Abstract
The complementary sex determination (csd) gene is the primary gene determining the gender of honey bees (Apis spp). In this study we analyzed the polymorphism of csd gene in six Apis mellifera subspecies. The genomic region 3 of csd gene in these six A. mellifera was cloned, and identified. A total of 79 haplotypes were obtained from these six subspecies. Analysis showed that region 3 of csd gene has a high level of polymorphism in all the six A. mellifera subspecies. The A. m. anatolica subspecies has a slightly higher nucleotide diversity (π) than other subspecies, while the π values showed no significant difference among the other five subspecies. The phylogenetic tree showed that all the csd haplotypes from different A. mellifera subspecies are scattered throughout the tree, without forming six different clades. Population differentiation analysis showed that there are significant genetic differentiations among some of the subspecies. The NJ phylogenetic tree showed that the A. m. caucasica and A. m. carnica have the closest relationship, followed by A. m. ssp, A. m. ligustica, A. m. carpatica and A. m. anatolica that were gathered in the tree in turn.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
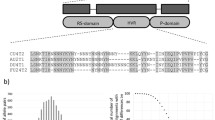
In honey bees, sex is controlled by a mechanism termed complementary sex determination (csd) [1, 2]. Haploids containing a single csd allele will develop into normal males (drones), while diploids, heterozygous for csd, become females. When csd genes are homozygous, a bee will develop into a diploid male, which is consumed at the larval stage by workers [3]. Therefore, nucleotide mutations leading to heterozygotes at csd gene are selectively favoured. The complementary sex determination mechanism of honey bee remained a hypothesis for a long time [4]. In 2003, Beye et al. cloned the csd gene from Apis mellifera by positional cloning [5]. They found that no transcription differences existed between the two sexes but suppression of csd in females with double stranded RNA for csd resulted in male phenotypes. Csd encodes an arginine serine-rich (SR) type protein, which contains an RS domain in the middle and a proline-rich region at its C terminus, between these two domains is a hypervariable region that differs highly between alleles, and has a variable number of asparagine/tyrosine repeats. The honey bee csd protein is homologous to the Drosophila Tra protein, which is involved in Drosophila sex determination [5].
Previous research has demonstrated that the csd genes of three honey bee species (A. mellifera, A. cerana and A. dorsata) have evolved under balancing selection, and several parts of the coding region are possible targets of selection [6–10]. Moreover, the polymorphic level is approximately seven times higher in the csd coding region than in the neutral regions [7].
Although much evolutionary research on the csd gene was conducted in A. mellifera, A. cerana and A. dorsata, no research on csd has yet been conducted in different subspecies of A. mellifera. A. mellifera includes several different geographic subspecies, which arose through long-term natural selection in different geographic environments, and it is worth investigating whether the csd gene is polymorphic among these subspecies.
In this study, we analyzed the polymorphism of csd gene in six A. mellifera subspecies, A. m. anatolica, A. m. caucasica, A. m. carnica, A. m. carpatica, A. mellifera ssp., A. m. ligustica. In A. mellifera, the csd gene contains nine exons, which form three clusters separated by two large introns [5]. The genomic region of the third cluster (from exon six to nine) has the highest polymorphism compared with the other two regions. Therefore, we chose region 3 to study its polymorphism.
Materials and methods
Sample collection
All samples were collected from the bee breeding apiary of the Honeybee Research Institute, Jilin province, China, with 50 workers for each subspecies. Every subspecies is reproductively isolated from other subspecies through artificial breeding. The samples were first collected into 95% ethanol and then stored frozen at −70°C until further use.
DNA extraction
Total genomic DNA was extracted from the cephalothorax of each sampled bee according to the protocol of the Animal Genomic DNA Extraction Kit (BEST ALL-HEAL LLC, NY, USA).
PCR and sequencing
The primers used for amplifying region 3 of the csd gene in this study were designed by consulting those reported by Cho et al. [9] with some modifications. The primers are 5′ AATTGGATTTATTAATATAATTTATTATTCAGG 3′ (forward) and 5′ ATYTCATTATTCAATATGTTNGCATCA 3′ (reverse). The high fidelity LA Tag DNA polymerase (BEST ALL-HEAL LLC, NY, USA) were used for all PCR reactions. PCR conditions were: denaturation at 94°C for 3 min, followed by 30 cycles of 94°C for 30 s, annealing at 54°C for 30 s and extension at 72°C for 2 min, with a final extension at 72°C for 10 min. PCR products were purified using DNA GEL EXTRACTION kits (Sangon, Shanghai, China) and cloned into the pEASY-T3 vector (Transgene, Beijing, China). To obtain as many csd alleles as possible, the genomic region 3 of the csd gene was cloned from cephalothorax of each sampled worker bee, and 1–3 clones of each cloned fragment were subjected to double-strand sequencing. Single-sequencing reads were assembled using the Seqman program in the DNAstar software [11].
Sequence analysis
The exons, introns and coding regions of our sequences were determined by consulting those sequences of the genomic region 3 of A. mellifera csd gene reported by Cho et al. [9] and cDNA sequences of A. mellifera csd gene reported by Hasselmann and Beye [10]. Nucleotide sequence alignments were performed with Clustal X version 1.8 [12], and alignment results were adjusted manually for obvious alignment errors.
DAMBE 4.1.19 [13] was used to identify haplotypes. Phylogenetic trees were constructed using MEGA version 4.0 program [14]. The minimal evolution (ME) method and Kimura’s 2-parameter distances were adopted to obtain an unrooted tree with 2,000 bootstrap replications. Nucleotide diversity (π) was calculated by using DNAsp5.0 program [15]. Two tailed Z test was adopted to detected significant difference between two π values. Fst distances were computed using ARLEQUIN 3.0 software [16]. Kimura’s 2-parameter genetic distance was calculated by MEGA 4.0.
Results
Polymorphism of the csd haplotypes in six A. mellifera subspecies
We cloned the genomic region 3 of the csd gene from six A. mellifera subspecies, A. m. anatolica, A. m. caucasica, A. m. carnica, A. m. carpatica, A. m. ssp., A. m. ligustica. After sequencing, we obtained 6, 10, 19, 14, 28 and 7 haplotypes from these subspecies (table 1), respectively. There are a total of 79 haplotypes, and five haplotypes exist in two subspecies. We compared the nucleotide diversity (π) of the csd gene of the six A. mellifera subspecies. As shown in Table 1, the csd gene has a high level of polymorphism in all the six subspecies. Of them, the π value of the A. m. anatolica subspecies is the highest among the six subspecies. It is significantly higher than that of the A. m. ssp. subspecies (two tailed Z test, P < 0.05), but failed to be significant compared with π values of the other four subspecies (two tailed Z test, P < 0.1). Except for the A. m. anatolica subspecies, there is no significant difference in π values among the other five subspecies (two tailed Z test, P > 0.05) Table 2.
Phylogenetic tree of all the csd haplotypes
A genealogy tree was constructed based on all the haplotypes from the six subspecies. As shown in Fig. 1, all the csd haplotypes from different A. mellifera subspecies mainly form two clades in the tree, and they are mixed in the genealogy tree while not forming different clades according to subspecies.
The gene genealogy of csd haplotypes in region 3 of six A. mellifera subspecies. The minimum evolution method and Kimura’s two parameter distances are used to construct the tree. Bootstrap percentages are shown on internal branches. The scale bar represents the number of nucleotide changes per site. Amcau A. m. caucasica, Amssp A. m. ssp., Amcan A. m. carnica, Amcap A. m. carpatica, Amlig A. m. ligustica, Amana A. m. anatolica
Hypervariable region of CSD proteins in six A. mellifera subspecies
We analyzed the amino acid sequence of the csd sequences in these subspecies, since this region contains a hypervariable region critical for determining the specificity of csd alleles. The exons, introns and coding sequences on the obtained csd haplotypes were determined by consulting sequences of A. mellifera csd gene reported by Cho et al. [9] and Hasselmann et al. [10]. Similar to that in A. mellifera, A. cerana and A. dorsata, the coding region of this part also contains an RS domain at the N terminal, a P-rich domain at the C terminal and a hypervariable region between these two domains. The hypervariable region is rich in asparagine (N) and tyrosine (Y), and they form a basic (N) 1–4Y repeats terminated mainly with KK, KQ in each haplotype (supplementary Fig. S1).
Genetic differentiation among six A. mellifera subspecies
The pairwise Fst values between different subspecies range from 0.02570 to 0.23848, and of them, Fst value between A. m. anatolica and A. m. carnica is the highest, while that between A. m. carpatica and A. m. caucasica is the lowest. From the Fst values, it indicated that the genetic differentiation levels between A. m. caucasica and all other five subspecies is very low. The genetic differentiation between A. m. carpatica and both A. m. ligustica and A. m. anatolica is also not significant (P > 0.05). Except for these subspecies pairs, the remaining pairs showed significant genetic differentiation (P < 0.001).
Genetic distance and phylogenesis of six A. mellifera subspecies
The kimura’s 2-parameter genetic distances between different subspecies range from 0.04216 to 0.06415. Of them, the genetic distance between A. m. anatolica and A. m. carnica is the largest, while that between A. m. ssp. and A. m. caucasica is the nearest.
When a NJ tree was constructed based on Kimura’s 2-parameter genetic distance, A. m. caucasica and A. m. carnica formed a clade first, followed by A. m. ssp, A. m. ligustica, A. m. carpatica and A. m. anatolica that were gathered in the tree in turn (Fig. 2).
Discussion
Previous studies have shown that the csd genes in A. mellifera, A. cerana and A. dorsata have a very high level of polymorphism [9, 10]. In this study we found that the csd gene in the six A. mellifera subspecies also shows a high level of polymorphism. This result further confirms that the complementary sex determination mechanism is common for all the honey bee species [1].
Cho and Hasselmann found that the (N)1–4Y repeats and (KHYN) 1–4 motif are two types of important repeat sequences in the hypervariable region of CSD protein, but the (KHYN)1–4 motif exists in A. cerana and A. dorsata while not in A. mellifera [8, 9]. In this study, we also did not find (KHYN)1–4 motif in any of the six A. mellifera subspecies—it may have existed in the ancient csd alleles, but was lost in A. mellifera during evolution, since the A. dorsata and A. cerana speciated prior to A. mellifera in their evolutionary history.
Some researchers have divided A. mellifera into four evolutionary clades based on the analysis of mitochondrial genes [17–22]. They are the African type, western and northern Europe type, eastern Mediterranean type, and the near Eastern type. According to these researchers, A. m. carnica, A. m. carpatica and A. m. ligustica belong to the Eastern Mediterranean type, A. m. caucasica, A. m. ssp. and A. m. anatolica belong to the near East type. In this study, the structure of the NJ tree of six A. mellifera subspecies based on the Kimura’s 2-parameter genetic distances did not match the above geographical division. For example, the A. m. anatolica subspecies did not group with the A. m. caucasica and A. m. ssp. subspecies to form a clade, although they are meant to belong to the same geographical type. One reason may be that other different strains were once introduced into this subspecies during artificial breeding, causing an increase in the polymorphism of csd gene in this subspecies, as well as a relatively distant relationship with the A. m. caucasica and A. m. ssp. subspecies.
Apis mellifera carpatica was once considered to be a strain belonging to A. m. carnica subspecies [23]. But our pairwise Fst analysis indicated that there is significant genetic differentiation between these two subspecies. Meanwhile, Liu also obtained significant genetic differentiation between them by analyzing the polymorphism difference of malate dehydrogenase between these two subspecies [24]. Therefore, the A. m. carpatica and A. m. carnica subspecies maybe really have a large genetic difference, despite their close geographical distribution.
Our analysis showed that the genetic differentiation level between A. m. caucasica and all other five subspecies is very low; this may suggest that A. m. caucasica is a transitional or intermediate subspecies among the six subspecies.
In conclusion, we found a high level of polymorphism of the csd gene in six A. mellifera subspecies by molecular analysis, and detected significant genetic differences between some of these subspecies pairs. The present study has further verified that gender in bees is determined by the csd gene, and has expanded our understanding of sex determination mechanism in bee subspecies.
References
Cook JM (1993) Sex determination in the hymenoptera: a review of models and evidence. Heredity 71:421–435. doi:10.1038/hdy.1993.157
Heimpel GE, De Boer JD (2008) Sex determination in the hymenoptera. Annu Rev Entomol 53:209–230. doi:10.1146/annurev.ento.53.103106.093441
Woyke J (1963) What happens to diploid drone larvae in a honeybee colony. J Apic Res 2:73–76
Whiting P (1943) Multiple alleles in complementary sex determination of Habrobracon. Genetics 28:365–382
Beye M, Hasselmann M, Fondrk MK et al (2003) The Gene csd is the primary signal for sexual development in the honeybee and encodes an SR-Type protein. Cell 114(4):419–429. doi:10.1016/S0092-8674(03)00606-8
Hasselmann M, Beye M (2004) Signatures of selection among sex-determining alleles of the honey bee. Proc Natl Acad Sci USA 101(14):4888–4893. doi:10.1073/pnas.0307147101
Hasselmann M, Beye M (2006) Pronounced differences of recombination activity at the sex determination locus of the honeybee, a locus under strong balancing selection. Genetics 174(3):1469–1480. doi:10.1534/genetics.106.062018
Charlesworth D (2004) Sex determination: balancing selection in the honey bee. Curr Biol 14:568–569. doi:10.1016/j.cub.2004.07.014
Cho S, Huang ZY, Green DR et al (2006) Evolution of the complementary sex-determination gene of honey bees: balancing selection and trans-species polymorphisms. Genome Res 16:1366–1375. doi:10.1101/gr.4695306
Hasselmann M, Vekemans X, Pflugfelder J et al (2008) Evidence for convergent nucleotide evolution and high allelic turnover rates at the complementary sex determiner gene of Western and Asian honeybees. Mol Biol Evol 25(4):696–708. doi:10.1093/molbev/msn011
Burland TG (2000) DNASTAR’s Lasergene sequence analysis software. Methods Mol Biol 132:71–91. doi:10.1385/1-59259-192-2:71
Thompson JD, Gibson TJ, Plewniak F et al (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. doi:10.1093/nar/25.24.4876
Xia X, Xie Z (2001) DAMBE: software package for data analysis in molecular biology and evolution. J Hered 92:371–373. doi:10.1093/jhered/92.4.371
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599. doi:10.1093/molbev/msm092
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25(11):1451–1452. doi:10.1093/bioinformatics/btp187
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Smith DR, Slaymaker A, Palmer M, Kaftanoglu O (1997) Turkish honey bees belong to the east Mediterranean mitochondrial lineage. Apidologie 28:269–274. doi:10.1051/apido:19970503
Franck P, Garnery L, Solignac M, Cornuet JM (1998) The origin of west European subspecies of honeybees (Apis mellifera): new insights from microsatellite and mitochondrial data. Evolution 52:1119–1134
Franck P, Garnery L, Solignac M, Cornuet JM (2000) Molecular confirmation of a fourth lineage in honeybees from Middle-East. Apidologie 31:167–180. doi:10.1051/apido:2000114
Franck P, Garnery L, Loiseau A et al (2001) Genetica diversity of the honeybee in Africa: microsatellite and mitochondrial data. Heredity 86:420–430. doi:10.1046/j.1365-2540.2001.00842.x
Colltet T, Ferreira KM, Arias MC et al (2006) Genetic structure of Africanized honeybee populations (Apis mellifera L.) from Brazil and Uruguay viewed through mitochondrial DNA CO I-CO II patterns. Heredity 97:329–335. doi:10.1038/sj.hdy.6800875
Cánovas F, La De, Ru′a P, Serrano J, Galia′n J (2008) Geographical patterns of mitochondrial DNA variation in Apis mellifera iberiensis (Hymenoptera: Apidae). Zool Syst Evol Res 46(1):24–30. doi:10.1111/j.1439-0469.2007.00435.x
Ruttner F (1986) Geographical variability and classification. In: Rinderer TE (ed) Bee genetics and breeding. Academic Press Inc, London, pp 49–50
Liu YH (2002) Genetic variability of MDHII gene in six subspecies of Apis mellifera. Acta Entomol Sinica 45(2):188–192 ISSN: 0454-6296.0.2002-02-006
Acknowledgments
We are grateful to Doctor Aung Si and Professor Shaowu Zhang for improving the english of the manuscript. This work was supported by the China Agriculture Research System (No. CARS-45), the National Natural Science Foundation of China (No. 31060327) and the China Postdoctoral Science Foundation (20100481191).
Author information
Authors and Affiliations
Corresponding author
Additional information
Zilong Wang and Zhiyong Liu contributed equally to this paper.
All the sequences obtained from this study have been submitted to GenBank under accession numbers HQ915780-HQ915863.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2011_1069_MOESM1_ESM.eps
Fig. S1. Amino acid sequence alignment of the hypervariable regions of csd haplotypes in six A. mellifera subspecies. ”-” indicates alignment gaps. (EPS 1918 kb)
Rights and permissions
About this article
Cite this article
Wang, Z., Liu, Z., Wu, X. et al. Polymorphism analysis of csd gene in six Apis mellifera subspecies. Mol Biol Rep 39, 3067–3071 (2012). https://doi.org/10.1007/s11033-011-1069-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-1069-7