Abstract
Study of retina specific genes would offer insights into retinal diseases and treatment. Based on the information from the gene expression profiles of mouse retinas, we here identified a mitochondria-localized glutamic acid-rich protein (MGARP/OSAP) as one of the highly expressed proteins in retina. Sequence analysis revealed that mouse and rat MGARPs have an extra insertion of four consecutive amino acid repeats at the C-terminus, while other homologues do not. MGARP was demonstrated to be localized to the mitochondria and overexpression of MGARP missing N-terminal region causes severe mitochondrial aggregation, implying an important role of MGARP in maintaining mitochondrial morphology. MGARP is highly expressed in mitochondria-rich layers, including inner segment of the photoreceptor, outer plexiform layer and ganglion cell layers of mouse retina. Far-UV CD spectrum analysis suggested that MGARP exhibits a large area of intrinsic disorder and the unusual position of its Tyr fluorescence suggested that Tyr residues in MGARP might form excimer and exist in an ionized state. These findings implied that MGARP be a good candidate for assembling certain ion channels on mitochondria membrane and have great potential to be involved in retinal energetic metabolism through mitochondria related pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Retina is a component of the eye tissue and plays an essential role in vision. Vision, as an enormously complex form of information processing, critically depends on the retina. Seeing is initiated when light passing through the pupil of the eye is focused by the lens onto the retina’s sensory neuroepithelium [1]. Retina diseases can cause vision loss or irreversible blindness. Age-related macular degeneration (AMD) and diabetic retinopathy (DR) are two typical causes of blindness that severely threaten people’s health [2–4]. AMD is the leading cause of blindness in the elderly, affecting 10 million people around the world [5, 6]. DR is also a severe syndrome of diabetes. Almost all diabetes patients suffer from retinal dysfunction in late-stage diabetes [7, 8]. Additionally, one major cause of blindness in the developed world is inherited retinal degeneration (RD). In recent years, research on AMD has achieved significant breakthroughs [9–12]. Evidence has shown that multiple genes and proteins are involved in AMD, DR and RD, but the etiology still remains unclear. Thus, annotation and characterization of retinal genes and proteins have become more important to understanding the detailed mechanism of the development of retinal disease and improving clinical interventions.
Mammalian photoreceptors are the most intensively studied type of sensory neuron [13]. Large scale screenings have been done to identify the full set of genes expressed by mammalian rods from mature and developing mouse retina, which has provided enormous information for researchers to further study the physiology of the retina and the development of retinal diseases [14, 15]. However, there are still a large number of novel genes remaining to be discovered, and their functions and properties need to be defined.
To investigate the gene expression profile of retinal genes at different developmental stages, DNA microarray analysis was carried out using mRNAs from E13.5, P1 and adult C57BL/6 mouse retina as described previously [16]. Here, we reported a novel gene obtained by the above DNA microarray screening in the same batch with MPP4 protein [16]. It is a mitochondria-localized glutamic acid-rich protein predominately expressed in the retina. Immunolocalization showed a positive staining in each layer of the retina, with particularly higher staining in the inner segment of the photoreceptor (IS), the outer plexiform layer (OPL) and the ganglion cell layer (GCL). Our study unveiled some distinct features of this protein and suggested the importance of further studying its functions, which would improve our understanding of the development and pathogenesis of some retinal diseases.
Materials and methods
Gene analysis and plasmids construction
Bioinformatic analysis of MGARP was carried out to analyze the basic features of this gene. Gene structure analysis, homologous alignment, signal peptide and protein conserved domain predictions were performed using the following sources: http://www.ncbi.nih.gov/BLAST/, http://www.cbs.dtu.dk/services/SignalP/, http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi, http://www.cbs.dtu.dk/services/TMHMM-2.0/, Bioedit and the DNA STAR software. Full-length cDNAs of mouse MGARP (mMGARP) and human MGARP (hMGARP) were obtained by Reverse Transcription-Polymerase Chain (RT-PCR) from mouse and human retina with the primers of mMGARP1/mMGARP2 and hMGARP1/hMGARP2, as shown in Table 1. RT-PCR was performed as described in the user’s manual (TaKaRa Biotechnology Co., Ltd). The following PCR conditions were used: 94°C for 4 min, 30 cycles of 94°C for 30 s, 60°C for 45 s, and 72°C for 1 min, followed by a final extension at 72°C for 5 min. The amplified full-length cDNAs of mMGARP and hMGARP were inserted into the vectors of pcDNA3.1/myc-His (Invitrogen) and pGEX-4T-1 (Amersham Pharmacia Biotech) at EcoRI and XhoI restriction sites to construct eukaryotic expression plasmids (pcDNA3.1/mMGARP/myc and pcDNA3.1/hMGARP/myc) and prokaryotic expression plasmids (pGST/mMGARP). mMGARP and hMGARP genes were then subcloned into EcoRI/ApaI sites of pEGFP-N2 (BD Biosciences Clontech) from pcDNA3.1/myc-His to generate pmMGARP/EGFP and phMGARP/EGFP expression plasmids. The cDNAs of mMGARP-deletion (ΔE for deleting 1–42aa and ΔET for deleting 1–67aa) and hMGARP-deletion (ΔE for deleting 1–41aa and ΔET for deleting 1–63aa) were obtained by PCR using the above constructed plasmids as templates and the primers of mMGARP-TM1/mMGARP2 for amplifying the fragment of mMGARP-ΔE, mMGARP-deletion1/mMGARP2 for amplifying mMGARP-ΔET, hMGARP-TM1/hMGARP2 for amplifying hMGARP-ΔE, and hMGARP-deletion1/hMGARP2 for amplifying hMGARP-ΔET (Table 1). All target fragments were cloned into pEGFP-N2 vectors using the same method as that for cloning the full-length cDNA. Thus, a series of MGARP-deletion expression plasmids were constructed, namely, pmMGARP-ΔE/EGFP, pmMGARP-ΔET/EGFP, phMGARP-ΔE/EGFP and phMGARP-ΔET/EGFP. All clones were confirmed by DNA sequencing.
Cell culture and imaging of scanning laser confocal microscopy
The HeLa cells were cultured in DMEM (GIBCO) medium containing 10% calf serum, 100 units/ml penicillin, 100 μg/ml streptomycin, and 2 mmol/L glutamine. Cells were plated on a 6-well culture plate or 35 mm-diameter dishes with glass coverslips at a density of 5 × 104 cells/ml and grown at 37°C for 2 days. About 3 μg per well of pmMGARP-ΔE/EGFP, pmMGARP-ΔET/EGFP, phMGARP-ΔE/EGFP and phMGARP-ΔET/EGFP plasmids were respectively delivered into the cells using VigoFect (Vigorous Biotechnology). The pEGFP-N2 alone was transfected into cells as control. Twenty-four hours after transfection, the cells were incubated with 100 nmol Mito Tracker Red (Molecular Probes) in PBS for 10 min at 37°C. Cells were then washed twice with PBS and fixed with 4% formaldehyde for 10 min at room temperature. The nuclei were counterstained with Hoechst 33342 (Sigma). The coverslips were mounted in glycerol and analyzed with a laser confocal Olympus fluoview 500 microscope.
Immunofluorescent staining of retinal sections
Retina of adult mouse was fixed with 10% formalin in PBS for 12 h, and then dehydrated gradually in alcohol. The tissues were embedded in paraffin and sectioned in a thickness of 5 μm. Paraffin sections were routinely dewaxed, rehydrated and washed with 0.01 M PBST (1/1000 Tween20 in PBS) 3 × 5 min. Sections were incubated with MGARP antibody (dilution concentration 1:10,000) for 1 h in room temperature, washed with 0.01 M PBST 3 × 5 min and then incubated with the appropriate secondary antibody (TRITC, dilution concentration 1:100) for 1 h at room temperature followed by washing by 0.01 M PBST 3 × 5 min and counterstained by antibody labeled by Hoechst (dilution concentration 1:2,000) for 1 h in 37°C. Control sections were incubated with anti-GST rabbit serum. Finally, all sections were sealed with glycerin and observed under fluorescence microscope and laser confocal microscope.
Tissue expression analysis
To study mMGARP expression pattern from different mouse tissues, total RNAs were isolated from the following tissues of adult mouse: muscle, liver, uterus, lung, blood, kidney, spleen, heart, testis, retina, fat, brain, intestine, and stomach. One microgram total RNAs were used as template for RT-PCR analysis (TaKaRa Biotechnology Co., Ltd) as described in the manual. The RT-PCR products were analyzed by 1% agarose gel electrophoresis. The expression level of β-Actin was used as loading and internal control.
Spectroscopic experiments
The protein was prepared as described as Supporting Methods and determined according to the Bradford method [17] using bovine serum albumin as a standard. The purity of the final products was identified by SDS–polyacrylamide gel electrophoresis (SDS–PAGE). The samples used for spectroscopic experiments were prepared by dissolving the protein in 20 mmol/l Tris–HCl buffer with a final protein concentration of 0.1 mg/ml. The fluorescence emission spectra were collected on a Hitachi F-2500 spectrofluorimeter using 1-cm-pathlength cuvettes. The intrinsic fluorescence was measured using an excitation wavelength of 280 nm and an emission spectral range of 300–400 nm. The far-UV circular dichroism (CD) spectra were recorded on a Jasco-715 spectrophotometer using 0.1-cm pathlength cells over a wavelength range of 190–250 nm. Spectra were scanned at a rate of 200 or 300 nm/min, a resolution of 0.2 nm and a bandwidth of 1 nm. The percentages of secondary structure types were determined by the CONTINLL, SELCON3 and CDSSTR algorithms within the CDPro analytical software [18]. All spectroscopic experiments were conducted at 25°C.
Results
Identification and sequence analysis of MGARP
Sequence analysis revealed that this novel gene contains an open reading frame of 852 bp encoding a protein of 283 amino acids and is mapped to the mouse chromosome 3qC. Its coding region spans four exons scattering 7.7 kb. Using RT-PCR, we also cloned the human counterpart cDNA from human retina and HeLa cells, which also contains four exons and is located on the human chromosome 4q28. The calculated molecular weights of the mouse and human gene products were 29.9 and 25.4 kDa, respectively. Amino acid analysis showed that the deduced proteins are rich in acidic amino residues, especially glutamic acids, which are present at a ratio of 17.3% (49 out of 283aa) for the mouse gene and 14.6% (35 out of 240aa) for the human gene. To investigate the similarity and evolutionary relationship of the homologues from different species, phylogenetic and homologous analyses were carried out. The results demonstrated that the homologous sequences could be found only in higher animals, including humans, macaques, cattle, dogs, rats and mice (Supplementary Fig. 1a). Interestingly, the same proteins have been previously identified as mouse ovary-specific acidic protein (OSAP) and human corneal endothelium specific protein (CESP) in large-scale screenings for tissue specific proteins [19, 20]. Considering the conservation of this gene among different species, the presence of a transmembrane domain and specific cellular localization, we therefore suggested the protein an accordant and universal name, mitochondrial-localized glutamic acid-rich protein (MGARP).
Primary structure analysis defines MGARP a novel protein family
MGAPR was predicted to have a transmembrane region. Further analysis indicated that the deduced amino acid sequences of MGARPs from different species share higher similarity at the N-terminal portion than the C-terminal portion and none have glutamic acid residues at their N-terminus (Supplementary Fig. 1b). The C-terminal regions of mouse and rat MGARPs have an insertion that human and other MGARPs do not have (Supplementary Fig. 1b). The insertion is actually of four consecutive repeats, R1–R4 (Fig. 1a). The repeats between mouse and rat share certain similarities. The mouse repeat motif is commonly characterized by a Glu-Gly-Ala-Asp-Thr-Ser-Gln septenary, except the fourth repeat harbors two conservative substitutions. For rat MGARP, the first two repeats are identical and characterized by a Ser-Lys-Glu-Gly-Ala-Glu-Thr septenary, and the last two by a Ser-Lys-Glu-(Glu/Ala)-(Ala/Gly)-(Glu/Gly)-Thr septenary (Fig. 1b). Among rat MGARP repeats, Ser-Lys-Glu and Thr seem to be the conserved residues. These indicate that amino acids in the repeats are highly conserved, but the sequences of these repeats are not conserved among different species. By looking at the above and what will be shown later, we conclude that the primary structure defines MGARP as a novel protein family.
Protein structural organization of MGARP. a Schematic diagram of the protein structure of human and mouse MGARPs. Mouse MGARP has four repeats (denoted R1, R2, R3 and R4). b Amino acid alignment of R1–R4 from mouse and rat MGARPs. Black boxes indicate the amino acid identities and grey boxes show conservative substitutions
The transmembrane domain is more required for MGARP’s localization
Cellular localization of a protein has a fundamental implication in its function. Analysis with the cellular localization prediction algorithm suggested that MGARP is likely to reside in the mitochondria. In order to verify this prediction, we used confocal microscopy to examine HeLa cells transfected with expression plasmids encoding MGARP/EGFP fusion protein and different deletion mutants (Fig. 2a). Green fluorescence analysis showed that EGFP alone was expressed evenly in both the nucleus and cytoplasm (Fig. 2b). In contrast, mMGARP/EGFP and hMGARP/EGFP were absent in the nucleus and significantly expressed in cytoplasm in a punctuate pattern that overlapped with the staining pattern of Mito Tracker, a fluorescent marker taken up specifically by mitochondria (Fig. 2b). This result was consistent with previous report [19]. To further determine the important region for MGARP mitochondrial localization, deletion mutants were generated as described in “Materials and methods”. Confocal imaging showed that the mutants missing the N-terminal region (mMGARP-ΔE/EGFP and hMGARP-ΔE/EGFP) could still be expressed in the cytoplasm and overlaid with Mito Tracker staining, but they caused remarkable mitochondria aggregation to the nuclear periphery (Fig. 2b), indicating that MGARP plays a potential role in maintaining mitochondrial morphology. Furthermore, the mutants missing both the N-terminal region and transmembrane domain (mMGARP-ΔET/EGFP and mMGARP-ΔET/EGFP) showed quite different staining patterns from those of the full-length and ΔE forms, and were no longer overlaid with Mito Tracker staining (Fig. 2b). mMGARP-ΔET/EGFP was expressed in the cytoplasm, while hMGARP-ΔET/EGFP was expressed in the cytoplasm and partially in the nucleus, indicating its expression was totally in disorder. These experiments clearly showed that MGARP is predominately localized to the mitochondria, and the transmembrane domain is more required for MGARP’s localization and translocation.
Cellular localization of MGARP. a Schematic diagram showing different deletion mutants of human and mouse MGARPs. ΔE: deletion of N-terminal region; ΔET: deletion of extracellular domain plus transmembrane domain. b Cellular localization of MGARPs. After transfection with different plasmids, HeLa cells were double-labeled for mitochondria (Mito Tracker, red/second lane) and nucleus (Hoechst, blue/third lane). The image was obtained by confocal microscopy
Distribution of endogenous MGARP
Expression pattern is a significant hint to understand the physiological functions of novel genes. To clarify the tissue distribution pattern of mMGARP, we performed semi-quantity RT-PCR by using the same amount of total RNA isolated from different mouse tissues, including muscle, liver, uterus, lung, blood, kidney, spleen, heart, testis, retina, fat, brain, intestine, and stomach. The β-Actin gene was as positive control. The result demonstrated that mMGARP was highly expressed in retina and testis (Fig. 3a). In addition to retina, mMGARP expression is also detected at lower levels in kidney, spleen and brain, suggesting that mMGARP expression is highly enriched in but not restricted to mouse retina.
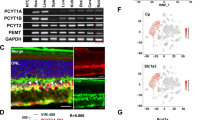
Tissue and spatial expression patterns of MGARP. MGARP distribution pattern. a Total RNAs were isolated from the following tissues of adult mouse: muscle (Mu), liver (Li), uterus (Ut), lung (Lu), blood (Bl), kidney (Ki), spleen (Sp), heart (He), testis (Te), retina (Re), fat (Fa), brain (Br), intestine (In), and stomach (St). The RT-PCR products were analyzed by 1% agarose gel electrophoresis. The expression level of β-Actin was used as loading and internal control. b The protein expression was detected with anti-GST/MGARP fusion protein antibody (marked as MGARP). c Anti-GST antibody (marked as GST) was used as negative control. The nuclei were labeled by Hoechst. All figures were amplified at ×400. OS outer segment of photoreceptor, IS inner segment of photoreceptor, ONL outer nuclear layer, OPL outer plexiform layer, INL inner nuclear layer, IPL inner plexiform layer, GCL ganglion cell layer
For further examine the protein expression pattern in retina, the retinal sections were stained with anti-mMGARP antibody. The result showed that the MGARP protein expression was detected in each layer of the retina, with especially higher signal intensity in the inner segment of the photoreceptor (IS), outer plexiform layer (OPL) and ganglion cell layer (GCL) (Fig. 3b). In addition, the staining clearly showed that MGARP was only expressed in the cytoplasm, consistent with the results of its localization analysis by confocal imaging in HeLa cells. The control stained with anti-GST antibody showed negative or non-specific staining (Fig. 3c).
MGARP exhibits a large area of intrinsic disorder
We next addressed what kind of secondary structure the MGARP protein may have, which can give us more information about its roles. For this purpose, spectroscopic experiments were further carried out. The percentages of the secondary structure types were evaluated by deconvolution of the CD spectrum using CDPro software according to the established procedures [18]. The far-UV CD spectrum analysis of native and fully-denatured hMGARP revealed that the protein was only partially structured (Fig. 4a). The three methods contained in the CDPro software returned well-coincided results, which yielded values of 17.9 ± 0.9% for the α-helix, 23 ± 2% for the β-sheet, 26 ± 2% for the β-turn and 33 ± 6% for unordered structures. This implied that about 40% of the amino acid residues in hMGARP were in folded structures, while about 60% of the others were unstructured. Furthermore, Tyr fluorescence is generally centered at 303–310 nm in the emission fluorescence spectrum, and Trp fluorescence is centered at 316–350 nm. It was a surprise to find that the maximum emission wavelength of hMGARP was centered at 333 nm (Fig. 4b), since hMGARP contains no Trp residue. The unusual positioning of the Tyr fluorescence of hMGARP suggested that Tyr residues in hMGARP might form excimer and/or exist in an ionized state [21]. This deduction was further supported by Tyr fluorescence analysis of GdnHCl-denatured protein, which contained two peaks centered at 306 and 343 nm. The 306 nm component was typically from Tyr residue fully exposed to solvent, while the 343 nm peak might be from its ionized state. Nevertheless, the dissimilarity in the shape of the intrinsic fluorescence suggested that hMGARP only contained partial ordered structures, which is consistent with the result from the CD analysis.
Biophysical properties of MGARP. a Far-UV CD spectra analysis of the native and denatured MGARP. The percentages of the specific secondary structures were evaluated by the algorithms included the CDPro software. b Intrinsic fluorescence of the native and denatured MGARP. The spectra were collected with an excitation wavelength of 280 nm. All samples were prepared in 20 mmol/l Tris–HCl buffer at a final concentration of 0.1 mg/ml
Discussion
In this report, we isolated and systemically characterized MGARP as a novel mitochondrial-localized glutamic acid-rich protein in retina. Phylogenetic analysis and multiple sequence alignment indicated that MGARP homologues exist in all examined vertebrate species and share higher similarities, especially for the amino acid sequences at the N-terminus. This suggests that the MGARP gene family is conserved among vertebrate species and the functions of the family members would be of evolutionary importance. Despite the existence of the above similarities, MGARP proteins among all species examined were quite different in their finer primary structures. Mouse and rat MGARPs have an insertion of four repeats at their N-terminus, whereas the MGARPs of other species do not. It remains to be clarified if this would be one of the reasons why mice and rats have a different visual capacity from that of other animals, especially primates.
From the retina, several specific glutamic acid-rich proteins (GARPs) have been identified. GARPs are characterized by extremely high content of glutamate residues. GARPs have two soluble forms, GARP1 and GARP2, and a third form as a large cytoplasmic domain (GARP’) of the B1 subunit of the cyclic GMP-gated channel [22–24]. It has been demonstrated that GARPs are natively unfolded due to their high abundance of acidic residues and they interact with phototransduction proteins at the rim of the disc membrane, playing a critical role in visual signal transduction [26, 27]. Comparison between MGARPs and the reported GARPs revealed that their amino acids and DNA sequences share very low homology (data not shown); however, they all are rich in glutamic acid residues, contain four repeats and a large unstructured region [25]. Interestingly, GARPs contain four repeats at the N-terminus, while mouse and rat MGARPs have their repeats at the C-terminus. The sequences among MGARP repeats share higher homology than those among GARP repeats. Moreover, MGARPs contain a single transmembrane domain at the N-terminus and locates to the mitochondria while GARPs do not have such a transmembrane domain and they tend to bind to or be a part of the cytoplasmic membrane associated protein, cyclic GMP-gated channel [25].
The abundant expression of MGARP protein in retina and the identification as a mitochondria-localized protein have significant physiological implications. Mitochondria are critical regulators of cell death, a key feature of neurodegeneration [28]. Many lines of evidence suggest that mitochondria have a central role in ageing-related neurodegenerative diseases. Mitochondria-associated diseases are increasingly recognized as a variety of many important clinical entities that develop as a consequence of abnormalities in energy supply, generators of reactive oxygen species and initiators of apoptotic processes [29]. Characterization of novel mitochondrial proteins has been shown to be critical to understanding the role of mitochondria in retinal diseases, and also important to explore the general mechanisms of mitochondrial dysfunction in other neurodegenerative disorders of complex etiology [2, 30, 31]. Our results clearly showed that MGARP is located in the mitochondria and removal of the N-terminal domain and transmembrane domain caused abnormal MGARP translocation, indicating that these parts are essential for MGARP localization and translocation to the mitochondria.
Further analyses by using spectroscopy, Tyr fluorescence and CD spectrometry indicated that Tyr residues in hMGARP might form excimer and/or exist in an ionized state. These results were consistent with MGARP being a good candidate as a component of the ion channel complex. The large area of unfolded region in MGARP would be a molecular basis to provide flexible spacers or linkers tethering channels in the mitochondrial membrane [22, 25, 27].
In summary, our findings on the properties of this novel retinal mitochondrial gene provide a fundamental advance to further understand the mechanisms of the development of retinal diseases caused by energy disorder.
References
Zrenner E (2002) Will retinal implants restore vision? Science 295:1022–1025
Fritsche LG, Loenhardt T, Janssen A, Fisher SA, Rivera A, Keilhauer CN, Weber BH (2008) Age-related macular degeneration is associated with an unstable ARMS2 (LOC387715) mRNA. Nat Genet 40:892–896
Mitchell P, Smith W, Attebo K, Wang JJ (1995) Prevalence of age-related maculopathy in Australia. The blue mountains eye study. Ophthalmology 102:1450–1460
Vingerling JR, Dielemans I, Hofman A, Grobbee DE, Hijmering M, Kramer CF, de Jong PT (1995) The prevalence of age-related maculopathy in the Rotterdam Study. Ophthalmology 102:205–210
Marx J (2006) Gene offers insight into macular degeneration. Science 314:405
Hollyfield JG, Bonilha VL, Rayborn ME, Yang X, Shadrach KG, Lu L, Ufret RL, Salomon RG, Perez VL (2008) Oxidative damage-induced inflammation initiates age-related macular degeneration. Nat Med 14:194–198
Hammes HP, Lin J, Renner O, Shani M, Lundqvist A, Betsholtz C, Brownlee M, Deutsch U (2002) Pericytes and the pathogenesis of diabetic retinopathy. Diabetes 51:3107–3112
Sivaprasad S, Jackson H (2007) Blood pressure control in type II diabetics with diabetic retinopathy. Eye 21:708–711
DeWan A, Liu M, Hartman S, Zhang SS, Liu DT, Zhao C, Tam PO, Chan WM, Lam DS, Snyder M, Barnstable C, Pang CP, Hoh J (2006) HTRA1 promoter polymorphism in wet age-related macular degeneration. Science 314:989–992
Edwards AO, Ritter R III, Abel KJ, Manning A, Panhuysen C, Farrer LA (2005) Complement factor H polymorphism and age-related macular degeneration. Science 308:421
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C, Henning AK, SanGiovanni JP, Mane SM, Mayne ST, Bracken MB, Ferris FL, Ott J, Barnstable C, Hoh J (2005) Complement factor H polymorphism in age-related macular degeneration. Science 308:385
Yang Z, Camp NJ, Sun H, Tong Z, Gibbs D, Cameron DJ, Chen H, Zhao Y, Pearson E, Li X, Chien J, Dewan A, Harmon J, Bernstein PS, Shridhar V, Zabriskie NA, Hoh J, Howes K, Zhang K (2006) A variant of the HTRA1 gene increases susceptibility to age-related macular degeneration. Science 314:992
Ebrey T, Koutalos Y (2001) Vertebrate photoreceptors. Prog Retin Eye Res 20:49–94
Blackshaw S, Fraioli RE, Furukawa T, Cepko CL (2001) Comprehensive analysis of photoreceptor gene expression and the identification of candidate retinal disease genes. Cell 107:579–589
Lord-Grignon J, Tétreault N, Mears AJ, Swaroop A, Bernier G (2004) Characterization of new transcripts enriched in the mouse retina and identification of candidate retinal disease genes. Invest Ophthalmol Vis Sci 45:3313–3319
Li M, Zhang SS, Barnstable CJ (2003) Developmental and tissue expression patterns of mouse Mpp4 gene. Biochem Biophys Res Commun 307:229–235
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Sreerama N, Woody RW (2000) Estimation of protein secondary structure from CD spectra: comparison of CONTIN, SELCON, and CDSSTR methods with an expanded reference set. Anal Biochem 287:252–260
Hennebold JD, Tanaka M, Saito J, Hanson BR, Adashi EY (2000) Ovary-selective genes I: the generation and characterization of an ovary-selective complementary deoxyribonucleic acid library. Endocrinology 141:2725–2734
Kinouchi R, Kinouchi T, Hamamoto T, Saito T, Tavares A, Tsuru T, Yamagami S (2006) Distribution of CESP-1 protein in the corneal endothelium and other tissues. Invest Ophthalmol Vis Sci 47:1397–1403
Lehrer SS, Fasman GD (1965) Excimer fluorescence in liquid phenol, p-ethylphenol, and anisole. J Am Chem Soc 87:4687–4691
Ardell MD, Bedsole DL, Schoborg RV, Pittler SJ (2000) Genomic organization of the human rod photoreceptor cGMP-gated cation channel beta-subunit gene. Gene 245:311–318
Colville CA, Molday RS (1996) Primary structure and expression of the human b-subunit and related proteins of the rod photoreceptor cGMP-gated channel. J Biol Chem 271:32968–32974
Grunwald ME, Yu W-P, Yu H-H, Yau K-W (1998) Identification of a domain on the b-subunit of the rod cGMP-gated cation channel that mediates inhibition by calcium-calmodulin. J Biol Chem 273:9148–9157
Körschen HG, Beyermann M, Müller F, Heck M, Vantler M, Koch KW, Kellner R, Wolfrum U, Bode C, Hofmann KP, Kaupp UB (1999) Interaction of glutamic-acid-rich proteins with the cGMP signalling pathway in rod photoreceptors. Nature 400:761–766
Batra-Safferling R, Abarca-Heidemann K, Körschen HG, Tziatzios C, Stoldt M, Budyak I, Willbold D, Schwalbe H, Klein-Seetharaman J, Kaupp UB (2006) Glutamic acid-rich proteins of rod photoreceptors are natively unfolded. J Biol Chem 281:1449–1460
Pentia DC, Hosier S, Cote RH (2006) The glutamic acid-rich protein-2 (GARP2) is a high affinity rod photoreceptor phosphodiesterase (PDE6)-binding protein that modulates its catalytic properties. J Biol Chem 281:5500–5505
Lin MT, Beal MF (2006) Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443:787–795
Zeviani M, Carelli V (2007) Mitochondrial disorders. Curr Opin Neurol 20:564–571
Wang X (2001) The expanding role of mitochondria in apoptosis. Gene Dev 15:2922–2933
Green DR, Reed JC (1998) Mitochondria and apoptosis. Science 281:1309–1312
Acknowledgments
This work was supported by the following grants: the National Basic Research Program (also called the 973 Program) of China (No. 2006CB705700), the National Natural Science Foundation of China (No. 30671036). We appreciate Dr. Shaoyong Chen (BIDMC, Harvard Medical School) for discussions and reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qi, S., Wang, Y., Zhou, M. et al. A mitochondria-localized glutamic acid-rich protein (MGARP/OSAP) is highly expressed in retina that exhibits a large area of intrinsic disorder. Mol Biol Rep 38, 2869–2877 (2011). https://doi.org/10.1007/s11033-010-9948-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-9948-x