a marker of resistance to particular treatments. Another group of investigators proposed genome scale expression profiling for defining proteasome inhibitor sensitive breast cancer sub-sets and described a proteasome gene module comprised of 50 genes including members of the proteasome core and regulatory particles and ubiquitination enzymes and regulators [127]. Breast cancer patients with increased expression of this module had poorer metastasis-free and overall survival than patients with low expression of genes of the module. It was also shown that breast cancer cell lines with high expression of the proteasome module were more sensitive to bortezomib in vitro [127]. These data, if confirmed, beg for a “re-development” of bortezomib specifically in the sub-set of breast cancer patients with the proteasome gene signature. The same drug development strategy based on marker-defined sub-sets of sensitive cancers could be used for the development of other drugs such as conjugating enzymes inhibitors. It represents a more rational way of development than the usual phase II model that includes all patients of a given cancer type or location.
A final strategy to explore in ubiquitin–proteasome system inhibitors development in breast cancer could be the rational combination with other targeted therapies. Such therapies already exist in breast cancer such as trastuzumab, lapatinib and pertuzumab for the Her2/Neu positive sub-type of the disease. Hormonal therapies also represent a form of targeted treatment as they inhibit specific targets, ERα itself or the aromatase enzyme. Recently PARP inhibitors have shown encouraging results in BRCA1 and 2 mutant cancers [128] illustrating the concept of synthetic lethality. Synthetic lethality refers to the development of dependence of a cell to a pathway if a lesion in a parallel pathway is present, so that if the first pathway is pharmacologically inhibited the cell dies [129]. In BRCA mutated breast cancer homologous recombination is dysfunctional and cells depend on the base excision repair pathway for DNA repair and are particularly sensitive to inhibition of this pathway by PARP inhibitors. Interestingly a state of “BRCAness” has been induced in myeloma cells by bortezomib treatment and has sensitized these cells to PARP inhibitor ABT-888 [130]. If the same effect could be produced by ubiqutin-proteasome system inhibition and PARP inhibition in BRCA wild type breast cancer remains to be investigated. Inhibitors of the specific E2 enzymes co-operating with BRCA1, if they were to be discovered and developed, would also be rational candidates for combinations with PARP inhibitors.
There are clearly ample opportunities for personalized drug development in the field of UPS in breast cancer and the future holds the hope that even metastatic breast cancer will become a curable disease using molecular targeted therapies alone or in combinations.
References
Harbeck N, Salem M, Nitz U, Gluz O, Liedtke C (2010) Personalized treatment of early-stage breast cancer: present concepts and future directions. Cancer Treat Rev 36:584–594
Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA (2010) Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol 11:174–183
Groettrup M, Pelzer C, Schmidtke G, Hofmann K (2008) Activating the ubiquitin family: UBA6 challenges the field. Trends Biochem Sci 33:230–237
Metzger MB, Hristova VA, Weissman AM (2012) HECT and RING finger families of E3 ubiquitin ligases at a glance. J Cell Sci 125:531–537
Hochstrasser M (2009) Origin and function of ubiquitin-like proteins. Nature 458:422–429
Voutsadakis IA (2010) Ubiquitin, ubiquitination and the ubiquitin-proteasome system in cancer. Atlas Genet Cytogen Oncol Haematol http://AtlasGeneticsOncologyorg/Deep/UbiquitinCancerID20083.httml
Kirkin V, Dikic I (2011) Ubiquitin networks in cancer. Curr Opin Genet Dev 21:21–28
Behrends C, Harper JW (2011) Constructing and decoding unconventional ubiquitin chains. Nat Struct Mol Biol 18:520–528
Michelle C, Vourc’h P, Mignon L, Andres CR (2009) What was the set of ubiquitin and ubiquitin-like conjugating enzymes in the eukaryote common ancestor? J Mol Evol 68(616):628
Huang DT, Hunt HW, Zhuang M, Ohi MD, Holton JM, Schulman BA (2007) Basis for a ubiquitin-like protein thioester switch toggling E1–E2 affinity. Nature 445:394–398
Wenzel DM, Stoll KE, Klevit RE (2011) E2 s: structurally economical and functionally replete. Biochem J 433:31–42
Eddins MJ, Pickart CM (2005) Ubiquitin-conjugating enzymes. In: Mayer J, Ciechanover A, Rechsteiner M (eds) Protein degradation, vol 1. Wiley-VCH Verlag, Weinheim, pp 102–134
Van Wijk SJL, Timmers HTM (2010) The family of ubiquitin-conjugating enzymes (E2s): deciding between life and death of proteins. FASEB J 24:981–993
Li W, Ye Y (2008) Polyubiquitin chains: functions, structures, and mechanisms. Cell Mol Life Sci 65:2397–2406
Tokgöz Z, Siepmann TJ, Streich F Jr, Kumar B, Klein JM, Haas AL (2012) E1–E2 interactions in ubiquitin and Nedd8 ligation pathways. J Biol Chem 287:311–321
Hochstrasser M (2006) Lingering mysteries of ubiquitin-chain assembly. Cell 124:27–34
Zheng N, Wang P, Jeffrey PD, Pavletich NP (2000) Structure of a c-Cbl-UbcH7 complex: RING domain function in ubiquitin-protein ligases. Cell 102:533–539
Huang DT, Kinnucan E, Wang G, Baudenon S, Howley PM, Huibregtse JM, Pavletich NP (1999) Structure of an E6AP-UbcH7 complex: insights into ubiquitination by the E2–E3 enzyme cascade. Science 286:1321–1326
Praefcke GJK, Hofmann K, Dohmen RJ (2012) SUMO playing tag with ubiquitin. Trends Biochem Sci 37:23–31
Anderson DD, Eom JY, Stover PJ (2012) Competition between Sumoylation and Ubiquitination of serine hydroxymethyltransferase 1 determines its nuclear localization and its accumulation in the nucleus. J Biol Chem 287:4790–4799
Karamouzis MV, Konstantinopoulos PA, Badra FA, Papavassiliou AG (2008) SUMO and estrogen receptors in breast cancer. Breast Cancer Res Treat 107:195–210
Le Romancer M, Poulard C, Cohen P, Sentis S, Renoir J-M, Corbo L (2011) Cracking the estrogen receptor’s posttranslational code in breast tumors. Endocrine Rev 32:597–622
Sentis S, Le Romancer M, Bianchin C, Rostan M-C, Corbo L (2006) Sumoylation of the estrogen receptor α hinge region regulates its transcriptional activity. Mol Endocrinol 19:2671–2684
Li S, Yang C, Hong Y, Bi H, Zhao F, Liu Y, Ao X, Pang P, Xing X, Chang AK, Xiao L, Zhang Y, Wu H (2012) The transcriptional activity of co-activator AIB1 is regulated by the SUMO E3 ligase PIAS1. Biol Cell 104:1–10
Turner N, Tutt A, Ashworth A (2004) Hallmarks of “BRCAness” in sporadic cancers. Nat Rev Cancer 4:1–6
Foulkes WD (2010) Traffic control by BRCA1. New Engl J Med 362:755–756
Morris JR, Boutell C, Keppler M, Densham R, Weekes D, Alamshah A, Butler L, Galanty Y, Pangon L, Kiuchi T, Ng T, Solomon E (2009) The SUMO modification pathway is involved in the BRCA1 response to genotoxic stress. Nature 462:886–890
Kim H, Chen J, Yu X (2007) Ubiquitin-binding protein RAP80 mediates BRCA1-dependent DNA damage response. Science 316:1202–1205
Yan J, Yang X-P, Kim Y-S, Joo JH, Jetten AM (2007) RAP80 interacts with the SUMO-conjugating enzyme UBC9 and is a novel target for sumoylation. Biochem Biophys Res Commun 362:132–138
Wang B (2012) BRCA1 tumor suppressor network: focusing on its tail. Cell Biosci 2:6
Xu J, Watkins T, Reddy A, Reddy ESP, Rao VN (2009) A novel mechanism whereby BRCA1/1a/1b fine tunes the dynamic complex interplay between SUMO-dependent/independent activities of Ubc9 on E2-induced ERα activation/repression and degradation in breast cancer cells. Int J Oncol 34:939–949
Ma Y, Fan S, Hu C, Meng Q, Fuqua SA, Pestell RG, Tomita YA, Rosen EM (2010) BRCA1 regulates acetylation and ubiquitination of estrogen receptor-α. Mol Endocrinol 24:76–90
Qin Y, Xu J, Aysola K, Begum N, Reddy V, Chai Y, Grizzle WE, Partridge EE, Reddy ESP, Rao VN (2011) Ubc9 mediates nuclear localization and growth suppression of BRCA1 and BRCA1a proteins. J Cell Physiol 226:3355–3367
Woeller CF, Anderson DD, Szebenyi DME, Stover PJ (2007) Evidence for small ubiquitin-like modifier-dependent nuclear import of the thymidylate biosynthesis pathway. J Biol Chem 282:17623–17631
Palczewska M, Casafont I, Ghimire K, Rojas AM, Valencia A, Lafarga M, Mellström B, Naranjo JR (2011) Sumoylation regulates nuclear localization of repressor DREAM. Biochim Biophys Acta 1813:1050–1058
Deng H, Lin Y, Badin M, Vasilcanu D, Strömberg T, Jernberg-Wiklund H, Sehat B, Larsoon O (2011) Over-accumulation of nuclear IGF-1 receptor in tumor cells requires elevated expression of the receptor and the SUMO-conjugating enzyme Ubc9. Biochem Biophys Res Commun 404:667–671
Sehat B, Tofigh A, Lin Y, Trocmé E, Liljedahl U, Lagergren J, Larsson O (2010) SUMOylation mediates the nuclear translocation and signalling of the IGF-1 receptor. Sci Signal 3:ra10
Park MA, Seok Y-J, Jeong G, Lee J-S (2008) SUMO1 negatively regulates BRCA1-mediated transcription, via modulation of promoter occupancy. Nucleic Acids Res 36:263–283
Chauhan D, Bianchi G, Anderson KC (2008) Targeting the UPS as therapy in multiple myeloma. BMC Biochem 9(Suppl 1):S1
Liu Q, Li J, Khoury J, Colgan SP, Ibla JC (2009) Adenosine signalling mediates SUMO-1 modification of IκBα during hypoxia and reoxygenation. J Biol Chem 284:13686–13695
Shifera AS (2010) Protein-protein interactions involving IKKγ (NEMO) that promote the activation of NF-Κb. J Cell Physiol 223:558–561
Zhou Y, Eppenberger-Castori S, Eppenberger U, Benz CC (2005) The NF-κB pathway and endocrine-resistant breast cancer. Endocr Relat Cancer 12:S37–S46
Montagut C, Tusquets I, Ferrer B, Corominas JM, Bellosillo B, Campas C, Suarez M, Febregat X, Campo E, Gascon P, Serrano S, Fernandez PL, Rovira A, Albanell J (2006) Activation of nuclear factor-κ B is linked to resistance to neoadjuvant chemotherapy in breast cancer patients. Endocr Relat Cancer 13:607–616
Buchholz TA, Garg AK, Chakravarti N, Aggarwal BB, Esteva FJ, Kuerer HM, Singletary SE, Hortobagyi GN, Pusztai L, Cristofanilli M, Sahin AA (2005) The nuclear transcription factor κB/bcl-2 pathway correlates with pathologic complete response to doxorubicin-based neoadjuvant chemotherapy in human breast cancer. Clin Cancer Res 11:8398–8402
Jones RL, Rojo F, A’Hern R, Villena N, Salter J, Corominas JM, Servitja S, Smith IE, Rovira A, Reis-Filho JS, Dowsett M, Albanell J (2011) Nuclear NF-κB/p65 expression and response to neoadjuvant chemotherapy in breast cancer. J Clin Pathol 64:130–135
Perkins ND (2012) The diverse and complex roles of NF-κB subunits in cancer. Nat Rev Cancer 12:121–132
Shostak K, Chariot A (2011) NF-κB, stem cells and breast cancer: the links get stronger. Breast Cancer Res 13:214
Yu F, Deng H, Yao H, Liu Q, Su F, Song E (2010) Mir-30 reduction maintains self-renewal and inhibits apoptosis in breast tumor-initiating cells. Oncogene 29:4194–4204
Zhu S, Sachdeva M, Wu F, Lu Z, Mo Y-Y (2010) Ubc9 promotes breast cell invasion and metastasis in a sumoylation-independent manner. Oncogene 29:1763–1772
Guo Y, Yang M-CW, Weissler JC, Yang Y-S (2008) Modulation of PLAGL2 transactivation activity by Ubc9 co-activation not SUMOylation. Biochem Biophys Res Commun 374:570–575
Chen S-F, Gong C, Luo M, Yao H-R, Zeng Y-J, Su F-X (2011) Ubc9 expression predicts chemoresistance in breast cancer. Chin J Cancer 30:638–644
Bremm A, Komander D (2011) Emerging roles for Lys11-linked polyubiquitin in cellular regulation. Trends Biochem Sci 36:355–363
Castro A, Vigneron S, Lorca T, Labbé J-C (2003) La mitose sous surveillance. Med Sci 19:309–317
Matyskiela ME, Morgan DO (2009) Analysis of activator-binding sites on the APC/C supports a cooperative substrate-binding mechanism. Mol Cell 34:68–80
Song L, Rape M (2011) Substrate-specific regulation of ubiquitination by the anaphase-promoting complex. Cell Cycle 10:52–56
Wickliffe KE, Williamson A, Meyer H-J, Kelly A, Rape M (2011) K11-linked ubiquitin chains as novel regulators of cell division. Trends Cell Biol 21:656–663
Mocciaro A, Rape M (2012) Emerging regulatory mechanisms in ubiquitin-dependent cell cycle control. J Cell Sci 125:255–263
Okamoto Y, Ozaki T, Miyazaki K, Aoyama M, Miyazaki M, Nakagawara A (2003) UbcH10 is the cancer-related E2 ubiquitin-conjugating enzyme. Cancer Res 63:4167–4173
van Ree JH, Jeganathan KB, Malureanu L, van Deursen JM (2010) Overexpression of the E2 ubiquitin-conjugating enzyme UbcH10 causes chromosome missegregation and tumor formation. J Cell Biol 188:83–100
Wagner KW, Sapinoso LM, El-Rifai W, Frierson HF Jr, Butz N, Mestan J, Hofmann F, Deveraux QL, Hampton GM (2004) Overexpression, genomic amplification and therapeutic potential of inhibiting the UbcH10 ubiquitin conjugase in human carcinomas of diverse anatomic origin. Oncogene 23:6621–6629
Fujita T, Ikeda H, Kawasaki K, Taira N, Ogasawara Y, Nakagawara A, Doihara H (2009) Clinicopathological relevance of UbcH10 in breast cancer. Cancer Sci 100:238–248
Berlingieri MT, Pallante P, Sboner A, Barbareschi M, Bianco M, Ferraro A, Mansueto G, Borbone E, Guerriero E, Troncone G, Fusco A (2007) UbcH10 is overexpressed in malignant breast carcinomas. Eur J Cancer 43:2729–2735
Psyrri A, Kalogeras KT, Kronenwett R, Wirtz RM, Batistatou A, Bournakis E, Timotheadou E, Gogas H, Aravantinos G, Christodoulou C, Makatsoris T, Linardou H, Pectasides D, Pavlidis N, Economopoulos T, Foutzilas G (2012) Prognostic significance of UBE2C mRNA expression in high-risk early breast cancer. A Hellenic Cooperative Oncology Group (HeCOG) study. Ann Oncol 23:1422–1427
Loussouarn D, Campion L, Leclair F, Campone M, Charbonnel C, Ricolleau G, Gouraud W, Bataille R, Jezéquel P (2009) Validation of UBE2C protein as a prognostic marker in node-positive breast cancer. Br J Cancer 101:166–173
Tedesco D, Zhang J, Trinh L, Lalehzadeh G, Meisner R, Yamaguchi KD, Ruderman DL, Dinter H, Zajchowski DA (2007) The ubiquitin-conjugating enzyme E2-EPF is overexpressed in primary breast cancer and modulates sensitivity to topoisomerase II inhibition. Neoplasia 9:601–613
Wu-Baer F, Lagrazon K, Yuan W, Baer R (2003) The BRCA1/BARD1 heterodimer assembles polyubiquitin chains through an unconventional linkage involving lysine residue K6 of ubiquitin. J Biol Chem 278:34743–34746
Polanowska J, Martin JS, Garcia-Muse T, Petalcorin MIR, Boulton SJ (2006) A conserved pathway to activate BRCA1-dependent ubiquitylation at DNA damage sites. EMBO J 25:2178–2188
Mittal MK, Singh K, Misra S, Chaudhuri G (2011) SLUG-induced elevation of D1 cyclin in breast cancer cells through the inhibition of its ubiquitination. J Biol Chem 286:469–479
Wang C, Fan S, Li Z, Fu M, Rao M, Ma Y, Lisanti MP, Albanese C, Katzenellenbogen BS, Kushner PJ, Weber B, Rosen EM, Pestell RG (2005) Cyclin D1 antagonizes BRCA1 repression of estrogen receptor α activity. Cancer Res 65:6557–6567
Chen L, Madura K (2005) Increased proteasome activity, ubiquitin-conjugating enzymes, and eEF1A translation factor detected in breast cancer tissue. Cancer Res 65:5599–5606
Saville MK, Sparks A, Xirodimas DP, Wardrop J, Stevenson LF, Bourdon J-C, Woods YL, Lane DP (2004) Regulation of p53 by the ubiquitin-conjugating enzymes UbcH5B/C in vivo. J Biol Chem 279:42169–42181
Gonen H, Bercovich B, Orian A, Carrano A, Takizawa C, Yamanaka K, Pagano M, Iwai K, Ciechanover A (1999) Identification of the ubiquitin carrier proteins, E2s, involved in signal-induced conjugation and subsequent degradation of IκBα. J Biol Chem 274:14823–14830
Bergink S, Jentsch S (2009) Principles of ubiquitin and SUMO modifications in DNA repair. Nature 458:461–467
Lyakhovich A, Shekhar MPV (2004) RAD6B overexpression confers chemoresistance: RAD6 expression during cell cycle and its redistribution to chromatin during DNA damage-induced response. Oncogene 23:3097–3106
Shekhar MPV, Gerard B, Pauley RJ, Williams BO, Tait L (2008) Rad6B is a positive regulator of β-catenin stabilization. Cancer Res 68:1741–1750
Voutsadakis IA (2012) The ubiquitin–proteasome system and signal transduction pathways regulating epithelial mesenchymal transition of cancer. J Biomed Sci 19:67
Shekhar MPV, Tait L, Gerard B (2006) Essential role of T-cell factor/β-catenin in regulation of Rad6B: a potential mechanism for Rad6B overexpression in breast cancer cells. Mol Cancer Res 4:729–745
Gerard B, Tait L, Nangia-Makker P, Shekhar MPV (2011) Rad6B acts downstream of Wnt signaling to stabilize β-catenin: implications for a novel Wnt/β-catenin target. J Mol Signal 6:6
Chen S, Wang D-L, Liu Y, Zhao L, Sun F-L (2012) RAD6 regulates the dosage of p53 by a combination of transcriptional and posttranscriptional mechanisms. Mol Cell Biol 32:576–587
Kumar B, LeCompte KG, Klein JM, Haas AL (2010) Ser120 of Ubc2/Rad6 regulates ubiquitin-dependent N-end rule targeting by E3α/Ubr1. J Biol Chem 285:41300–41309
Perry ME (2010) The regulation of the p53-mediated stress response by MDM2 and MDM4. Cold Spring Harb Perspect Biol 2:a000968
Shekhar MPV, Biernat LA, Pernick N, Tait L, Abrams J, Visscher DW (2010) Utility of DNA postreplication repair protein Rad6B in neoadjuvant chemotherapy response. Med Oncol 27:466–473
Waite KA, Eng C (2003) BMP2 exposure results in decreased PTEN protein degradation and increased PTEN levels. Hum Mol Genet 12:679–684
Verma S, Ismail A, Gao X, Fu G, Li X, O’Malley BW, Nawaz Z (2004) The ubiquitin-conjugating enzyme UBCH7 acts as a coactivator for steroid hormone receptors. Mol Cell Biol 24:8716–8726
Brzovic PS, Keeffe JR, Nishikawa H, Miyamoto K, Fox D III, Fukuda M, Ohta T, Klevit R (2003) Binding and recognition in the assembly of an active BRCA1/BARD1 ubiquitin-ligase complex. Proc Natl Acad Sci USA 100:5646–5651
Whitcomb EA, Taylor A (2009) Ubiquitin control of S phase: a new role for the ubiquitin conjugating enzyme, UbcH7. Cell Div 4:17
Zhang D, Zhang D-E (2011) Interferon-stimulated gene 15 and the protein ISGylation system. J Interf Cytok Res 31:119–130
Desai SD, Haas AL, Wood LM, Tsai Y-C, Pestka S, Rubin EH, Saleem A, Nur-E-Kamal A, Liu LF (2006) Elevated expression of ISG 15 in tumor cells interferes with the ubiquitin/26S proteasome pathway. Cancer Res 66:921–928
Bektas N, Noetzel E, Veeck J, Press MF, Kristiansen G, Naami A, Hartmann A, Dimmler A, Beckmann MW, Knüchel R, Fasching PA, Dahl E (2008) The ubiquitin-like molecule interferon-stimulated gene 15 (ISG15) is a potential prognostic marker in human breast cancer. Breast Cancer Res 10:R58
Desai SD, Reed RE, Burks J, Wood LM, Pullikuth AK, Haas AL, Liu LF, Breslin JW, Meiners S, Sankar S (2012) ISG15 disrupts cytoskeletal architecture and promotes motility in human breast cancer cells. Exp Biol Med 237:38–49
Malakhov MP, Kim KI, Malakhova OA, Jacobs BS, Borden EC, Zhang DE (2003) High-throughput immunoblotting. Ubiquitin-like protein ISG15 modifies key regulators of signal transduction. J Biol Chem 278:16608–16613
Malakhova OA, Yan M, Malakhov MP, Yuan Y, Ritchie KJ, Kim KI, Peterson LF, Shuai K, Zhang DE (2003) Protein ISGylation modulates the JAK-STAT signaling pathway. Genes Dev 17:455–460
Desai SD, Wood LM, Tsai Y-C, Hsieh T-S, Marks JR, Scott GL, Giovanella BC, Liu LF (2008) ISG15 as a novel tumor biomarker for drug sensitivity. Mol Cancer Ther 7:1430–1439
Tripathi MK, Chaudhuri G (2005) Down-regulation of UCRP and UBE2L6 in BRCA2 knocked-down human breast cells. Biochem Biophys Res Commun 328:43–48
Lopergolo A, Pennati M, Gandellini P, Orlotti NI, Poma P, Daidone MG, Folini M, Zaffaroni N (2009) Apollon gene silencing induces apoptosis in breast cancer cells through p53 stabilization and caspase-3 activation. Br J Cancer 100:739–746
Ren J, Shi M, Liu R, Yang Q-H, Johnson T, Skarnes WC, Du C (2005) The Birc6 (Bruce) gene regulates p53 and the mitochondrial pathway of apoptosis and is essential for mouse embryonic development. Proc Natl Acad Sci USA 102:565–570
Hao Y, Sekine K, Kawabata A, Nakamura H, Ishioka T, Ohata H, Katayama R, Hashimoto C, Zhang X, Noda T, Tsuruo T, Naito M (2004) Apollon ubiquitinates SMAC and caspase-9, and has an essential cytoprotection function. Nat Cell Biol 6:849–860
Pohl C, Jentsch S (2008) Final stages of cytokinesis and midbody ring formation are controlled by BRUCE. Cell 132:832–845
Ueki T, Park J-H, Nishidate T, Kijima K, Hirata K, Nakamura Y, Katagiri T (2009) Ubiquitination and downregulatio of BRCA1 by ubiquitin conjugating enzyme E2T overexpression in human breast cancer cells. Cancer Res 69:8752–8760
Kawakami T, Chiba T, Suzuki T, Iwai K, Yamanaka K, Minato N, Suzuki H, Shimbara N, Hidaka Y, Osaka F, Omata M, Tanaka K (2001) NEDD8 recruits E2-ubiquitin to SCF E3 ligase. EMBO J 20:4003–4012
Fan M, Bigsby RM, Nephew KP (2003) The NEDD8 pathway is required for proteasome-mediated degradation of human estrogen receptor (ER)-α and essential for the antiproliferative activity of ICI 182,780 in ERα-positive breast cancer cells. Mol Endocrinol 17:356–365
Nikseresht M, Seghatoleslam A, Monabati A, Talei A, Ghalati FB, Owji AA (2010) Overexpression of the novel human gene, UBE2Q2, in breast cancer. Cancer Genet Cytogenet 197:101–106
Seghatoleslam A, Nikseresht M, Shafiee SM, Monabati A, Namavari M, Talei A, Safaei A, Owji AA (2012) Expression of the novel human gene, UBE2Q1, in breast tumors. Mol Biol Rep 39:5135–5141
Seghatoleslam A, Zambrano A, Million R, Ganguli G, Argentini M, Cromer A, Abecassis J, Wasylyk B (2006) Analysis of a novel human gene, LOC92912, over-expressed in hypopharyngeal tumours. Biochem Biophys Res Commun 339:422–429
Hosey AM, Gorski JJ, Murray MM, Quinn JE, Chung WY, Stewart GE, James CR, Farragher SM, Mulligan JM, Scott AN, Dervan PA, Johnston PG, Couch FJ, Daly PA, Kay E, McCann A, Mullan PB, Harkin DP (2007) Molecular basis for estrogen receptor α deficiency in BRCA1-linked breast cancer. J Natl Cancer Inst 99:1683–1694
Gorski JJ, James CR, Quinn JE, Stewart GE, Crosbie Staunton K, Buckley NE, McDyer FA, Kennedy RD, Wilson RH, Mullan PB, Harkin DP (2010) BRCA1 transcriptionally regulates genes associated with the basal-like phenotype in breast cancer. Breast Cancer Res Treat 122:721–731
Bonéy-Montoya J, Ziegler YS, Curtis CD, Montoya JA, Nardulli AM (2010) Long-range transcriptional control of progesterone receptor gene expression. Mol Endocrinol 24:346–358
Katiyar P, Ma Y, Riegel A, Fan S, Rosen EM (2009) Mechanism of BRCA1-mediated inhibition of progesterone receptor transcriptional activity. Mol Endocrinol 23:1135–1146
Schramek D, Leibbrandt A, Sigl V, Kenner L, Pospisilik JA, Lee HJ, Hanada R, Joshi PA, Aliprantis A, Glimcher L, Pasparakis M, Khokha R, Ormandy CJ, Widschwendter M, Schett G, Penninger JM (2010) Osteoclast differentiation factor RANKL controls development of progestin-driven mammary cancer. Nature 468:98–102
Gonzalez-Suarez E, Jacob AP, Jones J, Miller R, Roudier-Meyer MP, Erwert R, Pinkas J, Branstetter D, Dougall WC (2010) RANK ligand mediates progestin-induced mammary epithelial proliferation and carcinogenesis. Nature 468:103–107
Al Saleh S, Al Mulla F, Luqmani YA (2011) Estrogen receptor silencing induces epithelial to mesenchymal transition in human breast cancer cells. PLoS ONE 6:e20610
Wang X, Belguise K, Kersual N, Kirsch KH, Mineva ND, Galtier F, Chalbos D, Sonenshein GE (2007) Oestrogen signalling inhibits invasive phenotype by repressing RelB and its target BCL2. Nat Cell Biol 9:470–478
Martins FC, De S, Almendro V, Gönen M, Park SY, Blum JL, Herlihy W, Ethington G, Schnitt SJ, Tung N, Garber JE, Fetten K, Michor F, Polyak K (2012) Evolutionary pathways in BRCA1-associated breast tumors. Cancer Discov 2:503–511
Wickenden JA, Watson CJ (2010) Signalling downstream of PI3 kinase in mammary epithelium: a play in 3 Akts. Breast Cancer Res 12:202
Frasor J, Weaver A, Pradhan M, Dai Y, Miller LD, Lin CY, Stanculescu A (2009) Positive cross-talk between estrogen receptor and NF-kappaB in breast cancer. Cancer Res 89:8918–8925
Kisselev AF, van der Linden WA, Overkleeft HS (2012) Proteasome inhibitors: an expanding army attacking a unique target. Chem Biol 19:99–115
Milano A, Perri F, Caponigro F (2009) The ubiquitin-proteasome system as a molecular target in solid tumors: an update on bortezomib. OncoTargets Ther 2:171–178
Cao B, Mao X (2011) The ubiquitin-proteasomal system is critical for multiple myeloma: implications in drug discovery. Am J Blood Res 1:46–56
Powers GL, Ellison-Zelski SJ, Casa AJ, Lee AV, Alarid ET (2010) Proteasome inhibition represses ERα gene expression in ER+ cells: a new link between proteasome activity and estrogen signaling in breast cancer. Oncogene 29:1509–1518
Wang M, Medeiros BC, Erba HP, DeAngelo DJ, Giles FJ, Swords RT (2011) Targeting protein neddylation: a novel therapeutic strategy for the treatment of cancer. Expert Opin Ther Targets 15:253–264
Liu G, Xirodimas DP (2010) NUB1 promotes cytoplasmic localization of p53 through cooperation of the NEDD8 and ubiquitin pathways. Oncogene 29:2252–2261
Broemer M, Tenev T, Rigbolt KTG, Hempel S, Blagoev B, Silke J, Ditzel M, Meier P (2010) Systematic in vivo RNAi analysis identifies IAPs as NEDD8-E3 ligases. Mol Cell 40:810–822
Dickens MP, Fitzgerald R, Fischer PM (2010) Small-molecule inhibitors of MDM2 as a new anticancer therapeutics. Semin Cancer Biol 20:10–18
Ceccarelli DF, Tang X, Pelletier B, Orlicky S, Xie W, Plantevin V, Neculai D, Chou Y-C, Ogunjimi A, Al-Hakim A, Varelas X, Koszela J, Wasney GA, Vedadi M, Dhe-Paganon S, Cox S, Xu S, Lopez-Girona A, Mercurio F, Wrana J, Durocher D, Meloche S, Webb DR, Tyers M, Sicheri F (2011) An allosteric inhibitor of the human Cdc34 ubiquitin-conjugating enzyme. Cell 145:1075–1087
Shiratori T, Shimada H, Kagaya A, Kuboshima M, Nabeya Y, Machida T, Goto K, Takiguchi M, Ochiai T, Hiwasa T (2007) Sensitization against anticancer drugs by transfection with UBE2I variant gene into ras-NIH3H3 mouse fibroblasts. Anticancer Res 27:3227–3234
Zhao M, Vuori K (2011) The docking protein p130Cas regulates cell sensitivity to proteasome inhibition. BMC Biol 9:73
Wong DJ, Nuyten DS, Regev A, Lin M, Adler AS, Segal E, van de Vijver MJ, Chang HY (2008) Revealing targeted therapy for human cancer by gene module maps. Cancer Res 68:369–378
Fong PC, Boss DS, Yap TA, Tutt A, Wu P, Mergui-Roelvink M, Mortimer P, Swaisland H, Lau A, O’Connor MJ, Ashworth A, Carmichael J, Kaye SB, Schellens JH, de Bono JS (2009) Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med 361:123–134
Chan DA, Giaccia AJ (2011) Harnessing synthetic lethal interactions in anticancer drug discovery. Nat Rev Drug Discov 10:351–364
Neri P, Ren L, Gratton K, Stebner E, Johnson J, Klimowicz A, Duggan P, Tassone P, Mansoor A, Stewart DA, Lonial S, Boise LH, Bahlis NJ (2011) Bortezomib-induced “BRCAness” sensitizes multiple myeloma cells to PARP inhibitors. Blood 118:6368–6379
Jiang F, Basavappa R (1999) Crystal structure of the cyclin-specific ubiquitin-conjugating enzyme from clam, E2-C, at 2.0 A resolution. Biochemistry 18:6471–6478
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Voutsadakis, I.A. Ubiquitin- and ubiquitin-like proteins-conjugating enzymes (E2s) in breast cancer. Mol Biol Rep 40, 2019–2034 (2013). https://doi.org/10.1007/s11033-012-2261-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2261-0