Abstract
Veneridae is a diverse, commercially important, and cosmopolitan family. Here we present the complete mitochondrial genome of the hard clam Meretrix meretrix (Bivalvia: Veneridae). The entire mitochondrial genome (mitogenome) sequence of M. meretrix is 19,826 bp in length, and contains 37 genes including 12 protein-coding genes, 2 ribosomal RNAs, and 23 tRNAs. All genes are encoded on the heavy strand. In contrast to the typical animal mitochondrial genome, it lacks the protein-coding gene ATP8, and has only one copy of the tRNASer gene, but three duplications of the tRNAGln, which is the first report among the present molluscan mtDNAs. We observed that the gene arrangement between M. meretrix and M. petechialis is same except one more tRNAGln gene in M. meretrix., and the sequence similarity is as high as 99%, indicating that M. petechialis and M. meretrix could be treated as a junior synonym of M. meretrix. Maximum Likelihood and Bayeslan analysis of 12 concatenated protein-coding amino acid sequences place the Unionidae as a sister group to other bivalves, which reflects the general opinion that the Unionidae deverged very early in Bivalvia evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The study of mitochondrial DNA (mtDNA) has always been a hot field in molecular biology. With their rapid evolutionary rate, and lack of recombination, fragments of mtDNA have been extensively used for studies of genetic structure, phylogenetics, and phylogeography at various taxonomic levels. Since complete mtDNA sequence can uncover more information about gene rearrangement and other variation at the genome level for all phyla, there have been significant increases in the number of mtDNA sequences available in recent years [1–7].
Metazoan mtDNA is a closed-circular molecule, except in some hydrozoan cnidarians where it is one or two linear molecules [8, 9]. The gene content of metazoan mtDNA typically consists of genes for 2 ribosomal subunit RNAs (small- and large-rRNA, s-rRNA and l-rRNA), for 22 tRNAs, and for 13 protein subunits (cytochrome c oxidase subunits I–III, COI-III, cytochrome b (Cytb), ATP synthase subunits 6 and 8 (ATP6 and ATP8), and NADH dehydrogenase subunits 1–6 and 4L (NDl-6, 4L). Metazoan mtDNAs range in size from 14 to 42 kb [10–12]. Typically, there are few intergenic nucleotides except for a single large non-coding region generally thought to contain elements that control the initiation of replication and transcription [13]. Size variation in mtDNA is usually due to differences in non-coding regions [14, 15], but occasionally it is due to duplications or multiplications of coding regions [10, 16–19].
Mollusca, the second largest animal phylum, exhibits much variation in the features and gene arrangement of their mitochondrial genomes. Some bivalve lineages (families Mytilidae, Unionidae, and Veneridae) have an unusual mode of inheritance for mtDNA, termed doubly uniparental inheritance [20–22]. The ATP6 and ATP8 are separated into the scaphopods and most gastropods. Most bivalves are lacking the ATP8 gene, only in Hiatella arctica and Lampsilis ornata ATP8 has been found to date. In addition, several duplicated gene have been observed in two squid mtDNAs, Watasenia scintillans and Todarodes pacificus [23]. Veneridae is a large and diverse family of Bivalvia, including over 800 extant, presumably valid, species [24], many of which are commercially important by virtue of their dominance in benthic communities [25]. Despite their colonization of all types of soft bottoms, from coastal to deep-sea areas, venerids generally exhibit few morphological differences connected with soft-tissue anatomy; this makes it difficult to identify cases of morphological parallelism among evolutionarily distant species and of shell diversification among closely related ones. The study of phylogenetic relationship is further complicated by the large number of shell polymorphisms observed in populations of the same species [25].
Hard clams, genus Meretrix (Veneridae), are commercially important bivalves, scattered through coastal areas of the Indian Ocean, Southeast Asia, China, Korean Peninsula and Japanese Archipelago. Traditionally hard clams are mainly identified based on visible shell characters. Because of the remarkable variation of shapes and patterns of shells, early researchers divided this genus into many species, such as M. meretrix, M. typical, M. petechialis, M. castanea, M. graphica, M. labiosa, M. inpudina, M. zonaria, M. lusoria, M. lyrata, M. lamarckii, M. mophina, M. exilis, etc. The taxon of the Meretrix is still in argument over several discrepancies among systematists in the world.
In the present study, we present the complete mitochondrial genome of the clam M. meretrix, a commercially important aquatic resource in China. The mtDNA features of the clam are described and compared with other molluscs.
Materials and methods
Sample collection and DNA isolation
Fifty hard clam specimens were collected from the coast of Panjin, Liaoning province, China. One of them was randomly chosen as the specimen for complete mitochondrial genome sequencing. A portion of the food muscle tissue was dissected and frozen at −80°C. Total genomic DNA was isolated using Genomic DNA Extracting Kit (TaKaRa, Dalian) following the manufacturer’s protocols and stored at −20°C.
Primer design, PCR amplification and DNA sequencing
To amplify the entire mitochondrial genome of M. meretrix, seven pairs of primers were designed based on seven partial sequences of mt genes obtained from the ESTs data from cDNA library of M. meretrix. All the PCR primers were designed that there are at least 100 bp overlapping between each fragment to ensure proper sequence assembly. The sequences of primers and PCR amplification conditions were shown in Table 1. The PCR reactions were performed with a Mastercycler gradient PCR machine (Eppendorf, Germany). The cycling was set up with an initial denaturation step at 95°C for 5 min, followed by 35 cycles comprising denaturatin at 94°C for 50 s, annealing at 60°C for 1 min and extension at 72°C for from 2 to 6 min depending on the expected length of the PCR products. The process was completed with a final extension at 72°C for 10 min.
Gene annotation, sequence analysis and genome arrangement
The sequences were assembled by using Sequence Analysis v3.4.1(Applied Biosystems), Seqman V5.05(DNASTAR) and the Blast Two Sequences program of NCBI(http://www.ncbi.nlm.nih.gov/BLAST/). Protein-coding genes, rRNA genes and the boundaries of each gene were identified with DOGMA [26], BLAST searches (http://www.ncbi.nlm.nih.gov/BLAST) and comparison with sequences of other published bivalve mitochondrial sequences including M. petechialis [27]. The tRNA genes were identified by tRNAscan-SE 1.21 by employing the cove-only search mode and the invertebrate mitochondrial genetic code for secondary structure prediction [28]. In addition, codon usage in the M. meretrix mitochondrial protein-coding genes was estimated with DnaSP 4.50.3 [29]. Mitogenome arrangements of all published species from Bivalvia class were compared. The tRNA genes were included and possible mitogenome arrangements were analyzed both manually and by utilizing the rearrangement program DERANGE II [30].
Phylogenetic analysis
Along with the complete mtDNA sequence from M. meretrix sequenced in this study, all currently available 29 bivalves mitogenome data from GenBank were used in phylogenetic analyses (Supplementary data file 2). The root of all trees was determined by using the data from Ascaris suum species as outgroup. The amino acid sequences of 12 concatenated protein-coding genes with the arrangement of COI–ND1–ND2–ND4L–COII–Cyt b–ND4–ATP6–ND3–ND5–ND6–COIII were aligned using Clustal X with the default setting [31]. Model selection for the amino acid dataset was done with ProtTest [32]. Protein maximum-likelihood tree was generated using the program PhyML, version 3.0 employing the the mtREV model of amino acid substitution for mtDNA-encoded proteins. Bayesian approach using MRBAYES 3.1.2 [33, 34] was employed to analyze the aligned data sets and trees were built using the mtREV matrix model of evolutionary change. An initial run of 30,000 generations was set to estimate the likelihood value convergence. The Markov Chain Monte Carlo analysis was run for 150,000 generations (sampling every 100 generations). After omitting the first 300 “burnin” trees, the remaining 1,200 sampled trees were used to estimate the 50% of majority rule consensus tree and the Bayesian posterior probabilities (BPP).
Results and discussion
Genome composition
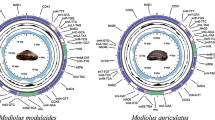
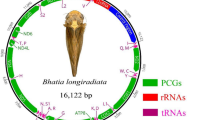
The complete mitogenome of M. meretrix is 19,826 bp in length, which is within the range of genome sizes for already sequenced molluscan mitogenomes. The mitogenome of M. meretrix contains 37 genes including 12 protein-coding genes, two ribosomal RNAs, and 23 tRNAs. All genes are encoded on the heavy strand, a common feature in marine bivalves. In contrast to the typical animal mitochondrial genome, it lacks the protein-coding gene ATP8, and has only one copy of the tRNASer gene, but three duplications of the tRNAGln gene (Fig. 1). The sequence has been deposited in GenBank under accession number NC_013188. The M. meretrix mtDNA has an overall 68.4% A + T content, and the coding regions are 16,369 bp in length accounting for 82.6% of the whole genome. There are two overlapping regions located between three adjacent tRNA genes in the genome (see Supplementary Data 1).
Gene map of the mitochondrial genome of the M. meretrix. Genes have standard abbreviations except for tRNAs, which are designated by the one-letter code for the corresponding amino acid. S2, L1, L2 designate genes for those tRNAs recognizing the codons UCN, CUN and UUR, respectively. Grey shaded parts represent non-coding regions. The grey shaded part flanked by G and Q is the putative control region
Protein-coding genes
Of the thirteen typical protein-coding genes (cox1–cox3, nad1–nad6, nad4L, cytb, atp6, and atp8), twelve were determined. No Atp8 was identified in the genome. All genes were transcribed on the same strand. These findings have been observed in all marine bivalve genomes so far published except for Hialella arctica (Family Hiatellidae), Cristaria plicata, Hyriopsis cumingii and Lampsilis ornata (Family Unionidae) [3, 7]. Thus, the placement of all coding genes on the same strand and the lack of an atp8 gene are the most distinctive features of marine bivalve mitochondrial genomes. Eight of twelve protein-coding genes of M. meretrix initiate with ATG start codon, while Cyt b, ND5 and ND6 start with ATA and COI with TTG. Eight protein-coding genes use TAA as stop codon, and the remaining four genes terminate with TAG (see Supplementary Data 1).
The bias of the base composition of an individual strand can be described by skewness [35], which measures the relative numbers of As to Ts and Gs to Cs, and is calculated as (A% − T%)/(A% + T%) and (G% − C%)/(G% + C%), respectively. The twelve protein-coding genes show a typical positive GC skew and negative AT skew (Fig. 2). Only Cyt b has a slightly skew of G vs C (GCskew = 0.238), other eleven protein-coding genes have strong skew of G vs C (GCskew = 0.288–0.538). In contrast to the AT skew of COII gene (AT skew = −0.192), a slightly negative one, the others have strong negative skew of A vs. T (AT skew = −0.244 to −0.421). The nucleotide compositions of protein-coding genes in M. meretrix are all skewed away from C in favor of G and from A in favor of T as in other bivalves [36].
The pattern of codon usage in M. meretrix mtDNA was also studied (Table 2). There are a total of 4014 codons in twelve mitochondrial protein-coding genes. Phenylalanine is the most frequent amino acid, Leucine is the second most frequent amino acid, and Glutamine is the least frequent. Individually, TTT (Phenylalanine) is definitely the most frequently used codon not only in M. meretrix, but also in other bivalves [27, 37].
Transfer and ribosomal RNA genes
The mitogenome of M. meretrix encodes 23 tRNA genes, ranging in size from 62(tRNAHis) to 71(tRNAAla) nucleotides, which can be folded into the typical clover-leaf secondary structures with several mismatch pairs within acceptor and anticodons (see Supplementary Data 2). Compared with M. petechialis, which has two duplications of tRNAGln, the M. meretrix mitogenome has three duplications of tRNAGln, which is the first report among the present molluscan mtDNAs. In the mitogenomes of metazoan, almost all amino acids codons but leucine and serine are decoded by only one tRNA each [38]. However, the duplication of tRNAAsp was observed in the mitogenome of Mizuhopecten yessoensis and it is a common phenomenon that mtDNA of most bivalves contain two tRNAMet [27, 36, 39].
Although putative gene boundaries for the two rRNA genes in the genome have been found, these cannot be accurately determined until transcript mapping is carried out. The lengths of lrRNA, located between cytb and nad4, and of srRNA, located between tRNAThr and tRNACys, are 1581 and 1187 bp, respectively; their overall A + T contents being 71% and 69.4%, respectively. The lrRNA is the largest rRNA locus reported so far within bivalves. The size and A + T content of lrRNA and srRNA in M. meretrix are same as those in M. petechialis, and larger than those in the rest bivalves.
Non-coding regions
As in most bivalves, the M. meretrix mtDNA contains a large number of unassigned nucleotides. There are as many as 29 non-coding regions, ranging in size from 2 to 1625 bp and totaling 3,457 bp, throughout the mitochondrial genome (see Supplementary Data 1). Metazoan mtDNAs usually have lengthy non-coding regions that vary in size [40, 41]. There were seven relatively large non-coding regions greater than 100 bp. The largest non-coding region with increased A + T composition was thought to contain the signals for replication and transcription, and hence was regarded to as the control region [11]. The largest non-coding region with length of 1625 bp (putative origin of replication) in M. meretrix located between tRNAGly and tRNAGln,which accounted for 47 percent of all non-coding regions. This region had a 69.9 percent of A + T content, which is slightly larger than that of the whole genome and protein-coding regions (68.4 and 66.9%). Interestingly, in the M. meretrix mitogenome, one tandem repeat comprising three nearly identical motifs was found in the area covering four adjacent non-coding regions. Each of the motifs contains one tRNAGln; it is thus obvious that the second and third copy of tRNAGln occurred as the motif duplicated. In the M. petechialis mitogenome, only two nearly identical motifs were found in the area covering three adjacent non-coding regions. Tandem repeats are common within the control region of animal mtDNAs [42]; they often form stable secondary structures and play an important role in the early stages of the replication and transcription process [43]. Tandem repeat units within non-coding regions have also been extensively found in molluscs [44, 45].
Gene arrangement
Gene arrangement comparisons may be a useful tool for phylogenetic studies. This is based on the fundamental assumption that shared gene arrangements imply common ancestry since it is highly unlikely that the same gene order would arise independently in separate lineages [46]. Thirty mitochondrial genomes available allow a comparison of the mitochondrial gene orders within bivalvia (Fig. 3). In the family Veneridae, the gene arrangement between M. petechialis and M. meretrix are the same except for one more tRNAGln gene in M. meretrix. Comparing the gene arrangement between two species of the genus Meretrix and Venerupis, they share three completely identical gene blocks. In the family Pectinidae, three conserved gene blocks were identified between Mizuhopecten yessoensis and Chlamys farreri. However, the Argopecten irradians is very different from M. yessoensis and G. farreri. Only four small conserved gene blocks could be found between A. irradians and C. farreri; The gene orders are most highly rearranged in Pectinidae. All species published of Ostridae except Crassostrea virginica shared the same mitochondrial gene arrangement. Compared with C. hongkongensis, there were some tRNA translocation and one tRNA loss in C. virginica. The overall genomic organization of C. hongkongensis is more similar to that of C. ariakensis than to C. virginica, maybe corresponding evidently to their closer genetic relationship. It is likely that gene arrangement is highly conserved among Mytilus species. Among M. galloprovincialis, M. edulis and M. trossulus, not only protein-coding genes arrangement, but also the order of tRNAs, ribosomal RNA, and control region are almost the same except for one more tRNAGln flanked by control region and tRNATyr in M. trossulus. Several genes inversion are seen in all three species published from Unionidae family, those genes encompass COI, COII, ND3, tRNAHis, ND5, ND4, ND4L, ATP8, tRNAAsp, ATP6 and COIII. Additionally, the gene arrangement of Cristaria plicata differs from that of Lampsilis ornate by only the loss of tRNAGlu. Comparing the gene arrangement between C. plicata and Hyriopsis cumingii, they share two completely identical gene blocks.
Linearized representation of gene arrangement for bivalve mitochondrial genomes. The gene arrangement of mt genomes for all species here is start from COI and all genes are transcribed from left to right except those indicated by underlining, which are transcribed from right to left. The bars show identical gene blocks and CR denote the control region. The non-coding regions are not presented. Genes for tRNAs are designated by s single letter for the corresponding amino acid: L1, L2, S1 and S2 denote tRNALeu(CUN), tRNALeu(UUR), tRNASer(AGN) and tRNASer(UCN), respectively
Among the species of Bivalvia, the ATP8 gene is absent except the species of family Unionidae. The gene orders are most highly rearranged in Bivalvia. It may because all of the genes of most species of Bivalvia are encoded on the heavy strand except the species of family Unionidae, and the asymmetric gene replication and transcription accelerate the gene rearrangement in the evolutionary process [27]. Investigation on many more bivalve mtDNAs will provide invaluable information for understanding the detailed evolutionary process of the bivalve mitogenomes.
Phylogenetic analysis
Bayesian and ML trees based on amino acid sequences of 12 concatenated protein-coding from thirty bivalves place the Unionidae as a sister group to other bivalves and reflects the general opinion that the Unionidae deverged very early in Bivalvia evolution. The Mytilidae places the grouping composed with Pectinidae and Ostridae as the sister group. The Mizuhopecten yessoensis and Chlamys gigas are closer than other Pectinidaes, and the Crassostrea angulata is closer with Crassostrea gigas than other Ostridae, which is the same as the gene rearrangement method. The Lucinidae is placed in a clade which places the grouping composed with Veneridae, Solenidae, Cardiidae and Saxicavidae as the sister group. However, the relationship among Veneridae, Solenidae, Cardiidae and Saxicavidae is not identified. The Mytilidae and Unionidae are divided into two different clades, respectively. In present, considering many questions in the phylogeny of Bivalvia remain unresolved, it is desirable to increase the resolution by adding more mitochondrial genomes. Further taxon sampling will be very useful for determining the phylogenetic relationships among the major lineages of Bivalvia (Fig. 4).
Phylogenetic tree derived from Bayesian analysis of M. meretrix relative to 29 other bivalve species. Ascaris suum was used as the out-group. The phylogenetic relationship was performed by Bayesian and ML methods. The first numbers are from Bayesian inferences and the second numbers are from maximum likelihood analysis. Tree topologies produced by the two methods were similar and only more than 50% bootstrap values are shown and others are represented by “–”
Taxonomic position of M. meretrix and M. petechialis
The comparison of mitogenome structures showed that M. petechialis and M. meretrix were most closely related species. The sequence similarity of between M. petechialis and M. meretrix is as high as 99% and their gene arrangement is same except one more tRNAGln gene in M. meretrix. Our results did not support the present taxonomic status of M. meretrix and M. petechialis, which were in accordance with viewpoint that M. petechialis and M. meretrix belong to different geographic subspecies of one species [47, 48]. M. meretrix and M. petechialis were previously considered as independent species in the genus Meretrix. It was shown that the species differed from M. petechialis by the following shell characteristics: anterior and ventral margins, posterior end of shell. The Meretrix species are widely scattered through coastal areas of the Indian Ocean, Southeast Asia, China, Korean Peninsula and Japanese Archipelago. Because of the remarkable variation of shapes and patterns of shells, early researchers divided this genus into many species, such as M. meretrix, M. typical, M. petechialis, M. castanea, M. graphica, M. labiosa, M. inpudina, M. zonaria, M. lusoria, M. lyrata, M. lamarckii, M. mophina, M. exilis, etc. To date, nine species are generally recognized in Meretrix [49]. Zhuang pointed out that Meretrix has a little species and only three species, M. meretrix, M. lusoria and M. lamarckii, have been found in China [50]. It was pointed out that M. meretrix is a species which experienced the greatest variation in the group of bivalves, and because of shades of shells and shell colors, it was wrongly divided into many species [51]. In summary, the views of taxonomic status of M. meretrix and M. petechialis are ambiguous based on morphology. The viewpoint that M. petechialis and M. meretrix should be treated as a junior synonym of M. meretrix.
Abbreviations
- Atp6 and 8:
-
Genes encoding ATPase subunits 6 and 8
- bp:
-
Base pairs
- coxI-III:
-
Genes encoding cytochrome oxidase subunits I–III
- Cyt b:
-
Gene encoding cytochrome b
- ND1-6 and 4L:
-
Genes encoding NADH dehydrogenase subunits 1–6 and 4L
- lrRNA and srRNA:
-
Large (16S) and small (12S) ribosomal RNA subunit (gene)
- tRNA:
-
Transfer RNA
- L1:
-
tRNALeu(CUN)
- L2:
-
tRNALeu(UUR)
- S2:
-
tRNASer(UCN)
- NR:
-
Non-coding region
References
Helfenbein KG, Brown WM, Boore JL (2001) The complete mitochondrial genome of the articulate brachiopod Terebratalia transversa. Mol Biol Evol 18:1734–1744
Giorgi CD, Martiradonna A, Lanave C, Saccone C (2002) Complete sequence of the mitochondrial DNA in the sea urchin Arbacia lixula: conserved features of the echinoid mitochondrial genome. Mol Phylogenet Evol 5:323–332
Serb JM, Lydeard C (2003) Complete mtDNA sequence of the North American freshwater mussel, Lampsilis ornata (Unionidae): an examination of the evolution and phylogenetic utility of mitochondrial genome organization in Bivalvia (Mollusca). Mol Biol Evol 20:1854–1866
Boore JL, Medina M, Rosenberg LA (2004) Complete sequences of the highly rearranged molluscan mitochondrial genomes of the scaphopod Graptacme eborea and the bivalve Mytilus edulis. Mol Biol Evol 21:1492–1503
Milbury CA, Gaffney PM (2005) Complete mitochondrial DNA sequence of the eastern oyster Crassostrea virginica. Mar Biotechnol 7:697–712
Maynard BT, Kerr LJ, McKiernan JM, Jansen ES, Hanna PJ (2005) Mitochondrial DNA sequence and gene organization in the Australia blacklip abalone Haliotis rubra (Leach). Mar Biotechnol 7:645–658
Dreyer H, Steiner G (2006) The complete sequence and gene organization of the mitochondrial genomes of the heterodont bivalves Acanthocardia tuberculata and Hiatella arctica—and the first record for a putative Atpase subunit 8 gene in marine bivalves. Front Zool 3:13
Warrior R, Gall J (1985) The mitochondrial DNAo f Hydra attenuata and Hydra littoralis consists of two linear molecules. Arch Sci 38:439–445
Bridge D, Cunningham CW, Schierwater B, DeSalle R, Buss LW (1992) Class-level relationships in the phylum Cnidaria: evidence from mitochondrial genome structure. Proc Natl Acad Sci USA 89:8750–8753
Moritz C, Brown WM (1987) Tandem duplications in animal mitochondrial DNAs: variation in incidence and gene content among lizards. Proc Natl Acad Sci USA 84:7183–7187
Wolstenholme DR (1992) Animal mitochondrial DNA: structure and evolution. Int Rev Cytol 141:173–216
Snyder M, Fraser AR, Laroche J, Gartner-Kepkay KE, Zouros E (1987) Atypical mitochondrial DNA from the deep-sea scallop Placopecten magellanicus. Proc Natl Acad Sci USA 84:7595–7599
Shadel GS, Clayton DA (1997) Mitochondrial DNA maintenance in vertebrates. Annu Rev Biochem 66:409–435
Brown WM (1985) The mitochondrial genome of animals. In: Macintyre RJ (ed) Molecular evolutionary genetics. Plenum Press, New York, pp 95–130
Harrison RG (1989) Animal mitochondrial DNA as a genetic marker in population and evolutionary biology. Trends Ecol Evol 4:6–11
Azevedo J, Hyman BC (1993) Molecular characterization of lengthy mitochondrial DNA duplications from the parasitic nematode Romanomermis culicivorax. Genetics 133:933–942
Fuller KM, Zouros E (1993) Dispersed length polymorphism of mitochondrial DNA in the scallop Placopecten magellanicus (Gmelin). Curr Genet 23:365–369
Moritz C, Brown WM (1986) Tandem duplications of D-loop and ribosomal RNA sequences in lizard mitochondrial DNA. Science 233:1425–1427
Zevering CE, Moritz C, Heideman A, Sturm RA (1991) Parallel origins of duplications and the formation of pseudogenes in mitochondrial DNA from parthenogenetic lizards (Heteronotia binoei : Gekkonidae). J Mol Evol 33:431–441
Zouros E, Ball AO, Saavedra C, Freeman KR (1994) An unusual type of mitochondrial DNA inheritance in the blue mussel Mytilus. Proc Natl Acad Sci USA 91:7463–7467
Liu HP, Mitton JB, Wu SK (1996) Paternal mitochondrial DNA differentiation far exceeds maternal mitochondrial DNA and allozyme differentiation in the freshwater mussels, Anodonta grandis grandis. Evolution 50:952–957
Passamonti M, Scalim V (2001) Gender-associated mitochondrial DNA heteroplasmy in the venerid clam Tapes philippinarum (Mollusca Bivalvia). Curr Genet 39:117–124
Akasaki T, Nikaido M, Tsuchiya K, Segawa S, Hasegawa M, Okada N (2006) Extensive mitochondrial gene arrangements in coleoid Cephalopoda and their phylogenetic implications. Mol Phylogenet Evol 38:648–658
Mikkelsen PM, Bieler R, Kappner I, Rawlings TA (2006) Phylogeny of Veneroidea (Mollusca: Bivalvia) based on morphology and molecules. Zool J Linn Soc 148:439–521
Canapa A, Schiaparelli S, Marota I, Barucca M (2003) Molecular data from the 16S rRNA gene for the phylogeny of Veneridae (Mollusca: Bivalvia). Mar Biol 142:1125–1130
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255
Ren J, Shen X, Sun M, Jiang F, Yu Y, Chi Z, Liu B (2009) The complete mitochondrial genome of the clam Meretrix petechialis (Mollusca: Bivalvia: Veneridae). Mitochondrial DNA 20:78–87
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Blanchette M, Kunisawa T, Sankoff D (1996) Parametric genome rearrangement. Gene 172: GC11–GC17
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21(9):2104–2105
Huelsenbeck JP, Ronquist F (2001) MRBAYES: bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Perna NT, Kocher TD (1995) Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J Mol Evol 410:353–358
Wu X, Xu X, Yu Z, Kong X (2009) Comparative mitogenomic analyses of three scallops(Bivalvia: Pectinidae) reveal high level variation of genomic organization and a diversity of transfer RNA gene sets. BMC Res Notes 2:69
Yu Z, Wei Z, Kong X, Shi W (2008) Complete mitochondrial DNA sequence of oyster Crassostrea hongkongensis—a case of “Tandem duplication-random loss” for genome rearrangement in Crassostrea? BMC Genomics 9:477
Podsiadlowski L, Braband A, Mayer G (2008) The complete mitochondrial genome of the onychophoran Epiperipatus biolleyi reveals a unique transfer RNA set and provides further support for the ecdysozoa hypothesis. Mol Biol Evol 25:42–51
Breton S, Burger G, Stewart DT, Blier PU (2006) Comparative analysis of gender-associated complete mitochondrial genomes in marine mussels (Mytilus spp.). Genetics 172:1107–1119
Jacobs HT, Elliott DJ, Math VB, Farquharson A (1988) Nucleotide sequence and gene organization of sea urchin mitochondrial DNA. J Mol Biol 202:185–217
Boyce TM, Zwick ME, Aquadro CF (1989) Mitochondrial DNA in the bark weevils: sizes, structure and heteroplasmy. Genetics 123:825–836
Lunt DH, Whipple LE, Hyman BC (1998) Mitochondrial DNA variable number tandem repeats (VNTRs): utility and problems in molecular ecology. Mol Ecol 7:1441–1455
Willkinson GS, Chapman AM (1991) Length and sequence variation in evening bat D-loop mtDNA. Genetics 128:607–617
Boore JL, Medina M, Rosenberg LA (2004) Complete sequences of the highly rearranged molluscan mitochondrial genome of the scaphopod Graptacme eborea and the bivalve Mytilus edulis. Mol Biol Evol 21:1492–1503
Tomita K, Yokobori S, Oshima T, Ueda T, Watanabe K (2002) The cephalopod Loligo bleekeri mitochondrial genome: multiplied noncoding regions and transposition of tRNA genes. J Mol Evol 54:486–500
Boore JL, Brown WM (1998) Big trees from little genomes: mitochondrial gene order as a phylogenetic tool. Curr Opin Genet Dev 8:668–674
Pan BP, Wu Q, Zhang SP, Song LS, Bu WJ (2006) Molecular phylogeny of Meretrix (Mollusca, Bivalvia) based on 16S rRNA and ITS1 sequences. Oceanologia Etlmnologia Snica 37(4):342–347
Ai-hui CHEN, Zhao-xia LI, Gong-neng FENG (2009) Phylogenetic relationships of the genus Meretrix (Mollusca:Veneridae) based on mitochondrial COI gene sequences. Zool Res 30(3):233–239
Ayako YY, Masashi Y, Hideyuki I (2008) Genetic relationships among species of Meretrix (Mollusca: Veneridae) in the Western Pacific Ocean. Pac Sci 62(3):385–394
Zhuang QQ (2001) Fauna sinica, phyllum Mollusca, class Bivalvia, family Veneridae [M]. Science Press, Beijing, pp 229–236
Prashad B (1932) The Lamellibranchia of the Siboga expedition [M]. Sysmatic part. Siboga-Expenditie, LIIIc, pp 212–264
Acknowledgments
This work was supported by National Nature Science Foundation of China (No.:30972246), Liaoning Key Project of Sci-Tech (No.: 2008203001) and Liaoning scientific research program of ocean and fisheries department (No.:200801). We give thanks to the two anonymous reviewers who provided constructive comments that significantly improved the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
He, Cb., Wang, J., Gao, Xg. et al. The complete mitochondrial genome of the hard clam Meretrix meretrix . Mol Biol Rep 38, 3401–3409 (2011). https://doi.org/10.1007/s11033-010-0449-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0449-8