Abstract
The transcription factor AP-1 plays an important role in cellular proliferation, transformation and death. In this study, we report a novel human gene, AC3-33 (GenBank name: c3orf33, FLJ31139), which encodes a secretory protein that can inhibit Elk1 transcriptional activity via ERK1/2 pathway. The AC3-33 mRNA encodes a protein of 251 amino acids, which is a classical secretory protein. Functional investigation reveals that overexpression of AC3-33 significantly inhibit AP-1 activity and DNA-binding ability. Further investigation indicated that overexpression of AC3-33 significantly inhibit transcriptional activity of Elk1 and c-jun, but not c-fos. As for the upstream of signaling pathway of Elk-1, our study demonstrated that overexpression of AC3-33 significantly down-regulates phosphorylation of ERK1/2, but not JNK/SAPK or p38 MAPK. These results clearly indicate that AC3-33 is a novel member of the secretory family and inhibits Elk1 transcriptional activity via ERK1/2 MAPK.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
After the completion of the human genome project, it is urgent to identify thousands of novel genes that have been deposited in the human Refseq and EST databases in GenBank, most of which with unknown or poorly understood functions. We have established several high-throughput functional gene-screening systems based on overexpression in human cell lines, including the cell-based assays using reporter genes and so on [1, 2].
The pathway of transcription factor activator protein-1 (AP-1) plays important roles in the regulation of cellular proliferation, transformation and death [3]. Because the disorder of these physiological processes has been linked with many tumorous processes, such as neoplastic transformation, tumor progression, metastasis, AP-1 has been described as a major culprit in cancer [4–6]. Particularly, it is well characterized that AP-1 is a double-edged sword in tumorigenesis, either antioncogenic by inducing apoptosis or oncogenic by signaling cell survival. The outcome of AP-1 activity in tumors seems to depend on AP-1 dimer composition as well as the cellular and genetic contexts [7]. Therefore, identification of key modulating genes is anticipated to be useful in thorough elucidation of AP-1 signal pathway, as well as the prevention and/or treatment of cancer.
AC3-33, a novel human gene, also known as chromosome 3 open reading frame 33, encodes 251 amino acids with a predicted molecular mass of 29.3 kDa. Expression of the AC3-33 gene was widely found in adrenal glands and cervix [8]. Bioinformatics analysis showed that the amino acid sequences of AC3-33 was highly conserved, but had no homology to other known proteins. Particularly, AC3-33 was predicted to be a secretory protein with a putative signal peptide (http://www.cbs.dtu.dk/services/SignalP/). To our knowledge, no functional study has been performed on this hypothetical gene.
Using dual-luciferase reporter assay system, we found AC3-33 significantly inhibited AP-1 transcriptional activity. In this paper, we described AC3-33, a novel secretory protein, and investigated its functional role in signal transduction. Functional studies indicated that overexpression of AC3-33 significantly inhibit AP-1 transcription activity. Moreover, AC3-33 significantly inhibited transcriptional activity of Elk1 and phosphorylation of ERK1/2. Data are presented suggesting that AC3-33 might be a novel human MAPK regulating protein.
Materials and methods
Reagents
Restricted endo-nucleases XhoI and BamHI were purchased from TaKaRa (Japan). PMA, monoclonal antibodies against β-actin were purchased from Sigma-Aldrich (USA). Cell lysis buffer (10×), polyclonal antibodies against phosphorylated ERK1/2, JNK/SAPK and P38 were purchased from Cell Signaling Technology (USA). IRDye™ 800-conjugated secondary antibodies against mouse and rabbit IgG were purchased from LI-COR Bioscience (USA). Ionomycin was from Santa Cruz Biotechnology (USA). Protease inhibitor cocktail tablets were obtained from Roche Applied Science (Switzerland).
cDNA cloning and vector construction
The full-length coding region of AC3-33 was amplified from the mixed Matchmaker™ human testis, brain and spleen cDNA Library (Clontech, USA) by PCR using the specific primers: upper (5′-GGCTGGAACCACAGAACATCAGCACTGGA-3′) and lower (5′-AACGGTCAAGAAATCTAGAGGTAACCTTCA-3′). The N-terminal truncated mutation, AC3-33-ΔSP, was amplified using the specific primers: upper (5′-AAACTCGAGACCATGTCGGATATTCCAGTAGAATTTA-3′) and lower (5′-GAGCGGATCCCACCCTTTTCTACGAAAGTTTA-3′). The purified PCR product was ligated into the pGEM-T Easy vector (Promega, USA). The insert was released by XhoI and BamHI, and subcloned into pcDNA.3.1/myc-his(-)B (Invitrogen, USA) to construct plasmids of PCDB–AC3-33 and PCDB–AC3-33-ΔSP (Fig. 1a). To construct PEGFP–AC3-33, the corresponding primers was used for fusion with N-terminal GFP tag in the PEGFP-C3 (Clontech, USA) vector with XhoI and BamHI. All clones were confirmed by sequencing using an ABI Prism® 3100 Genetic Analyzer (Applied Biosystems, USA). After confirmation, all plasmids were extracted and purified for transfection using EndoFree Plasmid Maxi Kit (Qiagen, USA).
Cell culture and transient transfections
HEK 293T (human embryonic kidney cell line, a kind gift from T. Matsuda, Japan) and HeLa (human cervical carcinoma cell line) cells were maintained in DMEM (LifeTechnologies, USA) supplemented with 10% fetal bovine serum (Hyclone, USA) and Lglutamine (2 mM) at 37°C in a humidified 5% CO2. DNA transfection was performed using VigoFect (Vigorous, USA), a non-liposomal cationic formula, according to the manufacturer’s instructions.
RNA isolation and real time reverse transcription-PCR
Total RNAs were extracted using TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. One microgram of RNA was subjected to reverse transcription using SuperscripII (Invitrogen). The reactions were incubated at 42°C for 50 min. The AC3-33 primers, upper (5′-CACTGCAACCTCCTGCTTC-3′) and lower (5′-TAGCTCACGCCTGTAATC-3′), and the GAPDH795/771 primers (as a control), upper (5′-GAGCCAAAAGGGTCATCATCT-3′) and lower (5′-AGGGGCCATCCACAGTCTTC-3′). Real time PCR was performed on Takara TP600 Real time PCR detection system. The cycling conditions were 95°C for 5 min followed by 40 cycles of 95°C for 15 s, 65°C for 35 s, 72°C for 30 s, then 72°C for 5 min, store at 4°C. Each sample was run in triplicate. Data were analyzed by Mylab Corporation.
Protein extraction and Western blot analysis
Immunoblot analysis was conducted as previously described [9] with a slight modification. Briefly, cells were washed twice with ice-cold phosphate-buffered saline (PBS) and lysed in cell lysis buffer (20 mM Tris–HCl, pH 7.5, 150 mM NaCl, 1 mM Na2 EDTA, 1 mM EGTA, 2.5 mM sodium pyrophosphate, 1 mM β-glycerophosphate, 1 mM Na3VO4, 1 mg/ml leupeptin, 1% Triton X-100, with freshly added proteinase inhibitor cocktail) for 30 min at 4°C. Cell lysate was clarified by centrifugation at 4°C at 16,000×g for 15 min. Protein concentration was determined using the BCA protein assay reagent (Pierce, USA). Equal amounts of protein were separated by 12.5% SDS-PAGE and transferred onto nitrocellulose membranes (Amersham Pharmacia, UK). Membranes were blocked in Tris-buffered saline containing 0.1% Tween-20 (TBS-T) and 5% nonfat milk for 2 h, and incubated overnight at 4°C with the appropriate primary antibody. After washing in TBST buffer, membranes were incubated for 1 h in the dark with the appropriate IRDye™ 800-conjugated secondary antibodies. Signals were detected on an Odyssey Infrared Imager (LI-COR Bioscience, USA).
Polyclonal anti-AC3-33 antibody preparation
Antibodies against AC3-33 were generated by immunization of rabbits with KLH-coupled peptide (YFSVNLNEEILRRGLGKTVLVKGLKYDSKIYWTVHRNLLK), which were synthesized by solid phase synthesis and purified by HPLC to 95% purity (Chinese Peptide, China). Rabbit polyclonal antibodies were purified using CNBR Sepharose 4B coupled with specific AC3-33 peptide. The antibodies were validated by ELISA and Western blot analysis according to standard procedures as above (data not shown).
Expression of mammalian recombinant protein and immunoprecipitation
PEGFP–AC3-33 plasmid (20 mg) containing a gfp-tag at the N terminus was transiently transfected into HeLa cells (3 × 106) by VigoFect as previously described. The transfected cells were cultured in the media without serum (Gibco, USA) for 24 h, and then harvested for purification. The transfected cell supernatant was combined with 3 μl anti-gfp antibody and protein G beads at 4°C, for overnight, followed by three washes with wash buffer (50 mM Tris, pH 8, 150 mM NaCl, 5 mM MgCl2, 0.4% Nonidet P-40). Bound proteins were analyzed by Western blot according to standard procedures as above.
Dual-luciferase reporter assay
AP-1 luciferase activity was measured using cis-reporting system. Approximately 1.0 × 104 293T or HeLa cells per well were seeded into a 96-well culture plate. After 24 h, the cells in each well were co-transfected with 80 ng of the PCDB–AC3-33 or PCDB–AC3-33ΔSP plasmids, 40 ng of the pAP-1-Luc plasmids containing the firefly luciferase reporter gene (PathDetect, Stratagene), and 4 ng of the pRL-TK plasmids as the internal control containing the Renilla luciferase gene (Promega, Madison, WI). The negative control was performed with PCDB plasmid. Each transfection experiment was performed in triplicate wells. At 24 h after transfection, the cells were stimulated with PMA (12-O-tetradecanoylphorbol-13-acetate) (50 mM, sigma) and Ionomycin (1 mM, Calbiochem, USA) for 6 h, and then lysed in standard lysis buffer. Using a GENios Pro reader (Tecan, Switzerland) the cell lysate were assayed for both firefly and renilla luciferase activities according to the manufacturer’s instructions. Each independent experiment was performed three times.
Elk1, c-jun and c-fos luciferase activities were measured using trans-reporting systems as previously described [9] with a slight modification, respectively. For Elk1 activity detection, approximately 1.0 × 104 293T cells/well was seeded into a 96-well culture plate. After 24 h, the cells in each well were cotransfected with 80 ng PCDB–AC3-33 or PCDB–AC3-33ΔSP plasmids, 40 ng pFR-Luc plasmid, 4 ng pFA-Elk1 fusion trans-activator plasmid (PathDetect, Stratagene) and 4 ng pRL-TK plasmids as the internal control (Promega, Madison, WI). For c-jun and c-fos activities detection, 4 ng pFA-c-jun and 4 ng pFA-c-fos were used to substitute pFA-Elk1, respectively. The negative control was performed with PCDB plasmid. Each transfection experiment was performed in triplicate wells. At 36 h after transfection, the cells were stimulated with PMA (50 mM) and ionomycin (1 mM) for 6 h. The cells were collected and the cell lysate were measured as above.
Electrophoretic mobility shift assay (EMSA)
24 h after transfection, cells were treated with PMA (50 mM) and ionomycin (1 mM) for 6 h. The treated cells (2.0 × 106) were trypsinized, washed twice with PBS and pelleted by centrifugation. The nuclear protein extracts were prepared with the NEPER Nuclear and Cytoplasmic Extraction Reagents (Pierce, USA) according to the protocols. The oligonucleotides for AP-1 consensus and AP-1 mutant were obtained from Santa Cruz Biotechnology and were end-labeled with biotin. Nuclear extracts (5 μg) were incubated with labeled DNA probes (50 fmol) for 30 min at room temperature in a total 20 μl binding reaction containing 10 × binding buffer, 50% glycerol, 1 μg/μl poly (dI–dC), according to the protocols of LightShift™ Chemiluminescent EMSA Kit (Pierce, USA). The DNA–protein complexes were separated from free DNA probe through 5% non-denaturing polyacrylamide gel electrophoresis (PAGE) in 0.5× TBE buffer. Then the gel was transferred to nylon membrane. The following steps were in accordance with the instructions of the above EMSA Kit (Pierce, USA). For competition experiments, 100 folds of unlabeled DNA were co-incubated with labeled probes and nuclear extracts.
Phosphorylation of MAPKs and immunoblot
Approximately 2.0 × 105 293T cells per well were seeded into a 6-well culture plate. After 24 h, the cells in each well were co-transfected with 5 μg of the PCDB–AC3-33 plasmids, and 5 μg of the PCDB–MEKK plasmids as the internal positive stimulant. The negative control was performed with PCDB plasmid. Each experiment was performed in triplicate wells. Following incubation (24 h, 37°C), transfected cells were harvested and lysated. Cell lysate were analyzed by Western blot as above.
Results
Cloning and bioinformatics analysis of human AC3-33
The full-length coding region of AC3-33 cDNA (GenBank accession No. NM_173657; C3orf33) was directly isolated from a mixed human testis, brain and spleen cDNA Library. It is 756 bp long with the putative ATG start codon and termination codon. The ORF encodes 251 amino acids with a predicted molecular mass of 29.3 kDa and a pI of 9.9 (http://ca.expasy.org/tools/pi_tool.html). The full-length cDNA and predicted amino acid sequences of AC3-33 were shown in our previous report [8]. Human AC3-33 is located on chromosome 3q25.31, and encompasses 6 exons and 5 introns. AC3-33 is conserved in Homo sapiens, Macaca mulatta, Rattus norvegicus, Mus musculus and Canis familiaris, etc., which suggests that AC3-33 play an important role in mammals. AC3-33 shares no obvious homology to any known genes or proteins. Particularly, bioinformatics analysis suggests that there is a putative signal peptide near the N terminus of the protein, with most likely cleavage site between pos. 22 and 23: TSS-SD.
AC3-33 is a classical secretory protein
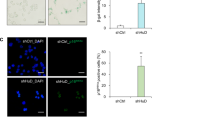
To confirm that AC3-33 is a secretory protein, we constructed the full-length coding PEGFPC3–AC3-33, and transiently transfected the construct to cells to express N-GFP-tagged human AC3-33 proteins. The cell supernatant was immunoprecipitated with anti-gfp antibody and protein G beads. The bound proteins and the cell lysate were performed Western blot analysis. We detected specific protein bands in cell lysate transfected with PEGFPC3–AC3-33, but not in cell lysate transfected with PEGFPC3. We also detected specific protein bands in cell culture supernatant transfected with PEGFPC3–AC3-33, but not in cell culture supernatant transfected with PEGFPC3 (Fig. 1b). Particularly, the results demonstrated that AC3-33 exhibited a single 29-kDa form in both transfected cell lysate and transfected cell culture supernatant, which is similar to the predicted AC3-33 protein in molecular weight.
Expression profile of AC3-33
The expression profile of AC3-33 was analyzed by real time PCR in various human tissues. Expression of AC3-33 on mRNA level can be detected in not only normal tissues but also abnormal tissues, such as colon tissue, ovary tissue, and breast tissue. Especially at a high level in the normal lleocecal tissue, kidney cancer tissue, endometrium cancer tissue, normal endometrium tissue (Fig. 2).
Over-expression of AC3-33 suppresses AP-1 activity
Using dual-luciferase cis-reporter assay system, we found that over-expression of either AC3-33 or AC3-33-ΔSP significantly suppressed AP-1 reporter gene activity in the absence or presence of AP-1 stimulus. PMA plus ionomycin was the potent activator of AP-1 signaling [10, 11]. As shown in Fig. 3a, PMA plus ionomycin treatment enhanced AP-1 luciferase activity markedly. The AP-1 activity of AC3-33 under stimulation was significantly inhibited compared with the PCDB empty vector (approximately 35% of that in the empty vector). The same luciferase results were observed in Hela cells (data not shown). Taken together, these data showed that over-expression of AC3-33 inhibited AP-1 activity.
Over-expression of AC3-33 suppresses AP-1 activity, as well as AP-1 translocation into the nucleus or binding with DNA. a AP-1 luciferase activity of AC3-33 or AC3-33-ΔSP overexpression was measured using cis-reporting AP-1 system. Cells were transiently cotransfected with the pAP-1-Luc plasmids and pRL-TK plasmid. Relative luciferase activity was normalized by co-transfection with pRL-TK plasmid (internal control). b Electrophoretic mobility shift assay (EMSA) analysis of AP-1 activation. The mobility of the shifted AP-1 consensus probe is shown by the arrow. Compared with the empty vector (pcDB), AC3-33 diminished AP-1 complex binding to the AP-1 probes in 293T cells
AC3-33 inhibits AP-1 translocation into the nucleus or binding with DNA
EMSA analysis was then used to assess the effect of AC3-33 on nuclear extract protein binding to AP-1 specific probes. As shown in Fig. 3b, PMA plus ionomycin treatment markedly induced an AP-1 complex specific retardation band in the empty vector (PCDB) or AC3-33 transfected cells. Free probe or excess unlabeled competitor probe, eliminates the AP-1 complex specific band, confirming the specificity of the interaction. Compared with the empty vector (PCDB), AC3-33 diminished AP-1 complex binding to the AP-1 probes in 293T cells. Similar result was obtained in HeLa cells (data not shown). These results in combination with the luciferase reporter assay suggest that over-expression of AC3-33 may inhibit AP-1 translocation into the nucleus or binding with DNA.
Over-expression of AC3-33 inhibits Elk1 and c-jun transcriptional activity, but not c-fos
To identify the specific transcription factor which mediate the process AC3-33 suppressed the AP-1 activity, dual-luciferase trans-reporter assay system were used. As shown in Fig. 4a, PMA plus ionomycin treatment enhanced Elk1, c-jun and c-fos luciferase activity markedly. The Elk1 and c-jun activity of AC3-33 under stimulation was significantly inhibited compared with the PCDB empty vector (approximately 45% and 68% of that in the empty vector, respectively, Fig. 4a, b), whereas c-fos activity had hardly changed (Fig. 4c). Furthermore, overexpressed truncated AC3-33-ΔSP (the putative signal peptide was deleted) could significantly inhibited Elk1 and c-jun activation in stimulated cells, but not c-fos. The same luciferase results were observed in HeLa cells (data not shown). These data suggest that AC3-33 might play a key role in the regulation of transcription factor Elk1.
Over-expression of AC3-33 inhibits Elk1 and c-jun transcriptional activity, but not c-fos. a Elk1 luciferase activity of AC3-33 or AC3-33-ΔSP overexpression was measured using trans-reporting Elk1 system. Cells were transiently cotransfected with the pFR-luc reporter plasmid, pFA-Elk1 fusion activator plasmid and pRL-TK plasmid. Relative luciferase activity was normalized by co-transfection with pRL-TK plasmid (internal control). The negative control was performed with PCDB plasmid. b c-jun luciferase activity was measured using trans-reporting c-jun system. pFA-c-jun pFA-c-fos were used to substitute pFA-Elk1. c c-fos luciferase activity was measured using trans-reporting c-fos system. pFA-c-fos were used to substitute pFA-Elk1
AC3-33 inhibits ERK1/2, but not SAPK/JNK or p38 MAPK signaling pathway
To define the mechanism by which AC3-33 inhibited PMA and ionomycin-induced Elk1 activation, we assessed the phosphorylation status of the upstream signaling pathway of Elk-1. Because there were three main subgroups in the Mammalian MAPK family that affected Elk1, ERK1/2, JNK/SAPK and p38 MAPK [12], we detected the protein levels of ERK1/2, JNK/SAPK and p38 MAPK as well as their phosphorylation levels. As shown in Fig. 4a, AC3-33-overexpressing cells had a lower phosphorylation level of ERK1/2 compared with cells transfected with pCDB. Conversely, AC3-33 had no effect on phosphorylation of JNK/SAPK (Fig. 5b) or p38 MAPK (Fig. 5c). These data indicated that the inhibition of AC3-33 on Elk1 transcriptional activity was uniquely mediated by the ERK1/2 MAPK.
Overexpression of AC3-33 inhibits ERK1/2, but not SAPK/JNK or p38 MAPK signaling pathway. AC3-33 inhibits the ERK1/2 pathway. The effect of AC3-33 on phosphorylation of ERK1/2 (a), SAPK/JNK (b) or p38 pathway (c). The presence of phosphorylated proteins was monitored by Western blot using phospho-specific antibodies as indicated. Loading of equal amounts of proteins was determined by restraining of the blots with antibodies directed against Erk1/2, JNK or p38. a Cells were transfected with PCDB as negative control; b co-transfected with 5 μg of the PCDB and PCDB–MEKK plasmids; c co-transfected with 5 μg of the PCDB–AC3-33 and PCDB–MEKK plasmids
Discussion
In our previous study, we identified AC3-33 as a potentially novel functional gene based on a search of EST sequences in GenBank and screening with dual-luciferase reporter assay system [8]. In present study, bioinformatics analysis indicated the presence of a potential signal peptide at the N terminus of AC3-33, suggesting that it might be a secretory protein. Using immunoprecipitation, we purified the proteins from cell culture supernatant, and Western blot analysis indicated that recombinant AC3-33 protein could be secreted into cell culture supernatant. Since many secretory proteins are thought to rely upon transmembrane cargo receptors for efficient ER-to-Golgi transport [13], further researches will be performed to validate whether the 22 amino acids at the N terminus act as a signal peptide.
AC3-33 was predicted to have an N-glycosylation site (position 231–234) and four N-myristoylation sites (position 77–82, 119–124, 134–139 and 204–209), so that modifications could be made during the protein passed through the ER and the Golgi apparatus [14].
We also studied the effect of AC3-33 on the transcriptional activity of AP-1. First, we demonstrated that overexpression of AC3-33 inhibited AP-1 activity, as well as AP-1 translocation into the nucleus or binding with DNA in 293T and HeLa cells. Previous studies indicate that the transcription activity of AP-1 is mainly mediated by c-jun, c-fos and/or Elk-1 [15]. In present study, AC3-33 was identified to exert its functional role in the unique Elk1 and c-jun signaling pathway, but not c-fos. Elk-1 (also known as p62 ternary complex factor, TCF), is an Ets-domain transcription factor activated by MAPK that binds to the serum response element (SRE) to induce immediate early gene transcription, in response to extracellular stimuli such as serum and growth factors [16].
Within the Elk1 signaling pathway, the evolutionarily conserved mitogen-activated protein kinase (MAPK) family consists of serine/threonine-specific protein kinases, involved in signal transduction pathways between the membrane and the nucleus [17]. Additionally, we illuminated that overexpression of AC3-33 significantly inhibited phosphorylation of ERK1/2, but not JNK/SAPK in 293T and HeLa cells. Interestingly, we also observed that overexpression of AC3-33 weakly inhibited phosphorylation of p38, further experiments should be need to validate the result. MAPKs transduce diverse extracellular stimuli (mitogenic growth factors, environmental stresses and proapoptotic agents) to the nucleus via kinase cascades to regulate proliferation, DNA synthesis arrest, differentiation, and apoptosis [18, 19]. MAPKs become activated through phosphorylation of specific threonines and tyrosines by an upstream dual specificity kinase. Regulation of the signaling pathway occurs via the sequential phosphorylation and activation of each member of the kinase cascade [20, 21].
AC3-33 could inhibit the MAPK pathway by two possible ways [22]. First, as a secretory protein, AC3-33 might inhibit MAPK phosphorylation via an unidentified receptor on the plasma membrane. There are many known growth factor receptors on the plasma membrane, which initiate biochemical cascades or signal transduction pathways [23–25]. Second, intracellular AC3-33 directly inhibited ERK1/2 pathway via unknown cascades or signal transduction. Further researches will be performed to elucidate the process.
In conclusion, the AC3-33 gene was detected through high-throughput functional screening systems, and was identified as being a secretory protein that played an important role in AP-1/Elk1 transcription regulation and MAPK signal transduction via the ERK1/2, but not JNK/SAPK or p38 MAPK, pathways. These results contribute to a better understanding of the complex mechanism of AC3-33 as well as AP-1 transcription regulation.
Abbreviations
- MAPK:
-
Mitogen-activated protein kinase
- MAPKK, MKK or MEK:
-
MAPK kinase
- MEKK:
-
MAPKK kinase or MEK kinase
- PMA:
-
Phorbolmyrlstate acetate
- AP-1:
-
Activation protein 1
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- ERK:
-
Extracellular signal regulated protein kinase
- Elk1:
-
The ETS-domain transcription factor
References
He PF, Peng Z, Luo Y, Wang L, Yu P, Deng WW, An YQ, Shi TP, Ma DL (2009) High-throughput functional screening for autophagy-related genes and identification of TM9SF1 as an autophagosome-inducing gene. Autophagy 5:52–60
Ma X, Wang X, Gao X, Wang L, Lu Y, Gao P, Deng W, Yu P, Ma J, Guo J, Cheng H, Zhang C, Shi T, Ma D (2007) Identification of five human novel genes associated with cell proliferation by cell-based screening from an expressed cDNA ORF library. Life Sci 81:1141–1151
Angel P, Karin M (1991) The role of Jun, Fos and the AP-1 complex in cell-proliferation and transformation. Biochim Biophys Acta 1072:129–157
Young MR, Li JJ, Rincón M, Flavell RA, Sathyanarayana BK, Hunziker R, Colburn N (1999) Transgenic mice demonstrate AP-1 (activator protein-1) transactivation is required for tumor promotion. Proc Natl Acad Sci USA 96:9827–9832
Li JJ, Westergaard C, Ghosh P, Colburn NH (1997) Inhibitors of both nuclear factor-kappaB and activator protein-1 activation block the neoplastic transformation response. Cancer Res 57:3569–3576
Bode AM, Dong Z (2000) Signal transduction pathways: targets for chemoprevention of skin cancer. Lancet Oncol 1:181–188
Eferl R, Wagner EF (2003) AP-1: a double-edged sword in tumorigenesis. Nat Rev Cancer 3:859–868
Liu P, Deng WW, Gao P, Lu Y, Sun B, Li M, Zhao J, Shi TP, Zhang XJ (2008) Molecular cloning and preliminary function study of a novel human gene AC3-33 related to suppress AP-1 activity. Yi Chuan (Chinese) 30:575–582
Ma X, Zhao HS, Shan JX, Long F, Chen YY, Chen YY, Zhang YM, Han X, Ma DL (2007) PDCD10 interacts with Ste20-related kinase MST4 to promote cell growth and transformation via modulation of the ERK pathway. Mol Biol Cell 18:1965–1978
Qi M, Elion EA (2005) MAP kinase pathways. J Cell Sci 118:3569–3572
Su B, Karin M (1996) Mitogen-activated protein kinase cascades and regulation of gene expression. Curr Opin Immunol 8:402–411
Yang S-H, Sharrocks AD, Whitmarsh AJ (2003) Transcriptional regulation by the MAP kinase signaling cascades. Gene 320:3–21
Andrea CB, Bin Z (2007) Receptor-mediated protein transport in the early secretory pathway. Trends Biochem Sci 32:381–388
Apweiler R, Hermjakob H, Sharon N (1999) On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim Biophys Acta 1473:4–8
Gallo A, Cuozzo C, Esposito I, Maggiolini M, Bonofiglio D, Vivacqua A, Garramone M, Weiss C, Bohmann D, Musti AM (2002) Menin uncouples Elk-1, JunD and c-Jun phosphorylation from MAP kinase activation. Oncogene 21:6434–6445
Yang SH, Sharrocks AD (2006) Convergence of the SUMO and MAPK pathways on the ETS-domain transcription factor Elk-1. Biochem Soc Symp 73:121–129
Cobb MH (1999) MAP kinase pathways. Prog Biophys Mol Biol 71:479–500
Turnier C, Hess P, Yang DD, Xu J, Turner TK, Nimnual A, Bar SD, Jones SN, Flavell RA, Davis RJ (2000) Requirement of JNK for stress-induced activation of the cytochrome c-mediated death pathway. Science 288:870–874
Karin M (1998) Mitogen-activated protein kinase cascades as regulators of stress responses. Ann N Y Acad Sci 851:139–146
Pearson EJ, Wilsbacher G, Swantek J, Karandikar J, Xu M, Cobb MH (1999) New insights into the control of MAP kinase pathways. Exp Cell Res 253:255–270
Chang L, Karin M (2001) Mammalian MAP kinase signalling cascades. Nature 410:37–40
Huang J, Shi T, Ma T, Zhang Y, Ma X, Lu Y, Song Q, Liu W, Ma D, Qiu X (2008) CCDC134, a novel secretory protein, inhibits activation of ERK and JNK, but not p38 MAPK. Cell Mol Life Sci 65:338–349
Simonsen A, Tooze SA (2009) Coordination of membrane events during autophagy by multiple class III PI3-kinase complexes. J Cell Biol 186:773–782
Fadeel B, Xue D (2009) The ins and outs of phospholipid asymmetry in the plasma membrane: roles in health and disease. Crit Rev Biochem Mol Biol 44:264–277
Söderberg SS, Karlsson G, Karlsson S (2009) Complex and context dependent regulation of hematopoiesis by TGF-beta superfamily signaling. Ann N Y Acad Sci 1176:55–69
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (30671092), the National High Technology Research and Development Program (863 Program) of China (2006AA02A305), the Natural Science Foundation of Hebei Province (C2009001260), the Central Public-Interest Scientific Institution Basal Research Fund (2009GJSSJKB03), and the Key National S&T Program—“Major New Drug Development” (2009ZX09503-004).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Dongxia Hao, Peng Gao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Hao, D., Gao, P., Liu, P. et al. AC3-33, a novel secretory protein, inhibits Elk1 transcriptional activity via ERK pathway. Mol Biol Rep 38, 1375–1382 (2011). https://doi.org/10.1007/s11033-010-0240-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0240-x