Abstract
Methionine synthase reductase (MTRR) gene is involved in tumorigenesis by regulating DNA methylation through activation of methionine synthase (MTR). MTRR is polymorphic at nucleotide 66 (A-to-G) and the resulting variant enzyme has a lower affinity for MTR. The reported associations of MTRR A66G polymorphism with cancer risk are contradictory. Therefore, we performed a meta-analysis to better assess the associations, including 18,661 cases and 27,678 controls from 35 studies. Crude ORs with 95% CIs were used to assess the strength of association between the MTRR A66G polymorphism and cancer risk. The pooled ORs were performed for homozygote model (GG vs. AA), heterozygote model (GG vs. GA), recessive genetic model (GG vs. GA + AA), and dominant genetic model (GG + GA vs. AA), respectively. Overall, results indicated that the G allele and GG variant genotypes were associated with a significantly increased cancer risk (G vs. A: OR, 1.039; 95% CI, 1.009–1.078; homozygote model: OR, 1.094; 95% CI, 1.006–1.191). In subgroup analysis by ethnicity, significant increased risks were found among Asians with G allele (G vs. A: OR, 1.063; 95% CI, 1.011–1.119; homozygote model: OR, 1.189; 95% CI, 1.055–1.341; recessive model: OR, 1.197; 95% CI, 1.068–1.341). For stratification analysis, the cancer types with fewer than three studies were categorized into “other cancers”, and the results indicated that there was a significant elevated cancer risk in “other cancers” in all genetic models, not in colorectal cancer, lymphoid leukemia or breast cancer. In summary, our study suggests that the MTRR A66G polymorphism is a potential biomarker for cancer risk.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One-carbon metabolism, also referred as folate-mediated one-carbon metabolism, is a network of biological reactions with a critical role in DNA methylation and synthesis, and an impact on both epigenetic and genetic pro-carcinogenic processes [1]. DNA methylation is critical for regulating gene expression. The mechanism by which abnormal DNA methylation leads to carcinogenesis is complex. Methionine is an essential amino acid and a precursor of S-adenosylmethionine, a universal methyl-group donor involved in methylation reactions, including DNA methylation [2]. Both methionine synthase reductase (MTRR) and methionine synthase (MTR) regulate the reaction that produces methionine through the irreversible transfer of a methyl group from 5-methyltetrahydrofolate. MTR is maintained in its active form by MTRR, an enzyme that regenerates a functional MTR via reductive methylation.
A genetic polymorphism at nucleotide 66 (A-to-G) of the MTRR gene is functional, but the variant enzyme has a lower affinity for MTR [3]. Several case–control studies have evaluated the association between the genetic polymorphism and cancer risk, with inconclusive or controversial results. For MTRR A66G, some studies have reported elevated homocysteine levels for carriers of the homozygote wildtype genotype (AA), compared to other genotypes [4, 5], while others have not [6]. However, in subsequent investigations, the 66GG genotype was associated with an increased risk of colorectal cancer among Japanese compared to the GA + AA genotype [7], and another study found an increased risk for the GG genotype among white populations only [8]. Moreover, it was shown that carrying the G allele was associated with a marginally decreased risk of acute lymphoblastic leukemia (ALL) [9], but another study suggested that the MTRR polymorphism was less clearly associated with susceptibility to ALL [10].
A large number of molecular epidemiological studies have been conducted to evaluate the role of MTRR polymorphisms in different kinds of neoplasm. However, the association between the polymorphisms and cancer risk is still controversial. To clarify this issue, we performed a meta-analysis with subgroup analysis from all eligible studies, to assess the association of the MTRR A66G polymorphism with cancer risk.
Materials and methods
Study identification and selection
Before the study, inclusion criteria were defined as: (a) articles evaluating the association between MTRR A66G polymorphism and cancer risk; (b) studies with case–control design; (c) sufficient data to estimate an odds ratio (OR) with its 95% confidence interval (95% CI).
A literature search of PubMed and China National Knowledge Infrastructure (updated to 2010/07/16) was conducted using the terms: “MTRR” or “methionine synthase reductase”, “polymorphism(s)”, and “cancer” or “carcinoma” or “neoplasm”, without restriction on language. All the searched studies were retrieved by two of the authors, and their bibliographies were checked for other relevant publications. Reference lists of reviews and retrieved articles were also searched to find additional eligible studies. When an article reported results on different racial descent subpopulations or tumor types, we treated each subpopulation or tumor as a separate comparison. Any disagreement was resolved by discussion between the two authors.
Data extraction
Information was carefully and independently extracted from all eligible publications by two of the authors, according to the inclusion criteria. For each study, the collected characteristics were: first author’s last name, journal, year of publication, ethnicity and country of study population (mixed or unknown populations were categorized as an “others” group), source of control groups (population- or hospital-based controls), demographics, numbers of genotyped cases and controls, methods for genotyping, MTRR polymorphism genotyping information.
Statistical analysis
The strength of association between the MTRR A66G and cancers was measured by OR with 95% CI. The statistical significance of the pooled OR was determined using the Z-test. The meta-analysis assessed association between allele G and cancer risk compared to allele A (G vs. A), as well as homozygote comparison (GG vs. AA), heterozygote comparison (GG vs. GA), recessive genetic model (GG vs. GA + AA), and dominant genetic model (GG + GA vs. AA) comparison. Stratification analysis was performed by cancer type (if one cancer type contained fewer than three individual studies, it was combined into an “other cancers” group), ethnicity and study designs (hospital-based studies and population-based studies).
A chi-square test was used to determine if the distribution of genotypes among controls conformed to Hardy–Weinberg equilibrium (HWE), with P value <0.05 signifying a departure from HWE. Weighted mean was used to calculate mean allele prevalence in controls using SPSS 13.0 software for Windows. The Q-test was used to investigate the degree of heterogeneity between studies, with a P value >0.05 indicating lack of heterogeneity. In cases of no statistical heterogeneity, a fixed-effects model was used to estimate the summery OR (Mantel–Haenszel method) [11]; otherwise, the random-effects model (the DerSimonian–Laird method) was used [12].
Potential publication bias was examined visually in a funnel plot of log [OR] against its standard error (SE), and the degree of asymmetry tested by Egger’s test (P < 0.05 was significant publication bias) [13]. Sensitivity analysis was performed by omitting each study in turn to assess the results stability. We also did cumulative meta-analysis to evaluate the trend of summary ORs (95% CIs) by year of publication. All statistical tests were two-sided. Software STATA version 10.0 (Stata Corporation, College Station, TX, USA) was used for all analyses.
Results
Study characteristics
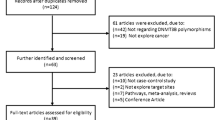
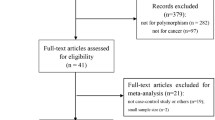
We obtained 61 articles after searching and screening based on our eligibility criteria. During data extraction, 26 articles were excluded because they did not provide allele frequencies needed for OR calculation, leaving 35 eligible studies that had assessed the association between MTRR A66G and cancer risk using human genomic DNA samples. Overall, 18,661 cancer patients and 27,678 controls were distributed among the 35 eligible studies.
We established a database from the extracted information from each eligible article (Table 1). All were case–control studies, including seven colorectal cancer studies, six lymphoid leukemia studies, and six breast cancer studies, with the rest in the “other cancers” group. Cancers were confirmed histologically or pathologically in most studies. Age-matching was performed by 29 articles and sex-matching in 28. Of the 35 studies, only 19 discussed quality control of genotyping, such as blindness to the case–control status, randomly repeated assays, or validation using a different genotyping method. In addition, 17 studies investigated interactions between polymorphisms and environmental factors or the combined effect with other genes.
Quantitative synthesis
Significant differences were found in the variant 66G allele frequency between the two major ethnicities (Asian, 28.9%; 95% CI, 28.4–29.5%; European, 53.0%; 95% CI, 52.4–53.6%; P = 0.001). Table 2 summarizes results of the MTRR A66G polymorphism and cancer risk. A significant association between the MTRR A66G polymorphism and cancer risk was found, for an overall OR for the allele G versus allele A of 1.039 (P = 0.043; 95% CI, 1.009–1.078; P heterogeneity = 0.021; Fig. 1). In addition, individuals carrying the MTRR 66GG genotype had a significantly increased cancer risk compared to individuals with the 66AA genotype (OR, 1.094; P = 0.037; 95% CI, 1.006–1.191; P heterogeneity = 0.004; Fig. 2).
Forest plot of cancer risk associated with the MTRR A66G polymorphism (G vs. A) in overall populations. The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the study-specific weight (inverse of the variance). Diamonds represent the pooled OR and 95% CI
Forest plot of cancer risk associated with the MTRR A66G polymorphism (GG vs. AA) in overall populations. The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the study-specific weight (inverse of the variance). The diamonds represent the pooled OR and 95% CI
We also performed subgroup analysis stratified by ethnicity, study design, and cancer type. By ethnicity, the G allele was associated with a significantly increased cancer risk in Asian populations (OR, 1.063; P = 0.018; 95% CI, 1.011–1.119; P heterogeneity = 0.328; Fig. 3). A marginally significant association between the A66G polymorphism and increased cancer risk was also detected in Asians under homozygote (GG vs. AA: OR, 1.189; P = 0.005; 95% CI, 1.055–1.341; P heterogeneity = 0.191), and recessive model comparison (GG vs. GA/AA: OR, 1.197; P = 0.002; 95% CI, 1.068–1.341; P heterogeneity = 0.106). By different study designs for Asian populations, the 66GG genotype led to a significantly increased cancer risk in population-based studies under allelic frequency (OR, 1.089; P = 0.011; 95% CI, 1.020–1.163; P heterogeneity = 0.508), homozygote model (OR, 1.283; P = 0.002; 95% CI, 1.096–1.501; P heterogeneity = 0.661), and recessive model comparison (OR, 1.263; P = 0.003; 95% CI, 1.085–1.471; P heterogeneity = 0.8). However, cancer risk decreased non-significantly in hospital-based studies under dominant model comparison (OR, 0.484; P = 0.001; 95% CI, 0.330–0.709; P heterogeneity = 0.001). In the subgroup analysis stratified by tumor type, the MTRR G allele was associated with an increased risk of “other cancers” compared to the A allele (OR, 1.08; P = 0.000; 95% CI, 1.035–1.127; P heterogeneity = 0.102). Also for “other cancers”, we found that the variant genotypes were associated with a significantly increased cancer risk using all genetic models (homozygote comparison: OR, 1.196, P = 0.001, 95% CI: 1.093–1.310, P heterogeneity = 0.079; heterozygote comparison: OR, 1.089, P = 0.015, 95% CI: 1.017–1.166, P heterogeneity = 0.193; dominant model: OR, 1.111, P = 0.001, 95% CI: 1.042–1.185, P heterogeneity = 0.077; recessive model: OR, 1.108, P = 0.007, 95% CI: 1.028–1.194, P heterogeneity = 0.199). No significant association was found for other tumor sites.
Forest plot of cancer risk associated with the MTRR A66G polymorphism (G vs. A) in Asian populations. The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the study-specific weight (inverse of the variance). The diamonds represent the pooled OR and 95% CI
Sensitivity analysis and cumulative meta-analysis
Pooled ORs were consistently significant in Asian populations or European populations by omitting one study or one tumor at a time under the homozygote and dominant genetic model comparison, suggesting robustness of our results (data not shown). In the cumulative meta-analysis, the pooled ORs tended to be stable and the associations tended toward significant associations with accumulation of more data over time.
Publication bias
Funnel plots were generated to assess publication bias. The Egger’s test was performed to statistically evaluate funnel plot symmetry. The results showed no evidence of publication bias (P = 0.58; Fig. 4).
Discussion
Similar to MTR, MTRR is a critical enzyme for the biosynthesis of methionine, which is the precursor for methylation reactions. MTRR is also involved in the regeneration of tetrahydrofolate for nucleotide biosynthesis. Changes in this enzyme may significantly influence DNA synthesis, methylation and repair. The A66G single nucleotide polymorphism at codon 22 is one of the most common polymorphisms in the MTRR gene, and the variant MTRR enzyme has a lower affinity for MTR [14], and is inconsistently associated with elevated blood or plasma homocysteine levels [15]. DNA hypomethylation is an early and consistent event in cancer development, marked by an elevation in homocysteine [16]. The MTRR variant G allele-bearing genotype has been significantly associated with an increased risk of hepatocellular carcinoma [17] and esophageal squamous cell carcinoma [18], however other studies suggested that this MTRR polymorphism was less clearly associated with susceptibility to cancer [10, 19, 20]. The lack of concordance across many of these studies reflects limitations in the studies, such as small sample sizes, ethnic differences, and poor research methodology. Meta-analysis is a powerful tool for summarizing the results from different studies with enhanced precision, by producing a single estimate of the major effects. It can overcome the problem of small sample size and inadequate statistical power in genetic studies of complex traits, and it can provide more reliable results than a single case–control study [21].
Our meta-analysis, including 18,661 cases and 27,678 controls from 35 published case–control studies, explored the association between the MTRR A66G polymorphism and cancer risk. Overall, we found evidence that the variant genotypes of MTRR were associated with a significant increase in overall cancer risk using a G vs. A, or a dominant model comparison. Interestingly, 66GG was associated with a significantly increased cancer risk in Asian, but not in European populations under the recessive genetic model and by homozygote comparison. Many factors may contribute to the finding that the same polymorphism has different impacts in different ethnic populations. This may be due to genetic trait differences, since the MTRR A66G polymorphism showed distinct frequencies among different ethnic groups. For example, the G-allele frequency among controls was 0.29 in Asian populations and 0.53 in European populations, suggesting a possible ethnic difference. Nonetheless, different linkage disequilibrium patterns usually exist in different populations. The MTRR A66G polymorphism may be in close linkage with different nearby causal variants in one ethnic population but not in others. Other factors such as selection bias and different matching criteria may also play an important role in the discrepancy. Finally, the influence of the genetic variant may be masked by the presence of other, as-yet unidentified causal genes involved in the tumor formation. Different populations may have differences in dietary intake of nutrients, some of which affect cancer development. Thus, further investigations are warranted to validate ethnic differences in the effect of this functional polymorphism on cancer risk, especially in Europeans.
Although the pooled results were robust in Asian populations, they should still be treated with caution because of different study designs. When stratified separately by population-based and hospital-based studies, inverse results were observed in Asian populations, that is, 66GG was associated with increased cancer risk in the population-based studies, and decreased cancer risk in the hospital-based studies. The pooled results in Asian populations may be a spurious finding, and larger population-based studies are required to further clarify the association between the MTRR A66G polymorphism and cancer susceptibility in Asian populations. These inconsistent results suggest that selection bias is an important issue for studies of the genetic cause of cancer. Hospital-based studies have a high risk of producing unreliable results because hospital-based controls may not always authentically represent the general population, especially when the genotypes under investigation are expected to affect disease conditions that might be seen in the hospital-based controls. Thus, the use of proper and representative population-based control participants is of great importance in reducing bias in such genotype association studies.
Subgroup analysis by cancer type showed an increased cancer risk for “other cancers” by homozygote or heterozygote comparison, as well as using the dominant and recessive models. However, we failed to find any significant association between the MTRR A66G polymorphism and colorectal cancer, breast cancer, or lymphoid leukemia in any comparison models. Although the reason for these discrepancies is not completely understood, many factors may be contributing. First, the MTRR A66G polymorphism might have a different role in different cancers. Second, studies with small sample sizes may be underpowered for detecting a small but real association. In the “other cancers” groups of our meta-analysis, only one or two studies were available for each specific cancer type, and they had very small case–control numbers, so larger studies are needed to confirm this relationship. Third, the result of each study might be influenced by gene–gene or gene–environment interactions, because environmental factors or other genes may predominate in the development of cancer. Consequently, additional prospective studies are needed to clarify whether the MTRR A66G polymorphism truly affects different types of cancer in different ways.
In this meta-analysis, 17 of 35 eligible studies investigated interactions between polymorphisms and environmental factors, and 15 studied gene–gene interactions. The results were conflicting. Not all analyzed the same genetic or environmental factors, which included folate-mediated one-carbon metabolism genes (MTHFR, TYMS, MTR, SHMT1, CBS), vitamin B6 intake, vitamin B12 intake, folate intake, methionine intake, cigarette consumption, and alcohol status. Environmental or genetic factors may have different effects on different cancer types or ethnicities. The obtained data may not reflect intake as accurately as other methods, such biological markers. Consequently, large-scale, well-designed, population-based studies are required to investigate gene–gene and gene–environment interactions on the MTRR A66G polymorphism and cancer risk.
Heterogeneity is a potential problem when interpreting the results of all meta-analysis. Significant between-study heterogeneity existed in homozygote (GG vs. AA), recessive genetic model (GG vs. GA + AA), and dominant genetic model (GG + GA vs. AA) comparisons. After subgroup analyses by ethnicity, the heterogeneity was effectively decreased or removed. The reason might be that differences of genetic backgrounds and the environment existed among different ethnicities.
Some limitations of this meta-analysis should be addressed. First, not having the original data for the reviewed studies limited our evaluation of potential gene–gene, gene–environment, or even different polymorphism loci of the same gene, which all may affect cancer risk. Second, because of the data limitation, we did not perform stratification analysis by age, sex, smoking status, drinking status, obesity, or other variables. This might have caused serious confounding bias. Third, although the funnel plot and Egger’s test showed no publication bias, selection bias may have occurred because only published studies were retrieved. The number of published studies was not sufficiently large for a comprehensive analysis, particularly for a specific cancer type. Nonetheless, advantages in our meta-analysis should also be acknowledged. First, a substantial number of cases and controls were pooled from different studies, which significantly increased the statistical power of the analysis. Second, no publication bias was detected, indicating that the pooled results should be reliable. Third, the quality of case–control studies included in the meta-analysis was satisfactory based on our selection criteria.
In summary, this meta-analysis identified evidence of an association between the MTRR A66G polymorphism and cancer risk, supporting the hypothesis that the MTRR A66G may cause an increased risk of cancer, especially in people of Asian descent. As the biological role of MTRR A66G SNP is still unclear, predicting the effect of MTRR A66G on cancer risk in European populations is difficult. Future studies with large sample sizes and tissue-specific biological characterization are required to investigate the biological mechanism and function of the MTRR A66G polymorphism.
References
Stern LL, Mason JB, Selhub J, Choi SW (2000) Genomic DNA hypomethylation, a characteristic of most cancers, is present in peripheral leukocytes of individuals who are homozygous for the C677T polymorphism in the methylenetetrahydrofolate reductase gene. Cancer Epidemiol Biomarkers Prev 9(8):849–853
Ma J, Stampfer MJ, Christensen B, Giovannucci E, Hunter DJ, Chen J, Willett WC, Selhub J, Hennekens CH, Gravel R, Rozen R (1999) A polymorphism of the methionine synthase gene: association with plasma folate, vitamin B12, homocyst(e)ine, and colorectal cancer risk. Cancer Epidemiol Biomarkers Prev 8(9):825–829
Olteanu H, Munson T, Banerjee R (2002) Differences in the efficiency of reductive activation of methionine synthase and exogenous electron acceptors between the common polymorphic variants of human methionine synthase reductase. Biochemistry 41(45):13378–13385
Gaughan DJ, Kluijtmans LA, Barbaux S, McMaster D, Young IS, Yarnell JW, Evans A, Whitehead AS (2001) The methionine synthase reductase (MTRR) A66G polymorphism is a novel genetic determinant of plasma homocysteine concentrations. Atherosclerosis 157(2):451–456
Geisel J, Zimbelmann I, Schorr H, Knapp JP, Bodis M, Hubner U, Herrmann W (2001) Genetic defects as important factors for moderate hyperhomocysteinemia. Clin Chem Lab Med 39(8):698–704. doi:10.1515/CCLM.2001.115
O’Leary VB, Parle-McDermott A, Molloy AM, Kirke PN, Johnson Z, Conley M, Scott JM, Mills JL (2002) MTRR and MTHFR polymorphism: link to Down syndrome? Am J Med Genet 107(2):151–155. doi:10.1002/ajmg.10121
Matsuo K, Hamajima N, Hirai T, Kato T, Inoue M, Takezaki T, Tajima K (2002) Methionine synthase reductase gene A66G Polymorphism is associated with risk of colorectal cancer. Asian Pac J Cancer Prev 3(4):353–359
Le Marchand L, Donlon T, Hankin JH, Kolonel LN, Wilkens LR, Seifried A (2002) B-vitamin intake, metabolic genes, and colorectal cancer risk (United States). Cancer Causes Control 13(3):239–248
Gast A, Bermejo JL, Flohr T, Stanulla M, Burwinkel B, Schrappe M, Bartram CR, Hemminki K, Kumar R (2007) Folate metabolic gene polymorphisms and childhood acute lymphoblastic leukemia: a case-control study. Leukemia 21(2):320–325. doi:10.1038/sj.leu.2404474
Koppen IJ, Hermans FJ, Kaspers GJ (2010) Folate related gene polymorphisms and susceptibility to develop childhood acute lymphoblastic leukaemia. Br J Haematol 148(1):3–14. doi:10.1111/j.1365-2141.2009.07898.x
Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22(4):719–748
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7(3):177–188
Egger M, Davey Smith G, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. Br Med J 315(7109):629–634
Brown CA, McKinney KQ, Kaufman JS, Gravel RA, Rozen R (2000) A common polymorphism in methionine synthase reductase increases risk of premature coronary artery disease. J Cardiovasc Risk 7(3):197–200
Feix A, Winkelmayer WC, Eberle C, Sunder-Plassmann G, Fodinger M (2004) Methionine synthase reductase MTRR 66A > G has no effect on total homocysteine, folate, and Vitamin B12 concentrations in renal transplant patients. Atherosclerosis 174(1):43–48. doi:10.1016/j.atherosclerosis.2003.12.036
Jones PA, Laird PW (1999) Cancer epigenetics comes of age. Nat Genet 21(2):163–167. doi:10.1038/5947
Kwak SY, Kim UK, Cho HJ, Lee HK, Kim HJ, Kim NK, Hwang SG (2008) Methylenetetrahydrofolate reductase (MTHFR) and methionine synthase reductase (MTRR) gene polymorphisms as risk factors for hepatocellular carcinoma in a Korean population. Anticancer Res 28(5A):2807–2811
Stolzenberg-Solomon RZ, Qiao YL, Abnet CC, Ratnasinghe DL, Dawsey SM, Dong ZW, Taylor PR, Mark SD (2003) Esophageal and gastric cardia cancer risk and folate- and vitamin B(12)-related polymorphisms in Linxian, China. Cancer Epidemiol Biomarkers Prev 12(11 Pt 1):1222–1226
Suzuki T, Matsuo K, Hasegawa Y, Hiraki A, Wakai K, Hirose K, Saito T, Sato S, Ueda R, Tajima K (2007) One-carbon metabolism-related gene polymorphisms and risk of head and neck squamous cell carcinoma: case-control study. Cancer Sci 98(9):1439–1446. doi:10.1111/j.1349-7006.2007.00533.x
Moore LE, Malats N, Rothman N, Real FX, Kogevinas M, Karami S, Garcia-Closas R, Silverman D, Chanock S, Welch R, Tardon A, Serra C, Carrato A, Dosemeci M, Garcia-Closas M (2007) Polymorphisms in one-carbon metabolism and trans-sulfuration pathway genes and susceptibility to bladder cancer. Int J Cancer 120(11):2452–2458. doi:10.1002/ijc.22565
Yong D, QingQing W, Hua L, Yang LX, QingLing Z, Ying H, QiaoJing Q, HanChao S (2006) Association of uteroglobin G38A polymorphism with IgA nephropathy: a meta-analysis. Am J Kidney Dis 48(1):1–7. doi:10.1053/j.ajkd.2006.03.048
Koushik A, Kraft P, Fuchs CS, Hankinson SE, Willett WC, Giovannucci EL, Hunter DJ (2006) Nonsynonymous polymorphisms in genes in the one-carbon metabolism pathway and associations with colorectal cancer. Cancer Epidemiol Biomarkers Prev 15(12):2408–2417. doi:10.1158/1055-9965.EPI-06-0624
Theodoratou E, Farrington SM, Tenesa A, McNeill G, Cetnarskyj R, Barnetson RA, Porteous ME, Dunlop MG, Campbell H (2008) Dietary vitamin B6 intake and the risk of colorectal cancer. Cancer Epidemiol Biomarkers Prev 17(1):171–182. doi:10.1158/1055-9965.EPI-07-0621
Otani T, Iwasaki M, Hanaoka T, Kobayashi M, Ishihara J, Natsukawa S, Shaura K, Koizumi Y, Kasuga Y, Yoshimura K, Yoshida T, Tsugane S (2005) Folate, vitamin B6, vitamin B12, and vitamin B2 intake, genetic polymorphisms of related enzymes, and risk of colorectal cancer in a hospital-based case-control study in Japan. Nutr Cancer 53(1):42–50. doi:10.1207/s15327914nc5301_5
Steck SE, Keku T, Butler LM, Galanko J, Massa B, Millikan RC, Sandler RS (2008) Polymorphisms in methionine synthase, methionine synthase reductase and serine hydroxymethyltransferase, folate and alcohol intake, and colon cancer risk. J Nutrigenet Nutrigenomics 1(4):196–204. doi:10.1159/000136651
Hazra A, Wu K, Kraft P, Fuchs CS, Giovannucci EL, Hunter DJ (2007) Twenty-four non-synonymous polymorphisms in the one-carbon metabolic pathway and risk of colorectal adenoma in the Nurses’ Health Study. Carcinogenesis 28(7):1510–1519. doi:10.1093/carcin/bgm062
Kim HN, Kim YK, Lee IK, Yang DH, Lee JJ, Shin MH, Park KS, Choi JS, Park MR, Jo DY, Won JH, Kwak JY, Kim HJ (2009) Association between polymorphisms of folate-metabolizing enzymes and hematological malignancies. Leuk Res 33(1):82–87. doi:10.1016/j.leukres.2008.07.026
Gemmati D, Ongaro A, Scapoli GL, Della Porta M, Tognazzo S, Serino ML, Di Bona E, Rodeghiero F, Gilli G, Reverberi R, Caruso A, Pasello M, Pellati A, De Mattei M (2004) Common gene polymorphisms in the metabolic folate and methylation pathway and the risk of acute lymphoblastic leukemia and non-Hodgkin’s lymphoma in adults. Cancer Epidemiol Biomarkers Prev 13(5):787–794
Petra BG, Janez J, Vita D (2007) Gene-gene interactions in the folate metabolic pathway influence the risk for acute lymphoblastic leukemia in children. Leuk Lymphoma 48(4):786–792. doi:10.1080/10428190601187711
Gra OA, Glotov AS, Kozhekbaeva Z, Makarova OV, Nasedkina TV (2008) Genetic polymorphism in GST, NAT2, and MTRR and susceptibility to childhood acute leukemia. Mol Biol (Mosk) 42(2):214–225
de Jonge R, Tissing WJ, Hooijberg JH, Jansen G, Kaspers GJ, Lindemans J, Peters GJ, Pieters R (2009) Polymorphisms in folate-related genes and risk of pediatric acute lymphoblastic leukemia. Blood 113(10):2284–2289. doi:10.1182/blood-2008-07-165928
Lissowska J, Gaudet MM, Brinton LA, Chanock SJ, Peplonska B, Welch R, Zatonski W, Szeszenia-Dabrowska N, Park S, Sherman M, Garcia-Closas M (2007) Genetic polymorphisms in the one-carbon metabolism pathway and breast cancer risk: a population-based case-control study and meta-analyses. Int J Cancer 120(12):2696–2703. doi:10.1002/ijc.22604
Shrubsole MJ, Gao YT, Cai Q, Shu XO, Dai Q, Jin F, Zheng W (2006) MTR and MTRR polymorphisms, dietary intake, and breast cancer risk. Cancer Epidemiol Biomarkers Prev 15(3):586–588. doi:10.1158/1055-9965.EPI-05-0576
Suzuki T, Matsuo K, Hirose K, Hiraki A, Kawase T, Watanabe M, Yamashita T, Iwata H, Tajima K (2008) One-carbon metabolism-related gene polymorphisms and risk of breast cancer. Carcinogenesis 29(2):356–362. doi:10.1093/carcin/bgm295
Kotsopoulos J, Zhang WW, Zhang S, McCready D, Trudeau M, Zhang P, Sun P, Narod SA (2008) Polymorphisms in folate metabolizing enzymes and transport proteins and the risk of breast cancer. Breast Cancer Res Treat 112(3):585–593. doi:10.1007/s10549-008-9895-6
Xu X, Gammon MD, Zhang H, Wetmur JG, Rao M, Teitelbaum SL, Britton JA, Neugut AI, Santella RM, Chen J (2007) Polymorphisms of one-carbon-metabolizing genes and risk of breast cancer in a population-based study. Carcinogenesis 28(7):1504–1509. doi:10.1093/carcin/bgm061
Sangrajrang S, Sato Y, Sakamoto H, Ohnami S, Khuhaprema T, Yoshida T (2010) Genetic polymorphisms in folate and alcohol metabolism and breast cancer risk: a case-control study in Thai women. Breast Cancer Res Treat. doi:10.1007/s10549-010-0804-4
Rouissi K, Ouerhani S, Oliveira E, Marrakchi R, Cherni L, Ben Othman F, Ben Slama MR, Sfaxi M, Ayed M, Chebil M, Amorim A, Prata MJ, Benammar Elgaaied A (2009) Polymorphisms in one-carbon metabolism pathway genes and risk for bladder cancer in a Tunisian population. Cancer Genet Cytogenet 195(1):43–53. doi:10.1016/j.cancergencyto.2009.06.007
Kim HN, Kim YK, Lee IK, Lee JJ, Yang DH, Park KS, Choi JS, Park MR, Jo DY, Kim HJ (2007) Polymorphisms involved in the folate metabolizing pathway and risk of multiple myeloma. Am J Hematol 82(9):798–801. doi:10.1002/ajh.20967
Lima CS, Ortega MM, Ozelo MC, Araujo RC, De Souza CA, Lorand-Metze I, Annichino-Bizzacchi JM, Costa FF (2008) Polymorphisms of methylenetetrahydrofolate reductase (MTHFR), methionine synthase (MTR), methionine synthase reductase (MTRR), and thymidylate synthase (TYMS) in multiple myeloma risk. Leuk Res 32(3):401–405. doi:10.1016/j.leukres.2007.06.001
Kim HN, Lee IK, Kim YK, Tran HT, Yang DH, Lee JJ, Shin MH, Park KS, Shin MG, Choi JS, Kim HJ (2008) Association between folate-metabolizing pathway polymorphism and non-Hodgkin lymphoma. Br J Haematol 140(3):287–294. doi:10.1111/j.1365-2141.2007.06893.x
Zhang FF, Terry MB, Hou L, Chen J, Lissowska J, Yeager M, Zatonski W, Chanock S, Morabia A, Chow WH (2007) Genetic polymorphisms in folate metabolism and the risk of stomach cancer. Cancer Epidemiol Biomarkers Prev 16(1):115–121. doi:10.1158/1055-9965.EPI-06-0513
Zhang Z, Shi Q, Liu Z, Sturgis EM, Spitz MR, Wei Q (2005) Polymorphisms of methionine synthase and methionine synthase reductase and risk of squamous cell carcinoma of the head and neck: a case-control analysis. Cancer Epidemiol Biomarkers Prev 14(5):1188–1193. doi:10.1158/1055-9965.EPI-04-0501
Suzuki T, Matsuo K, Hiraki A, Saito T, Sato S, Yatabe Y, Mitsudomi T, Hida T, Ueda R, Tajima K (2007) Impact of one-carbon metabolism-related gene polymorphisms on risk of lung cancer in Japan: a case control study. Carcinogenesis 28(8):1718–1725. doi:10.1093/carcin/bgm104
Shi Q, Zhang Z, Li G, Pillow PC, Hernandez LM, Spitz MR, Wei Q (2005) Polymorphisms of methionine synthase and methionine synthase reductase and risk of lung cancer: a case-control analysis. Pharmacogenet Genomics 15(8):547–555
Bethke L, Webb E, Murray A, Schoemaker M, Feychting M, Lonn S, Ahlbom A, Malmer B, Henriksson R, Auvinen A, Kiuru A, Salminen T, Johansen C, Christensen HC, Muir K, McKinney P, Hepworth S, Dimitropoulou P, Lophatananon A, Swerdlow A, Houlston R (2008) Functional polymorphisms in folate metabolism genes influence the risk of meningioma and glioma. Cancer Epidemiol Biomarkers Prev 17(5):1195–1202. doi:10.1158/1055-9965.EPI-07-2733
Suzuki T, Matsuo K, Sawaki A, Mizuno N, Hiraki A, Kawase T, Watanabe M, Nakamura T, Yamao K, Tajima K, Tanaka H (2008) Alcohol drinking and one-carbon metabolism-related gene polymorphisms on pancreatic cancer risk. Cancer Epidemiol Biomarkers Prev 17(10):2742–2747. doi:10.1158/1055-9965.EPI-08-0470
Marchal C, Redondo M, Reyes-Engel A, Perea-Milla E, Gaitan MJ, Machuca J, Diaz F, Caballero J, Carnero J (2008) Association between polymorphisms of folate-metabolizing enzymes and risk of prostate cancer. Eur J Surg Oncol 34(7):805–810. doi:10.1016/j.ejso.2007.09.008
Tong SY, Lee JM, Song ES, Lee KB, Kim MK, Yun YM, Lee JK, Son SK, Lee JP, Kim JH, Hur SY, Kwon YI (2010) The effects of polymorphisms in methylenetetrahydrofolate reductase (MTHFR), methionine synthase (MTR), and methionine synthase reductase (MTRR) on the risk of cervical intraepithelial neoplasia and cervical cancer in Korean women. Cancer Causes Control 21(1):23–30. doi:10.1007/s10552-009-9430-z
Acknowledgments
The project was supported by The National High Technology R&D Program of China [2009AA022701] and The National Basic Research Program of China [2010CB534901]. We would like to thank the anonymous reviewer and the editor for their constructive comments on revising this manuscript.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
Dong Han and Chao Shen contributed equally to this work.
Rights and permissions
About this article
Cite this article
Han, D., Shen, C., Meng, X. et al. Methionine synthase reductase A66G polymorphism contributes to tumor susceptibility: evidence from 35 case–control studies. Mol Biol Rep 39, 805–816 (2012). https://doi.org/10.1007/s11033-011-0802-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0802-6