Abstract
Ginseng (Panax ginseng C. A. Mey.) is widely used as a major medicinal herb and as a feedstock for the medicine, beverage, food, cosmetic, etc. industries, in China and several other Asian countries. However, limited research has been accomplished into its genetics, genomics and breeding. To clone, characterize and utilize the genes of economic importance in the species, we have developed a large-insert plant-transformation-competent binary bacterial artificial chromosome (BIBAC) library for Jilin ginseng cv. Damaya. The library contains 141,312 clones, with an average insert size of 110 kb, each likely containing approximately 20–30 genes. The clones of the library have all been arrayed in 384-well microplates and permanently archived. We screened the library and identified BIBAC clones containing nine genes likely involved in the biosynthesis pathway of ginsenosides—the major medicinally effective compounds of ginseng—with approximately four BIBACs per gene. This result further verified the quality of the library and demonstrated its utility in cloning, characterization and utilization of economically important genes in ginseng. Furthermore, since the library is cloned in a plant-transformation-competent BIBAC vector (pCLD04541) that can be directly transformed in a variety of plants via both the Agrobacterium-mediated method and the particle bombardment method, we have also demonstrated the stability of large-insert ginseng DNA BIBACs in different Agrobacterium strains, which is crucial to large-insert BIBAC transformation in plants. Therefore, the Jilin ginseng BIBAC library provides resources and tools useful for functional genomics research, and cloning, characterization and utilization of economically important genes in the species.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Ginseng (Panax ginseng C. A. Mey.) is known as the “king of herbs” in China, Korea, Japan and several other eastern Asian countries. It has been cultivated as a medicinal herb for over 2,000 years in China. Because ginseng has been shown to have numerous bioactive effects on human health, such as immune system stimulation, anti-carcinogenic activity, alleviation of fatigue stress and reduction of blood glucose levels, it has been widely used as a herbal medicine and feedstock, especially its major medically effective component, ginsenosides, in a number of industries, including medicine, beverage, food, cosmetic, etc. Most of the world’s ginseng is produced in northeast Asia, 70 % in the Changbai Mountains, Jilin, China. Currently, the annual business revenue stimulated by ginseng in Jilin, China exceeds US $3.0 billion. It is estimated that this number will approach US $15.0 billion by 2020.
However, ginseng research in the past has been focused on chemical components and their medical bioactivities (Popovich et al. 2012). Research into modern genetics, genomics and breeding of the species is limited. This limits ginseng genetic improvement and the isolation, characterization and utilization of genes economically important in ginseng, such as those involved in the biosynthesis pathway of ginsenosides. To facilitate such research, hundreds of DNA markers including randomly amplified polymorphic DNA, inter-simple sequence repeat, amplified fragment length polymorphism, restricted fragment length polymorphism and simple sequence repeat (Bang et al. 2004; In et al. 2005; Kim et al. 2005; Ma et al. 2007; van Dan et al. 2010), over 10,000 expressed sequence tags (ESTs) (Jung et al. 2003) and a large-insert bacterial artificial chromosome (BAC) library (Hong et al. 2004) have been developed. These DNA markers, ESTs and BAC library have provided useful resources and tools for modern genomics, genetics and breeding research in ginseng, but they are not sufficient for all research purposes and are underdeveloped, in comparison with other economically important plant species such as rice and tomato. Moreover, most of these researches were carried out with Korean ginseng; few genome resources and tools have been developed for Chinese Jilin ginseng, even though it contributes over 70 % of the world’s ginseng to ginseng industries. Finally, the existing ginseng BAC library (Hong et al. 2004) only has a genome coverage of 3.34× that is not well-suited for many genomic studies and is not competent for direct transformation through either Agrobacterium or biolistic bombardment, which is necessary for functional analysis and use of economically important genes for ginseng breeding and molecular farming through genetic engineering.
Ginseng is perennial, usually taking 3–4 years from planting to seed setting, and special soils and climates are required for its growth and production. Moreover, research and technical practices for breeding of the species are limited, even though it has been cultivated for over 2,000 years. These limitations significantly restrict ginseng genetic improvement and production. It has been demonstrated in several plant species that the binary bacterial artificial chromosome (BIBAC) technology promises to provide efficient tools for enhanced plant genetic improvement, molecular breeding, and isolation, characterization and utilization of economically important genes for agricultural production (Liu et al. 1999; Song et al. 2004; Chang et al. 2011). Compared with BACs, BIBACs have several advantages in their use for genomics research, especially functional and post-genomics research, and molecular breeding. This is because BIBACs not only are capable of cloning and stably maintaining DNA fragments of over 300 kb in bacteria as BACs (Hamilton et al. 1996; Tao and Zhang 1998), but also can be directly used for high-molecular-weight (HMW) DNA transformation in plants by the particle bombardment (Chang et al. 2011) and Agrobacterium-mediated methods (Hamilton et al. 1996; Tao and Zhang 1998; Liu et al. 1999). The plant transformation competence of BIBACs expedites many functional genomics, molecular breeding and molecular farming researches, including economically-important gene and quantitative trait locus (QTL) cloning and genetic engineering (Zhang et al. 2012b). Moreover, a BIBAC having an insert size of 100–150 kb potentially contains 10–30 genes (Chang et al. 2011). Transforming such a large-insert BIBAC allows transfer of the genes or gene clusters and regulatory elements involved in a single pathway or biological process (Song et al. 2004), such as the genes involved in the biosynthesis of ginsenosides in ginseng. Furthermore, large-insert BIBAC transformation is well suited for engineering genes from a distantly or unrelated species: for instance, engineering the genes involved in the biosynthesis of ginsenosides from ginseng to tractably cultivated crop plants such as carrot and tomato for molecular farming. This is because the native regulatory sequences and neighboring genes of the donor species are crucial to the expression of transgenes in host species (Song et al. 2004; Chang et al. 2011). Therefore, large-insert BIBAC transformation will more likely produce transgenics that express the target genes in an appropriate spatial and temporal manner due to the reduced problems of gene silencing that are frequently encountered in the traditional gene transformation method.
Techniques for large-insert BIBAC transformation have been well established in plants, either via Agrobacterium-mediated transformation (Hamilton et al. 1996; Liu et al. 1999, 2002; Frary and Hamilton 2001; He et al. 2003) or by particle bombardment (Chang et al. 2011). Large-insert BIBACs transformed into plants are inherited stably (Hamilton et al. 1996; Frary and Hamilton 2001; Song et al. 2004; Chang et al. 2011), and the transgenes contained in large-insert BIBACs are actively expressed, yielding new or varied traits in host plants (Song et al. 2004; Chang et al. 2011). Therefore, BIBACs are emerging as an important tool for gene/QTL cloning (Hamilton et al. 1996; Zhang 2007) and large-scale genome functional analysis (Song et al. 2004; Chang et al. 2011). A large number of large-insert plant-transformation-ready BIBAC libraries have been constructed for a variety of plant species (e.g., Liu et al. 1999, 2002; Men et al. 2001; He et al. 2003; Chang et al. 2001; Meksem et al. 2000; Xu et al. 2005; Zhang et al. 2010, 2012b, c). Nevertheless, no large-insert BIBAC library has yet been available for ginseng.
In this study, we constructed a large-insert BIBAC library from Chinese Jilin ginseng cv. Damaya, a major cultigen of ginseng in northeastern China. To characterize the library, further validate its genome coverage and to identify the clones containing the genes potentially involved in the biosynthesis pathway of ginsenosides, we printed the library on nylon membranes in high clone density and screened it using overgo probes designed from nine genes potentially involved in the pathway. We isolated a total of 36 positive BIBAC clones, with an average of 4.0 positive clones per gene probe. Moreover, we also tested the transformability and stability of ginseng BIBAC clones in Agrobacterium, because they are crucial to BIBAC transformation in plants (Song et al. 2003; Chang et al. 2011). Therefore, this BIBAC library provides resources and tools for modern genome research in several aspects, particularly ginseng molecular breeding, and isolation, characterization and utilization of its economically important genes in molecular farming.
Materials and methods
Plant materials
Chinese Jilin ginseng cv. Damaya was used as the source DNA of the BIBAC library. Fresh root systems were collected from the plants growing in the field and used as materials for preparation of megabase-sized nuclear DNA.
Preparation of megabase-sized nuclear DNA
Megabase-sized nuclear DNA was prepared according to Zhang et al. (1995, 2008, 2012a). Because ginseng roots are abundant in starch and polysaccharides that potentially influence the digestion of the resultant DNA embedded in agarose, nuclei were washed for two additional times to minimize the contamination of these metabolites. The nuclei were suspended at a concentration of about 1.5 × 107 nuclei/ml for preparation of low-melting-point agarose plugs, thus making approximately 5 μg nuclear DNA per 100-μl plug.
Preparation of BIBAC vector
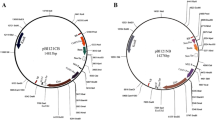
The BIBAC library was constructed in the widely-used BIBAC vector pCLD04541 (Tao and Zhang 1998; Meksem et al. 2000; Wu et al. 2000; Men et al. 2001; Tao et al. 2002; Fang et al. 2004; Ortiz-Vázquez et al. 2005; Feng et al. 2006; Zhang et al. 2010; Chang et al. 2011). The isolation and purification of the vector DNA and vector preparation followed Zhang et al. (2012b).
Library construction
The BIBAC library was constructed as described by Zhang et al. (2012b). We first conducted partial digestion tests using a series of amounts of BamHI per reaction to determine the optimal condition, particularly the amount of restriction enzyme (BamHI) per reaction and digestion incubation time. The partial digestion tests indicated that 20 units of BamHI per reaction containing three of the nine slices derived from a 100-μl megabase-sized DNA plug and 8-min incubation at 37 °C generated the fragments with most being between 80 and 200 kb. Therefore, the condition was selected for large-scale partial digestion for the BIBAC library construction.
We partially digested 15 100-μl megabase-sized DNA plugs with 20 units of BamHI per reaction at 37 °C for 8 min to construct the BIBAC library. The partially digested megabase-sized DNA was selected on a 1 % agarose gel by pulsed-field gel electrophoresis. The fragments ranging from 80 to 200 kb were selected, electroeluted, dialyzed and ligated into the dephosphorylated pCLD04541 vector (Zhang et al. 2012b). The ligated DNA was transformed into Escherichia coli strain DH10B (Invitrogen, USA) by electroporation using the Cell Porator™ Device (Gibco BRL, USA) as described by Zhang et al. (2012b).
Library characterization and identification of clones containing genes important in ginseng
A random sample of 124 clones was analyzed to estimate the insert size of the library according to Zhang et al. (2012b). BIBAC DNA was isolated, digested with NotI and run on a 1 % agarose gel by pulsed-field gel electrophoresis. The insert size of each clone was estimated by adding up all insert band(s) of each clone lane using the lambda ladder PFG marker (New England BioLabs, USA) as the molecular weight standard.
The BIBAC library was double-spotted onto 22.5 × 22.5 cm Hybond N+ membrane (Amersham-Pharmacia, USA) in a 4 × 4 format using the GeneTAC Robotic Workstation (Genomic Solutions, USA). Each membrane contained a total of 18,432 (48 × 384) double-printed clones and the entire library was spotted on 7.5 membranes (Zhang et al. 1996a; Zhang 2000). The library was screened by hybridization with overgo probes designed from the single-copy regions of nine genes potentially involved in the biosynthesis pathway of ginsenosides, using a two-step procedure. The library membranes were first hybridized with a probe prepared from a mixture of nine gene-specific overgos, then, the positive clones were re-arrayed into a 96-well plate, printed onto nylon membrane and re-hybridized with probes prepared from individual overgos of the genes. The hybridization was carried out at 60 °C overnight. After the hybridization, the membranes were washed in 1× SSC, 0.1 % (w/v) SDS at 60 °C twice, 15 min each time, followed by 0.5× SSC, 0.1 % SDS (Sambrook et al. 1989) at 60 °C for 10 min.
Stability of large-insert ginseng genomic DNA BIBACs in Agrobacterium
Panax ginseng has a large complex genome (3,300 Mb/1C, Hong et al. 2004), which may affect the stability of its large-insert BIBACs in Agrobacterium (Song et al. 2003). To explore the feasibility of transforming the ginseng genomic DNA large-insert BIBACs in plants via Agrobacterium for ginseng molecular breeding and ginseng gene molecular farming through genetic engineering, we randomly selected two large-insert BIBACs from the ginseng library with insert sizes of approximately 100 and 150 kb, respectively. The two clones were transformed into Agrobacterium tumefaciens strains COR308 and C1C58, respectively, by electroporation using the Cell Porator™ Device (Gibco BRL, USA) and tested for their stability in the strains. The conditions of electroporation were 4-kΩ resistance for the Voltage Booster setting; and low-ohm impedance, 330-μF capacitance and 360-V for the Cell-Porator settings. Two–five Agrobacterium BIBAC clones were selected from each BIBAC Agrobacterium transformation and grown in LB medium containing appropriate antibiotics at 28 °C, 250 rpm for 100 generations. DNA of the clone Agrobacterium cultures was isolated and re-transformed into E. coli strain DH10B (Invitrogen, USA) by electroporation as described above. Random bacterial clones were selected from the re-transformed BIBACs, and DNA of the clones was isolated and analyzed against the original BIBAC DNA used in the Agrobacterium transformation by pulsed-field gel electrophoresis (Zhang et al. 2012b).
Results and discussion
Construction and characterization of the BIBAC library
We constructed a BIBAC library from the megabase-sized nuclear DNA of Jilin ginseng cv. Damaya partially digested with BamHI in the BIBAC vector pCLD04541 (Jones et al. 1992; Tao and Zhang 1998). This vector has been widely used to construct BIBAC libraries for a variety of plant species and its transformability demonstrated in Agrobacterium and plants (Meksem et al. 2000; Wu et al. 2000; Men et al. 2001; Tao et al. 2002; Fang et al. 2004; Ortiz-Vázquez et al. 2005; Feng et al. 2006; Zhang et al. 2010; Chang et al. 2011). We isolated megabase-sized nuclear DNA from ginseng roots. Because they are abundant in metabolic substances such as starch and polysaccharides that may influence large-insert DNA library construction, we washed the nuclei extensively to minimize them, thus enhancing the DNA clonability. Moreover, we also conducted pre-size selection on the megabase-sized DNA embedded in agarose plugs to further minimize the metabolic substances and to remove the small fragments in the plugs (Zhang et al. 2012b). These measures significantly facilitated the construction of the BIBAC library.
The ginseng BIBAC library contains 143,312 clones arrayed in 368 384-well microtiter plates. To determine the insert sizes of the library clones, we analyzed a random sample of 124 clones from the library (Fig. 1a). The result showed that the clones of the library had insert sizes ranging from 45 to 200 kb, with an average insert size of 110 kb (Fig. 1b). Of the 124 clones analyzed, 97 (78.23 %) had insert sizes larger than 80 kb, each thus likely containing 10–30 genes. Therefore, the BIBACs could be possibly used to engineer clusters of genes involved in a biological process or pathway such as those for the biosynthesis of ginsenosides. Since ginseng is estimated to have a genome size of some 3,300 Mb/1C (Hong et al. 2004), the BIBAC library has a haploid genome coverage of 4.8×, with a probability of >99 % of obtaining at least one positive clone from the library using a single-copy probe (Zhang et al. 1996b; Wu et al. 2004). Therefore, the library provides useful resources and tools for genomics and genetics research on the species. Furthermore, since the library was constructed in a BIBAC vector that is competent in plant transformation with either the widely-used Agrobacterium-mediated method or the widely-used particle bombardment method (Meksem et al. 2000; Wu et al. 2000; Men et al. 2001; Tao et al. 2002; Fang et al. 2004; Ortiz-Vázquez et al. 2005; Feng et al. 2006; Zhang et al. 2010; Chang et al. 2011), this has added additional utilities to the library for ginseng genome research, particularly molecular breeding and molecular farming.
Identification of BIBACs containing genes important in ginseng
To validate the genome coverage of the library, test its utility for genome research and identify the BIBACs containing genes potentially involved in ginsenoside biosynthesis, we searched Genbank for the genes potentially involved in the pathway (Jung et al. 2003). As a result, we found nine genes likely involved in the biosynthesis pathway of ginsenosides (Table 1). We then designed single-copy overgos from the genes and used them as probes to screen the library. The library screening with the nine gene-specific overgos resulted in a total of 36 positive clones, with each gene probe having 1–7 positive clones and an average of 4.0 positive clones (Fig. 2; Table 1). Interestingly, we found that two of the genes, DS and OSC, shared five positive clones, suggesting that these two genes are physically clustered in the ginseng genome. The results further confirmed the genome coverage of the library estimated above by statistical analysis and suggested that the BIBAC library is suited for isolation and characterization of genes important in ginseng. The positive clones identified provide tools useful for ginseng molecular breeding and molecular farming via genetic engineering.
Stability of ginseng genomic DNA large-insert BIBACs in Agrobacterium
Previous studies have demonstrated that large-insert BIBACs can be readily transformed and engineered using the traditional Agrobacterium-mediated and particle bombardment methods in plants (Hamilton et al. 1996; Liu et al. 1999; Frary and Hamilton 2001; Liu et al. 2002; He et al. 2003; Chang et al. 2011). The key to the successful transformation of large-insert BIBACs into plants through Agrobacterium was shown to be their stability in Agrobacterium (Song et al. 2003; Chang et al. 2011). Therefore, to test the utility of the ginseng large-insert BIBACs in genetic engineering through genetic transformation, we selected two random clones having insert sizes of 100 and 150 kb, respectively, named Pg100 and Pg150, from the library and tested their stability in two Agrobacterium strains COR308 and C1C58. Analysis of two random clones of Pg100 transformed into Agrobacterium strain C1C58 showed that both of them had identical NotI restriction pattern to that of the original BIBAC DNA after being grown in the Agrobacterium strain for 100 generations (Fig. 3a). Analysis of four random clones of Pg150 transformed into Agrobacterium strain COR308 showed that three of them had identical NotI restriction patterns to that of the original BIBAC DNA while one was clearly rearranged after growing in the Agrobacterium strain for 100 generations (Fig. 3b). These results indicated that the ginseng large-insert BIBACs could be readily transformed into Agrobacterium by electroporation and were largely stable in Agrobacterium, even though their stability may vary among different Agrobacterium strains. Further studies are needed to test the utility of the library in ginseng gene genetic engineering in different plant species.
Stability of large-insert ginseng DNA BIBACs in widely-used Agrobacterium strains. a BIBAC Pg100 (100 kb) digested with NotI and fractionated on a pulsed-field gel. Lane 1 BIBAC DNA before transformed into Agrobacterium, lanes 2 and 3 DNA of the BIBAC transformed into Agrobacterium strain C1C58 and grown in the Agrobacterium strain for 100 generations. b BIBAC Pg150 (150 kb) digested with NotI and fractionated on a pulsed-field gel. Lane 1 BIBAC DNA before transformed into Agrobacterium, lanes 2, 3, 4 and 5 DNA of the BIBAC transformed into Agrobacterium strain COR308 and grown in the Agrobacterium strain for 100 generations
Abbreviations
- BIBAC:
-
Binary bacterial artificial chromosome
- BAC:
-
Bacterial artificial chromosome
- EST:
-
Expressed sequence tag
- QTL:
-
Quantitative trait locus
- SSR:
-
Simple sequence repeat
References
Bang KH, Lee SW, Hyun DY, Cho JH, Cha SW, Seong NS, Huh MK (2004) Molecular authentication and genetic polymorphism of Korean ginseng (Panax ginseng C. A. Meyer) by inter-simple sequence repeats (ISSRs) markers. J Life Sci 14:425–428
Chang Y-L, Tao Q, Scheuring C, Meksem K, Zhang H-B (2001) An integrated map of Arabidopsis thaliana for functional analysis of its genome sequence. Genetics 159:1231–1242
Chang Y-L, Chuang H-W, Meksem K, Wu F-C, Chang C-Y, Zhang MP, Zhang H-B (2011) A plant-transformation-ready large-insert BIBAC library of Arabidopsis and bombardment transformation and expression of its large-insert BIBACs in tobacco. Genome 54:437–447
Fang X, Gu S, Xu Z, Chen F, Guo D, Zhang H-B, Wu N (2004) Construction of a binary BAC library for an apomictic monosomic addition line of Beta corolliflora in sugar beet and identification of the clones derived from the alien chromosome. Theor Appl Genet 108:1420–1425
Feng J, Vick BA, Lee M-K, Zhang H-B, Jan CC (2006) Construction of BAC and BIBAC libraries from sunflower and identification of linkage group-specific clones by overgo hybridization. Theor Appl Genet 113:23–32
Frary A, Hamilton CM (2001) Efficiency and stability of high molecular weight DNA transformation: an analysis in tomato. Transgenic Res 10:121–132
Hamilton CM, Frary A, Lewis C, Tanksley SD (1996) Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc Natl Acad Sci USA 93:9975–9979
He R-F, Wang Y, Shi Z, Ren X, Zhu L, Weng Q, He G-C (2003) Construction of a genomic library of wild rice and Agrobacterium-mediated transformation of large-insert DNA linked to BPH resistance locus. Gene 321:113–121
Hong CP, Lee SJ, Park JY, Plaha P, Park YS, Lee YK, Choi JE, Kim KY, Lee JH, Lee J, Jin H, Choi SR, Lim YP (2004) Construction of a BAC library of Korean ginseng and initial analysis of BAC-end sequences. Mol Gen Genomics 271:709–716
In DS, Kim YC, Bang KH, Chung JW, Kim OT, Hyun DY, Cha SW, Kim TS, Seong NS (2005) Genetic relationships of Panax species by RAPD and ISSR analyses. Korean J Med Crop Sci 13:249–253
Jones JDG, Shlumukov L, Carland F, English J, Scofield SR, Bishop GJ, Harrison K (1992) Effective vectors for transformation, expression of heterologous genes, and assaying transposon excision in transgenic plants. Transgenic Res 1:285–297
Jung JD, Park HW, Hahn Y, Hur CG, In DS, Chung HJ, Liu JR, Choi DW (2003) Discovery of genes for ginsenoside biosynthesis by analysis of ginseng expressed sequence tags. Plant Cell Rep 22:224–230
Kim BB, Jeong JH, Jung SJ, Yun DW, Yoon ES, Choi YE (2005) Authentication of Korean Panax ginseng from Chinese Panax ginseng and Panax quinquefolius by AFLP analysis. J Plant Biotechnol 7:81–86
Liu Y-G, Shirano Y, Fukaki H, Yanai Y, Tasaka M, Tabata S, Shibata D (1999) Complementation of plant mutants with large genomic DNA fragments by a transformation-competent artificial chromosome vector accelerates positional cloning. Proc Natl Acad Sci USA 96:6535–6540
Liu Y-G, Liu H, Chen L, Qiu W, Zhang Q, Wu H, Yang C, Su J, Wang Z, Tian D, Mei M (2002) Development of new transformation-competent artificial chromosome vectors and rice genomic libraries for efficient gene cloning. Gene 282:247–255
Ma KH, Dixit A, Kim YC, Lee DY, Kim TS, Cho EG, Park YJ (2007) Development and characterization of new microsatellite markers for ginseng (Panax ginseng C. A. Meyer). Conserv Genet 8:1507–1509
Meksem K, Ruben E, Zobrist K, Hyten D, Tao Q, Zhang H-B, Lightfoot DA (2000) Two large-insert soybean genomic libraries constructed in a binary vector: applications in chromosome walking and genome-wide physical mapping. Theor Appl Genet 101:747–755
Men AE, Meksem K, Kassem MA, Lohar D, Stiller J, Lightfoot D, Gresshoff PM (2001) A bacterial artificial chromosome library of Lotus japonicus constructed in an Agrobacterium tumefaciens-transformable vector. Mol Plant Microbe Interact 14:422–425
Ortiz-Vázquez E, Kaemmer D, Zhang H-B, Muth J, Rodríguez-Mendiola M, Arias-Castro C, James A (2005) Construction and characterization of a plant transformation-competent BIBAC library of the black Sigatoka-resistant banana Musa acuminata cv. Tuu Gia (AA). Theor Appl Genet 110:706–713
Popovich DG, Yeo C-R, Zhang W (2012) Ginsenosides derived from Asian (Panax ginseng), American ginseng (Panax quinquefolius) and potential cytoactivity. Int J Biomed Pharm Sci 6:56–62
Sambrook J, Fritsch EE, Maniatis T (1989) Molecular cloning: laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Song J, Bradeen JM, Naess SK, Helgeson JP, Jiang J (2003) BIBAC and TAC clones containing potato genomic DNA fragments larger than 100 kb are not stable in Agrobacterium. Theor Appl Genet 107:958–964
Song R, Segal G, Messing J (2004) Expression of the sorghum 10-member Kafirin gene cluster in maize endosperm. Nucleic Acids Res 32:e189
Tao Q, Zhang H-B (1998) Cloning and stable maintenance of DNA fragments over 300 kb in Escherichia coli with conventional plasmid-based vectors. Nucleic Acids Res 26:4901–4909
Tao Q, Wang A, Zhang H-B (2002) One large-insert plant-transformation-competent BIBAC library and three BAC libraries of japonica rice for genome research in rice and other grasses. Theor Appl Genet 105:1058–1066
van Dan N, Ramchiary N, Choi SR, Uhm TS, Yang TJ, Ahn IO, Lim YP (2010) Development and characterization of new microsatellite markers in Panax ginseng (C. A. Meyer) from BAC end sequences. Conserv Genet 11:1223–1225
Wu Y, Tulsieram L, Tao Q, Zhang H-B, Rothstein SJ (2000) A binary vector-based large insert library for Brassica napus and identification of clones linked to a fertility restorer locus for Ogura cytoplasmic male sterility (CMS). Genome 43:102–109
Wu C, Xu Z, Zhang H-B (2004) DNA libraries. In: Meyers RA (ed) Encyclopedia of molecular cell biology and molecular medicine, vol 3, 2nd edn. Wiley, Weinheim, pp 385–425
Xu Z, van den Berg M, Scheuring C, Covaleda L, Lu H, Santos FA, Uhm T, Lee M-K, Wu C, Liu S, Zhang H-B (2005) Genome-wide physical mapping from large-insert clones by fingerprint analysis with capillary electrophoresis: a robust physical map of Penicillium chrysogenum. Nucleic Acids Res 33:e50
Zhang H-B (2000) Manual: construction and manipulation of large-insert bacterial clone libraries. Texas A&M University, College Station, Texas
Zhang H-B (2007) Map-based cloning of genes and quantitative trait loci. In: Kole C, Abbott AG (eds) Principles and practices of plant genomics. Genome mapping, vol 1. Science, New Hampshire, pp 229–267
Zhang H-B, Zhao XP, Ding XD, Paterson AH, Wing RA (1995) Preparation of megabase-sized DNA from plant nuclei. Plant J 7:175–184
Zhang H-B, Choi S, Woo SS, Li ZK, Wing RA (1996a) Construction and characterization of two rice bacterial artificial chromosome libraries from the parents of a permanent recombinant inbred mapping population. Mol Breed 2:11–24
Zhang H-B, Woo SS, Wing RA (1996b) BAC, YAC and cosmid library construction. In: Foster G, Twell D (eds) Plant gene isolation: principles and practice. Wiley, England, pp 75–99
Zhang MP, Li Y, Zhang H-B (2008) Isolation of megabase-sized DNA fragments from plants. In: Liu D (ed) Handbook of nucleic acid purification. Taylor & Francis Group, LLC, FL, pp 513–524
Zhang X, Scheuring CF, Zhang MP, Dong JJ, Zhang Y, Huang JJ, Lee M-K, Abbo S, Sherman A, Shtienberg D, Chen W, Muehlbauer F, Zhang H-B (2010) A BAC/BIBAC-based physical map of chickpea, Cicer arietinum L. BMC Genomics 11:501
Zhang H-B, Scheuring CF, Zhang MP, Zhang Y, Wu C-C, Dong JJ, Li Y (2012a) Construction of BIBAC and BAC libraries from a variety of organisms for advanced genomics research. Nat Protoc 7:479–499
Zhang MP, Zhang Y, Huang JJ, Lee M-K, Zhang XJ, Stelly DM, Zhang H-B (2012b) Physical mapping of polyploid genomes: a BIBAC physical map of allotetraploid upland cotton. PLoS One 7:e33644
Zhang MP, Zhang Y, Scheuring CF, Wu C-C, Dong JJ, Zhang H-B (2012c) Preparation of megabase-sized DNA from a variety of organisms using the nuclei method for advanced genomics research. Nat Protoc 7:467–478
Acknowledgments
This study was supported by grants from the City of Changchun Bureau of Science and Technology (2007GH31 and 08GH09) and the Province of Jilin Bureau of Science and Technology (20080715).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhai, J., Wang, Y., Sun, C. et al. A plant-transformation-competent BIBAC library of ginseng (Panax ginseng C. A. Meyer) for functional genomics research and characterization of genes involved in ginsenoside biosynthesis. Mol Breeding 31, 685–692 (2013). https://doi.org/10.1007/s11032-012-9826-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-012-9826-4