Abstract
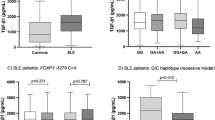
It has been reported that stromal cell-derived factor-1 (SDF1), currently also designated CXCL12, plays a significant role in the development of nephritis and death in the lupus mice model. Using restriction length fragment polymorphism (RFLP) analysis we assessed the frequencies of SDF1-3′ G801A (rs 1801157) polymorphic variants between systemic lupus erythematosus (SLE) patients (n = 150) and controls (n = 300). There were no significant differences in the prevalence of SDF1-3′ G801A polymorphic variants in SLE patients and healthy individuals. However, we observed that the SDF1-3′ A/A and G/A genotypes (recessive model) contributed to renal manifestations of SLE OR = 3.042 (95% CI = 1.527–6.058, P = 0.002), and the p value stayed statistically significant after Bonferroni correction (p corr = 0.032) in SLE patients. We also found an association of the SDF1-3′ A/A and G/A genotypes (recessive model) with dermal manifestations of SLE OR = 2.510 (95% CI = 1.247–5.052, P = 0.0122), (p corr = 0.1952) but this did not remain statistically significant after Bonferroni correction. Our observations suggest that the SDF1-3′ G801A genotype may be associated with some clinical manifestations in patients with SLE.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Systemic lupus erythematosus (SLE) is a chronic, multiorgan, and systemic autoimmune disorder [1]. SLE is characterized by defective functioning of the immune system, resulting in autoantibody production and inflammatory manifestations in several organs [1]. Immune cells from SLE patients display various aberrations, including skewed cytokine production, defective function of CD4+ T cells, abnormal activation of B cells, and reduction of cytotoxic T cell function [2–6]. Occupational exposure, drugs, chemicals, food, viruses and other infectious factors may contribute to this alteration in the immune system [7, 8]. Genetic factors also play a significant role in the susceptibility to SLE incidence [9–11]. In particular, numerous genes encoding disparate proteins regulating the immune system pathways are candidates to be the susceptibility locus in the development of SLE [9].
Chemokines encompass a collection of small molecular weight (8–14 kDa) chemotactic cytokines, which bind to specific G-protein-coupled plasma cell membrane receptors [12]. Chemokines recruit hematopoietic and immune cells to sites of differentiation and inflammation, respectively [12, 13]. Stromal cell-derived factor-1 (SDF1), which is currently also designated CXCL12, is a chemokine involved in organogenesis, lymphopoiesis, and myelopoiesis [14, 15].
In the lupus model NZB/W mice, SDF1 contributes to the chemotaxis, proliferation, and survival of PerB1a lymphocyte subpopulations that express a self-reactive repertoire of antibodies [16, 17]. Balabanian et al. [17] showed that SDF1 played a significant role in the production of anti-dsDNA antibodies, nephritis, and death in NZB/W mice.
It has been demonstrated that SDF1 has a G801A transition at position 801 in the 3′-untranslated region of the transcript, known as SDF1-3′A (rs 1801157) [18, 19]. This polymorphism may have an important regulatory function via an increase in the biosynthesis of SDF1 protein [18, 19]. We investigated the prevalence of SDF1-3′ G801A genotypes and alleles in patients with SLE (n = 150) and controls (n = 300) from a Polish cohort. We also examined the association of SDF1-3′ G801A genotypes with clinical manifestations and the presence of autoantibodies in patients with SLE.
Patients and methods
Patients and controls
One hundred and fifty consecutive SLE patients (all women) enrolled at the Institute of Rheumatology in Warsaw, Poland were included in the present study (Table 1). Patients fulfilled at least four of the American College of Rheumatology 1982 revised criteria for SLE [20, 21]. Both patients and control groups were of Polish Caucasian origin. The control group included three hundred healthy women. The mean age of healthy individuals was 37.6 ± 8.6 years. The protocol of the investigation was approved by the Local Ethical Committee of Poznań University of Medical Sciences. Written informed consent was signed by all participating individuals.
Genotyping
DNA was obtained from peripheral blood leucocytes by salt extraction. Polymorphic variants of SDF1-3′ G801A (rs 1801157) were identified using PCR with the primer pair 5′-TTATTGTACTTGCCTTATTAGAG-3′ and 5-′GTAGTTCACCCCAAAGGACC-3′; enzyme digestion followed the identification process. The PCR-amplified fragments of SDF1 that were 732 bp in length were isolated and subjected to digestion with MspI (C/CGG). The SDF1-3′G allele was digested into 456 and 276 bp fragments, whereas the SDF1-3′A allele stayed uncut at a size of 732 bp. DNA fragments were separated by electrophoresis on 3% agarose gel and visualized by ethidium bromide staining. Polymorphism confirmation was performed by sequencing analysis.
Statistical analysis
The distribution of genotypes in all groups was tested for deviation from Hardy–Weinberg equilibrium. Fisher exact test was used to determine differences in the genotypic and allelic distribution between patients and controls. Moreover, the Odds Ratio (OR) and 95% Confidence Intervals (CI) were calculated. A P value <0.05 was considered statistically significant. Associations between clinical manifestations, the production of autoantibodies, and polymorphism distribution in patients with SLE were determined by Fisher exact test.
Results
Genotype assessment of SDF1 G801A polymorphisms disclosed no significant deviation from Hardy–Weinberg equilibrium in any group.
We found a significant association between SDF1-3′ A/A and G/A genotypes (recessive model) and, respectively, renal manifestations of the disease OR = 3.042 (95% CI = 1.527–6.058, P = 0.002), and the P value stayed statistically significant after Bonferroni correction (p corr = 0.032) (Table 1). However, the statistically significant association of SDF1-3′ A/A and G/A genotypes (recessive model) with dermal manifestations OR = 2.510 (95% CI = 1.247–5.052, P = 0.0122), (p corr = 0.1952) did not remain statistically significant after Bonferroni correction (Table 1).
We did not observe a significant variation in the distribution of SDF1 G801A polymorphic variants in SLE patients and controls (Table 2). OR for SLE patients with SDF1-3′ A/A genotype was 2.034 (95% CI = 0.5796–7.142, P = 0.3123) and OR of the SDF1-3′ A/A and A/G genotypes was 1.309 (95% CI = 0.8660–1.978), P = 0.2029 (Table 2). We also did not find a correlation between the SDF1-3′ A allele or AA genotype with SLE disease activity.
Discussion
The human genome project disclosed more than ten million single nucleotide polymorphisms. Despite this vast number of polymorphisms, the role of most of them in the incidence of various disorders remains elusive [22].
The human SDF1 gene is located at 10q11.1 and its transcription produces alpha and beta alternative splice variants [23]. Translation of both transcripts produces the SDF1 chemokine, which binds to only one receptor, CXCR4 [24]. SDF1 is a chemoattractant for lymphocytes, megakaryocytes, endothelial cells, and stem cells [24–26], and is involved in the development of neuronal, cardiac, vascular, and craniofacial systems [27].
To date, the SDF1-3′A gene variant has been considered a factor in increased susceptibility to lymphoma, oral and squamous carcinomas, and cancers of the breast, lung, and prostate [28–33]. Schröppel et al. [34] observed that the SDF1-3′A variant is significantly associated with higher mortality in liver allograft recipients. Moreover, the SDF1-3′A allele has been associated with the microvascular involvement in systemic sclerosis and with age-at-onset of type 1 diabetes mellitus in the Japanese population [35, 36].
We did not observe a difference in the contribution of the SDF1-3′A gene variant between SLE patients and healthy individuals. These findings are consistent with Lima et al. [37], who also did not find differences between SDF1-3′ G801A genotypes and allele prevalence in SLE patients and controls. However, they found that SLE patients with antiphospholipid syndrome exhibited a higher distribution of SDF1-3′ A/A genotype frequencies compared with patients without this syndrome [37].
We did not observe a contribution of SDF1-3′ G801A genotypes to antiphospholipid syndrome (results not shown). However, we observed a significant association of SDF1-3′ AA and GA genotypes with renal manifestations of SLE. There was also a higher distribution of SDF1-3′ A/A and A/G genotypes in patients with dermal manifestations compared with patients without dermal manifestations. The discrepancies between Lima’s and our findings can be due to differences in the racial structure of these investigated groups. Moreover, exposure to varying environmental factors in patients with distinct SDF1-3′ G801A genotypes can also have a disparate effect on SLE manifestation [7].
The SDF1 receptor, CXCR4, is also considered a co-receptor for entry of the human immunodeficiency virus (HIV) into immune cells [38]. The genetic association analysis of 2,857 patients disclosed that homozygous SDF1-3′A/A individuals had delayed onset of AIDS [18]. Since SDF1 competes with HIV for entry into target cells, this delay in the onset of AIDS may suggest that SDF1-3′A variants are responsible for higher SDF1 biosynthesis [18].
Balabanian et al. [17] observed an increased production of SDF1 in the podocytes of glomeruli of NZB/W mice. They also reversed lupus nephritis in NZB/W mice following SDF1 neutralization [17]. Moreover, Robak et al. [39] indicated that patients with SLE exhibited a higher median serum level of SDF1 than healthy individuals.
Our observations suggest that the SDF1-3′A variant, which may lead to a higher production of the SDF1 chemokine, may be associated with renal manifestations of SLE. The SDF1-3′A can also be associated with a gene with which SDF1 interacts and is not in linkage disequilibrium but this gene contributes to renal SLE manifestations. However, to more precisely determine the significance of the SDF1-3′A G801A polymorphism in SLE manifestation or incidence, additional examination of the prevalence of these variants in other cohorts is required.
References
Sekigawa I, Naito T, Hira K et al (2004) Possible mechanisms of gender bias in SLE: a new hypothesis involving a comparison of SLE with atopy. Lupus 13:217–222
Crispín JC, Tsokos GC (2008) Novel molecular targets in the treatment of systemic lupus erythematosus. Autoimmun Rev 7:256–261
Januchowski R, Wudarski M, Chwalińska-Sadowska H et al (2008) Prevalence of ZAP-70, LAT, SLP-76, and DNA methyltransferase 1 expression in CD4(+) T cells of patients with systemic lupus erythematosus. Clin Rheumatol 27:21–27
Stohl W, Metyas S, Tan SM et al (2003) B lymphocyte stimulator overexpression in patients with systemic lupus erythematosus: longitudinal observations. Arthritis Rheum 48:3475–3486
Gröndal G, Gunnarsson I, Rönnelid J et al (2000) Cytokine production, serum levels and disease activity in systemic lupus erythematosus. Clin Exp Rheumatol 18:565–570
al-Janadi M, al-Balla S, al-Dalaan A et al (1993) Cytokine profile in systemic lupus erythematosus, rheumatoid arthritis, and other rheumatic diseases. J Clin Immunol 13:58–67
Jönsen A, Bengtsson AA, Nived O et al (2007) Gene–environment interactions in the aetiology of systemic lupus erythematosus. Autoimmunity 40:613–617
Love LA (1994) New environmental agents associated with lupus-like disorders. Lupus 3:467–471
Wong M, Tsao BP (2006) Current topics in human SLE genetics. Springer Semin Immunopathol 28:97–107
Hom G, Graham RR, Modrek B et al (2008) Association of systemic lupus erythematosus with C8orf13-BLK and ITGAM-ITGAX. N Engl J Med 358:900–909
International Consortium for Systemic Lupus Erythematosus Genetics (SLEGEN), Harley JB, Alarcón-Riquelme ME, Criswell LA et al (2008) Genome-wide association scan in women with systemic lupus erythematosus identifies susceptibility variants in ITGAM, PXK, KIAA1542 and other loci. Nat Genet 40:204–210
Moser B, Loetscher P (2001) Lymphocyte traffic control by chemokines. Nat Immunol 2:123–128
Zlotnik A, Yoshie O (2000) Chemokines: a new classification system and their role in immunity. Immunity 12:121–127
Zou YR, Kottmann AH, Kuroda M et al (1998) Function of the chemokine receptor CXCR4 in a haematopoiesis and in cerebellar development. Nature 393:595–599
Onai N, Zhang Y, Yoneyama H et al (2000) Impairment of lymphopoiesis and myelopoiesis in mice reconstituted with bone marrow-hematopoietic progenitor cells expressing SDF-1-intrakine. Blood 96:2074–2080
Herzenberg LA (2000) B-1 cells: the lineage question revisited. Immunol Rev 175:9–22
Balabanian K, Couderc J, Bouchet-Delbos L et al (2003) Role of the chemokine stromal cell-derived factor 1 in autoantibody production and nephritis in murine lupus. J Immunol 170:3392–3400
Winkler C, Modi W, Smith MW et al (1998) Genetic restriction of AIDS pathogenesis by an SDF-1 chemokine gene variant. Science 279:389–393
Watanabe MA, de Oliveira Cavassin GG, Orellana MD et al (2003) SDF-1 gene polymorphisms and syncytia induction in Brazilian HIV-1 infected individuals. Microb Pathog 35:31–34
Tan EM, Cohen AS, Fries JF et al (1982) The 1982 revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum 25:1271–1277
Hochberg MC (1997) Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum 40:1725
International HapMap Consortium, Frazer KA, Ballinger DG, Cox DR et al (2007) A second generation human haplotype map of over 3.1 million SNPs. Nature 449:851–861
Shirozu M, Nakano T, Inazawa J et al (1995) Structure and chromosomal localization of the human stromal cell-derived factor 1 (SDF1) gene. Genomics 28:495–500
Bleul CC, Fuhlbrigge RC, Casasnovas JM et al (1996) A highly efficacious lymphocyte chemoattractant, stromal cell-derived factor 1 (SDF-1). J Exp Med 184:1101–1109
Yong K, Fahey A, Reeve L et al (1999) Cord blood progenitor cells have greater transendothelial migratory activity and increased responses to SDF-1 and MIP-3beta compared with mobilized adult progenitor cells. Br J Haematol 107:441–449
Hamada T, Möhle R, Hesselgesser J et al (1998) Transendothelial migration of megakaryocytes in response to stromal cell-derived factor 1 (SDF-1) enhances platelet formation. J Exp Med 188:539–548
McGrath KE, Koniski AD, Maltby KM et al (1999) Embryonic expression and function of the chemokine SDF-1 and its receptor, CXCR4. Dev Biol 213:442–456
Teng YH, Liu TH, Tseng HC et al (2009) Contribution of genetic polymorphisms of stromal cell-derived factor-1 and its receptor, CXCR4, to the susceptibility and clinicopathologic development of oral cancer. Head Neck. doi:10.1002/hed.21094
Khademi B, Razmkhah M, Erfani N et al (2008) SDF-1 and CCR5 genes polymorphism in patients with head and neck cancer. Pathol Oncol Res 14:45–50
de Oliveira Cavassin GG, De Lucca FL, Delgado Andre N et al (2004) Molecular investigation of the stromal cell-derived factor-1 chemokine in lymphoid leukemia and lymphoma patients from Brazil. Blood Cells Mol Dis 33:90–93
Razmkhah M, Talei AR, Doroudchi M et al (2004) Stromal cell-derived factor-1 (SDF-1) alleles and susceptibility to breast carcinoma. Cancer Lett 225:261–266
Razmkhah M, Doroudchi M, Ghayumi SM et al (2005) Stromal cell-derived factor-1 (SDF-1) gene and susceptibility of Iranian patients with lung cancer. Lung Cancer 49:311–315
Hirata H, Hinoda Y, Kikuno N et al (2007) CXCL12 G801A polymorphism is a risk factor for sporadic prostate cancer susceptibility. Clin Cancer Res 13:5056–5062
Schröppel B, Fischereder M, Ashkar R et al (2002) The impact of polymorphisms in chemokine and chemokine receptors on outcomes in liver transplantation. Am J Transplant 2:640–645
Manetti M, Liakouli V, Fatini C et al (2009) Association between a stromal cell-derived factor 1 (SDF-1/CXCL12) gene polymorphism and microvascular disease in systemic sclerosis. Ann Rheum Dis 68:408–411
Ide A, Kawasaki E, Abiru N (2003) Stromal-cell derived factor-1 chemokine gene variant is associated with type 1 diabetes age at onset in Japanese population. Hum Immunol 64:973–978
Lima G, Soto-Vega E, Atisha-Fregoso Y et al (2007) Llorente LMCP-1, RANTES, and SDF-1 polymorphisms in Mexican patients with systemic lupus erythematosus. Hum Immunol 68:980–985
Bleul CC, Farzan M, Choe H et al (1996) The lymphocyte chemoattractant SDF-1 is a ligand for LESTR/fusin and blocks HIV-1 entry. Nature 382:829–833
Robak E, Kulczycka L, Sysa-Jedrzejowska A et al (2007) Circulating proangiogenic molecules PIGF, SDF-1 and sVCAM-1 in patients with systemic lupus erythematosus. Eur Cytokine Netw 18:181–187
Acknowledgements
This study is supported by a Grant No. 502-01-01124182-07474 Poznań University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Warchoł, T., Lianeri, M., Łącki, J.K. et al. SDF1-3′ G801A polymorphisms in Polish patients with systemic lupus erythematosus. Mol Biol Rep 37, 3121–3125 (2010). https://doi.org/10.1007/s11033-009-9890-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-009-9890-y