Abstract
Toll-like receptors (TLRs) are one of the key components of innate or non-specific immunity. Among various types of TLRs, TLR3 recognizes dsRNA, the genetic material or replicative intermediate of many RNA viruses and triggers TIR-domain-containing adapter-inducing interferon-β dependent signalling pathway to induce type I interferon (IFN) and pro-inflammatory cytokines. In this study, we cloned and characterized full-length TLR3 cDNA in rohu (Labeo rohita), that comprised of 2,619 bp nucleotides encoding a putative protein of 873 amino acid with the estimated molecular mass of 98.57 kDa. The constitutive expression of TLR3 gene was detected in all embryonic developmental stages and in various organs/tissues of rohu fingerlings. In vivo tissue specific modulation of TLR3, type I IFN, Mx (myxovirus-resistant protein) and pro-inflammatory cytokines (TNF-α and IL-1β) gene expression were analysed by quantitative real-time PCR following intravenous injection of polyinosinic-polycytidylic acid (poly I:C), a synthetic analogue of viral dsRNA. A significant relationship of TLR3 induction, and type I IFN, Mx, IL-1β and TNF-α gene expression were observed in majority of the treated fish tissues, as compared to their control. Together, these data highlight the important role of TLR3 in recognizing dsRNA, and in augmenting the innate immunity in fish in response to viral infections.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Innate or non-specific immunity is an evolutionary ancient part of the defense mechanism in all multi-cellular organisms [1]. It is the first line of defense against infection and is regarded as the primeval and hence the universal form of host defense [2]. A major component of the innate immune system is contributed by the pattern-recognizing receptors (PRRs) that recognize conserved microbial structures like peptidoglycan (PGN), lipoteichoic acid (LTA), zymosan, flagellin, lipopolysaccharides (LPS), lipoproteins, heat shock protein (hsp), CpG-DNA, and nucleic acids of the micro-organisms (pathogen/microbes-associated molecular patterns, PAMPs/MAMPs) [3]. The various types of PRRs, notably toll-like receptors (TLRs), NOD-like receptors (NLRs) and RIG-I like receptors (RLRs) which are mostly expressed on the cell surface, in intracellular compartments, or secreted into the blood stream and tissue fluids recognize different PAMPs and activates their signaling cascades to induce innate immunity [4].
Toll-like receptors (TLRs) are the type I trans-membrane proteins having an N-terminal ectodomain (ECD), a single trans-membrane domain (TM), and a C-terminal cytoplasmic TIR (Toll/IL-1) domain [5, 6]. Depending upon their primary sequence similarity and types of ligands (PAMPs) recognition, TLRs are grouped into six major families viz, TLR1, TLR3, TLR4, TLR5, TLR7 and TLR11 [7]. TLR3 is a member of the TLR family that plays an essential role in inducing the innate immune response against viral infections by recognizing the double-stranded RNA (dsRNA). The dsRNA is the molecular signature in many viruses that are produced either as an intermediate of the viral replication or as part of the viral RNA genome, secreted by the necrotic cells, generated by in vitro transcription, or mimicked by synthetic polyinosinic-polycytidylic acid (poly I:C) [8]. In TLR3, the leucine rich repeat (LRR) region recognizes dsRNA, and upon recognition and binding, the adaptor protein TRIF (TIR-domain-containing adapter-inducing interferon-β) is recruited to the TIR domain in TLR3 [9–11]. In one hand, TRIF interacts with RIP1 (receptor-interacting protein1) and activates NF-κB to induce pro-inflammatory cytokines [12]. On the other hand, TRIF activates TANK (TRAF-family member associated NF-κB activator) binding kinase1 (TBK1 also known as NAK or T2K) through TRAF6. TBK1 activation leads to the phosphorylation of IRF3 and IRF7, enabling the formation of their homodimer, translocation into the nucleus, binding to the respective motifs on the DNA resulting in the induction of type I IFNs (IFN-α/β).
The identification of TLR3 and elucidation of its dsRNA ligand recognition profile and signaling in mammalian species [12, 13] have been followed by identification and characterization of homologs in several teleost species, including zebrafish (Danio rerio) [14–16], fugu (Takifugu rubripes) [10] channel catfish and hybrid catfish (Ictalurus sp.) [17–19], rainbow trout (Oncorhynchus mykiss) [20], rare minnow (Gobiocypris rarus) [21], grass carp (Ctenopharyngodon idella) [22], common carp (Cyprinus carpio) [23], large yellow croaker (Pseudosciaena crocea) [24] and Japanese flounder (Paralichthys olivaceus) [25].
The modulation of TLR3 gene expression was previously been reported in zebrafish [26] following infection with viral haemorrhagic septicemia virus (VHSV), a single stranded RNA (ssRNA) virus, and in grass carp, by the grass carp reovirus (GCRV), a dsRNA virus. The polymorphism and haplotype of TLR3 gene in grass carp was reported to be associated with the susceptibility or resistance to GCRV infection [27]. In rohu and other cyprinid fish, spring viremia of carp (SVC) is a fatal viral disease caused by the spring viremia of carp virus (SVCV), a linear ssRNA virus [28, 29]. The polyinosinic: polycytidylic acid (poly I:C) is a chemically synthesized dsRNA, with one strand having a polymer of inosinic acid and the other a polymer of cytidylic acid. It mimics the structure of viral dsRNA and functions as a ligand of TLR3 [12]. Treatment with poly I:C was shown to induce the anti-viral activity in salmon [30] and protection against VHSV infection in Japanese flounder [31]. These data together, highlighted the important role of TLR3 in fish.
The fish production in India is the third largest in the world [32], and among the freshwater cultured fish species, rohu (Labeo rohita) is the most commercially important and highly favored fish species. Innate immunity contributed by various PRRs play the pivotal role in protecting fish, and TLR3 is one of them. In spite of being the most important fish species in the Indian sub-continent, there is no information on TLR3 in rohu. Therefore, this work was undertaken to clone and characterize TLR3 in rohu, and to postulate its role during viral infections by studying its property following the administration of poly I:C, a synthetic dsRNA which is likely to mimic the dsRNA like structure of GCRV, VHSV and SVCV. Here we report the full-length cDNA sequence of rohu TLR3 (rTLR3), its in vivo mRNA expression kinetics and down-stream signaling cascades in response to poly I:C treatment.
Materials and methods
Fish
Rohu (L. rohita) weighing ~50 g, was obtained from the Central Institute of Freshwater Aquaculture (CIFA), and was stocked in 500 l aerated tanks (water temperature 25–28 °C; pH 7.4–7.6) with each tank containing 50 fish. During acclimatization for 3-weeks, the fishes were fed twice a day with commercial carp diet with daily two-third water exchange.
Cloning of rohu TLR3 (rTLR3)
To clone rTLR3, total RNA was extracted from gill and cDNA was prepared following the protocol as described later. For PCR amplification of TLR3 gene, TLR3-FW and RV primers (Table 1a) were designed based on the nucleotide sequences of the conserved regions of common carp (DQ885910.1), grass carp (DQ864497.1) and zebrafish (AY616582.1) TLR3 gene. PCR was carried out in a 50 μl reaction volume, utilizing 1 μl of the cDNA as a template under the following conditions: one cycle of initial denaturation at 94 °C for 2 min, 45 cycles of 94 °C for 30 s, 50 °C for 30 s, 72 °C for 1 min and a final extension at 72 °C for 5 min. In a 2 % agarose gel, 10 μl of the PCR product was analyzed, and from the single specific band, DNA was purified with agarose gel DNA extraction kit (Roche, Germany). The eluted DNA was cloned in pGEM-T Easy vector (Promega, Madison, USA) and both strand DNA sequencing was carried out with T7 and SP6 primer following Sanger sequencing protocol in 96 capillary high through put sequencer, ABI 3730 XL (Xcelris Labs Ltd, Ahmedabad, India). The obtained DNA sequence was confirmed as partial cDNA of rTLR3 by BLAST search [33] and was submitted to the GenBank (GU248414.1). To obtain the full-length rTLR3 cDNA sequence, 5′ and 3′ RACE (rapid amplification of cDNA ends) were carried out using the SMARTer RACE cDNA amplification kit (Clontech, USA). The rTLR3 gene specific forward (TLR3-GSP-FW-3′) and reverse (TLR3-GSP-RV-5′) primers were designed from the partial rTLR3 cDNA sequence (GU248414.1). Touchdown-PCR was carried out for 3′ and 5′ RACE using TLR3-GSP-FW-3′/UPM and TLR3-GSP-RV-5′/UPM primer sets (Table 1a) respectively under the following conditions: one cycle of initial denaturation at 94 °C for 2 min, followed by five cycles of 94 °C/30 s, 72 °C/3 min; next five cycles 94 °C/30 s, 70 °C/30 s, 72 °C/3 min; next 27 cycles 94 °C/30 s, 68 °C/30 s, 72 °C/3 min; and one cycle at 72 °C/5 min. Nested PCR for 3′ and 5′ RACE were performed using TLR3-NP-FW-3′/NUP for 3′ and TLR3-NP-RV-5′/NUP for 5′ nested primer sets (Table 1a). For nested PCR, 1 μl of primary RACE-PCR product was used as a template with the following conditions: initial one cycle of 94 °C/2 min, then 30 cycle of 94 °C/30 s, 68 °C/30 s and 72 °C/3 min followed by one cycle of 72 °C/5 min. The PCR products were cloned in pGEM-T Easy vector, sequenced and validated through BLAST search and overlapping sequences from the fragments were combined together to get the full-length cDNA of rTLR3 and was submitted to the GenBank.
Analysis of rTLR3 cDNA sequence
Various structural features such as LRR, trans-membrane and TIR domains in rTLR3 were predicted using SMART (http://smart.embl-heidelberg.de). The signal peptide was predicted by SMART and SignalP 3.0 server [34], and the percentages of similarity and identity of rTLR3 amino acids with other species were calculated by MatGAT program [35]. The phylogenetic tree of TLR3 among various species was constructed by the neighbor-joining method of MEGA4 program [36], and the branches were validated by bootstrap analysis from 1,000 replications using default parameter.
In vivo expression of rTLR3 gene
To investigate TLR3 gene expression during various developmental stages of rohu, fertilized eggs were collected from CIFA hatchery, and were maintained at ambient temperature (28–29 °C) for embryogenesis. Periodical observation was carried out under the inverted microscope to identify various developmental stages from fertilized eggs to hatchlings. At 0 h (fertilized egg), 4, 8, 12, 16, and hatchings at 24 and 48 h, samples were collected separately in TRIzol reagent. Total RNA was extracted and cDNA was prepared using oligo-dT primer following the protocol as described later. Quantitative real-time PCR (qRT-PCR) was carried out to analyze TLR3 and β-actin (internal control) gene expressions among the samples.
To examine basal TLR3 expression, gill, liver, kidney, spleen, intestine, skin, heart, brain, blood, muscle and eye were collected separately from three rohu fingerlings (~50 g), total RNA was isolated from each sample and cDNA was prepared following the protocol as described later. The qRT-PCR was carried out to analyze rTLR3 expression, keeping β-actin as an internal control and the mean value was graphically presented.
Poly I:C injection into the rohu fingerlings
Healthy rohu fingerlings (~50 g) were divided into control and treated groups keeping three fish in each group. Purified poly I:C (Sigma, USA) was diluted in DEPC treated water at 10 mg/ml, and 100 μl of DEPC-treated water containing 300 μg of diluted poly I:C was injected through intravenous (i.v.) route into three fish (the treated fish group). The control fish group (three fish) was i.v. injected with 100 μl of DEPC-treated water. After 4 h, control and treated group of fish were sacrificed and tissues were collected separately in TRIzol reagent, for RNA extraction.
RNA isolation and 1st strand cDNA synthesis
From the TRIzol treated samples, viz, gill, liver, kidney, spleen, intestine, skin, heart, brain, blood, muscle and eye, total RNA was extracted following the standard protocol (Invitrogen, USA). The concentration of the total RNA was measured by UV-spectrophotometer (Eppendorf, India) and the integrity of the RNA was assessed by observing the band intensity of 28 and 18S ribosomal RNA on 1 % agarose gel. To prepare 1st strand cDNA, 1 μg of total RNA was treated with 1 U of DNase I (MBI, Fermentas, USA) and reverse transcription was carried out using oligo-dT primer and RevertAid 1st strand cDNA synthesis kit (MBI, Fermentas, USA). The confirmation of cDNA synthesis was carried out with PCR-amplification of β-actin gene and the synthesized cDNA was kept at −80 °C till further analysis.
Real-time PCR analysis
Quantitative real-time RT-PCR (qRT-PCR) of the target genes: TLR3, TNF-α, IL-1β, IFN-1, Mx and the reference gene β-actin were carried out in 10 μl reaction volume, containing 1.0 μl of cDNA, 0.25 μl of FW and RV primers (2.5 μM each; Table 1b), 5 μl of 2X lightCycler®480 SYBR Green I master mix (Roche, Germany) and 3.5 μl of PCR grade H2O in LightCycler®480 II-real time PCR detection system (Roche, Germany). The qRT-PCR was performed in triplicate wells under following conditions: initial denaturation at 95 °C for 10 min followed by 45 cycles of 94 °C/10 s, 58 °C (for TLR3, IL-1β and β-actin) or 51 °C (for TNF-α) or 62 °C (for Mx and IFN-1) for 10 s and 72 °C/10 s. Negative control reaction was carried out without the template (cDNA). The PCR efficiencies were determined by analyzing the serial dilutions of cDNA, and efficiencies were almost 100 % allowing the use of 2−ΔΔCT method [37] for calculation of relative gene expression of the target genes TLR3, TNF-α, IL-1β, IFN-1, Mx with that of reference gene, β-actin. The correct size and single band amplification was determined by analyzing 8 μl of the real time PCR products in ethidium bromide-stained 2 % agarose gel. The relative expression ratios were obtained by normalizing expression of the target gene, as determined by mean crossing point (cp) deviation by that of a non-regulated reference gene, β-actin following 2−ΔΔCT method. The data obtained from qRT-PCR analysis was expressed as mean of three individual experiments ± standard error (s.e.), and the significant difference between the control and treated groups at each time point was determined by the Student’s t test using Microsoft Excel 2010 with p < 0.05 as significance level.
Results
Cloning and characterization of TLR3 in rohu
The cDNA sequence encoding the open reading frame (ORF) of rTLR3 was derived from the gill and was submitted to the GenBank (JN886779). The ORF consisted of 2,619 bp nucleotides that encoded a putative 873 amino acid (aa) residue with an estimated molecular mass of 98.57 kDa. The secondary structure prediction by SMART revealed the presence of signal peptide (1–22 aa), LRR regions (53–698 aa), trans-membrane (TM) domain (707–729 aa) and the TIR domain (757–873 aa). The domain organization in rTLR3, in other fish species and human were shown in Fig. 1.
Schematic representation of TLR3 domains in various fish species and human. In TLR3, secondary structures were predicted by SMART and TMHMM programs. Signal peptide (small solid box), leucine-rich repeat (LRR) domains (rectangular box), C-terminal LRR domain (oval shaded box), transmembrane domain (elongated box) and the TIR domain (rectangular shaded box) were shown with their respective positions
Phylogenetic relationship of rTLR3 with other species
The percentages of similarity and identity of rTLR3 amino acids with other species were calculated in MatGAT, and it showed highest similarity (90.6 %) and identity (83.5 %) with common carp. With other fish species, the similarity and identity % of rTLR3 amino acids were ~70–80 and ~50–80 respectively, and as expected less identity (~40 %) and similarity (~60 %) were observed with higher vertebrates (Table 2).
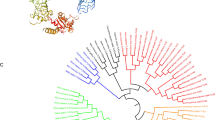
To understand the evolutionary relationship, phylogenetic tree was constructed with full-length TLR3 protein of various species, and it revealed two major clusters, supported by high bootstrapping values. Cluster-I represented various fish species, and cluster-II denoted the TLR3 of higher vertebrates (Fig. 2). Among various fish species, TLR3 in the cyprinidae family (D. rerio, L. rohita, M. amblycephala, G.rarus, C.carpio. C.auratus and C.idella) formed a separate group, and within the cyprinidae family, TLR3 of gold fish and grass carp were most closely related with 99.7 % aa similarity and identity.
Phylogenetic relationship of rTLR3 with other species. TLR3 amino acid sequences were aligned by CLUSTALW program within DNASTAR, and the unrooted phylogenetic tree was generated by the neighbor-joining method within the MEGA4 program. The tree was boosted 1,000 times and the percentages are shown. The full-length TLR3 amino acids sequences used for tree construction, were retrieved from following database: gorilla, BAG55031.1; human, ABC86908.1; gibbon ACC68077.1; monkey ABY64988.1; common marmoset, ACC68086.1; elephant, ABC95781.1, cattle ABN71665.1; buffalo, ADY18594.1; goat, ADZ17150.1; chinese hamster, XP_003514272.1; mouse, AAH99937.1; rat, NP_942086.1; zebra finch, XP_002190888.1; chicken, ABL74502.1; nile tilapia XP_003449776.1; orange-spotted grouper; ADZ76423.1; fugu, AAW69373.1; rainbow trout, AAX68425.1; catfish, AEI59664.1; zebrafish, AAT37633.1; Gobiocypris rarus, ABL11471.1; wuchang bream, ABI83673.1; common carp, ABL11473.1; goldfish, ABC86865.1, grass carp, ABI64155.1
Ontogenic expression of rTLR3
The ontogenic expression of TLR3 in rohu was analyzed by quantitative real-time PCR (qRT-PCR) assay. The results showed that TLR3 gene expression was higher in the fertilized eggs (0 h) as compared to other embryonic developmental stages. At 8–12 h post fertilization, TLR3 expression reached the lowest level, and thereafter it steadily increased in the hatchlings at 16 and 24 h and in the larvae at 48 h (Fig. 3).
TLR3 gene expression in various embryonic developmental stages of rohu. Total RNA was extracted from different stages of development (denoted by hours) and quantitative real-time PCR (qRT-PCR) was carried out to analyze the expression of TLR3 gene. Expressions of TLR3 gene were represented as a ratio relative to β-actin (internal control) levels in the same samples. Gene expression at 0 h was chosen as calibrator (1) and the relative expression of TLR3 at different hours of development was represented as fold changes from the calibrator. The results were expressed as mean ± standard error (bars in the graph) from three samples (n = 3)
Tissue specific expression of rTLR3
Basal expression of TLR3 in various organs/tissues of three rohu fingerlings were analyzed by qRT-PCR and their mean value were presented graphically in Fig. 4. Among the examined tissues, TLR3 expression was at detectable levels in all organs/tissues but the level of expression varied among them. The lowest TLR3 expression was observed in the brain (calibrator tissue; one fold), and as compared to it, the highest expression was detected in the spleen (three fold), followed by eye (~2.7-fold) and intestine (~2.3-fold). In blood, skin, muscle, heart and kidney, basal expression of TLR3 was ~1.7-fold, and in gill and liver the expression was equivalent to the brain.
Basal expression of TLR3 in various tissues Total RNA was extracted from spleen, eye, intestine, blood, skin, muscle, heart, kidney, gill, liver and brain and were analysed by quantitative real-time PCR (qRT-PCR) to examine the expression of TLR3 genes among the tissues. Expression of TLR3 transcripts were represented as a ratio relative to β-actin (internal control) levels in the same samples. Brain was chosen as calibrator (1) and the relative expression of TLR3 in other tissues was represented as fold changes from the calibrator. The results were expressed as mean ± standard error (bars in the graph) from three fish (n = 3)
Inductive expression of TLR3
In vivo modulation of TLR3 gene transcripts were analyzed by qRT-PCR following i.v. injection of poly I:C in rohu fingerlings. As shown in Fig. 5, all tested tissues except skin, significant (p < 0.05) induction of TLR3 gene expression was observed as compared to their control. Among the tissues, highest induction of TLR3 was observed in the liver (~250-fold), followed by the intestine (~50-fold). As compared to the control, TLR3 expression was ~3.5-fold up-regulated in the poly I:C treated fish kidney, heart and blood and ~2.5-fold in the gill.
Modulation of TLR3 expressions by poly I:C. In rohu fingerlings, poly I:C (300 μg/fish) was i.v. injected, and after 4 h, tissues were collected and were used for total RNA extraction and real-time PCR was analysed. The relative expression of TLR3 gene was normalized to the expression of β-actin (internal control) and expressed as fold changes relative to the un-treated control group. The mean value of three fish (n = 3) was shown and the bars indicated standard error. Significant difference (p < 0.05) between control and treated fish group was indicated with asterisks
Modulation of type I IFN expression
In TLR3-signaling pathway, type I IFNs were expected to be induced by dsRNA (poly I:C). To investigate it, we analyzed the gene transcripts of type I IFN by qRT-PCR assay, in the gill, liver, kidney, intestine, heart, spleen, skin and blood of the poly I:C treated fish, and compared them with the control fish (Fig. 6a). At 4 h post poly I:C treatment, except in the intestine and blood, inductive expression of type I IFN gene transcripts was observed in all other tested tissues with varied intensity. Liver was most responsive to induce type I IFN (~4.5-fold, p < 0.05), followed by skin, spleen and kidney (~2.0-fold) and the least induction was observed in the gill (~1.3-fold). However, there was down-regulation of type I IFN in the intestine and blood of the treated fish as compared to the control fish.
Modulation of IFN and Mx expressions by poly I:C. In rohu fingerlings, poly I:C (300 μg/fish) was i.v. injected, and after 4 h, various tissues were collected and were used for total RNA extraction and real-time PCR analysis. The relative expression of IFN and Mx gene was normalized to the expression of β-actin (internal control) and was expressed as fold changes relative to the un-treated control group. The mean value of three fish (n = 3) was shown and the bars indicated standard error. Significant difference (p < 0.05) between control and treated fish group was indicated with asterisks. a Type-I IFN. b Mx gene expression
Modulation of Mx gene expression
Mx is the type I IFN induced protein, and is expected to be induced by poly I:C through the activation of TLR3-signaling pathway. To explore this possibility, we analyzed poly I:C treated samples for Mx gene expression by qRT-PCR assay. As shown in Fig. 6b, at 4 h post poly I:C treatment, a highly significant (p < 0.05) induction of Mx gene expression was observed in all tested tissues (gill, liver, kidney, intestine, heart, spleen, skin and blood) as compared to control fish. Among the tested tissues, liver showed highest induction of Mx gene (~600-fold), followed by kidney (~200-fold), spleen (~200-fold), heart (~150-fold), gill (~50-fold), skin (~50-fold) and blood (~40-fold). The intestine showed lowest but significant induction of Mx gene (~5-fold, p < 0.05) as compared to its control.
Inductive expression of pro-inflammatory cytokines, TNF-α and IL-1β
In higher eukaryotes including human, the activation of TLR3-signaling by dsRNA resulted in the recruitment of the adaptor TRIF to the TIR domain. Following binding to the TLR3-TIR domain, TRIF interacts with RIP1 to activate NF-κB resulting in the induction of pro-inflammatory cytokines. We investigated in vivo induction of pro-inflammatory cytokines viz, TNF-α and IL-1β in rohu fingerlings by qRT-PCR assay following poly I:C treatment. As shown in Fig. 7a, poly I:C treatment induced TNF-α in majority of the tested tissues as compared to their control. The highest induction of TNF-α (~50-fold) was observed in blood followed by spleen (~3.5-fold), kidney (~2.5-fold), liver (~2.3-fold) and skin (~2-fold). However, in gill and intestine, significant down-regulation of TNF-α was observed as compared to their control.
Modulation of TNF-α and IL-1β expressions by poly I:C. In rohu fingerlings, poly I:C (300 μg/fish) was i.v. injected, and after 4 h, various tissues were collected and were used for total RNA extraction and real-time PCR analysis. The relative expression of TNF-α and IL-1β gene was normalized to the expression of β-actin (internal control) and was expressed as fold changes relative to the un-treated control group. The mean value of three fish (n = 3) was shown and the bars indicated standard error. Significant difference (p < 0.05) between control and treated fish group was indicated with asterisks. a TNF-α. b IL-1β gene expression
We next analyzed IL-1β gene transcripts in the same samples, and we noted the significant (p < 0.05) induction of this gene in most of the tissues (Fig. 7b). Among the treated tissues, skin showed highest induction (~7-fold) of IL-1β followed by liver (~5-fold), as compared to their respective control tissues. Intestine, heart and spleen revealed almost equal induction (~2.5-fold) of IL-1β. In contrast to the other tissues, IL-1β expression in gill and in blood was significantly down-regulated.
Discussion
This study describes the identification, cloning and characterization of TLR3 in rohu (L. rohita), the most important freshwater cultured fish species in the Indian subcontinent. In rohu, the putative TLR3 protein comprised of 17 LRR domains, and are expected to be involved in recognizing various PAMPs. The spans of LRR regions in rTLR3 were almost comparable with other fish species and higher vertebrates (Fig. 1) signifying the functionally capability of rTLR3 in responding to the mammalian and other fish TLR3 agonists. In human, localization of TLR3 on the cell surface was previously been reported [38]. In rTLR3, the presence of 22 aa signal peptide and the occurrence of 23 aa transmembrane domain suggested it’s localization on the cell surface and supported the similar finding in common carp TLR3 [23]. The existence of a highly conserved TIR domain at the C-terminal end of rTLR3 revealed its ability to transfer the signal by interacting with the down-stream adaptor molecule, TRIF.
The molecular tree of vertebrate TLR3 demonstrated that TLR3 in fish and higher eukaryotes formed two separate clusters (Fig. 2). As expected, within the fish group (cluster-I), TLR3 in rohu was closely related to the other members in the cyprinidae family, and was distinct from the catfish TLR3. The evolutionary close relations of TLR3 among the cyprinid fish species might predict their functional similarities.
The importance of innate immunity in protecting fish embryo was previously been reported in zebrafish [39], carps [40] and channel catfish [19]. Recently, we reported the ontogenic expression of TLR2 [41] and TLR5 [42] gene in mrigal, and NOD1 [43], NOD2 [44] and TLR2 [45] gene in rohu. In addition to these innate defense factors, we detected the constitutive expression of TLR3 during the embryogenesis of rohu, and this is the first report in the Indian major carp. Among the tested developmental stages, a relatively higher expression of TLR3 was detected in fertilized eggs. The enhanced expressions of TLR3 in the fertilized eggs were likely to be contributed by the brood fish. The embryonic expression of TLR3 suggested that along with other innate immune factors, TLR3 could be involved in protecting fish eggs, hatched embryos and larvae during viral infections.
We noted wide expression of TLR3 in various organs of rohu fingerlings, and among the organs, highest expression of TLR3 was observed in the spleen, followed by eye, intestine and blood. The pattern of tissue specific TLR3 expression in rohu greatly differed from previously reported fish species, viz., fugu [10], large yellow croaker [24], Japanese flounder [25], common carp [23] and catfish [46]. The wide expression of TLR3 in various organs of fish may suggest its key role in the immune surveillance system in various organs of the host.
Several reports in mammals and in higher vertebrates showed that TLR3 recognizes dsRNA through its LRR domains and transmits signal through its TIR domain involving TRIF that resulted in the induction of type I IFN and various cytokines. Structurally, poly I:C mimics viral dsRNA, and induced anti-viral activity in salmon [30] and Japanese flounder [25, 31]. In grass carp, inductive expression of Mx gene was reported following dsRNA containing GCRV infection [27]. In rohu, we injected poly I:C through intravenous (i.v.) and intraperitoneal (i.p.) route, and analyzed TLR3 gene expression at various time course. TLR3 induction was much higher and quicker in i.v. than i.p. (results not shown), indicating the better efficiency of ligand delivery through i.v. route. In response to poly I:C, a significant (p < 0.05) up regulation of TLR3, type I IFN and Mx genes expressions were observed in majority of the tissues. These data in rohu was in agreement with the previously reported fish species. Additionally, a significant correlation of TLR3 induction, and type I IFN and Mx gene expressions were observed in most of the tissues. These results indicated the activation of TLR3 signaling pathway by dsRNA (poly I:C). However, to elucidate the detail mechanism further works employing either TLR3 gene silencing or TLR3 overexpression are required.
In the TLR3 signaling cascades, NF-κB functions as the down-stream signaling molecule. In Japanese flounder, poly I:C induced TLR3 and activated NF-κB regulated gene expression [12]. In rohu, poly I:C also induced TNF-α and IL-1β which were known to be NF-κB regulated gene [47, 48], and are expected to be involved in innate immunity. Together, these data in rohu, along with salmon, Japanese flounder and common carp highlighted the important role of TLR3 in fish to trigger a rapid innate immune response during viral invasion to the host.
In conclusion, we report the cloning of TLR3, its constitutive expression during embryonic developments, and in various organs/tissues in the Indian major carp, rohu. In in vivo studies, the expression of type I IFN, Mx, TNF-α and IL-1β were investigated following poly I:C stimulation, showing that the activation of TLR3 signaling could play an important role in inducing the anti-viral immunity to the host.
References
Janeway C, Medzhitov R (2002) Innate immune recognition. Annu Rev Immunol 20:197–216
Aoki T, Takano T, Santos MD et al (2008) Molecular innate immunity in Teleost fish: Review and future perspectives. In: 5th world fisheries congress, pp 263–276
Strandskog G, Skjæveland I, Ellingsen T et al (2008) Double-stranded RNA- and CpG DNA-induced immune responses in Atlantic salmon: comparison and synergies. Vaccine 26(36):4704–4715
Medzhitov R, Preston-Hurlburt P, Janeway J (1997) A human homologue of the Drosophila toll protein signals activation of adaptive immunity. Nature 388:394–397
Akira S, Takeda K (2004) Toll-like receptor signalling. Nat Rev Immunol 4:499–511
Iwasaki A, Medzhitov R (2004) Toll-like receptor control of the adaptive immune responses. Nat Immunol 5:987–995
Roach JC, Glusman G, Rowen L et al (2005) The evolution of vertebrate toll-like receptors. Proc Natl Acad Sci USA 102:9577–9582
Nishiya T, Kajita E, Miwa S et al (2005) TLR3 and TLR7 are targeted to the same intracellular compartments by distinct regulatory elements. J Biol Chem 280:37107–37117
Hoebe K, Du X, Georgel P et al (2003) Identification of Lps2 as a key transducer of MyD88-independent TIR signalling. Nature 424:743–748
Oshiumi H, Tsujita T, Shida K et al (2003) Prediction of the prototype of the human toll-like receptor gene family from the pufferfish, Fugu rubripes, genome. Immunogenetics 54:791–800
Yamamoto M, Sato S, Hemmi H et al (2003) Role of adaptor TRIF in the MyD88-independent toll-like receptor signaling pathway. Science 301:640–643
Alexopoulou L, Holt AC, Medzhitov R et al (2001) Recognition of double-stranded RNA and activation of NF-κB by toll like receptor 3. Nature 413:732–738
Matsumoto M, Funami K, Oshiumi H et al (2004) Toll-like receptor 3: a link between toll-like receptor, interferon and viruses. Microbiol Immunol 48:147–154
Meijer AH, Gabby Krens SF, Medina Rodriguez IA et al (2004) Expression analysis of the toll-like receptor and TIR domain adaptor families of zebrafish. Mol Immunol 40:773–783
Jault C, Pichon L, Chluba J (2004) Toll-like receptor gene family and TIR-domain adapters in Danio rerio. Mol Immunol 40:759–771
Phelan PE, Mellon MT, Kim CH (2005) Functional characterization of full-length TLR3, IRAK-4, and TRAF6 in zebrafish (Danio rerio). Mol Immunol 42:1057–1071
Bilodeau AL, Waldbieser GC (2005) Activation of TLR3 and TLR5 in channel catfish exposed to virulent Edwardsiella ictaluri. Dev Comp Immunol 29:713–721
Bilodeau AL, Peterson BC, Bosworth BG (2006) Response of toll like receptors, lysozyme, and IGF-I in back-cross hybrid [F1male (blue × channel) × female channel] catfish challenged with virulent Edwardsiella ictaluri. Fish Shellfish Immunol 20:29–39
Peterson BC, Bosworth BG, Bilodeau AL (2005) Differential gene expression of IGF-I, IGF-II, and toll-like receptors 3 and 5 during embryogenesis in hybrid (channel × blue) and channel catfish. Comp Biochem Physiol A Mol Integr Physiol 141:42–47
Rodriguez MF, Wiens GD, Purcell MK et al (2005) Characterization of toll-like receptor 3 gene in rainbow trout (Oncorhynchus mykiss). Immunogenetics 57:510–519
Su J, Zhu Z, Wang Y et al (2008) Toll-like receptor 3 regulates Mx expression in rare minnow Gobiocypris rarus after viral infection. Immunogenetics 60:195–205
Su J, Jang S, Yang C et al (2009) Genomic organization and expression analysis of toll-like receptor 3 in grass carp (Ctenopharyngodon idella). Fish Shellfish Immunol 27:433–439
Yang C, Su J (2010) Molecular identification and expression analysis of toll-like receptor 3 in common carp Cyprinus carpio. J Fish Biol 76(8):1926–1939
Huang XN, Wang ZY, Yao CL (2011) Characterization of toll-like receptor 3 gene in large yellow croaker Pseudosciaena crocea. Fish Shellfish Immunol 31(1):106–998
Hwang SD, Ohtani M, Hikima JI et al (2012) Molecular cloning and characterization of toll-like receptor 3 in Japanese flounder Paralichthys olivaceus. Dev Comp Immunol 37(1):87–96
Novoa B, Romero A, Mulero V et al (2006) Zebrafish (Danio rerio) as a model for the study of vaccination against viral haemorrhagic septicemia virus (VHSV). Vaccine 24:5806–5816
Heng J, Su J, Huang T et al (2011) The polymorphism and haplotype of TLR3 gene in grass carp (Ctenopharyngodon idella) and their associations with susceptibility/resistance to grass carp reovirus. Fish Shellfish Immunol 30:45–50
Zhang NZ, Zhang LF, Jiang YN et al (2009) Molecular analysis of spring viraemia of carp virus in china: a fatal aquatic viral disease that might spread in east Asian. PLoS ONE 4(7):e6337
Khiabanian Asl AH, Azizzadeh M, Bandehpour M et al (2008) The first report of SVC from Indian carp species by PCR and histopathologic methods in Iran. Pak J Biol Sci 11:2675–2678
Eaton WD (1990) Anti-viral activity in four species of salmonids following exposure to poly inosinic:cytidylic acid. Dis Aquat Org 9:193–198
Takami I, kwon SR, Nishizawa T et al (2010) Protection of Japanese flounder Paralichthys olivaceus from viral hemorrhagic septicemia (VHS) by Poly (I:C) immunization. Dis Aquat Org 89:109–115
FAO, NACA (2003) Quarterly aquatic animal disease report (Asia and Pacific region)
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bendtsen JD, Nielsen H, von Heijne G et al (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795
Campanella JJ, Bitincka L, Smalley J (2003) MatGAT: an application that generates similarity/identity matrices using protein or DNA sequences. BMC Bioinf 4:29
Tamura K, Dudley J, Nei M et al (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C (T)) Method. Methods 25:402–408
Matsumoto M, Funami K, Tanabe M et al (2003) Subcellular localization of toll-like receptor 3 in human dendritic cells. J Immunol 171:3154–3162
Kanther M, Rawls JF (2010) Host-microbe interactions in the developing zebrafish. Curr Opin Immunol 22:10–19
Huttenhuis HB, Grou CP, Taverne-Thiele AJ et al (2006) Carp (Cyprinus carpio L.) innate immune factors are present before hatching. Fish Shellfish Immunol 20:586–596
Basu M, Swain B, Sahoo BR et al (2012) Induction of toll-like receptor (TLR) 2, and MyD88-dependent TLR signaling in response to ligands stimulation and bacterial infections in the Indian major carp, mrigal (Cirrhinus mrigala). Mol Biol Rep 39(5):6015–6028
Basu M, Swain B, Maiti NK et al (2012) Inductive expression of toll-like receptor 5 (TLR5) and associated downstream signaling molecules following ligand exposure and bacterial infection in the Indian major carp, mrigal (Cirrhinus mrigala). Fish Shellfish Immunol 32(1):121–131
Swain B, Basu M, Samanta M (2012) Molecular cloning and characterization of nucleotide binding and oligomerization domain-1 (NOD1) receptor in the Indian Major Carp, rohu (Labeo rohita), and analysis of its inductive expression and down-stream signalling molecules following ligands exposure and Gram-negative bacterial infections. Fish Shellfish Immunol 32(5):899–908
Swain B, Basu M, Sahoo BR et al (2012) Molecular characterization of nucleotide binding and oligomerization domain (NOD)-2, analysis of its inductive expression and down-stream signalling following ligands exposure and bacterial infection in rohu (L. rohita). Dev Comp Immunol 36(1):93–103
Samanta M, Swain B, Basu M et al (2012) Molecular characterization of toll-like receptor 2 (TLR2), analysis of its inductive expression and associated down-stream signaling molecules following ligands exposure and bacterial infection in the Indian major carp, rohu (Labeo rohita). Fish Shellfish Immunol 32(3):411–425
Baoprasertkul P, Peatman E, Somridhvei B et al (2006) Toll-like receptor 3 and TICAM genes in catfish: species-specific expression profiles following infection with Edwardsiella ictaluri. Immunogenetics 58(10):817–830
Engelsma MY, Stet RJM, Schipper H, Verburg-van Kemenade BM (2001) Regulation of interleukin 1 beta RNA expression in the common carp Cyprinus carpio L. Dev Comp Immunol 25(3):195–203
Zoysa MD, Nikapitiya C, Oh C et al (2010) Molecular evidence for the existence of lipopolysaccharide-induced TNF-alpha factor (LITAF) and Rel/NF-kB pathways in disk abalone (Haliotis discus discus). Fish Shellfish Immunol 28:754–763
Acknowledgments
This study was financially supported by the grant of National Agricultural Innovation Project (NAIP) of the Indian Council of Agricultural Research (ICAR) (Project code C4-C30018). We thank Dr. P. Routray, Senior Scientist, Aquaculture Production and Environment Division, CIFA, for hatchery facility, expert consultations and various suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Samanta, M., Basu, M., Swain, B. et al. Molecular cloning and characterization of toll-like receptor 3, and inductive expression analysis of type I IFN, Mx and pro-inflammatory cytokines in the Indian carp, rohu (Labeo rohita). Mol Biol Rep 40, 225–235 (2013). https://doi.org/10.1007/s11033-012-2053-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2053-6