Abstract
Subtractive hybridization cDNA library (SHL) is one of the powerful approaches for isolating differentially expressed genes. Using this technique between mouse heart and skeletal muscle (skm) tissues, we aimed to construct a cDNA-library that was specific to heart tissue and to identify the potential candidate genes that might be responsible for the development of cardiac diseases or related pathophysiological conditions. In the first step of the study, we created a cDNA-library between mouse heart and skm tissues. The homologies of the randomly selected 215 clones were analyzed and then classified by function. A total of 146 genes were analyzed for their expression profiles in the heart and skm tissues in published mouse microarray dataset. In the second step, we analyzed the expression patterns of the selected genes by Northern blot and RNA in situ hybridization (RISH). In Northern blot analyses, the expression levels of Myl3, Myl2, Mfn2, Dcn, Pdlim4, mt-Co3, mt-Co1, Atpase6 and Tsc22d1 genes were higher in heart than skm. For first time with this study, expression patterns of Pdlim4 and Tsc22d1 genes in mouse heart and skm were shown by RISH. In the last step, 43 genes in this library were identified to have relationships mostly with cardiac diseases and/or related phenotypes. This is the first study reporting differentially expressed genes in healthy mouse heart using SHL technique. This study confirms our hypothesis that tissue-specific genes are most likely to have a disease association, if they possess mutations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The developmental and pathophysiological conditions in several tissues cause qualitative and quantitative differences in gene expression. There are several polymerase chain reaction (PCR)-based techniques used in comparative gene expression analysis. These techniques involve many different approaches regarding the source of genetic material (DNA or mRNA) and the subtraction techniques (solid substrates, magnetic beads, chromatography, polyacrilamide gel systems etc.) [1–6]. Subtractive hybridization cDNA library (SHL) technique is one of the powerful approaches for identifying and isolating genes that are differentially expressed in defined tissues or cell lines. This method has been previously used to distinguish the genes that are differentially expressed in various tissues [1–3, 7–11]. Recent technological developments in large-scale expression analysis have made it possible to rapidly assess many genes for their transcriptional responses in biological and pathological events [12]. The applications of these techniques have various strengths and weaknesses. As a result, the most appropriate method should be determined according to the existing facilities and the experimental design of the planned study.
In a previous article reviewed by Nanni et al. [13] it was pointed out that many studies using microarray technologies have focused on the transcriptional differences on human tissue samples, and also cellular and animal models in pathological events related with several genetic and multifactorial cardiac disorders. However, the differentially expressed genes in normal heart have not been fully identified yet. Heart as a whole, with its cellular heterogeneity, is a complex organ. Therefore, the global transcriptional changes of heart in human are not possible to investigate. However, it becomes possible to identify human homologues of the novel heart-specific genes identified and isolated using gene expression analysis techniques such as SHL on rodent models. In this study, we aimed to construct a cDNA-library that was specific to heart tissue by using SHL technique between heart and skeletal muscle (skm) of mouse (BALB/c). Thus, novel and rare genes that are expressed in the heart could be isolated and determined as remarkable candidates for cardiac diseases or pathophysiological conditions.
Materials and methods
Animals and tissue preparation
The animal application procedures were approved by the Local Ethics Committee for Experimental Animals, Institute for Experimental Medicine, Istanbul University. Tissues were obtained from adult healthy male BALB/c mice with a minimum age of 8-week. Heart and skm were dissected, rinsed briefly twice in ice cold RNase free phosphate buffered saline (PBS) and then immediately frozen in liquid nitrogen and stored at −80 °C until RNA extraction. For SHL and Northern Blot analysis, total RNA was extracted from heart and skm. For in situ hybridization study, heart and skm were fixed with 4 % paraformaldehyde in PBS for 2 h, washed with PBS for 30 min and equilibrated with 30 % sucrose in PBS at +4 °C for overnight, then embedded in OCT medium onto the cryostat holder.
RNA isolation
Total RNAs were extracted from heart and skm tissues using the ToTALLY RNA™ Total RNA Isolation Kit (Ambion® CA, USA). The quantity and quality of each sample were determined spectrophotometrically by A260 and A260/280 ratio, and evaluated by visualizing the 28S and 18S ribosomal bands with ethidium bromide following electrophoretic separation on formaldehyde denaturing 1.2 % agarose gel.
Subtractive cDNA hybridization and cloning
For the subtraction of the heart specific transcripts, we used a modified technique reported by Hara et al. [1] which is an efficient method for subtractive cDNA cloning using oligo(dT)30-Latex and PCR. The subtraction was performed by hybridization of cDNA-oligo(dT)30-Latex of skm and the sense strand DNA synthesized from cDNA-oligo(dT)30-Latex of H. In this present study, we designed degenerative primers (EcoRI(dG)14N) to be used in an asymmetric PCR for the syntheses of the sense strand DNA. Subsequently, subtractive cDNAs were amplified by twenty rounds of PCR using primers (EcoRI(dG)14N and XhoI(dT)19N) that were specific for the adapter sequences. All PCR products were inserted into pGEM-T Easy plasmid vector and transformed into the competent cells (Promega, Madison, WI, USA) according to the manufacturer’s instructions.
Subtraction efficiency was assessed by comparing of beta-actin (Actb, accession number: M12481) and alpha-cardiac myosin heavy chain (MHC, accession number: M76601) PCR-products in the subtracted and unsubtracted cDNA aliquots after every four rounds of hybridizations. The forward (5′-GACCCAGATCATGTTTGAGACC-3′) and reverse (5′-TGGTGGTGAAGCTAGCC-3′) Actb primers and the forward (5′-GGAAGAGTGAGCGGCGCATCAAGG-3′) and reverse (5′-CTGCTGGAGAGGTTATTCCTCG-3′) alpha-cardiac MHC primers were used.
Screening of subtracted clones and sequencing
A total of 250 insert positive clones were randomly selected from this cDNA library and analyzed for their insert sizes. Plasmid isolation was performed using the Qiagen Plasmid Isolation Kits (Qiagen, CA, USA). Plasmid DNAs were purified and confirmed by PCR and agarose gel electrophoresis. Successfully cloned cDNA fragments were sequenced with universal primers (Sp6 and T7) by QIAGEN sequencing service (Germany).
Data analysis
The sequences of 224 insert cDNAs were determined, homology searches were performed using the BLAST program, and the functional classifications of the genes were done using Gene database (http://www.ncbi.nlm.nih.gov/gene) of the National Center of Biotechnology Information (GenBank®). The experimental set of Gene Expression Omnibus (GEO) data (http://www.ncbi.nlm.nih.gov/sites/geo) in GenBank was used for the evaluation of the expression levels of the genes in normal mouse heart and skm tissues. The numbers of the used dataset and platform were GDS3052 and GPL1261 (Affymetrix Mouse Genome 430 2.0 Array) respectively. The information of the GEO association IDs and described reporters in this dataset is given in Supplementary Table 1 (Online Resource 1). Furthermore, the relationships of the known genes with diseases were analyzed using Human Gene Mutation Database (HGMD) of Institute of Medical Genetics in Cardiff University (http://www.hgmd.cf.ac.uk).
Northern blot analysis
Selected cDNAs were analyzed for their tissue expression patterns using Northern blot analysis. cDNA fragments were labeled with PCR using Sp6/T7 primers and Dig-11-dUTP (Roche Applied Sciences, Germany). The detailed information of the labeled probes is shown in Supplementary Table 2 (Online Resource 2). 10 μg of total RNA was separated on Northern gel. Preparation of Northern gels and electrophoresis were performed using the Ambion NorthernMax Kit (Ambion, CA, USA). After the RNAs were blotted overnight on a nitrocellulose filter and ultraviolet cross-linked, hybridization was performed with specific probes overnight at 42–44 °C. After hybridization, filters were washed with 2× and 0.5× SSC containing 0.1 % SDS. Detection was carried with CSPD by using DIG Nucleic Acid Detection Kit (Roche Applied Sciences, Germany). Then, the filters were exposed to X-ray film for autoradiography.
RNA in situ hybridization (RISH)
The plasmid DNAs containing the Tpm1, Myl2, Tsc22d1, or Pdlim4 cDNA fragments were linearized either with SpeI or SacII restriction enzymes. DIG-labeled sense or antisense riboprobes (Online Resource 2) were obtained by in vitro transcription with DIG RNA Sp6/T7 Labeling Kit (Roche Applied Sciences, Germany). Frozen 10-μm tissue sections on RNase-free TESPA subbed glass slides were incubated in PBS containing 1 μg/ml of proteinase K for 15 min and then treated with hybridization buffer (50 % deionized formamide, 1.3× SSC, 5 mM EDTA, 50 μg/ml yeast RNA, 0.2 % Tween 20, 0.5 % CHAPS, 100 μg/ml heparin; pH 5.0 with citric acid) at 65 °C for 2–4 h. The hybridization reaction was carried out at 65 °C for overnight with 50 μl of hybridization mix (with 50 ng/100 μl riboprobe) on each section. After hybridization, slides were washed and incubated with anti-DIG antibody diluted 1:2,000 in a blocking buffer (Roche Applied Sciences, Germany) for 4 h. Following the washes in PBS containing 0.2 % Tween 20, sections were counterstained with NBT/BCIP (Roche Applied Sciences, Germany) at 4 °C for overnight. The analyses of the tissue sections for each probe were done at least in triple. Slides were observed under a light microscope (Leica DM4000B, Leica Microsystems, Germany) and the images were captured using a digital camera (Leica DC160, Leica Microsystems, Germany).
Results
Classification of SHL clones revealed relation to energy metabolism and myocyte structure
A total of 250 clones were randomly selected from this SHL and analyzed for their insert sizes. The average size of cDNA fragments was found to be 500 base pairs. In the next step, 224 out of 250 cDNA fragments were selected according to their sequencing results because 26 of those had insignificant smaller inserts. The homology screening was performed using the BLAST program in GenBank. We found that nine clones had no homology to any sequence in GenBank. The homology results of the remaining 215 clones are given in Table 1. Furthermore, 142 of the clones showed homologies to 117 different known genes and ESTs and next, they were classified according to functional information given in Gene database of GenBank. Although these genes and ESTs appeared to have relationship with many metabolisms and cellular components, most of them were related to energy metabolism and structural compartments of the myocyte (Table 1). Thirty-six of the clones showed homology to 29 different known genes or ESTs, however, their functions were unknown. Moreover, 37 of the clones showed homology to 28 different undetermined ESTs. Additionally, the occurrence numbers of cDNA fragments in the library are also indicated in Table 1.
Almost 50 % of the known genes were expressed in heart tissue
In the following step, we analyzed the known genes (n = 146) for their expression profiles in the heart and skm tissues in the published mouse microarray dataset of GEO database (Online Resource 1). It was not surprising that genes had higher expression levels in heart tissue (47.3 %, n = 69). In this dataset, we determined that 40.4 % of genes (n = 59) had higher expression levels in skm than heart whereas 7.5 % of genes (n = 11) had equal expression levels in both tissues. In addition, there were no indicated expression values for some genes (4.8 %, n = 7). However, there were dubious (not detected or up to one different values) expression values in this dataset and these values (n = 22) are labeled in Supplementary Table 1 (Online Resource 1).
Northern blot analysis confirmed higher expression in heart tissue for most of the analyzed genes
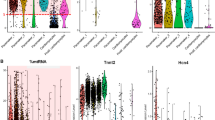
In the next step, a total of 18 known genes or ESTs selected cDNA fragments were analyzed for their tissue expression patterns using Northern blot analysis. Fifty percent were confirmed to have increased expression in heart, however, 28 % had higher expression in skm, and 22 % had equal expression in both tissues. Northern blot results showed that expression levels of Myl3, Myl2, Mfn2, Dcn, Pdlim4, mt-Co3, mt-Co1, Atpase6 and Tsc22d1 genes were relatively higher in heart than skm (Fig. 1a, b). We observed different band patterns for mt-Co3, mt-Co1, Atpase6 and Tsc22d1 in Northern blot results (Fig. 1b). With Northern hybridization analysis, it was not clearly confirmed that there existed two Tsc22d1 transcripts for heart tissue (approximately 1.8 kb, Tsc22d1-isoform 2) and for skm tissue (approximately 4.8 kb, Tsc22d1-isoform 1). Furthermore, the expression levels of mt-Co2, Rpl4, 1810027O10Rik and Tspyl1 genes were similar in both tissues (Fig. 1c). However, Chchd1, Tpm1, St3gal6, 2810428I15Rik and Actb showed higher expression in skm than the heart tissue (Fig. 1d). These Northern blot results were compared with the results of mouse microarray dataset except for the mitochondrial genes (mt-Co3, mt-Co1, Atpase6 and mt-Co3). In the result of this comparison, the expression profiles of ten genes were consistent whereas results of Pdlim4 and Rpl4 were relatively different.
Northern blot analysis of the selected mRNAs in mouse adult heart and skm. The blots were visualized by autoradiography (upper panels). Each lane contained 10 μg of total RNA from the heart (H) and skeletal muscle (skm). The lower panels display the ethidium bromide staining of the 18S rRNA as control. a higher expression levels in heart, b different band patterns in heart, c similar expression levels in both tissue, and d higher expression levels in skm
RISH analyses revealed the tissue expression patterns of selected genes
In this study, Pdlim4 and Tsc22d1 genes were selected for the determination of the heart and skm expression patterns in tissue sections to ensure integrity with the results of previous studies. As shown in Fig. 2, the tissue expression patterns of Pdlim4, Tsc22d1, Tpm1 and Myl2 genes in heart and skm were analyzed with RISH technique (Fig. 2). The Tpm1-antisense probe for both tissues and Myl2-antisense probe for heart were used as positive controls. Sense probes for four genes and Myl2-antisense probe for skm were used as negative controls, produced no distinct signals in all slides. Pdlim4 antisense transcript was localized in heart and skm and the localization patterns appeared to be uniform. In heart, Tsc22d1 antisense transcript was mainly localized in atrium in comparison to the ventricular site. Additionally, as shown in Fig. 2, Tsc22d1 isoform 2 in atrial myocytes and Tsc22d1 isoform 1 in skm had more dense expressions.
Sections in situ localization of tpm1, myl2, pdlim4 and tsc22d1 mRNAs in heart and skm. In situ hybridization of tpm1 (control for both tissues), myl2 (control for heart), pdlim4 and tsc22d1 mRNAs was performed with tissue cryosections (10 μm) of adult BALB/c mice. H heart, skm skeletal muscle, a atrium, v ventricle, as antisense probe, and s sense probe. NBT/BCIP staining was performed for overnight. Magnifications: ×40 and ×200
The relationships of known genes with diseases and/or cardiac phenotypes
In the scope of this study, we also analyzed the relationships of known genes detected in this SHL with cardiac diseases and/or phenotypes in human. Overall, mutations of the 43 genes were determined to cause several diseases and/or cardiac phenotypes depending on the data of HGMD and other published references (Table 2). However, there is no published information about the relationships of other genes in this library with cardiac diseases or pathological situations yet.
Discussion
The present study provides novel information on differentially expressed genes in healthy heart tissue and also data for remarkable candidate genes for cardiac diseases. Of the 215 clones analyzed within this study, we determined several novel ESTs using SHL technique between heart and skm of mouse. Not surprisingly, the functional classification showed that 117 different known transcripts from 142 clones were mostly related to energy metabolism and structural compartments of the myocyte. In addition, we showed that the 43 known genes in present cDNA library were mostly associated with cardiac diseases and/or related phenotypes.
The SSH technique combines equalization of abundantly expressed cDNAs and enrichment of (over 1,000-fold) rare sequences [2]. In most of the previous studies, higher success rates (84–94 %) were reported [3, 8, 9]. However, a high false positive rate has been reported, resulting in the identification of <20 % of clones and validated differential expression of approximately 2 % of the candidates [30]. In another study, using SSH, 56 candidates of 4,000 clones (1.4 %) have been identified [7]. In our study, differential expressions of 146 different genes were analyzed in published microarray dataset and 47.3 % of these genes were found to have higher expression levels in the mouse hearts when compared to skm. On the other hand, nine of the randomly selected 18 genes, namely Myl3, Myl2, Mfn2, Dcn, Pdlim4, mt-Co3, mt-Co1, Atpase6 and Tsc22d1, were confirmed to have differentiated expression in heart by Northern blot analysis (50 % in random).
The similarity of the gene expression pattern of the other genes in both tissues and analyses of the randomly selected small number of clones may explain the low success rate of our study; however, this rate does not reflect the overall success rate. The limitation of the present study is that the expression levels of all transcripts in this library have not been experimentally confirmed yet. The valid success rate of SHL techniques may be determined with next generation sequencing approaches in the future.
Similar to our Northern blot results, recent studies reported prominent expression of Pdlim4 and Tsc22d1-isoform 2 (short transcript) in heart [31, 32]. The role of PDZ and LIM domain protein 4 (Pdlim4; also known as ril), a member of PDZ–LIM protein family, in cardiac disease is unknown. However, another member of this family, PDLIM3 (also known as ALP) gene has been previously associated with dilated cardiomyopathy [33, 34]. On the other hand, Tsc22d1 (TGFβ-stimulated clone 22; also known as TSC-22) was reported to significantly enhance CNP (C-type natriuretic peptide) promoter activity in human aortic endothelial cells by the stimulation of transforming growth factor β [31]. Pdlim4 and Tsc22d1 mRNAs have been described to have roles as transcriptional regulator or interaction with other proteins in important mechanisms. In this study, we investigated the tissue expression profiles of these genes with RISH technique and showed their expression in all parts of the heart and also skm. Furthermore, we found that Tsc22d1 was predominantly expressed in atrium. This finding suggests that Tsc22d1 and the genes regulated by Tsc22d1 in heart tissue might be potential candidates for atrial fibrillation or other related phenotypes. A previous study reporting 4.0-fold upregulated expression of Tsc22d1 in porcine atria with pacing-induced fibrillation [35] strongly supports our suggestion.
Genes that are specifically or predominantly expressed in heart are likely to be important for the cardiac function. The subtracted genes can lead to uncover novel physiological functions of the heart in healthy conditions and also in diseases. Furthermore, this data has potential value in the identification of the novel candidate genes for cardiac diseases. Although, SSH was successfully used to identify differentially upregulated myocardial genes in idiopathic dilated cardiomyopathy [36], and ventricular septal defect [8], heart as a whole, with its cellular heterogeneity is a complex organ. Gene expression differences in normal heart tissue comparing with skm have not been studied before. This approach enabled us to compare the transcripts of the two muscle origined tissues. Thus, by using this approach, we identified several differentially expressed candidate genes whose malfunction might be responsible for the development of cardiac diseases or related pathophysiological conditions. These genes are mostly involved in energy metabolism and structural compartments of the cell in heart tissue. In addition, mutations or variants in some of these candidate genes were determined to cause cardiac diseases and/or related phenotypes [HGMB and 14–29]. However, other differentially expressed genes or functionally active genes in heart might be important in the determination of the novel candidate genes for cardiac diseases.
In our study, we utilized mice for creating cDNA library, because to obtain postmortem hearts are rather difficult. Mice and other animal models are being used in several experimental studies to explore mechanisms underlying pathophysiological conditions [13]. After identifying a mouse-specific gene as an effector, one can select the corresponding human counterpart of that particular gene. In fact, the human homologies of 43 genes in our mouse cDNA-library were identified to have relationship mostly with cardiac diseases and/or related phenotypes. For the rest, large-scale gene expression and functional analyses are needed. As a consequence of limitations, differentially gene expression analyses between heart and skm using subtractive hybridization library, Northern Blot and RISH techniques were applied in this study. This only allowed us to analyze a limited number of genes.
Our preliminary findings suggest that unknown genes and ESTs in this SHL are crucial candidates that might have a role in cardiac physiology. Moreover, further studies of heart specific subtracted genes will provide insights into their functional roles in heart, even provide new knowledge for the etiology of the cardiovascular diseases. This is a useful approach for tissue-specific genes related with pathophysiological conditions.
References
Hara E, Yamaguchi T, Tahara H, Tsuyama N, Tsurui H, Ide T, Oda K (1993) DNA–DNA subtractive cDNA cloning using oligo(dT)30-Latex and PCR: identification of cellular genes which are overexpressed in senescent human diploid fibroblasts. Anal Biochem 214:58–64. doi:10.1006/abio.1993.1456
Diatchenko L, Lau YF, Campbell AP, Chenchik A, Moqadam F, Huang B, Lukyanov S, Lukyanov K, Gurskaya N, Sverdlov ED, Siebert PD (1996) Suppression subtractive hybridization: a method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc Natl Acad Sci USA 93:6025–6030
von Stein OD, Thies WG, Hofmann M (1997) A high throughput screening for rarely transcribed differentially expressed genes. Nucleic Acids Res 25:2598–2602
Kozian DH, Kirschbaum BJ (1999) Comparative gene-expression analysis. Trends Biotechnol 17:73–78. doi:10.1016/S0167-7799(98)01292-X
Suzuki Y, Sato N, Tohyama M, Wanaka A, Takagi T (1996) Efficient isolation of differentially expressed genes by means of a newly established method, ‘ESD’. Nucleic Acids Res 24:797–799
Pathak RU, Kanungo MS (2007) Subtractive differential display: a modified differential display technique for isolating differentially expressed genes. Mol Biol Rep 34:41–46. doi:10.1007/s11033-006-9010-1
Wouters M, De Laet A, Donck LV, Delpire E, van Bogaert PP, Timmermans JP, de Kerchove d’Exaerde A, Smans K, Vanderwinden JM (2006) Subtractive hybridization unravels a role for the ion cotransporter NKCC1 in the murine intestinal pacemaker. Am J Physiol Gastrointest Liver Physiol 290:G1219–G1227. doi:10.1152/ajpgi.00032
Zhang H, Zhou L, Yang R, Sheng Y, Sun W, Kong X, Cao K (2006) Identification of differentially expressed genes in human heart with ventricular septal defect using suppression subtractive hybridization. Biochem Biophys Res Commun 342:135–144. doi:10.1016/j.bbrc.2006.01.113
Liang G, Zhang XD, Wang LJ, Sha YS, Zhang JC, Miao SY, Zong SD, Wang LF, Koide SS (2004) Identification of differentially expressed genes of primary spermatocyte against round spermatid isolated from human testis using the laser capture microdissection technique. Cell Res 14:507–512. doi:10.1038/sj.cr.7290254
Lee SW, Tomasetto C, Sager R (1991) Positive selection of candidate tumor-suppressor genes by subtractive hybridization. Proc Natl Acad Sci USA 88:2825–2829
Morozov G, Verlinsky O, Rechitsky S, Ivakhnenko V, Goltsman E, Gindilis V, Strom C, Kuliev A, Verlinsky Y (1999) Construction and analysis of subtraction complementary DNA libraries from human preimplantation embryos. J Assisr Reprod Genet 16:212–215. doi:10.1023/A:1020368908134
Stanton LW (2001) Methods to profile gene expression. Trends Cardiovasc Med 11:49–54. doi:10.1016/S1050-1738(01)00085-8
Nanni L, Romualdi C, Maseri A, Lanfranchi G (2006) Differential gene expression profiling in genetic and multifactorial cardiovascular diseases. J Mol Cell Cardiol 41:934–948. doi:10.1016/j.yjmcc.2006.08.009
Wang Z, Liu Y, Liu J, Liu K, Wen J, Wen S, Wu Z (2011) HSG/Mfn2 gene polymorphism and essential hypertension: a case–control association study in Chinese. J Atheroscler Thromb 18:24–31
Bolling MC, Pas HH, de Visser M, Aronica E, Pfendner EG, van den Berg MP, Diercks GF, Suurmeijer AJ, Jonkman MF (2010) PLEC1 mutations underlie adult-onset dilated cardiomyopathy in epidermolysis bullosa simplex with muscular dystrophy. J Invest Dermatol 130:1178–1181. doi:10.1038/jid.2009.390
Levitas A, Muhammad E, Harel G, Saada A, Caspi VC, Manor E, Beck JC, Sheffield V, Parvari R (2010) Familial neonatal isolated cardiomyopathy caused by a mutation in the flavoprotein subunit of succinate dehydrogenase. Eur J Hum Genet 18:1160–1165. doi:10.1038/ejhg.2010.83
Zaragoza MV, Brandon MC, Diegoli M, Arbustini E, Wallace DC (2011) Mitochondrial cardiomyopathies: how to identify candidate pathogenic mutations by mitochondrial DNA sequencing, MITOMASTER and phylogeny. Eur J Hum Genet 19:200–207. doi:10.1038/ejhg.2010.169
Chen J, Hattori Y, Nakajima K, Eizawa T, Ehara T, Koyama M, Hirai T, Fukuda Y, Kinoshita M, Sugiyama A, Hayashi J, Onaya T, Kobayashi T, Tawata M (2006) Mitochondrial complex I activity is significantly decreased in a patient with maternally inherited type 2 diabetes mellitus and hypertrophic cardiomyopathy associated with mitochondrial DNA C3310T mutation: a cybrid study. Diabetes Res Clin Pract 74:148–153. doi:10.1016/j.diabres.2006.03.024
Zifa E, Theotokis P, Kaminari A, Maridaki H, Leze H, Petsiava E, Mamuris Z, Stathopoulos C (2008) A novel G3337A mitochondrial ND1 mutation related to cardiomyopathy co-segregates with tRNALeu(CUN) A12308G and tRNAThr C15946T mutations. Mitochondrion 8:229–236. doi:10.1016/j.mito.2008.04.001
Chamkha I, Mkaouar-Rebai E, Aloulou H, Chabchoub I, Kifagi C, Fendri-Kriaa N, Kammoun T, Hachicha M, Fakhfakh F (2011) A novel m.3395A>G missense mutation in the mitochondrial ND1 gene associated with the new tRNA(Ile) m.4316A>G mutation in a patient with hypertrophic cardiomyopathy and profound hearing loss. Biochem Biophys Res Commun 404:504–510. doi:10.1016/j.bbrc.2010.12.012
Tang S, Batra A, Zhang Y, Ebenroth ES, Huang T (2010) Left ventricular noncompaction is associated with mutations in the mitochondrial genome. Mitochondrion 10:350–357. doi:10.1016/j.mito.2010.02.003
Zhu HY, Wang SW, Martin LJ, Liu L, Li YH, Chen R, Wang L, Zhang ML, Benson DW (2009) The role of mitochondrial genome in essential hypertension in a Chinese Han population. Eur J Hum Genet 17:1501–1506. doi:10.1038/ejhg.2009.63
Tang Z, Diamond MA, Chen JM, Holly TA, Bonow RO, Dasgupta A, Hyslop T, Purzycki A, Wagner J, McNamara DM, Kukulski T, Wos S, Velazquez EJ, Ardlie K, Feldman AM (2007) Polymorphisms in adenosine receptor genes are associated with infarct size in patients with ischemic cardiomyopathy. Clin Pharmacol Ther 82:435–440. doi:10.1038/sj.clpt.6100331
Yamada Y, Kato K, Oguri M, Fujimaki T, Yokoi K, Matsuo H, Watanabe S, Metoki N, Yoshida H, Satoh K, Ichihara S, Aoyagi Y, Yasunaga A, Park H, Tanaka M, Nozawa Y (2008) Genetic risk for myocardial infarction determined by polymorphisms of candidate genes in a Japanese population. J Med Genet 45:216–221. doi:10.1136/jmg.2007.054387
Omura T, Yoshiyama M, Yoshida K, Nakamura Y, Kim S, Iwao H, Takeuchi K, Yoshikawa J (2002) Dominant negative mutant of c-Jun inhibits cardiomyocyte hypertrophy induced by endothelin 1 and phenylephrine. Hypertension 9:1–6. doi:10.1161/hy0102.100783
Shastry S, Delgado MR, Dirik E, Turkmen M, Agarwal AK, Garg A (2010) Congenital generalized lipodystrophy, type 4 (CGL4) associated with myopathy due to novel PTRF mutations. Am J Med Genet A 152A:2245–2253. doi:10.1002/ajmg.a.33578
Desai PP, Bunker CH, Ukoli FA, Kamboh MI (2002) Genetic variation in the apolipoprotein D gene among African blacks and its significance in lipid metabolism. Atherosclerosis 163:329–338. doi:10.1016/S0021-9150(02)00012-6
Iolascon A, De Falco L, Borgese F, Esposito MR, Avvisati RA, Izzo P, Piscopo C, Guizouarn H, Biondani A, Pantaleo A, De Franceschi L (2009) A novel erythroid anion exchange variant (Gly796Arg) of hereditary stomatocytosis associated with dyserythropoiesis. Haematologica 94:1049–1059. doi:10.3324/haematol.2008.002873
Karst ML, Herron KJ, Olson TM (2008) X-linked nonsyndromic sinus node dysfunction and atrial fibrillation caused by emerin mutation. J Cardiovasc Electrophysiol 19:510–515. doi:10.1111/j.1540-8167.2007.01081.x
Rebrikov DV, Britanova OV, Gurskaya NG, Lukyanov KA, Lukyanov SA (2000) Mirror orientation selection (MOS): a method for eliminating false positive clones from libraries generated by suppression subtractive hybridization. Nucleic Acids Res 28:E90
Ohta S, Shimekake Y, Nagata K (1996) Molecular cloning and characterization of a transcription factor for the C-type natriuretic peptide gene promoter. Eur J Biochem 242:460–466. doi:10.1111/j.1432-1033.1996.460rr.x
Kiess M, Scharm B, Aguzzi A, Hajnal A, Klemenz R, Schwarte-Waldhoff I, Schäfer R (1995) Expression of ril, a novel LIM domain gene, is down-regulated in Hras-transformed cells and restored in phenotypic revertants. Oncogene 10:61–68
Arola AM, Sanchez X, Murphy RT, Hasle E, Li H, Elliott PM, McKenna WJ, Towbin JA, Bowles N (2007) Mutations in PDLIM3 and MYOZ1 encoding myocyte Z line proteins are infrequently found in idiopathic dilated cardiomyopathy. Mol Genet Metab 90:435–440. doi:10.1016/j.ymgme.2006.12.008
Pashmforoush M, Pomiès P, Peterson KL, Kubalak S, Ross J Jr, Hefti A, Aebi U, Beckerle MC, Chien KR (2001) Adult mice deficient in actinin-associated LIM-domain protein reveal a developmental pathway for right ventricular cardiomyopathy. Nat Med 7:591–597. doi:10.1038/87920
Chen CL, Lin JL, Lai LP, Pan CH, Huang SK, Lin CS (2007) Altered expression of FHL1, CARP, TSC-22 and P311 provide insights into complex transcriptional regulation in pacing-induced atrial fibrillation. Biochim Biophys Acta 1772:317–329. doi:10.1016/j.bbadis.2006.10.017
Haase D, Lehmann MH, Körner MM, Körfer R, Sigusch HH, Figulla HR (2002) Identification and validation of selective upregulation of ventricular myosin light chain type 2 mRNA in idiopathic dilated cardiomyopathy. Eur J Heart Fail 4:23–31. doi:10.1016/S1388-9842(01)00226-4
Acknowledgments
We thank Sema Bilgic (PhD) and Mehves Poda (PhD) for their contributions in the application of RISH technique also Neslihan Coban (PhD student) for her contributions in selections of positive clones from cDNA library and isolations of plasmid DNA. This study was supported by State Planning Organization of Turkey and Scientific Research Projects Coordination Unit of Istanbul University (Project numbers: T–1062/19022001, T–901/02062006, BYP-3135, ACIP-3107).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2012_1653_MOESM1_ESM.doc
Supplementary Table 1 (Online Resource 1). The determinations of differential expressed genes in heart according to skeletal muscle using published mouse microarray dataset in GEO database. Supplementary material 1 (DOC 355 kb)
11033_2012_1653_MOESM2_ESM.doc
Supplementary Table 2 (Online Resource 2). The detailed information of the probes in Northern Blot and RISH techniques. Supplementary material 2 (DOC 39 kb)
Rights and permissions
About this article
Cite this article
Komurcu-Bayrak, E., Ozsait, B. & Erginel-Unaltuna, N. Isolation and analysis of genes mainly expressed in adult mouse heart using subtractive hybridization cDNA library. Mol Biol Rep 39, 8065–8074 (2012). https://doi.org/10.1007/s11033-012-1653-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1653-5