Abstract
Phosphorus (P) is an essential macronutrient for plant growth and development. P deficiency could affect rubber tree productivity seriously, and understanding the mechanism responses of the rubber tree under the P deficiency will be helpful to improving rubber tree productivity. The molecular mechanism by which the rubber trees respond to a P-deficiency is a complex network involving many processes. To identify the genes differentially expressed in that response, we constructed subtractive suppression hybridization libraries for roots of plants growing under deficient or sufficient conditions. We identified 94 up-regulated genes from the forward library and 45 down-regulated from the reverse library. These differentially expressed genes were categorized into eight groups representing functions in metabolism, transcription, signal transduction, protein synthesis, transport, stress responses, photosynthesis, and development. We also performed quantitative real-time PCR to investigate the expression profiles of eight randomly selected clones. Our results provide useful information for further study of the molecular mechanism for adaptations to a P-deficiency in this species. Further characterization and functional analysis of these differentially expressed genes will help us improve its phosphorus utilization and overall productivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a major mineral nutrient, phosphorus (P) is essential for many key physiological and biochemical processes in plants, including energy metabolism, biosynthesis of nucleic acids, and membrane development [1]. In soils, P is easily fixed by organic compounds, iron, or aluminum oxides into forms that cannot be utilized by plants. Because the concentration of available-P usually cannot satisfy demands, a low supply can be a critical factor that limits crop production [2, 3]. To cope with this lack of inorganic P, plants employ tightly controlled mechanisms to maintain phosphate homeostasis, e.g., via P-uptake, storage and remobilization, and optimization of metabolic processes that require this nutrient [4, 5].
Extensive studies have revealed the molecular mechanisms for plant responses to P-deficiency. For example, Misson et al. [6] have reported on the deficiency-induced changes in expression of Arabidopsis genes that function in various metabolic pathways, ion transport, signal transduction, transcriptional regulation, and other growth processes. Hammond et al. [7] have shown that 1,659 genes in potatoes are significantly and differentially expressed after P is withdrawn, including those that encode proteins involved in the metabolism of lipids, proteins, and carbohydrates. Li et al. [8] have identified rice genes with altered expression in response to low-P stress. These are active in transportation, the production of phosphatase and other enzymes, and both primary and secondary metabolism. Meanwhile, researchers have cloned and characterized many important genes involved in the sensing, signaling, uptake, transport, and efficient use of P in various other plants [9–20].
Nevertheless, the molecular mechanism underlying P-efficiency by the rubber tree (Hevea brasiliensis Muell. Arg.) is still poorly understood. This tropical perennial species, native to the Amazon Basin, is the world’s major source of natural rubber. Although 10 million tons of rubber were produced in 2010, the ever-increasing, worldwide demand calls for enhanced tree productivity. A deficiency in available phosphorus could have serious effects. Thus, understanding the mechanism by which the rubber tree responds to such stress would be helpful in efforts to improve its productivity. Here, we applied suppression subtractive hybridization (SSH) technology to identify genes that are differentially expressed in the roots. We also compared transcript abundances under both deficient- and sufficient-P conditions.
Materials and methods
Plant materials
An elite clone of Hevea brasiliensis, Reyan7-33-97, was cultivated at the experimental plantation of the Rubber Research Institute of the Chinese Academy of Tropical Agricultural Sciences (Hainan, China). Three-month-seedlings were submerged into Hoagland’s solution containing either 0 mg L−1 P for the deficient treatment or 31 mg L−1 P for the sufficient (control) treatment. Roots exposed to these P conditions (three seedlings per sampling event) were harvested after 1, 2, 4, 6, or 8 d of treatment and stored at −70 °C.
Total RNA extraction
Total RNA was isolated using TRIzol (Invitrogen, USA) according to the manufacturer’s recommendations. RNA quantity and quality were assessed on a Nanodrop spectrophotometer (Nanodrop Technologies, Montchanin, DE, USA) and by agarose gel electrophoresis. Equal amounts (400 ng) of total RNA from the five sampling events were pooled for use in library construction.
Suppression subtractive hybridization and library construction
Two SSH cDNA libraries were constructed according to the manufacturer’s protocol. In the forward subtractive library, seedlings grown under the P-deficiency served as the ‘‘tester’’ while control seedlings were the ‘‘driver’’. For constructing the reverse library, the sufficient control was the tester and the deficient seedlings were the driver. Briefly, cDNA was synthesized from mRNA with a Clontech PCR-Select cDNA Subtraction Kit, using an oligo-dT primer that allowed us to amplify the complete mRNA population contained in each sample. After adaptor-ligation and two subtractive hybridizations, the PCR products were cloned into the pMD18-T vector (TaKaRa, Japan) and transformed, by electroporation, into Escherichia coli DH5a (TaKaRa). The clones were cultured overnight at 37 °C on an LB media plate containing ampicillin and X-Gal/IPTG. White clones were selected for generating a subtractive library and were stored in 384-well plates at −70 °C.
Subtractive library detection
The white clones were randomly picked out from the transformation plates, then placed in a liquid LB medium containing Amp and incubated at 37 °C for 8 h. As template, 1 μL of the medium was removed. Primers included the nested 1 (5′-TCGAGCGGCCGCC CGGG CAGGT-3′) and 2R (5′-AGCGTGGTCGCGGCCGAG GT-3′). The length of the subtractive cDNA fragment was then determined. PCR amplifications were performed with a PCR Thermal Cycler (Biometra) under conditions of pre-denaturation at 95 °C for 3 min; followed by 36 cycles of 95 °C/30 s, 54 °C/30 s, and 72 °C/1 min; and a final step at 72 °C for 10 min. The products were electrophoresed on a 1.0 % agarose gel to confirm their quality and quantity.
Sequencing and sequence analysis
Plasmids containing cDNA fragments that were differentially expressed were sequenced with universal M13 forward and reverse primers homologous to the vector sequence. EST identities and potential functions were determined by comparing sequences against the non-redundant GenBank database via BLASTN and BLASTX (http://www.ncbi. nlm. hih.gov). Functional categorization of those genes was done manually, based on the functional catalog (FunCat) from MIPS (Munich Information Center for Protein Sequences) [21].
Real-time quantitative RT-PCR
To monitor their expression patterns, we randomly selected the following clones for confirmation by RT-PCR: 9211496 (AP2 domain-containing transcription factor), Contig100 (fructose-bisphosphate aldolase), 9252418 (E3 ubiquitin protein ligase), Contig202 (cytochrome P450-like TBP protein), Contig50 (ribosomal protein L24-like protein), Contig211 (NADP-dependent malic enzyme), 9252382 (auxin response factor), and Contig225 (acyl-CoA oxidase). Total RNA was subjected to DNase I-digestion (TaKaRa, Japan) to remove any contaminating genomic DNA. First-strand cDNA was synthesized from 2 μg of total RNA per sample with AMV reverse transcriptase (TaKaRa, Japan), according to the supplier’s instruction manual. The rubber tree 18s rRNA gene was used as the inner control for RT-PCR analysis. All primers for the candidate genes and 18s are shown in Table 1. PCR was conducted in a total volume of 25 μL that contained 0.2 μM of each primer plus 1X SYBR Green PCR Master Mix (Takara). Reactions were amplified in a Light-Cycler 2.0 System (Roche Diagnostics, Germany) as follows: 95 °C for 30 s; then 45 cycles of 94 °C/5 s, 60 °C/20 s, and 72 °C/20 s. The 2−△△ct method [22] was used for calculating relative changes in expression.
Results
RNA isolation
The total RNA from rubber tree roots growing under P-deficient or -sufficient conditions produced two strong 28S and 18S bands (Fig. 1). Their OD260/OD280 values ranged from 1.9 to 2.0, indicating that the RNA was of good quality for library construction.
Detection by SSH
Differentially expressed cDNA was enriched via first- and second-round PCR, with primers outside and inside of the adaptor 1 and adaptor 2R sequences. Results are depicted in Fig. 2. The product of the first round had a band range of 100–1,000 bp; the product of the second round was enriched from 300 to 500 bp.
Quality of the subtractive library
The purified second-round PCR products of the subtractive libraries were ligated into the pMD18-T vector. In all, 731 and 216 clones were obtained from the forward and reverse libraries, respectively. Some clones were then randomly selected to detect their insert size by PCR, which revealed lengths of 200–800 bp. More than 90 % of the clones carried a single exogenous fragment (Fig. 3). This demonstrated that the libraries were of high quality.
Sequence analysis and annotation
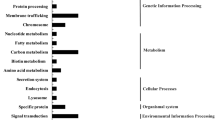
Sequence analysis indicated that 94 unigenes were obtained from the forward library and 45 from the reverse library. These were classified into functional groups according to MIPS. All of the differentially expressed genes were then grouped according to function (Fig. 4). The majority were involved in metabolism (11 %), transcription (5 %), signal transduction (5 %), protein synthesis (22 %), transport (5 %), stress responses (18 %), photosynthesis (13 %), and development (18 %).
Differentially expressed genes are presented in Tables 2 and 3. In the forward subtractive library (from roots under a P-deficiency), Seven of the more important genes that were up-regulated included 9211496 (AP2 domain-containing transcription factor), Contig100 (fructose-bisphosphate aldolase), 9252418 (E3 ubiquitin protein ligase), Contig202 (cytochrome P450-like TBP protein), Contig211 (NADP-malic enzyme), 9252382 (auxin response factor), and Contig225 (acyl-CoA oxidase). Another four clones for ribosomal proteins were also up-regulated under deficient conditions—9211568 (Arabidopsis thaliana 60S ribosomal protein L18A), 9211779 (Zea mays 60S ribosomal protein L18a), Contig33 (30S ribosomal protein S1), and Contig50 (Solanum tuberosum ribosomal protein L24-like protein).
Real-time RT-PCR
As shown in Fig. 5, the transcript level of 9211496 was increased significantly under the P-deficiency, and reached a peak on day 2. The maximum transcript level for Contig100 was 2.6-fold higher than the control after 2 days of treatment. The expression pattern was similar to that of 9211496. Transcript levels for 9252418 and Contig202 were at a maximum on day 2, with a second, small peak on day 8. Contig50 and Contig211 transcripts were highest after 4 d of treatment. Those of 9252382 and Contig225 reached a maximum level on day 6 and day 8, respectively. Based on their expression patterns, transcripts for these eight selected clones were significantly up-regulated in the roots under a P-deficiency.
Real-time quantitative RT-PCR analysis for randomly selected clones. a 9211496, AP2 domain-containing transcription factor; b Contig100, fructose-bisphosphate aldolase; c 9252418, E3 ubiquitin protein ligase; d Contig202, cytochrome P450-like TBP protein; e Contig50, ribosomal protein L24-like protein; f Contig211, NADP-malic enzyme; g 9252382, auxin response factor. h Contig225, acyl-CoA oxidase
Discussion
The molecular mechanism by which rubber trees respond to a phosphorus deficiency in the soil is a complex network of numerous processes. We used SSH techniques to identify 94 and 45 genes that were up- and down-regulated, respectively, in roots exposed to such a deficiency. The three major functional groups were metabolism, protein synthesis, and stress responses. Similarly, experiments with Arabidopsis have shown that, when P levels are reduced, genes associated with metabolism are up-regulated [23]. This has also been reported by Guo et al. [24] in soybean. Misson et al. [6] have demonstrated that a P-deficiency induces changes in the expression of Arabidopsis genes involved in various metabolic pathways, ion transport, signal transduction, transcriptional regulation, and other processes related to growth and development. Based on those earlier studies and our current results, we might conclude that the rubber tree has a similar adaptive mechanism for responding to a deficient supply of P.
For RT-PCR analysis, we selected eight clones that were up-regulated by P-deficient treatment. AP2 transcription factors play a crucial role in the transcriptional regulation of a variety of biological processes related to development and in response to biotic and abiotic stress conditions. Kitomi et al. [25] have shown that CRL5(an AP2/ERF transcription factor) promotes the development of crown roots in rice, being expressed in the stem region where this occurs. Kim et al. [26] have reported that the AP2/ERF transcription factor RAP2.11 modulates the plant response to low-potassium conditions. In this study, expression of 9211496 (an AP2 transcription factor) was up-regulated under a P-deficiency. Because AP2 transcription factor is a member of a transcription factor family has been implicated in P responses, our observation here suggests that this member plays a similarly important role in the reaction by rubber trees. As encoded by Contig100, fructose-1,6-bisphosphate aldolase (FBA) is a key metabolic enzyme that catalyzes the cleavage of β-fructose-1,6-phosphate to D-glyceraldehyde-3-phosphate and dihydroxyacetone phosphate. Overexpression of FBA enhances plant growth and biomass yields by increasing the rate of photosynthesis [27]. This gene is also up-regulated under high salinity and drought [28], thus indicating its role in responding to abiotic stress. Therefore, we conclude that Contig100 is involved in the response by rubber tree roots to reduced phosphorus.
Ubiquitination plays crucial roles in plant development and in response to biotic and abiotic stresses. Hur et al. [29] have noted that expression of OsUPS (E3 ligase) in rice is up-regulated in both plants and cell cultures in the absence of P. Here, expression of 9252418 (E3 ubiquitin protein ligase) was induced under a P-deficiency in the roots. Therefore, we believe it has an important role in response to P-deficiency through ubiquitin system. Cytochrome P450-like TBP is active in oxidative metabolism and ROS production. The family member CYP704B2 catalyzes the ω-hydroxylation of fatty acids and is required for anther cutin biosynthesis and pollen exine formation in rice [30]. Cytochrome P450-like TBP is also reportedly involved in biotic and abiotic environmental responses [31]. Here, expression of Contig202 (for cytochrome P450-like TBP protein) was induced under a P-deficiency. We suggest that it could contribute to protein activity during cell metabolism and growth.
Ribosomal proteins are essential for plant growth and development. In soybean, the expression of three ribosomal protein genes can begin to increase 3 days after low-temperature treatment is applied[32]. We found that Contig50 (ribosomal protein L24-like protein) was up-regulated in P-deficienct roots, meaning that the translation machinery is possibly involved in rubber tree in this stress response in the rubber tree. NADP-malic enzyme (NADP-ME) is widely distributed, functioning in many different pathways. Two such genes play an important role in the response by wheat to stresses associated with abscisic or salicylic acids, low temperatures, high salt, darkness, or drought [33]. In maize, ZmCytNADP-ME is specifically expressed in embryos and emerging roots [34]. Overexpression of rice NADP-ME genes in Arabidopsis can increase root lengths and enhance plant tolerance to salt stress [35]. Of all the vegetative and reproductive tissues, the roots exhibit the highest activity by NADP-ME [36]. Here, Contig211 (for NADP-dependent malic enzyme) was up-regulated in rubber tree roots by our deficiency treatment, possibly improving their growth in response to the reduced availability of P.
Auxin signaling is important for growth and development processes such as apical dominance, lateral root formation, vascular differentiation, embryo patterning, and shoot elongation [37]. Auxin response factors (ARFs) are critical regulators. For example, the OsARF12 knockout mutant in rice has shorter primary roots than do wild-type plants [38]. Optimizing the architecture of a root system can overcome limitations on crop yields caused by water or nutrient shortages [39]. We found that 9252382 (for ARF) was up-regulated when P was lacking. This clone may mediate the developmental processes of rubber tree roots under such stress. Acyl-CoA oxidase (ACX) is a core enzyme involved in peroxisomal β-oxidation. Acyl-coA oxidases are highly expressed in mature and germinating pollen, stem epidermal cells, and other tissues in which jasmonate-signaled processes occur [40]. Rice acyl-CoA oxidase isoenzyme(OsACX2) is expressed predominantly in the seeds [41]. Those reports suggest that OsACX2 helps to provide germinating seeds with sugar and energy. In this study, expression of Contig225, (acyl-CoA oxidase) was induced in rubber tree roots under P-deficiency, implying that it is involved in enhancing the supply of sugar and energy when P is lacking.
In summary, the identifications made here of genes differentially expressed in rubber tree roots provide valuable information for future studies of the molecular mechanism by which that species adapts to a P-deficiency. Moreover, the utilization of phosphorus might be improved as a result of through further characterization and functional analysis of those genes.
References
Marschner H (1995) Mineral nutrition of higher plants, Ed 2. Academic Press/Harcourt Brace, London
Raghothama KG (1999) Phosphate acquisition. Annu Rev Plant Physiol Plant Mol Biol 50:665–693
Vance CP, Uhde-Stone C, Allan DL (2003) Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytol 157:423–447
Poirier Y, Bucher M (2002) Phosphate transport and homeostasis in Arabidopsis. In: Somerville CR, Meyerowitz EM (eds) The Arabidopsis book. American Society of Plant Biologists, Rockville MD
Rouached H, Arpat AB, Poirier Y (2010) Regulation of phosphate starvation responses in plants: signaling players and cross-talks. Mol Plant 3(2):288–299
Misson J, Raghothama KG, Jain A, Jouhet J, Block MA, Bligny R et al (2005) A genome-wide transcriptional analysis using Arabidopsis thaliana Affymetrix gene chips determined plant responses to phosphate deprivation. Proc Natl Acad Sci USA 102(33):11934–11939
Hammond JP, Broadley MR, Bowen HC, Spracklen WP, Hayden RM, White PJ (2011) Gene expression changes in phosphorus deficient potato (Solanum tuberosum L.) leaves and the potential for diagnostic gene expression markers. PLoS ONE 6(9):e24606
Li L, Liu C, Lian X (2010) Gene expression profiles in rice roots under low phosphorus stress. Plant Mol Biol 72(4–5):423–432
Ai P, Sun S, Zhao J, Fan X, Xin W, Guo Q, Yu L, Shen Q, Wu P, Miller AJ et al (2009) Two rice phosphate transporters, OsPht1;2 and OsPht1;6, have different functions and kinetic properties in uptake and translocation. Plant J 57:798–809
Jia H, Ren H, Gu M, Zhao J, Sun S, Chen J, Wu P, Xu G (2011) Phosphate transporter gene, OsPht1;8, is involved in phosphate homeostasis in rice. Plant Physiol 156:1164–1175
Nagarajan VK, Jain A, Poling MD, Lewis AJ, Raghothama KG, Smith AP (2011) Arabidopsis Pht1;5 mobilizes phosphate between source and sink organs and influences the interaction between phosphate homeostasis and ethylene signaling. Plant Physiol 156(3):1149–1163
Veljanovski V, Vanderbeld B, Knowles VL, Snedden WA, Plaxton WC (2006) Biochemical and molecular characterization of AtPAP26, a vacuolar purple acid phosphatase up-regulated in phosphate-deprived Arabidopsis suspension cells and seedlings. Plant Physiol 142(3):1282–1293
Liang C, Tian J, Lam HM, Lim BL, Yan X, Liao H (2010) Biochemical and molecular characterization of PvPAP3, a novel purple acid phosphatase isolated from common bean enhancing extracellular ATP utilization. Plant Physiol 152(2):854–865
Hur YJ, Jin BR, Nam J, Chung YS, Lee JH, Choi HK, Yun DJ, Yi G, Kim YH, Kim DH (2010) Molecular characterization of OsPAP2: transgenic expression of a purple acid phosphatase up-regulated in phosphate-deprived rice suspension cells. Biotechnol Lett 32(1):163–170
Rubio V, Linhares F, Solano R, Martin AC, Iglesias J, Leyva A, Paz-Ares J (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Gonzalez E, Solano R, Rubio V, Leyva A, Paz-Ares J (2005) PHOSPHATE TRANSPORTER TRAFFIC FACILITATOR1 is a plant-specific SEC12-related protein that enables the endoplasmic reticulum exit of a high affinity phosphate transporter in Arabidopsis. Plant Cell 17:3500–3512
Yi K, Wu Z, Zhou J, Du L, Guo L, Wu Y, Wu P (2005) OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol 138(4):2087–2096
Wang C, Ying S, Huang H, Li K, Wu P, Shou H (2009) Involvement of OsSPX1 in phosphate homeostasis in rice. Plant J 57:895–904
Chen J, Liu Y, Ni J, Wang Y, Bai Y, Shi J, Gan J, Wu Z, Wu P (2011) OsPHF1 regulates the plasma membrane localization of low- and high-affinity inorganic phosphate transporters and determines inorganic phosphate uptake and translocation in rice. Plant Physiol 157(1):269–278
Zhou J, Jiao F, Wu Z, Li Y, Wang X, He X, Zhong W, Wu P (2008) OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol 146(4):1673–1686
Mewes HW, Frishman D, Guldener U, Mannhaupt G, Mayer K et al (2002) MIPS: a database for genomes and protein sequences. Nucleic Acids Res 30:31–34
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−△△ct method. Methods 25:402–408
Morcuende R, Bari R, Gibon Y, Zheng W, Pant BD, Bläsing O, Usadel B et al (2007) Genome-wide reprogramming of metabolism and regulatory networks of Arabidopsis in response to phosphorus. Plant Cell Environ 30(1):85–112
Guo W, Zhang L, Zhao J, Liao H, Zhuang C, Yan XL (2008) Identification of temporally and spatially phosphate-starvation responsive genes in Glycine max. Plant Sci 175:574–584
Kitomi Y, Ito H, Hobo T et al (2011) The auxin responsive AP2/ERF transcription factor CROWN ROOTLESS5 is involved in crown root initiation in rice through the induction of OsRR1, a type-A response regulator of cytokinin signaling. Plant J 67:472–484
Kim MJ, Ruzicka D, Shin R, Schachtman DP (2012) The Arabidopsis AP2/ERF transcription factor RAP2.11 modulates plant response to low-potassium conditions. Mol Plant. doi:10.1093/mp/sss003
Uematsu K, Suzuki N, Iwamae T, Inui M, Yukawa H (2012) Increased fructose 1,6-bisphosphate aldolase in plastids enhances growth and photosynthesis of tobacco plants. J Exp Bot 63(8):3001–3009
Fan W, Zhang Z, Zhang Y (2009) Cloning and molecular characterization of fructose-1,6-bisphosphate aldolase gene regulated by high-salinity and drought in Sesuvium portulacastrum. Plant Cell Rep 28:975–984
Hur YJ, Yi YB, Lee JH et al (2012) Molecular cloning and characterization of OsUPS, a U-box containing E3 ligase gene that respond to phosphate starvation in rice (Oryza sativa). Mol Biol Rep 39(5):5883–5888
Li H, Pinot F, Sauveplane V, Werck-Reichhart D, Diehl P, Schreiber L et al (2010) Cytochrome P450 family member CYP704B2 catalyzes the {omega}-hydroxylation of fatty acids and is required for anther cutin biosynthesis and pollen exine formation in rice. Plant Cell 22(1):173–190
Harvey PJ, Campanella BF, Castro PML, Harms H, Lichtfouse E et al (2002) Phytoremediation of polyaromatic hydrocarbons, anilines and phenols. Environ Sci Pollut Res Int 9:29–47
Kim KY, Park SW, Chung YS et al (2004) Molecular cloning of low-temperature-inducible ribosomal proteins from soybean. J Exp Bot 55(399):1153–1155
Fu ZY, Zhang ZB, Hu XJ, Shao HB, Ping X (2009) Cloning, identification, expression analysis and phylogenetic relevance of two NADP-dependent malic enzyme genes from hexaploid wheat. C R Biol 332(7):591–602
Detarsio E, Maurino VG, Alvarez CE, Müller GL, Andreo CS, Drincovich MF (2008) Maize cytosolic NADP-malic enzyme (ZmCytNADP-ME): a phylogenetically distant isoform specifically expressed in embryo and emerging roots. Plant Mol Biol 68(4–5):355–367
Liu S, Cheng Y, Zhang X, Guan Q, Nishiuchi S, Hase K, Takano T (2007) Expression of an NADP-malic enzyme gene in rice (Oryza sativa L) is induced by environmental stresses; over-expression of the gene in Arabidopsis confers salt and osmotic stress tolerance. Plant Mol Biol 64(1–2):49–58
Müller GL, Drincovich MF, Andreo CS, Lara MV (2008) Nicotiana tabacum NADP-malic enzyme: cloning, characterization and analysis of biological role. Plant Cell Physiol 49(3):469–480
Davies PJ (1995) The plant hormones: their nature, occurrence and functions. In: Davies PJ (ed) Plant hormones: physiology, biochemistry and molecular biology. Kluwer Academic Publishers, Dordrecht
Qi Y, Wang S, Shen C, Zhang S, Chen Y, Xu Y, Liu Y, Wu Y, Jiang D (2012) OsARF12, a transcription activator on auxin response gene, regulates root elongation and affects iron accumulation in rice (Oryza sativa). New Phytol 193(1):109–120
Werner T, Nehnevajova E, Köllmer I, Novák O, Strnad M et al (2010) Root-specific reduction of cytokinin causes enhanced root growth, drought tolerance, and leaf mineral enrichment in Arabidopsis and tobacco. Plant Cell 22:3905–3920
Schilmiller AL, Koo AJ, Howe GA (2007) Functional diversification of acyl-coenzyme A oxidases in jasmonic acid biosynthesis and action. Plant Physiol 143(2):812–824
Kim MC, Kim TH, Park JH et al (2007) Expression of rice acyl-CoA oxidase isoenzymes in response to wounding. J Plant Physiol 164(5):665–668
Acknowledgments
This research was supported by grants from the Fundamental Research Funds for Rubber Research Institute, CATAS (1630022011017, YWFZX2010-10(N)). We thank Dr. Yuwei Hua for providing the seedlings of Reyan7-33-97.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
He, P., Qin, H., Wu, M. et al. Identification of genes differentially expressed in the roots of rubber tree (Hevea brasiliensis Muell. Arg.) in response to phosphorus deficiency. Mol Biol Rep 40, 1397–1405 (2013). https://doi.org/10.1007/s11033-012-2183-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2183-x