Abstract
Acyl coenzyme A long-chain 1 synthetase (ACSL1) plays a key role in animal fat synthesis and fatty acid β-oxidation. In order to research the function of the ACSL1 gene in pig, we analyzed the mRNA expression in liver, backfat and longissimus dorsi muscle by quantitative real-time PCR in Tibet pig (TP, n = 10), Diannan small ear pig (DSP, n = 10) and large white pig (LW, n = 10). The results showed that the mRNA expressions of the ACSL1 gene in liver and longissimus dorsi muscle of DSP and TP were significant higher than that of LW (P < 0.01). However, the expression in backfat of LW was significant higher than that of TP (P < 0.01) and DSP (P < 0.05). In addition, four SNPs located in 5′ flanking region (T-1191C), exon 6(G173A), exon 14(C36T) and exon 17(T46C) were identified, and the allele frequencies of the four SNPs were significant different in indigenous and introduced pig breeds. The results indicated that the ACSL1 gene might be relative to the capacity of fat deposition and meat quality in pig breeds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lipid deposition in pig is a very complex trait that is likely to be controlled by many genes [1] and is determined by a complex balance between lipogenesis and lipolysis [2–4]. In the balance, fatty acids are intermediate products and must be activated to acyl coenzyme A before participating in most catabolic and anabolic reactions [5]. Through investigation in last decades, it was confirmed that the acyl coenzyme A synthetase (ACS) was the main enzyme to catalyze the activation reactions. ACS family includes multiple isoforms classified by their substrate specificities for fatty acids of varying chain length [6]. In mammals, long-chain acyl-CoA synthetase (ACSL) catalyzes the ATP-dependent acylation of fatty acids into long-chain acyl CoAs, which is the first step in lipid metabolism after fatty acid entry into the cell [7]. The ACSL isoform, an important ACS family member, plays an essential role in both lipid biosynthesis and fatty acid degradation [8], and also plays regulatory roles in numerous reactions, including, for example, protein modification [9], intracellular protein transport [10], protein kinase C activation [11], nuclear thyroid hormone receptor modulation [12], and cell proliferation [13].
There are five cloned isoforms of ACSL: ACSL1, ACSL3, ACSL4, ACSL5, and ACSL6 [9]. These isoforms are expressed not only specially in some certain tissues, but also differently at some certain developing ages. ACSL1, ACSL4, and ACSL5 are all found in liver and adipocytes [14–16], whereas both of ACSL3 and ACSL6 are expressed in brain [17, 18]. ACSL4 expression is most abundant in steroidogenic tissues and ACSL5 in intestine [14, 15]. After birth of rats, just the ACSL1 expression increased by fourfold in heart, whereas the ACSL3 decreased and the other ACSL isoforms remained stable [19]. Furthermore, during 3T3-L1 adipocyte differentiation, the expression of ACSL1 gene increases by about 160-fold, while the expression of other isoforms is unchanged [20]. Transgenic mice with overexpressed ACSL1 in heart increased TAG content [21]. Furthermore, overexpression of ACSL1 mRNA and protein by more than fivefold over controls caused a twofold increase in TAG content in mouse liver [22]. So the ACSL1 overexpression has convincingly been thought to increase TAG accumulation.
The pig ACSL1 gene has been mapped on chromosome 15, near the SW1989 microsatellite [23]. It is split into 20 exons and the resulting cDNA encompass 3,133 bp, of which 2,097 bp correspond to the coding region. However, it is unclear whether the trait of fat accumulation in pigs is relative to the ACSL1 gene expression in some organs. We proposed a hypothesis that the varying phenotype of fat traits in pig might be due to different quantity of the ACSL1 expression or polymorphism of SNPs in ACSL1 gene region. The aim of this study was to investigate the differences of ACSL1 expression and genome polymorphisms in several pig breeds that have different intramuscular fat (IMF) contents, and to provide basic molecular information for the further research on the function of the ACSL1 gene in pig.
Materials and methods
Experimental materials
This experiment covered three breeds of pig: Tibet pig (TP) from Tibet Agricultural and Animal Husbandry College, Diannan small ear pig (DSP) and large white (LW) from Xishuang Banna city, Yunnan province, China. 30 castrated boars (10 each group) were slaughtered when they were 6-month age. Tissue samples were collected from liver, backfat, longissimus dorsi muscle at the last rib. The samples were immediately frozen in liquid nitrogen, and then were stored at −80 °C. Ear samples were collected from populations of TP (n = 67), DSP (n = 54) and LW (n = 56) which was used to detect SNPs in the region of ACSL1 gene.
DNA, RNA extraction and cDNA preparation
Genomic DNA were isolated from ear tissues with the extraction procedure as described [24], dissolved in TE solution and preserved in −20 °C refrigerator.
Total RNA was isolated from the liver, backfat and longissimus dorsi muscle tissues with TRIZOL® Reagent (Invitrogen, San Diego, CA, USA) using the method of the manufacturer’s instructions. The RNA solutions were checked for concentration and purity using an NanoDrop 2000 Biophotometer (Thermo scientific, USA) at 260/280 nm absorbance ratio (range 1.8-2.0 indicates a pure RNA sample) and in a 1 % agarose gel to verify its integrity.
After treatment with DNase I, the total RNA was reverse transcribed to cDNA in a reaction volume of 20 μL containing 2 μg total RNA, 50 μmol oligo-d(T) 15 as a primer and 10 nmol dNTP mix. These mixtures were heated at 70 °C for 5 min and incubated on ice for 2 min. After that, 200 U ImProm-II™ reverse transcriptase (Promega, USA), 40 U RNase inhibitor (Promega Biotech Co., Ltd.) and 4 μL 5× reaction buffer were added to the each mixture and was incubated at 42 °C for 60 min, and then inactivated by heating at 70 °C for 15 min.
Quantitative analysis of ACSL1 mRNA expression
The expression quantity of ACSL1 gene was measured using real-time PCR. Primers were designed using Primer Premier 5.0 software spanning one intron to avoid genomic DNA contamination. ACSL1 (NM_001167629) primers were: 5′-GCA GGC ATT TCT CAT AGC G-3′ and 5′-TCC CTC CCC AGT CTC AGC AT-3′. We selected the housekeeping gene glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as the internal standard [25]. GAPDH primers were 5′-GGT CAC CAG GGC TGC TTT TA-3′ and 5′-CCT TGA CTG TGC CGT GGA AT-3′. Real-time PCR amplification was conducted using Bio-Rad CFX96 System (Bio-Rad, USA). The gene expression quantity was calculated with the method of 2−ΔΔCt (ΔΔCt = ΔCttarget gene − ΔCt housekeeping gene) [26]. A cDNA pool of all samples was used as a calibration.
SNP screening
Primers for SNP identification were designed using Primer Premier 5.0 software allowing the amplification of a region of 5′ flanking region and coding region (from exon 1 to 20). The targeted regions, primer sequence and the amplicon size are shown in Table 1. The PCR products amplified from 10 pigs each group were pooled and sequenced directly to identify SNPs. Chromas Pro and DNAMAN6.0 were used to analyze the sequencing results.
SNP genotyping
For SNPs found in sequence alignment, the software of NEBcutter V2.0 (http://tools.neb.com/NEBcutter2/) was used online to search for special restriction enzymes. The HinfI, HhaI, PvuII, Tasl were selected for genotyping loci of 5′ flanking T-1191C, exon 6 G173A, exon 14 C36T and exon 17 T46C respectively. The reaction conditions were shown in Table 2.
Statistical analysis
The expression levels were analyzed by one-way ANOVA with repeated measures using SAS9.1 software (SAS Inst. Inc., Cary, NC). The results are presented as mean ± standard error. Significant and extreme differences were set at P < 0.05 and P < 0.01, respectively. χ2 test was used to analyze the distribution of genotypes and the differences in alleles frequencies.
Results
ACSL1 mRNA expression level in the three tissues among the three breeds of pig
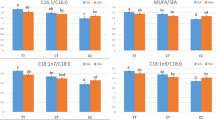
The ACSL1 mRNA expression level was different among breeds and tissues as shown in Fig. 1. In both DSP and TP, the ACSL1 gene expression was maximal in liver tissue and mid in backfat and minimal in longissimus dorsi muscle, and the differences among the three tissues were extremely significant differences (P < 0.01). But in LW, the expression difference between liver and backfat tissue was not significant (P > 0.05). The ACSL1 expression level in liver tissue of DSP and TP was significantly higher than that of LW (P < 0.01), and there was no significant differences of the gene expression in liver between DSP and TP (P > 0.05). However, the gene expression in backfat tissue of LW was higher than that of DSP and TP (P < 0.05), and no significant difference was seen in backfat of DSP and TP (P > 0.05). The expression level in longissimus dorsi muscle was the highest in DSP, and was higher than in TP and LW (P < 0.05). And the expression was significantly higher in TP than in LW (P < 0.05).
The ACSL1 mRNA expression level in three tissues of the three pig breeds. Note Error bars represent SE. Letters on bars denote the difference of expression level with significantly difference (P < 0.05). Break range on Y axis is 0.1–0.5 being omitted. DSP diannan small ear pig (n = 10). TP Tibet pig (n = 10), LW large white (n = 10)
SNP identification
Four SNPs, 5′ flanking region (T-1191C), exon 6 (G173A), exon 14 (C36T), and exon 17 (T46C), were identified by sequencing. The mutations of exon 6 G173A, exon 14 C36T and exon 17 T46C caused no amino acid changes, so they were all synonymous mutations. No restriction enzymes were found to identify the locus of 5′ flanking region T-1191C, so we designed a forward mismatched primer for identify the mutation using the dCAPS Finder 2.0 software (http://helix.wustl.edu/dcaps/dcaps.html). The mismatched primer sequence was 5′-CAA AAA TAT CAT CTC AAC TTA GAG T-3′, which was used for PCR-RFLP analyzing for the T-1191C with the reverse primer of 5′-FR2 (listed in Table 1). PCR product size of the primers was 218 bp, which could be digested by restriction enzyme Hinf I on the loci of T-1191C. The PCR products amplified using the primers ACSL-P5 and ACSL-P12 listed in Table 1 respectively were used to be digested by HhaI and Pvu II to genotype of the locus of exon 6 G173A and exon 14 C36T. Another forward primer was designed for amplification of exon 17 region with the reverse primer of ACSL-P15 listed in Table 1, and the sequence was 5′-ACA GGC TCA CTT CGC AGG TAG AT-3′. And the PCR product (292 bp) was digested by Tasl to detect the genotype of the exon 17 T46C.
SNP genotype frequency
The genotype frequency and allele frequency of each site of ACSL1 gene in TP (n = 67), DSP (n = 54) and LW (n = 56) breeds were shown in Table 3. The results showed that polymorphisms of T-1191C, G173A, and C36T were only identified in TP and DSP populations, whereas polymorphism of T46C was found in the three breed populations. The χ2 test showed that four locus were staying in Hardy–Weinberg equilibrium (P > 0.05) in TP population. But in DSP and LW populations, only the T46C site was staying in Hardy–Weinberg equilibrium (P > 0.05). The C allele frequency of T-1191C site and the T allele frequency of C36T site in TP population were higher than those in DSP (P > 0.05), and the G allele frequency of G173A site in TP population was significantly higher than that in DSP (P < 0.01). In T46C site, the C allele frequencies of TP and DSP populations were extremely significant higher than that of LW (P < 0.01).
Discussions
The native breeds such as TP in the Qinghai-Tibet Plateau and DSP in south of Yunnan province of China have good adaptation to the harsh conditions and have superior meat quality [27–29]. The two indigenous pigs have high IMF content, whereas LW, a introduced pig breed, possesses good performances in growth rate and lean ratio, but has a low IMF [29, 30]. The ACSL1 expression level was 1.8- and 2.2-fold higher in livers of TP and DSP respectively, and was 4- and 6-fold higher in longissimus dorsi muscles of TP and TSP respectively, comparing with LW. The data in vitro and in vivo had indicated that the ACSL1 was linked to the storage pathway of lipid metabolism in liver and that it might act to channel fatty acids into triglyceride synthesis rather than into β-oxidation and energy production [22]. Our data suggest that the DSP and TP breeds with higher capacity of adipose deposition need more ACSL1 to synthesize long-chain acyl-coenzyme A for the synthesis of triglyceride, which may be the reason of higher IMF content in TP and DSP breeds than in LW. Otherwise, the expression level in backfat of LW was higher than that in TP and DSP. Those results suggested that the expression of ACSL1 in different tissues may be regulated by different transcription factors. For instance, ACSL1 in liver is regulated by PPARα [31, 32] that mainly expressed in liver [33, 34], whereas it is regulated by PPARγ [35] in adipose tissue [36, 37]. Otherwise, it is also possible that more ACSL1 needed for β-oxidation in backfat of LW than the other breeds. The partitioning of intracellular fatty acids between storage pathways and β-oxidation is also controlled by regulation of the mitochondrial acyl-CoA transporter carnitine palmitoyltransferase-1 [38].
In this study, we identified four SNPs in the region of pig ACSL1 gene. The 5′ flanking region T-1191C, exon 6 G173A and exon 14 C36T sites had not been reported in previous works and the polymorphisms of the three locis were only found in Chinese indigenous pig breeds. For the 5′ flanking T-1191C, the binding sites of transcription factors were predicted by the software CONSITE online (http://asp.ii.uib.no:8090/cgi-bin/CONSITE/consite). The results indicated that the mutation of T → C led sites for combining with SOX17 and broad-complex-1 transcription factors. The SOX17 transcription factor mainly acts as a regulator for sustaining self-renewal of hematopoietic stem cells [39, 40]. The broad-complex-1 is a zinc-finger protein found in insects such as Bombyx mori, Drosophila melanogaster, and it has not been found the homologisation sequences in mammals. No polymorphisms of the exon 6 G173A and exon 14 C36T sites were found in LW and also in Duroc, Landrace, Piétain, LW and Iberian populations [41]. So, we suggested that the mutations of the two sites may be particular to the TP and DSP populations. The polymorphism of exon 17 T46C was detected in the three pig breeds and in other breeds such as Duroc, Landrace [41], but the mutation in exon 17 region was not found in Piétain and Iberian populations [41].
In conclusion, the pig ACSL1 gene expresses differently in different tissues and pig breeds. The acyl coenzyme A activated by ACSL1 can enter any metabolic both lipid biosynthesis and fatty acid degradation [5], the mechanism of higher IMF content in fatty-type pigs may be due to more ACSL1 needed for lipogenesis in liver and longissimus dorsi muscle tissues, and the thinner of backfat thickness in lean-type pigs may be due to more ACSL1 needed for β-oxidization in backfat. We found four mutant loci in the two indigenous pigs, but only one of them was multiple in LW populations. The allele frequencies of the four sites were significantly different in indigenous and introduced pig breeds. Those four mutations may be relative to fat deposition in backfat and IMF. However, further studies will be needed such as correlation analysis between ACSL1 expression level and the traits of backfat thickness and IMF content, genotypes of SNPs and lipid deposition traits.
References
Li L (2009) Genomic characterization and polymorphism analysis of genes involved in lipid- and energy metabolism in swine. Ph D Thesis, Technische Universität München, Germany
Ding ST, Schinkel AP, Weber TE, Mersmann HJ (2000) Expression of porcine transcription factors and genes related to fatty acid metabolism in different tissues and genetic populations. J Anim Sci 78:2127–2134
Reiter SS, Halsey CHC, Stronach BM, Bartosh JL, Owsley WF, Bergen WG (2007) Lipid metabolism related gene-expression profiling in liver, skeletal muscle and adipose tissue in crossbred Duroc and Pietrain pigs. Comp Biochem Phys D 2:200–206
Scott RA, Cornelius SG, Mersmann HJ (1981) Effects of age on lipogenesis and lipolysis in lean and obese swine. J Anim Sci 52:505–511
Watkins PA (1997) Fatty acid activation. Prog Lipid Res 36:55–83
Steinberg SJ, Morgenthaler J, Heinzer AK, Smith KD, Watkins PA (2000) Very long-chain acyl-CoA synthetases: human “bubblegum” represents a new family of proteins capable of activating very long-chain fatty acids. J Biol Chem 275:35162–35169
Mashek DG, Bornfeldt KE, Coleman RA, Berger J, Bernlohr DA, Black P, DiRusso CC, Farber SA, Guo W, Hashimoto N, Khodiyar V, Kuypers FA, Maltais LJ, Nebert DW, Renieri A, Schaffer JE, Stahl A, Watkins PA, Vasiliou V, Yamamoto TT (2004) Revised nomenclature for the mammalian long chain acyl-CoA synthetase gene family. J Lipid Res 45:1958–1961
Mercade A, Estelle J, Perez-Enciso M, Varona L, Silio L, Noguera JL, Sanchez A, Folch JM (2006) Characterization of the porcine acyl-CoA synthetase long-chain 4 gene and its association with growth and meat quality traits. Anim Genet 37:219–224
Grand RJ (1989) Acylation of viral and eukaryotic proteins. Biochem J 258:625–638
Glick BS, Rothman JE (1987) Possible role for fatty acyl-coenzyme A in intracellular protein transport. Nature 326:309–312
Prentki M, Corkey BE (1996) Are the beta-cell signaling molecules malonyl-CoA and cystolic long-chain acyl-CoA implicated in multiple tissue defects of obesity and NIDDM? Diabetes 45:273–283
Li QL, Yamamoto N, Inoue A, Morisawa S (1990) Fatty acyl-CoAs are potent inhibitors of the nuclear thyroid hormone receptor in vitro. J Biochem (Tokyo) 107:699–702
Tomoda H, Igarashi K, Cyong J-C, Omura S (1991) Evidence for an essential role of long chain acyl-CoA synthetase in animal cell proliferation: inhibition of long chain acyl-CoA synthetase by triacsins caused inhibition of Raji cell proliferation. J Biol Chem 266:4214–4219
Kang MJ, Fujino T, Sasano H, Minekura H, Yabuki N, Nagura H, Iijima H, Yamamoto TT (1997) A novel arachidonate-preferring acyl-CoA synthetase is present in steroidogenic cells of the rat adrenal, ovary, and testis. Proc Natl Acad Sci USA 94:2880–2884
Oikawa E, Iijima H, Suzuki T, Sasano H, Sato H, Kamataki A, Nagura H, Kang MJ, Fujino T, Suzuki H, Yamamoto TT (1998) A novel acyl-CoA synthetase, ACS5, expressed in intestinal epithelial cells and proliferating preadipocytes. J Biochem (Tokyo) 124:679–685
Suzuki H, Kawarabayasi Y, Kondo J, Abe T, Nishikawa K, Kimura S, Hashimoto T, Yamamoto T (1990) Structure and regulation of rat longchain acyl-CoA synthetase. J Biol Chem 265:8681–8685
Fujino T, Kang MJ, Suzuki H, Iijima H, Yamamoto T (1996) Molecular characterization and expression of rat acyl-CoA synthetase 3. J Biol Chem 271:16748–16752
Fujino T, Yamamoto T (1992) Cloning and functional expression of a novel long-chain acyl-CoA synthetase expressed in brain. J Biochem (Tokyo) 111:197–203
de Jong H, Neal AC, Coleman RA, Lewin TM (2007) Ontogeny of mRNA expression and activity of long-chain acyl-CoA synthetase (ACSL) isoforms in Mus musculus heart. Biochim Biophys Acta 1771:75–82
Marszalek JR, Kitidis C, Dararutana A, Lodish HF (2004) Acyl CoA synthetase 2 (ACS2) over-expression enhances fatty acid internalization and neurite outgrowth. J Biol Chem 279:23882–23891
Chiu HC, Kovacs A, Ford DA, Hsu FF, Garcia R, Herrero P, Saffitz JE, Schaffer JE (2001) A novel mouse model of lipotoxic cardiomyopathy. J Clin Invest 107:813–822
Parkes HA, Preston E, Wilks D, Ballesteros M, Carpenter L, Wood L, Kraegen EW, Furler SM, Cooney GJ (2006) Overexpression of acyl-CoA synthetase-1 increases lipid deposition in hepatic (HepG2) cells and rodent liver in vivo. Am J Physiol Endocrinol Metab 291:E737–E744
Vidal O, Amills M (2004) Assignment of the fatty acid coenzyme A ligase, long chain 2 (FACL2) gene to porcine chromosome 15. Anim Genet 35:245
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Malathi B, Aryamani B, Robert AT, Leo SL, James DT (2008) Evaluation and validation of housekeeping genes in response to ionizing radiation and chemical exposure for normalizing RNA expression in real-time PCR. Mutat Res 649:126–134
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C (T)) method. Methods 25:402–408
Cheng P (1984) Livestock breeds of China-animal production and health paper, vol 46. FAO, Rome, p 217 E, F, S
Jian-jun G, Zhi-ping H, Zheng que L, Xue-bin L, San-cheng Y, Xiao-hui C (2007) Investigation on fattening and carcass traits in Tibetan pig and its combinations. Southwest China J Agric Sci (Chinese Article) 20:1109–1112
Pan PW, Zhao SH, Yu M, Xiong TA, Li K (2003) Identification of differentially expressed genes in the longissimus dorsi tissue between Duroc and Erhualian pigs by mRNA differential display. Asian-Aust J Anim Sci 16:1066–1070
Plastow GS, Carrión D, Gil M, Garía-Regueiro JA, Furnols MFI, Gispert M, Oliver MA, Velarde A, Guàrdia MD, Hortós M, Rius MA, Sárraga C, Díaz I, Valero A, Sosnicki A, Klont R, Dornan S, Wilkinson JM, Evans G, Sargent C, Davey G, Connolly D, Houeix B, Maltin CM, Hayes HE, Anandavijayan V, Foury A, Geverink N, Cairns M, Tilley RE, Mormède P, Blott SC (2005) Quality pork genes and meat production. Meat Sci 70:409–421
Aoyama T, Peters JM, Iritani N, Nakajima T, Furihata K, Hashimoto T, Gonzalez FJ (1998) Altered constitutive expression of fatty acid-metabolizing enzymes in mice lacking the peroxisome proliferator-activated receptor alpha (PPARalpha). J Biol Chem 273:5678–5684
Djouadi F, Brandt JM, Weinheimer CJ, Leone TC, Gonzalez FJ, Kelly DP (1999) The role of the peroxisome proliferator-activated receptor alpha (PPAR alpha) in the control of cardiac lipid metabolism. Prostaglandins Leukot Essent Fatty Acids 60:339–343
Braissant O, Foufelle F, Scotto C, Dauça M, Wahli W (1996) Differential expression of peroxisome proliferator-activated receptors (PPARs): tissue distribution of PPAR-α, -β, and -γ in the adult rat. Endocrinology 137:354–366
Escher P, Braissant O, Basu-Modak S, Michalik L, Wahli W, Desvergne B (2001) Rat PPARs: quantitative analysis in adult rat tissues and regulation in fasting and refeeding. Endocrinology 142:4195–4202
Martin G, Schoonjans K, Lefebvre AM, Staels B, Auwerx J (1997) Coordinate regulation of the expression of the fatty acid transport protein and acyl-CoA synthetase genes by PPAR alpha and PPAR gamma activators. J Biol Chem 272:28210–28217
Hoekstra M, Kruijt JK, Van Eck M, Van Berkel TJ (2003) Specific gene expression of ATP-binding cassette transporters and nuclear hormone receptors in rat liver parenchymal, endothelial, and Kupffer cells. J Biol Chem 278:25448–25453
Peters JM, Rusyn I, Rose ML, Gonzalez FJ, Thurman RG (2000) Peroxisome proliferator-activated receptorα is restricted to hepatic parenchymal cells, not Kupffer cells: implications for the mechanism of action of peroxisome proliferators in hepatocarcinogenesis. Carcinogenesis 21:823–826
McGarry JD (2002) Banting lecture 2001: dysregulation of fatty acid metabolism in the etiology of type 2 diabetes. Diabetes 51:7–18
Chhabra A, Mikkola KAH (2011) Return to youth with Sox17. Genes Dev 25:1557–1562
He S, Kim I, Lim MS, Morrison SJ (2011) Sox17 expression confers self-renewal potential and fetal stem cell characteristics upon adult hematopoietic progenitors. Genes Dev 25:1613–1627
Vidal O, Sánchez A, Amills M, Noguera JL (2007) Nucleotide sequence and polymorphism of the pig acyl coenzyme A synthetase long-chain 1 (ACSL1) gene. Anim Biotechnol 18:117–122
Acknowledgments
This work was supported by the National Major Special Project on New Varieties Cultivation for Transgenic Organisms (No. 2011ZX08009-003-006) the National Natural Science Foundation of China (U1036604 and 31160441). And we thank the Yunnan Agricultural University and Tibet Agricultural and Animal Sciences College of the help for the samples collecting.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Q., Tao, Z., Shi, L. et al. Expression and genome polymorphism of ACSL1 gene in different pig breeds. Mol Biol Rep 39, 8787–8792 (2012). https://doi.org/10.1007/s11033-012-1741-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1741-6