Abstract
Viral infections generally cause disease symptoms by interfering with the microRNA (miRNA)-mediated regulation of gene expression of host plants. In tomato leaves, the accumulation levels of eleven miRNAs and ten target mRNAs were enhanced by different degrees upon Cucumber mosaic virus (CMV)-Fny and Tomato aspermy virus (TAV)-Bj infections. The ability of CMV-Fny to interfere with miRNA pathway was dramatically suppressed in the addition of the benign satellite (sat) RNA variant (satYn12), but was slightly affected when CMV-Fny was co-inoculated with the aggressive satRNA variant (satT1). In plants harboring the infection of CMV-FnyΔ2b (a CMV-Fny 2b-deletion mutant), the unaltered miRNAs and target mRNAs levels compared with mock inoculated plants indicated that 2b ORF was essential for perturbation of miRNA metabolism. When the amounts of viral open reading frames (ORFs) in these infections were quantified, we found satYn12 caused a higher reduction of CMV-Fny accumulation levels than satT1. These results indicate the complex mechanism by which satRNAs participate in CMV-tomato interaction, and suggest that the severity of disease symptoms positively correlates to some extent with the perturbation of miRNA pathway in tomato.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are an abundant subset of the plant small RNA population. They are defined by precise, dicer-like enzyme (DCL1)-catalyzed excision from the helical stems of hairpin-forming single stranded precursor RNAs [1]. Mature miRNAs are incorporated into an RNA-induced silencing complex (RISC), guide the cleavage or translational repression of endogenous target mRNAs in a sequence-specific manner. In plants, most of miRNA target genes are important transcription factors, which regulate multiple developmental processes such as organ polarity and morphogenesis, meristem identity, hormone signaling, stress responses and nutrient homeostasis [2–4].
Virus infections frequently stimulate the antiviral immune system of host plants. Similar to miRNA pathway, this is also a sequence-specific RNA degradation pathway but directed by small interfering RNAs (siRNAs), and its primary function is to restrict the accumulation and spread of exogenous virus invaders [5]. To overcome the siRNA-directed destruction of viral RNAs, most plant viruses have evolved suppressor proteins to counteract host RNA silencing [6–8]. As there are so many common features shared between the siRNA-directed RNA degradation and miRNA-mediated mRNA regulation, it is assumed the activity of viral silencing suppressors can alter miRNA metabolism through attacking common elements of the two pathways [9, 10].
Both Cucumber mosaic virus (CMV) and Tomato aspermy virus (TAV) are type species of the Cucumovirus, in the family Bromoviridae [11]. Their genomes are constituted by three single-stranded, messenger sense RNAs (RNAs 1, 2, and 3), and two subgenomic RNAs 4 and 4A, which encode five proteins designated 1a, 2a, 2b, 3a and 3b [11]. In addition, some isolates of CMV encapsidate a satellite RNA (satRNA), which is a small RNA molecular that is dependent on the virus for its replication, encapsidation and spread. CMV satRNAs exist as different variants, ranging from 330 to 405 nt in size, can usually attenuate or aggravate disease symptoms induced by the helper virus. CMV 2b is one of the first identified silencing suppressors [12]. Previous studies have reported the developmental abnormalities and perturbation of miRNA pathway in 2b-transgenic Arabidopsis are resulted from the direct interaction between 2b and Argonaut 1 (AGO1) protein, and the different effects of CMV-Fny versus CMV-Q on the miRNA pathway may be explained by differences between the two 2b proteins with respect to their stability in vivo [13, 14].
In the present work, wild type CMV-Fny, CMV-FnyΔ2b (a CMV-Fny 2b-deletion mutant), CMV-Fny-satYn12 (CMV-Fny plus a benign satRNA), CMV-Fny-satT1 (CMV-Fny plus an aggressive satRNA), and the naturally isolated TAV-Bj were inoculated on tomato, to investigate the role of different satRNAs in perturbing miRNA pathways in the course of infections. We found CMV-Fny, CMV-Fny-satT1 and TAV-Bj infections significantly altered the normal miRNA-mediated gene expression regulation of host plants. However, the ability of CMV-Fny to interfere with miRNA pathway was dramatically reduced in the addition of satYn12. In combination with the levels of CMV open reading frames (ORFs) monitored during pathogenic processes, we proposed a new model by which satRNAs participate in CMV-tomato interactions, and discussed the correlation between severity of disease symptoms and the perturbation of miRNA pathway in tomato.
Materials and methods
Plants, virus inoculation, and RNA extraction
Seedlings of tomato (Solanum lycopersicum cv. Hezuo903) were grown in a greenhouse at 16 h day/8 h night (22–28°C). CMV-Fny, the typical strain of CMV subgroup ΙA, was obtained by in vitro transcription of infectious cDNA clones pFny109, pFny209 and pFny309 [15]. CMV-FnyΔ2b, a mutant of CMV-Fny in which most (nt 2419–2713 of RNA 2) of the 2b ORF was deleted, was constructed as described by Du et al. [16]. CMV-Fny-satYn12 and CMV-Fny-satT1 were reconstituted by co-inoculation of the corresponding in vitro transcripts of satYn12 or satT1 with CMV-Fny, respectively, [17]. TAV-Bj was isolated from chrysanthemum in Beijing, China, and confirmed by ELISA (Agdia, Elkhart, IN, USA). All of the viruses were maintained on Nicotiana tabacum, and transferred 10–12 days before mechanically inoculating the first true leaves of 10-day old tomato seedlings. Mock-treated plants were inoculated with sodium phosphate buffer (0.1 M, pH 7.2).
At 7, 14, 21 and 28 dpi, 0.2 g tissues were sampled from the upper leaves of virus or mock inoculated plants. Total RNAs were extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA), followed by RNase-free DNase treatment (Takara, Dalian, China).
To assess the genetic stability of these CMV variants, viral double-stranded RNAs (dsRNAs) were extracted at 35 days postinoculation (dpi), using 25 g fresh leaf tissues from CMV-FnyΔ2b infected plants, or 1–2 g tissues from other infections. Subsequently, these dsRNAs were electrophoresed by procedures described in our previous publication [17].
Primer design and real-time quantitative RT-PCR
To date, 30 tomato miRNAs have been documented in miRNA Registry database (http://miRNA.sanger.ac.uk, Release 10, Sept 2010). Among them, miR162 and miR168 are believed to participate in miRNAs biogenesis. MiR164, miR165/166, miR167 and miR319, with their respective targets mRNAs, are essential regulators of meristem initiation and maintenance, axillary meristem differentiation, and leaf morphology. MiR156, miR159 and miR171 are reported to be implicated in promoting floral transitions and regulate flowering time. Therefore, the expression alterations of these miRNAs and target mRNAs upon virus infections were determined by quantitative real-time RT-PCR (qRT-PCR) in this study.
For miRNAs quantification, their mature sequences were downloaded from miRNA Registry database, and the stem-loop RT primers, forward primers and reverse primers were designed according to criteria mentioned by Tang et al. [18]. Tomato U6 small nuclear RNA (U6snRNA) was used as the reference gene for miRNA qRT-PCR. For target mRNAs, the design of primers was based on sequences obtained from clones of ARF8, AGO1-1 and AGO1-2 in our previous work [19], or from public databases. For qRT-PCR of CMV RNAs, the gene specific primers were designed according to the sequence of each CMV-Fny ORF [15]. The 18S rRNA sequence was chosen as the reference endogenous gene for both viral RNA and mRNA relative quantification. All primer pairs were optimized and validated as per our previous work [19], and summarized in Table 1.
The reverse transcription reactions were performed using stem-loop RT primers for miRNAs, or random primer (6 mer, Takara) for host mRNAs and CMV ORFs. A 1.0 μl aliquot of DNase-treated total RNA was inoculated with 1.0 μl of a solution containing 10 μM of each primer. The mixture was heated at 80°C for 5 min to denature the RNA, and then incubated at 60°C for 5 min to anneal the primers. After cooling to room temperature, the remaining reagents (5×buffer, dNTPs, RNase inhibitor, and M-MLV) were added according to the experimental protocol and the pulsed RT reaction proceeded for 30 min at 16°C, followed by 60 cycles at 20°C for 30 s, 42°C for 30 s, and 50°C for 1 s [18]. Finally, the reactions were heated at 85°C for 5 min to inactivate the reverse transcriptase.
PCR volumes were set up to 10 μl that contained 5 μl 2× SYBR Green PCR master mix (Takara), 1 μl of a 1:10 dilution of the cDNA template, and 800 nM each of the corresponding forward and reverse primers (as summarized in Table 1). The cycling profile was 95°C for 10 s, followed by 40 or 50 cycles of 10 s at 95°C and 30 s at 60°C. All reactions were performed in triplicate, and the controls with no template or no reverse transcription were included for each gene. Immediately after the final PCR cycle, a melting curve analysis was done to determine the specificity of each reaction. The threshold cycle (C T) values were determined automatically by Applied Biosystem’s 7300 Sequence Detection System, and the fold changes of each gene were calculated as relative quantity (RQ) values using the comparative C T (2−ΔΔCt) method as per our previous work [19].
Results
Symptom expression and viral RNA accumulation in tomato infected with CMV/satRNA combinations
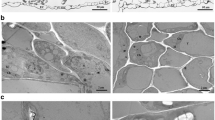
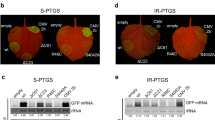
In tomato seedlings, the disease symptoms were monitored between 7 and 35 dpi. As shown in Fig. 1, CMV-Fny infection severely altered leaf morphogenesis, inducing the typical reduction of leaflet blades and of whole-plant growth. CMV-Fny-satYn12 led to an asymptomatic infection, with plants growing with vigor and leaf size comparable to healthy controls. In contrast, the symptoms were exacerbated in CMV-Fny-satT1 infection, which induced chlorosis and even necrosis of older tomato leaves, with severe mosaic and leaf distortion. CMV-Fny∆2b infected plants exhibited slight mottle at early stages of infection but then led to general growth promotion, with increased plant height, leaf area, root length and number of lateral root than mock inoculated plants. TAV-Bj induced severe leaf mosaic, stunting and obvious shortening of internodal distances. At 35 dpi, the genetic stability of these CMV variants was confirmed by results of agarose gel electrophoresis (Fig. 2).
Agarose gel electrophoresis of viral double-stranded RNAs (dsRNAs) extracted from CMV-Fny and its variants infected tomato plants at 35 days postinoculation (dpi). Marker GeneRuler™ 1 kb DNA ladder (MBI fermentas); Lane 1 CMV-Fny, Lane 2 CMV-FnyΔ2b, Lane 3 CMV-Fny-satT1, Lane 4 CMV-Fny-satYn12, Lane 5 mock inoculation
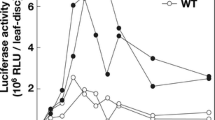
During the pathogenic processes, the accumulation of viral RNAs in infected tomato leaves was monitored at 7, 14, 21 and 28 dpi. As the results shown in Fig. 3, the abundance of viral RNAs in CMV-Fny-satT1 or CMV-Fny-satYn12 infected plants was reduced to 0.17- to 0.83-fold of those in CMV-Fny infection at 21 dpi, respectively. For each viral RNA, the average level of four detection time points in each infection was compared and summarized in Table 2, from which we found 2a and 2b ORFs were the most significantly affected in the addition of satYn12 and satT1, respectively. In the case of CMV-Fny∆2b infection, viral RNAs levels were extremely low throughout the detection time points, almost similar to mock inoculated plants (data not shown).
The relative accumulation levels of viral open reading frames (ORFs) in CMV-Fny, CMV-Fny-satT1 and CMV-Fny-satYn12 infected tomato leaves at 21 days postinoculation (dpi), determined by quantitative real-time RT-PCR. For each ORF, the expression level in CMV-Fny was set as 1, and the relative quantity (RQ) in CMV-Fny-satT1 and CMV-Fny-satYn12 was made relative to it. 18S rRNA was chosen as an endogenous control. The results represent mean values from three biological replicates and vertical bars indicate standard errors
Accumulation levels of miRNAs are differentially altered in CMV and TAV infected tomato plants
To investigate the interference of viral infections with miRNA pathways in tomato, the expression patterns of eleven selected miRNAs were quantified at 7, 14, 21 and 28 dpi, respectively.
As shown in Fig. 4, the expression of tested miRNAs was altered with evident strain specific differences upon different infections. Compared with mock inoculated plants, miR162, miR164 and miR168 were most significantly increased in CMV-Fny, CMV-Fny-satT1 and TAV-Bj infected plants at 7 dpi (RQ between 5.4 and 23.1). Besides them, the abundance of miR167 was also apparently increased in these infections at 14 dpi (RQ between 4.4 and 11.2). At 21 dpi, although miR162 and miR168 remained at relatively higher levels, miR164 and miR167 in CMV-Fny-satT1 infection and miR167 in CMV-Fny infection were reduced to 4.7-, 2.1- and 5.8-fold of those in mock inoculated plants. Meanwhile, CMV-Fny and CMV-Fny-satT1 induced a clear increase in miR165/166 levels (RQ > 7.0). MiR159, miR160 and miR169 were also increased in CMV-Fny or TAV-Bj infected plants to levels of approximately RQ = 5.0 at 21 dpi. At 28 dpi, the levels of miR162, miR165/166 and miR168 in CMV-Fny infection were decreased to 5.8- and 13.6-fold of those in mock, while they remained at constant levels in TAV-Bj infection. For CMV-Fny-satYn12 and CMV-Fny∆2b infected plants, the expression levels of all tested miRNAs were not or only slightly altered throughout the time course.
Expression levels of eleven microRNAs (miRNAs) modulated by different virus infections in tomato plants at 7, 14, 21, 28 days postinoculation (dpi). According to the comparative method (RQ = 2−∆∆Ct), the expression level of each miRNA was first normalized to 18S rRNA (reference gene), and then made relative to the amount of corresponding miRNAs in mock-inoculated sample, representing the calibrator. Columns represent mean fold change from three biological replicates and vertical bars indicate standard errors
Accumulation levels of miRNA-regulated genes are differentially altered in CMV and TAV infected tomato plants
Following, the transcript levels of ten miRNA-regulated mRNAs were also investigated using the same RNA preparations. Figure 5 indicated the abundance of most target mRNAs was variably enhanced after CMV-Fny, CMV-Fny-satT1 and TAV-Bj infections.
Expression levels of ten target mRNAs modulated by different virus infections in tomato plants at 7, 14, 21, 28 days postinoculation (dpi). According to the comparative method (RQ = 2−∆∆Ct), the expression level of each target mRNA was first normalized to 18S rRNA (reference gene), and then made relative to the amount of corresponding mRNAs in mock-inoculated sample, representing the calibrator. Columns represent mean fold change from three biological replicates and vertical bars indicate standard errors
In accordance with miR162 and miR168, the levels of DCL1, AGO1-1 and AGO1-2 were apparently increased in CMV-Fny infected plants at 7 dpi (RQ between 6.2 and 8.1). Other mRNAs did not show remarkable overaccumulation at this time point. At 14 dpi, CMV-Fny, CMV-Fny-satT1 and TAV-Bj induced a significant increase in transcript levels of NAC1 (RQ > 12.0), whereas DCL1, AGO1-1 and AGO1-2 in these infections were only 2.3- to 5.9-fold higher than those of in mock inoculated plants. At 21 dpi, besides NAC1, the levels of MYB, HD-ZIP and TCP4 were also notably increased in CMV-Fny, CMV-Fny-satT1 and TAV-Bj infected plants (RQ > 6.0). At 28 dpi, the abundance of TCP4 and MYB was decreased to 5.0- and 2.0-fold of those in mock inoculated plants, whereas NAC1 and HD-ZIP remained at relatively constant levels (approximately RQ = 9.0).
Taken together, we found besides AGO1 and DCL1, the expression levels of TCP4, NAC1, HD-ZIP and MYB in CMV-Fny, CMV-Fny-satT1 and TAV-Bj infected plants were also significantly altered during the pathogenic processes. When miRNAs and mRNAs were considered together, the abundance of most target mRNAs varied with the same tendency of their corresponding miRNAs, with higher values due to CMV-Fny, CMV-Fny-satT1 and TAV-Bj infections, but no or very limited changes induced by CMV-Fny-satYn12 and CMV-Fny∆2b compared with mock inoculated plants. Our data demonstrate that although with some exceptions, there is a substantial correlation between the accumulation levels of target mRNAs and the corresponding miRNA species. Moreover, these results also indicate that the addition of benign and aggressive satRNAs in CMV-Fny inoculum would differentially perturb miRNA-guided regulation of gene expression.
Discussion
It is well documented that the presence of satRNA results in a depression of the accumulation of CMV to a degree that varies with the isolate of CMV and the variant of satRNA that interact [11, 20]. In our study, we found satYn12 caused a higher reduction of CMV-Fny accumulation levels than satT1. These data completely match those of Cillo et al. [21], who reported that the benign satRNA variant induced much less CMV RNAs than the aggressive satRNAs in tomato. However, the substantial down-regulation of CMV 2b ORF in both satYn12 and satT1 co-inoculation indicated that the attenuated disease symptoms in the addition of satYn12 could not simply attribute to the low levels of 2b gene, and consequently the reduced amounts 2b proteins, then the reduced ability to interfere with miRNA pathway, as the mechanism proposed by Cillo et al. [22] based on the study of satTfn (a mild satRNA variant). It would be more appropriate to presume that the high levels of satRNAs in these plants might affect the function of 2b protein through certain mechanisms, thus 2b could not effectively disrupt siRNA-directed defensive pathway of host plants, and then the replication of viral RNAs was inhibited. But obviously, more directly evidences are required to support our opinion.
In our study, the interference of viral infection with miRNA pathways in tomato was indicated by the alteration of miRNAs and target mRNA expression levels. Based on the obtained results, we found the severity of CMV disease symptoms correlated with the extent to which miRNA pathway was disrupted in the co-inoculation of satYn12. But in the addition of satT1, although disease symptoms were obviously aggravated, no obvious differences were found between the miRNAs or mRNAs expression levels between CMV-Fny and CMV-Fny-satT1 infections. As the chlorosis and necrosis phenotype appeared on the older leaves of CMV-Fny-satT1 infected plants, whereas the samples were harvested from newly developed leaves in our study, thus further studies are needed to investigate if these results will be affected by the position of the leaves that are sampled.
CMV 2b is shown to be a multifunctional protein involved in symptoms induction, viral movement, suppression of RNA silencing, and as an antagonist of the salicylic acid-mediated defense response [16, 23]. In our study, we found the deletion of 2b coding sequence not only effectively prevented the systemic spread of this CMV-Fny mutant in tomato, disease symptoms and viral RNAs accumulation levels (data not shown) were substantially suppressed as well. Meanwhile, the expression levels of tested miRNAs and mRNAs were almost unaltered throughout the detection time points in CMV-Fny∆2b infected plants. These data substantiate earlier studies demonstrating the role of 2b in perturbing miRNA pathway and in virulence determination again.
Although different disease symptoms were induced in tomato seedlings infected with CMV-Fny and TAV-Bj, no apparent differences were observed between the expression alterations of tested miRNAs and target mRNAs in the two infections. Thus, we can not conclude if the two viruses have a similar mechanism in perturbing miRNA pathway.
The mRNAs quantification results have shown that, three genes with the known role in tomato leaf morphogenesis, NAC1, HD-ZIP and TCP4, were up-regulated during the pathogenic processes after CMV-Fny, CMV-Fny-satT1 and TAV-Bj infections. NAC1 is regulated by miR164, it has been reported that NAC1 controls lamina outgrowth on a grander scale than that of serrations [24]. Reduction of NAC gene function in a wide range of compound leaved plants reduces the number of leaflet and results in leaflet fusions as well as suppressing serrations [25, 26]. MiR165/166-regulated HD-ZIP is important for establishing the adaxial cell fate during leaf dorso-ventral patterning. MiR319-regulated TCP4 is also indispensable for proper leaf morphogenesis and leaf senescence [24, 27]. These data are in agreement with the observed leaf-altered morphology caused by CMV-Fny, CMV-Fny-satT1 and TAV-Bj infections.
In our study, the miR164 and NAC1 levels were noticeably increased in leaves of CMV-Fny, CMV-Fny-satT1 and TAV-Bj infected tomato. The up-regulation of miR164/NAC1 levels were also observed in root and leaf tissues of CMV-Fny 2b transgenic Arabidopsis plants [13]. As miR164/NAC1 are believed to involve in phytohormone response pathways, this probably occurred because tomato auxin signaling pathway was perturbed by virus infections, resulting in modulated root and leave phenotypes. In a very recent article, Lewsey et al. [28] reported the disruption of Arabidopsis salicylic acid (SA) and jasmonic acid (JA)-mediated defensive signaling pathways by CMV 2b protein, which was consistent with our speculation. However, they suggested the 2b protein may not be the only CMV-encoded factor that inhibits JA responses in Arabidopsis [28]. But according to our results and the reports of Cillo et al. [22], the effects of other viral factors on miRNAs or mRNAs expression were not apparent, which probably due to the limited transcripts studied.
We found the accumulation levels of ARF8 and SCL were not significantly altered in any of the five infections in tomato leaves. These results are consistent with their functions, as ARF8 is reported to regulate the development of lateral root through auxin signaling pathway, and SCL is regarded to control the development of floral tissues [2, 29]. However, the abundance of MYB, which is believed to regulate flowering time and anther development, was significantly increased in CMV-Fny, CMV-Fny-satT1 and TAV-Bj infected leaves at 21 dpi. As the mechanism of up-regulation appears to be non-miRNA mediated, we suspect these may result from spatio-temporal variation in transcription of individual mRNA [30, 31].
In conclusion, this study monitored the accumulation of viral ORFs, and quantified the expression changes of several miRNAs and their target mRNAs during the pathogenic processes, upon different CMV variants and TAV-Bj infections. Based on our observation, we find the miRNA pathway is implicated in the modification of disease symptoms in the presence of CMV satRNAs. It is expected this studies will broaden our understanding of the relationship between CMV infection, satRNA effects, miRNA pathway and symptom modifications in plants.
References
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136:669–687
Chuck G, Candela H, Hake S (2009) Big impacts by small RNAs in plant development. Curr Opin Plant Biol 12:81–86
Lu XY, Huang XL (2008) Plant miRNAs and abiotic stress responses. Biochem Biophys Res Co 368:458–462
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on plant transcription. Dev Cell 8:517–527
Mlotshwa S, Pruss GJ, Vance V (2008) Small RNAs in viral infection and host defense. Trends Plant Sci 13(7):375–382
Chapman EJ, Prokhnevsky AI, Gopinath K, Dolja VV, Carrington JC (2004) Viral RNA silencing suppressors inhibit the microRNA pathway at an intermediate step. Genes Dev 18:1179–1186
Dunoyer P, Lecellier C, Parizotto EA, Himber C, Voinnet O (2004) Probing the microRNAs and small interfering RNA pathways with virus-encoded suppressors of RNA silencing. Plant Cell 16:1235–1250
Levy A, Dafny-Yelin M, Tzfira T (2008) Attacking the defenders: plant viruses fight back. Trends Microbiol 16(5):194–197
Bazzini AA, Hopp HE, Beachy RN, Asurmendi S (2007) Infection and coaccumulation of tobacco mosaic virus proteins alter microRNA levels, correlating with symptom and plant development. Proc Natl Acad Sci USA 104:12157–12162
Berkhout B, Haasnoot J (2006) The interplay between virus infection and the cellular RNA interference machinery. FEBS Lett 580:2896–2902
Palukaitis P, Garcia-Arenalt F (2003) Cucumoviruses. Adv Virus Res 62:241–323
Ruiz-Ferrer V, Voinnet O (2007) Viral suppression of RNA silencing: 2b wins the Golden Fleece by defeating Argonaute. BioEssays 29:319–323
Lewsey M, Robertson FC, Canto T, Palukaitis P, Carr JP (2007) Selective targeting of miRNA-regulated plant development by a viral counter-silencing protein. Plant J 50:240–252
Zhang X, Yuan YR, Pei Y, Lin SS, Tuschl T, Patel DJ, Chuan NH (2006) Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute 1 cleavage activity to counter plant defense. Genes Dev 20:3255–3268
Feng JL, Chen SN, Tang XS, Ding XF, Du ZY, Chen JS (2006) Quantitative determination of cucumber mosaic virus genome RNAs in virions by real-time reverse transcription polymerase chain reaction. Acta Biochem Biophys Sin 38(10):669–676
Du ZY, Chen FF, Zhao ZJ, Liao QS, Palukaitis P, Chen JS (2008) The 2b protein and C-terminus of the 2a protein of Cucumber mosaic virus subgroup I strains both play a role in viral RNA accumulation and induction of symptoms. Virology 380:363–370
Liao QS, Zhu LP, Du ZY, Zeng R, Chen JS (2007) Satellite RNA-mediated reduction of cucumber mosaic virus genomic RNAs accumulation in Nicotiana tabacum. Acta Biochem Biophys Sin 39(3):217–223
Tang F, Hajkova P, Barton SC, Lao K, Surani MA (2006) MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res 34:e9
Feng JL, Wang K, Liu X, Chen SN, Chen JS (2009) The quantification of tomato microRNAs response to viral infection by stem-loop real-time RT-PCR. Gene 437:14–21
Escriu F, Fraile A, Garcia-Arenal F (2000) Evolution of virulence in nature populations of the satellite RNA of cucumber mosaic virus. Phytopathology 90:480–485
Cillo F, Pasciuto MM, De Giovanni C, Finetti-Sialer MM, Ricciardi L, Gallitelli D (2007) Response of tomato and its wild relatives in the genus Solanum to cucumber mosaic virus and satellite RNA combinations. J Gen Virol 88:3166–3176
Cillo F, Mascia T, Pasciuto MM, Gallitelli D (2009) Differential effects of mild and severe cucumber mosaic virus strains in the perturbation of microRNA-regulated gene expression in tomato map to the 3′ sequence of RNA 2. Mol Plant-Microbe Interact 22:1239–1249
Ji LH, Ding SW (2001) The suppressor of transgene RNA silencing encoded by cucumber mosaic virus interferes with salicylic acid-mediated virus resistance. Mol Plant-Microbe Interact 14:715–724
Kinder CA (2010) The many roles of small RNAs in leaf development. J Genet Genomics 37:13–21
Berger Y, Harpaz-Saad S, Brand A, Melnik H, Sirding N, Alvarez JP, Zinder M, Samach A, Eshed Y, Ori N (2009) The NAC-domain transcription factor GOBLET species leaflet boundaries in compound tomato leaves. Development 136:823–832
Blein T, Pulido A, Vialette-Guiraud A, Nikovics K, Morin H, Hay A, Johansen IE, Tsiantis M, Laufs P (2008) A conserved molecular framework for compound leaf development. Science 322:1835–1839
Garcia D (2008) A miracle in plant development: Role of microRNAs in cell differentiation and patterning. Semin Cell Dev Biol 19:586–595
Lewsey MG, Murphy AM, MacLean D, Dalchau N, Westwood JH, Macaulay K, Bennet MH, Moulin M, Hanke DE, Powell G, Smith AG, Carr JP (2010) Disruption of two defensive signaling pathways by a viral RNA silencing suppressor. Mol Plant-Microbe Interact 7:835–845
Yang T, Xue L, An L (2007) Functional diversity of miRNA in plant. Plant Sci 172:423–432
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/Hc-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 4:205–217
Valoczi A, Varallyay E, Kauppinen S, Burgyan J, Havelda Z (2006) Spatio-temporal accumulation of microRNAs is highly coordinated in developing plant tissues. Plant J 47:140–151
Acknowledgments
This work was supported by the grants from the National Natural Science Foundation of China (30800716) and the Science Foundation of Zhejiang Sci-Tech University (ZSTU) under Grant No. 1016816-Y.
Author information
Authors and Affiliations
Corresponding author
Additional information
Junli Feng and Leiyu Lai contributed equally to this work.
Rights and permissions
About this article
Cite this article
Feng, J., Lai, L., Lin, R. et al. Differential effects of Cucumber mosaic virus satellite RNAs in the perturbation of microRNA-regulated gene expression in tomato. Mol Biol Rep 39, 775–784 (2012). https://doi.org/10.1007/s11033-011-0798-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0798-y