Abstract
The TATA binding protein-associated factor 5 (TAF5) is a subunit of TFIID and SAGA complexes involved in RNA polymerase II transcription initiation and histone acetylation. Although members of the putative SAGA complex in Arabidopsis such as GCN5 and ADA2b have important roles in plant development and abiotic stress responses, the function of other components of the Arabidopsis putative SAGA complex, like TAF5, is unknown. We used reverse genetics to elucidate the biological role of TAF5 in Arabidopsis thaliana. The absence of homozygote taf5 mutants indicated that AtTAF5 is an essential gene for the plant viability. Genetic approaches also revealed that AtTAF5 plays a critical role in regulatory mechanisms involved in male gametogenesis and pollen tube growth. Moreover, Arabidopsis taf5 heterozygous mutants displayed terminal flower-like phenotype, suggesting that TAF5 could be involved in molecular mechanisms that regulate indeterminate inflorescence meristems. Therefore, this work suggests that Arabidopsis TAF5 is necessary and sufficient for a complete plant life cycle.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The general transcription factor TFIID plays a pivot role in the recognition of core promoter elements and is required for transcription initiation of the majority of eukaryotic genes by RNA pol II (Hahn 1998; Lee and Young 1998). TFIID is a multiprotein complex composed of ΤΑΤΑ box-binding protein (TBP) and 14 other TBP-associated factors (TAFs) that have been conserved throughout evolution (Tora 2002). TAFs subunits serve multiple functions within the TFIID holocomplex. For instance, TAF1 and TAF2 have been shown to bind to the transcription initiator, whereas TAF6 and TAF9 have been shown to interact with the promoter elements (Chalkley and Verrijzer 1999; Verrijzer et al. 1994). Furthermore, several TAFs have been shown to interact with transcription activators or other general transcription factors to stabilize the pre-initiation complex (Roeder 1996). In yeast, 13 TAFs are required for viability (Green 2000). In vitro experiments in humans and Drosophila show that TAFs are necessary for transcription initiation in TATA-less promoters (Wright et al. 2006). In Caenorhabditis elegans, TAF5 protein is important for transcription during embryogenesis (Walker and Blackwell 2003), whereas in Drosophila melanogaster the TAF5 homolog protein, called cannonball, is found only in male germ cells and is required for normal transcription in primary spermatocytes (Hiller et al. 2001, 2004).
Several TAFIIs are also components of histone acetyltransferases (HAT) complexes such as Spt-Ada-Gcn5-Acetyltransferase (SAGA), which functions in nucleosomal histone acetylation and chromatin-associated transcriptional activation or repression (Grant et al. 1998; Pray-Grant et al. 2002). The catalytic subunit of the SAGA complex is the protein GCN5; the complex also include ADA proteins and many TAFs, like TAF5, 6, 9, 10, 12 (Daniel and Grant 2006). The SAGA complex is also required for transcription elongation, mRNA export and nucleotide excision repair (Baker and Grant 2007). In Arabidopsis, mutations in GCN5/HAG1 and ADA2b/PRZ1 cause pleiotropic alterations in pattern formation and organogenesis, also manifested as defects in the interpretation of signals relevant to plant development (Bertrand et al. 2003; Sieberer et al. 2003; Vlachonasios et al. 2003; Benhamed et al. 2006; Long et al. 2006; Cohen et al. 2009; Kornet and Scheres 2009; Anzola et al. 2010; Servet et al. 2010). Furthermore, ADA2b and GCN5 are implicated in abiotic stress response mechanisms (Vlachonasios et al. 2003; Hark et al. 2009).
TBP and TAF-like proteins have been identified in Arabidopsis (Gasch et al. 1990; Lago et al. 2004; Bertrand et al. 2005). An extensive yeast two-hybrid study was conducted to map the interactions of putative TFIID components in Arabidopsis (Lawit et al. 2007). The results indicated that plant TFIID might posses some unique features. Alterations in the levels of Arabidopsis TBP or specific TAFs are implicated in plant development affecting the organization of shoot apical meristems, leaf development, formation of floral organs and leaves, fertility, pollen tube growth and light responses (Gurley et al. 2006). The TAF1 family of histone acetyltransferases in Arabidopsis consists of two members, HAF1 and HAF2/TAF1 (Bertrand et al. 2005; Benhamed et al. 2006). While mutations in HAF1 showed no visible phenotype, mutations in HAF2/TAF1 regulate light-responsive genes by controlling acetylation of histones H3 and H4 at the target promoters (Bertrand et al. 2005; Benhamed et al. 2006). Arabidopsis has two TAF6 genes that are distantly related (Lago et al. 2005). AtTAF6a is essential for plant viability since loss of function mutants are lethal, whereas heterozygous plants displayed a drastic reduction in pollen tube growth rate (Lago et al. 2005), suggesting that TAF6 plays an important role in pollen tube function. Arabidopsis mutants with reduced expression of TAF10 are more sensitive to salt stress, whereas overexpression of TAF10 increased seed germination rate upon osmotic stress (Gao et al. 2006), suggesting that TAF10 is involved in osmotic stress responses. When TAF10 from Flavenia trivernia was overexpressed in Arabidopsis, a terminal flower-like phenotype resulted, characterized by limited growth of an indeterminate inflorescence, chimeric floral organs and curled leaves (Furumoto et al. 2005). A mutation in an Arabidopsis TFIID-interacting transcription factor TAF12B/EER4 results in failure to induce a subset of ethylene-regulated genes in etiolated seedlings (Robles et al. 2007).
In this report, we explore the biological role of TAF5 in Arabidopsis. Our data suggests that TAF5 is necessary and sufficient for plant viability.
Materials and methods
Plant material and growth conditions
Five mutants bearing a T-DNA insertion in the Arabidopsis TAF5 gene were identified; taf5-1 (SALK_021380), taf5-2 (SALK_052284) and taf5-3 (SALK_036395) were found from the Salk Institute collections (Alonso et al. 2003). while taf5-4 (SAIL_274-A04) and taf5-5 (SAIL_752-BO9) were found from Syngenta collections (Sessions et al. 2002). Seeds from those mutants were obtained from the Nottingham Arabidopsis Stock Center (NASC). Seeds were surface-sterilized with 20% sodium hypochlorite solution for 10 min, washed three times with sterile water and plated onto Petri dishes containing Gamborg’s B5 medium (Duchefa) supplemented with vitamins, 1% (w/v) sucrose, and 0.8% agar, and adjusted to pH 5.7. Seeds were stratified for 3 days at 4°C in the dark and then grown in long-day conditions (16-h light/8-h dark at 21°C) for 14 days. The wild-type, ecotype Columbia-0 (Col), and the taf5 mutant plants were then grown in soil at 18–22°C in long-day conditions (16-h light/8 h dark).
Molecular characterization of taf5 mutants
The genotypes of mutant plants were confirmed by PCR using gene-specific and T-DNA border-specific primers (Electronic Supplementary Material Table 1S). PCR was carried out using EX-Taq™ DNA polymerase (Takara). taf5-1 was genotyped using KB165-166 primers to test for the wild-type gene and KB112 and LBa1 to test for the T-DNA insertion. taf5-2 was genotyped using KB165-166 primers to test for the wild-type gene and KB166 and LBa1 to test for the insertion. taf5-3 was genotyped using KB163-164 primers to test for the wild-type gene and KB164 and LBa1 to test for the insertion. taf5-4 was genotyped using KB163-164 primers to test for the wild-type gene and KB164 and KB162 to test for the insertion and taf5-5 was genotyped using KB229-230 primers to test for the wild-type gene and KB230 and KB162 to test for the insertion.
Total mRNA from rosette leaves or from flower inflorescences was isolated using the Nucleospin Plant RNA kit (Macherey–Nagel, Duren, Germany). DNase treatment was carried out by adding 5 μl 10x buffer and 1 μl DNase (10 U) to 60 μl of RNA solution followed by incubation for 2 h at 37°C. Samples were extracted using an equal volume of phenol, chloroform, isoamylic alcohol (25:24:1 v/v) and RNA was precipitated using a half volume of 7.5 M ammonium acetate and two volumes of 100% ethanol. The resulted RNA pellet was washed with 75% ethanol and resuspended in 50 μl nuclease-free water. Levels of specific mRNAs were assayed using reverse transcription (RT) followed by PCR and quantitated using real-time PCR. cDNA was synthesized using the Reverse Transcription System Kit (Promega, Madison, WI, USA). RT reactions were carried out using 0.25 μg of DNAse-treated RNA. The final reaction volume was 40 μl. After the reaction the samples were diluted five times in nuclease-free water. Real-time PCR was carried out in reactions using 1 × buffer containing KAPA SYBR® FAST qPCR Master Mix ABI Prism™ (Kapa Biosystems, Boston, MA, USA). In each 10 μl reaction, 2 μl of cDNA sample was used. The gene-specific primers used for RT-qPCR are listed in Table 1S. Gene-specific primers for At4g26410 or ACT2 open reading frame were used as references. The Ct values obtained from TAF5 or AP1 genes were normalized to the values obtained from At4g26410 or ACT2, respectively. The values were expressed as reference gene- normalized levels of the target genes. Student’s t test was used to compare the expression of the target genes between wild-type and mutants plants to determine whether the difference was significant at P ≤ 0.05. Semi-quantitative RT-PCR was performed to identify transcripts that terminate in the T-DNA of taf5-1 heterozygous mutants. PCR was applied using KB112 and LBa1 primers and the products were subjected to gel electrophoresis and visualized via ethidium bromide staining. PDF2 was used as reference gene.
Pollen development, germination and pollen tube growth
Sufficient mature pollen was obtained by placing 3–4 open flowers from wild-type and heterozygous taf5-1 and taf5-4 mutants in an Eppendorf tube containing 300 μl H2O. After brief vortexing and centrifugation, the pollen pellet was diluted in 2-μl of 4′,6-diamino-2-phenylindole dihydrochloride (DAPI) staining solution (0.4 μg/ml DAPI in antifade solution which consists of PBS solution containing 50% glycerol and 2–7 mM phenylenediamine, pH 8.0, Millipore, USA) for at least 2 h in the dark and then transferred to a microscope slide. Pollen was viewed by a light and UV epi-fluorescent inverted microscope, Nikon Eclipse TE 2000-S (Nikon Corporation, Tokyo, Japan) and images were captured with a digital camera, Nikon DS-L1 (Nikon Corporation). For the analysis of spores at earlier stages, single anthers were dissected from isolated buds. Anthers were disrupted on microscope slides using dissecting needles and gently squashed in DAPI staining solution (0.4 μg/ml) under a coverslip.
Pollen germination experiments were conducted according to the protocol of Boavista and McCormick (2007). The pollen grains, from flowers that had just opened from 1 week after bolting inflorescence, were applied to microscope slides with the pollen germination medium (5 mM KCl, 5 mM CaCl2, 1 mM MgSO4, 1.5 mM H3BO3, 10% w/v sucrose and 1% w/v agar) adjusted to pH 8. The slides were then placed onto moist paper in empty tip racks (Johnsson-Brousseau and McCormick 2004) and incubated into a dark room at 22°C for 16 h. Approximately 650–7,100 pollen grains collected from at least three different flowers were examined from each mutant and wild-type plant. Pollen germination and pollen tube growth was examined under the Zeiss Axiostar Plus (Jena, Germany) microscope with DIC optics. Images were captured using a camera (Canon USA, Inc.). The pollen grains were counted with the ImageJ software (http://rsb.info.nih.gov/nih-image/index.html), using the cell counter application.
Bioinformatics tools
The amino-acid sequence of TAF5 proteins from different organisms were retrieved from the National Center of Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov) and from Phytozone (http://www.phytozone.net/). The amino-acid sequence alignment was conducted using Clustal W2 (http://www.ebi.ac.uk/Tools/clustalw2/index.html) using default parameters. The alignment was graphically enhanced using ESPript software (Gouet et al. 1999). Phylogenetic relationships were analyzed using MEGA 4.1 (http://www.megasoftware.net/).
Results
Plants have one TAF5 homolog
Although members of the putative SAGA complex in Arabidopsis such as GCN5 and ADA2b have important roles in plant development and abiotic stress responses (Bertrand et al. 2003; Vlachonasios et al. 2003; Benhamed et al. 2006), the function of other components of the Arabidopsis putative SAGA complex is unknown. Therefore we searched for an Arabidopsis ortholog of TAF5, a component of SAGA and TFIID complex in yeast and other organisms. A BLAST search using the yeast TAF5 amino-acid sequence identified one Arabidopsis ortholog, At5g25150, which encodes for a 74.4-kDa protein. Alignment of Arabidopsis TAF5 with homologs from Homo sapiens (Hs), Saccharomyces cerevisiae (Sc) and D. melanogaster (Dm) are presented in Figure 1Sa. Two regions of AtTAF5 share a high degree of sequence identity with the other TAF5 proteins. The first conserved domain, the NTD2 domain, was found in the N-terminus of AtTAF5. This alpha-helical domain is homodimerized and could play an important role in organizing the TFIID structure (Bhattacharya et al. 2007). The other conserved domain was found in the C-terminus of AtTAF5, the WD40 motif, which is repeated five times and has 40% identity with the yeast TAF5 (Fig. 1Sa and 1Sb). The last four repeats are common in yeast, human and Drosophila but the first one is observed only in plants (Fig. 1Sb). The WD repeats form a closed beta-propeller structure (Smith et al. 1999). The beta-propeller domains are critical for the structural role of TAF5 in the TFIID complex (Leurent et al. 2004) and have been shown to mediate protein–protein interactions (Tao et al. 1997). Genes that encode TAF5 proteins from other plants are also found (Fig. 1Sc). Phylogenetic relationships between Arabidopsis TAF5 and the other plant TAF5 proteins show a distinct division between monocotyledonous and dicotyledonous plants as well as from mosses, Prasinophyceae and green algae lower plants (Fig. 1Sc). Interestingly, Populus trichocarpa and Glycine max have two and three orthologs of TAF5, respectively, that could arise from genome duplication.
Molecular characterization of taf5 mutants in Arabidopsis
In order to study the biological function of the Arabidopsis TAF5 gene, a search in Arabidopsis T-DNA insertion mutant collections was performed. Five mutants bearing a T-DNA insertion were identified. TAF5 mutants taf5-1, taf5-2, taf5-3, and taf5-4 disrupt the predicted coding region at 838, 1,057, 3,271, and 3,962 bp in the genomic DNA and downstream of the initiation codon, respectively (Fig. 1). T-DNA insertion in the taf5-5 mutant is located 641 bp upstream of the initiation codon (Fig. 1). The genotyping of the taf5 mutants were performed by PCR using gene-specific primers and T-DNA primers (Table 1S). The analysis revealed the presence of taf5-1, taf5-2, taf5-3, and taf5-4 heterozygotes and only the presence of taf5-5 homozygote plants (Fig. 2S). The progeny of a self-pollinated for taf5-1 (+/−), taf5-2 (+/−), taf5-3 (+/−), or taf5-4 (+/−) mutants was analyzed by PCR using gene-specific and T-DNA primers. PCR analysis of approximately 100 T3 mutant plants showed that 24–32 of them were taf5 (+/−) and 76–69 wild-type (+/+) (Table 1). The segregation ratio from each heterozygote plant was approximately 25% heterozygous taf5 (+/−) and 75% wild-type (+/+) plants. No homozygous individuals were identified in taf5-1, taf5-2, taf5-3, and taf5-4 mutants. This data strongly suggests that the Arabidopsis taf5 mutants are lethal. Furthermore, the segregation of wild-type (+/+) and heterozygous (±) individuals was in a ratio of approximately 3:1, which is in disagreement with the expected 1:2:1 (−/−, ±, +/+) according to Mendelian genetics (Table 1). These segregation ratios suggest a haplo-insuffiency character where the single wild-type TAF5 allele of the heterozygous locus is not sufficient to restore wild-type function and as a result wild-type TAF5 has a semi-dominant character. The loss of homozygous taf5 mutant plants could have occurred early during seed development or before fertilization, suggesting that TAF5 might play important role in Arabidopsis embryo development and/or male/female gametophyte formation.
The level of TAF5 gene expression was determined in taf5 mutants using quantitative real-time RT-PCR. In taf5-1 (+/−), taf5-2 (+/−), taf5-3 (+/−), and taf5-4 (+/−) mutants the level of TAF5 was reduced to approximately 50–60% in comparison to wild-type plants, since we predicted that only one copy of the TAF5 transcript was made (Fig. 2a). In the homozygous taf5-5 mutant the level of TAF5 expression was reduced to 25% in comparison to wild-type plants (Fig. 2b), suggesting that taf5-5 is a hypomorph. Moreover, RT-PCR analysis using primers from the N-terminus of the TAF5 gene and the T-DNA revealed that several splice variant transcripts are present in the heterozygous taf5-1 mutant plants (Fig. 2c).
TAF5 gene expression in taf5 mutants. Total mRNA was extracted from rosette leaves of wild-type and taf5 mutants. a Reverse transcriptase-qRCP was used to quantitate TAF5 expression in taf5-1 (+/−), taf5-2 (+/−), taf5-3 (+/−) and taf5-4 (+/−) mutant plants. The Ct values obtained for TAF5 were normalized to the Ct values obtained for At4g26410, and fold induction of TAF5 was calculated by normalizing values from wild-type plants with those from taf5 mutants. For each mutant and wild-type plants, triplicate samples in each experiment were assayed twice. Three independent experiments were performed; error bars represent SE, where n = 4. Asterisks mark values that are significantly different from the wild-type (t test, P = 0.001). b Reverse transcriptase-qPCR analysis to assess TAF5 expression in taf5-5 homozygous mutant plants; triplicates for taf5-5 and duplicates for wild-type plants were used; error bars represent SE, where n = 4. c RT-PCR analysis to assess TAF5 transcripts from the N-terminus of TAF5; wild-type and duplicates for taf5-1 heterozygous plants were used. PCR was performed with KB112 and LBa1 primers and PDF2 was used as control. Three splice variants ranging from approximately 500–700 bp were detected in the heterozygous taf5-1 plants
Phenotypical analysis of taf5 mutants
Although homozygous mutations in the coding region of TAF5 were lethal, we checked whether the heterozygous taf5 (+/−) mutants exhibited any developmental phenotypes. All the heterozygous taf5 (+/−) mutants that disrupt the TAF5 open reading frame displayed a terminal flower-like phenotype (Fig. 3a–d) in comparison to wild-type plants (Fig. 3e). This phenotype is characterized by suppressed growth of the main inflorescence and lack of indeterminate growth. Interestingly, the secondary inflorescences and the stem branches of the heterozygous taf5 mutants developed normally without showing terminal flower-like phenotypes. Homozygous taf5-5 plants showed no obvious morphological differences from wild-type plants at various stages of growth and development (Fig. 4f), suggesting that reduction of TAF5 gene expression to 25% was not sufficient to affect developmental processes in Arabidopsis. These differences between taf5-5 homozygous and the rest of taf5 heterozygous mutants could be explained by the fact that several truncated TAF5 transcripts are made at least in taf5-1 (+/−) (Fig. 3c). The terminal flower-like phenotype observed in heterozygous taf5-1, taf5-2, taf5-3, and taf5-4 mutants could be the result of the truncated transcripts that terminated inside the T-DNA, at least in taf5-1 (+/−) (Fig. 2c). These transcripts might generate peptides that could dimerize with the wild-type TAF5 protein, and this interaction could form a deleterious complex affecting the function of TAF5 during this developmental stage. The fact that TAF5 is a single gene in Arabidopsis, and taf5 (+/−) mutants have reduced expression of the TAF5 gene and displayed developmental problems, further suggests that taf5 mutants displayed a semi-dominant character.
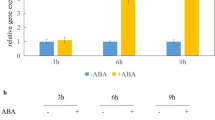
Phenotypic characterization of taf5 mutants. a Heterozygous taf5-1 mutant plant after 30 days of growth. Bar = 1 cm. b Heterozygous taf5-2 mutant plant after 30 days of growth. Bar = 1 cm. c Heterozygous taf5-3 mutant plant after 30 days of growth. Bar = 1 cm. d Heterozygous taf5-4 mutant plant after 30 days of growth. Bar = 1 cm. e Wild-type (Col-0) plants after 30 days of growth. Bar = 1 cm. The white arrows indicate the terminal flower phenotype observed. f Homozygous taf5-5 and wild-type (Col-0) plants after 25 days of growth. Bar = 1 cm. g AP1 expression in flower of taf5 mutant in comparison to wild-type plants. Total mRNA was isolated from the first flower of taf5-1 (+/−), taf5-3 (+/−) mutants, and wild-type plants. RT-qPCR was used to quantitate the expression of AP1 gene. The Ct values obtained for AP1 were normalized to the Ct values obtained for ACT2, and fold induction of AP1 was calculated by normalizing values from wild-type plants with those from taf5 mutants. For each mutant and wild-type plant, triplicate samples in each experiment were assayed twice. Three independent experiments were performed; error bars represent standard error of mean, where n = 3
Analysis of pollen development in taf5 (+/−) mutants. Light (a, c, e) and fluorescence images (b, d, f) of DAPI-stained mature pollen from wild-type (a, b), heterozygous taf5-1 (c, d) and heterozygous taf5-4 (e, f) plants. Light (g, i, k, m, o, q) and fluorescence images (h, j, l, n, p, r) of DAPI-stained microspores at uninucleate stage from wild-type (g–j), heterozygous taf5-1 (k–n), and heterozygous taf5-4 (o–r) plants. Arrows indicate non-DAPI-stained pollen in taf5-4 heterozygous pollen. Bar = 10 μm
In Arabidopsis, the terminal flower phenotype is known to arise from the function of two antagonist genes TERMINAL FLOWER 1 (TFL1) and APETALA 1 (AP1). Overexpression of AP1 is sufficient to convert the inflorescence meristem to a terminal flower (Mandel and Yanofsky 1995). Therefore, we examined AP1 expression in the terminal flower of taf5-1 and taf5-3 heterozygote mutants in comparison with wild-type plants. The flowers were in stage 13. As we expected, in both taf5 (+/−) mutants AP1 expression was increased more than two-fold in comparison to wild-type flowers (Fig. 3g), suggesting that the terminal flower observed in the TAF5/taf5 mutants could arise from the higher expression of AP1.
TAF5 is required for male gametophyte development
If only the taf5 homozygous developing seeds were not viable, then the progeny of taf5 (+/−) plants would segregate 2:1 for the heterozygous (±) and wild-type (+/+) locus, respectively. However, the percentage of heterozygous seedlings was much less than 67%, with only approximately 25% heterozygous at the TAF5 locus (Table 1). This deviation from a standard inheritance pattern implies the death of more than just the double-null developing seed. The effect of AtTAF5 mutations on male or female gametophytic development was further examined by back-crossing heterozygous taf5-2 mutants to wild-type plants. When the stigma of the taf5-2 mutant was pollinated with wild-type pollen, a normal Mendelian segregation ratio (1:1, wild-type:heterozygous) was observed. However, when pollen of taf5-2 was used to pollinate wild-type plants, then only 11 out of 75 individuals were heterozygous, indicating a strong deviation from the expected 1:1 segregation. These results indicate that there is a defect in the transmission of the mutated AtTAF5 allele through the male gametophyte, which suggests that pollen development is affected by the taf5-2 mutation. Since male gametophyte formation could be affected in taf5 mutants, we examined the viability of pollens of the taf5-2 mutants in comparison to wild-type plants. Pollens collected from wild-type and taf5-2 mutants were stained with Alexander solution. The pollens examined from wild-type and taf5-2 mutant plants were viable (data not shown).
We further analyzed male gametophyte development in taf5-1 and taf5-4 heterozygous mutants by fluorescence microscopy. In Arabidopsis, the male gametophyte is a three-cell organism that is derived by stereotypical cell divisions (McCormick 2004). During microsporogenesis, meiosis of the microspore mother cell produces a tetrad of cells. After release from the tetrad, during microgametogenesis, each microspore goes through an asymmetric mitosis, to produce a bicellular pollen grain containing a generative cell and a much larger vegetative cell. The larger vegetative cell does not divide again but eventually will form the pollen tube. The smaller generative cell undergoes a second round of cell division, pollen mitosis, to form the two sperm cells (McCormick 1993, 2004). Pollen grain collected from open flowers of both wild-type and heterozygous taf5-1 and taf5-4 mutants were examined by staining with DAPI that specifically stains DNA. When wild-type flowers were open, the mature pollen is characterized by two brightly stained sperm cell nuclei and a faintly stained vegetative cell nucleus (Fig. 4b). Similar images were obtained in heterozygous taf5-1 and taf5-4 pollen (Fig. 4d, f). However, in taf5-1 heterozygous plants the percentage of non-DAPI-stained pollen was approximately double that of wild-type, whereas the percentage of bicellular and tricellular pollen in taf5-1 heterozygous plants was lower in comparison to wild-type pollen. Nevertheless, this observation was not found in taf5-4 heterozygous pollen. In the early stages of microsporogenesis, microspores of wild-type plants (Fig. 4g–j) were indistinguishable from taf5-1 and taf5-4 heterozygous microspores (Fig. 4k–p). Moreover, tetrads are formed indicating that meiosis takes place normally (Fig. 4m–p). In a few cases, and only in taf5-4 heterozygous plants, we observed aborted and non-DAPI-stained pollen grains (Fig. 4q, r). These results indicate that, at least in taf5-1 heterozygous mutants, microspores are arrested in the bicellular and tricellular pollen stages.
We then studied the effect of TAF5 in pollen tube development. Pollen was collected from taf5-2, taf5-4, and taf5-5 mutants as well as from wild-type flowers from secondary inflorescence after 1 week of bolting, and germinated in vitro. At least three flowers per genotype and more than 200 pollen grain per flower were analyzed. The germinated pollen grains were classified into four groups, according to Lago et al. 2005: (1) germinated pollen that developed a pollen tube with length of more than three times the diameter of the pollen grain; (2) germinated pollen that has a pollen tube with a length of 1–3 times the pollen grain diameter; (3) germinated pollen that developed only a pollen tube primordia smaller than the diameter of the pollen grain; and (4) non-germinated pollen (Fig. 5a). In these experiments, the pollen germination percentage was rather low (15–25%) but there was no significant difference between the genotypes. However, differences were observed between the three classes (Fig. 5b). In the wild-type plants the majority of pollen tube observed belong to group I followed by the pollen tube of group II and group III (74, 17, and 9% respectively). In contrast, the number of class I pollen tubes was significantly reduced in the heterozygote taf5-2 and taf5-4 plants (19 and 38%, respectively) when compared to wild-type plants, whereas the number of pollen tubes of class II and III was significantly increased in the heterozygote taf5-2 (45 and 36%) and taf5-4 plants (35 and 27%) in comparison to wild-type plants. The homozygote taf5-5 plants exhibited a similar pattern of pollen tube classes to the wild-type plants. The highest percentage was in group I (83%) followed by group II (14%) and group III (3%). The reduction of class I pollen tubes is expected when considering that 50% of the pollen taken from heterozygous taf5-2 and taf5-4 mutant plants carries the wild-type allele These results suggest that the pollen tube growth rate was reduced in the heterozygous taf5 mutants, indicating that the pollen from wild-type TAF5 was germinated and formed a pollen tube that was more competent to reach the ovule.
Pollen tube growth of taf5 mutants. a Four classes of germinated pollen grains were examined: (I) germinated pollen that developed a pollen tube with length more than 3 times the diameter of the pollen grain; (II) germinated pollen with pollen tube length of 1–3 times the pollen grain diameter; (III) germinated pollen with a pollen tube primordia smaller than the diameter of the pollen grain; (IV) non-germinated pollen. Size bars are 100 μm for (I) and 50 μm for (II) (III), and (IV). b. The effect of taf5 mutants on pollen tube growth. Pollen from taf5-2 (+/−), taf5-4 (+/−) and taf5-5 homozygous mutants as well as from wild-type (Col-0) was collected from flowers 1 week after bolting and germinated on solidified media at 22°C. Bars represent means ± SE (n = 3). Asterisks mark values that are significantly different from the wild-type (t test, P = 0.001)
Discussion
Arabidopsis TAF5 is important for plant viability
Disruption mutations of the Arabidopsis ADA2b and GCN5 genes resulted in striking phenotypes that affect plant growth and development (Vlachonasios et al. 2003). In this study we explore the biological role of another component of the putative SAGA complex in Arabidopsis, TAF5. Unlike ada2b and gcn5 mutants, loss-of function mutation in Arabidopsis TAF5 affects plant viability, suggesting that TAF5 is required for the complete plant life cycle. These differences could arise from the fact that TAF5 is not only a component of the SAGA complex but also a component of the TFIID complex which is involved in RNA polymerase II transcription initiation (Burley and Roeder 1996). Similarly, yeast TAF5 is also required for yeast viability (Poon et al. 1995) and in particular for cell cycle progression through G2/M (Apone et al. 1996). Site-directed mutations in the WD40 repeats of yeast TAF5 cause a defect in the ability of TAF5 to interact with the TFIID and SAGA complexes and as a result TAF5 is required for the transcription of a large number of genes in yeast (Durso et al. 2001).
Alternative forms of the TAFs have been also proposed to play a role in selective activation of cell-type-specific gene expression programs during cellular differentiation (Freiman 2009). In Arabidopsis, our results suggest that TAF5 is required for male gametophyte development, probably by affecting transcription of genes involved in male gametophyte differentiation. The importance of TAF5 in spermatogenesis implies that it is conserved throughout evolution. For instance, in C. elegans TAF5 is essential during early development since the development of taf-5 (RNAi) embryos was arrested at the first 100 cells without signs of differentiation, suggesting a broad defect in zygotic transcription (Walker and Blackwell 2003). The cannonball (can) gene of Drosophila encodes a homolog of a dTAF5 protein expressed only in spermatocytes which is required for normal transcription of genes involved in spermatid differentiation (Hiller et al. 2001).
A particularly interesting aspect of our findings is the remarkable similarity between the taf5 mutant phenotype and the previously described effects of inhibiting expression of TAF6 in Arabidopsis (Lago et al. 2005) The null taf6 in Arabidopsis is also lethal, affecting male gametophyte development and pollen tube growth (Lago et al. 2005). This phenotype similarity supports the model that TAF5 and TAF6 are functionally linked. Further support of this model comes from the yeast two-hybrid experiments where AtTAF5 interacts with the histone-fold domain of TAF6 (Lawit et al. 2007).
Gametogenesis depends on coordination of sporophytic and gametophytic gene expression (McCormick 2004; Yadegari and Drews 2004). Expression profiling of male gametophytes has provided a list of genes that may be under gametophytic selection (Honys and Twell 2004; Pina et al. 2005). Studies on the sperm cell transcriptome (Borges et al. 2008), transcriptome changes associated with pollen germination and tube growth (Wang et al. 2008; Qin et al. 2009), and the recent discovery of several male gametophytic regulators of cell cycle progression and differentiation, together with the collection of male gametophyte development Ds insertional mutants (Kim et al. 2008; Boavista et al. 2009; Chen et al. 2009), serve as a foundation to decipher the regulatory mechanisms of male gametogenesis and delivery. It would be interesting to learn which of these regulatory mechanisms are dependent on the action of TAF5. The reduced expression of these genes in taf5 mutants could be responsible for male gametophyte defects and reduced pollen tube growth.
TAF5 is involved in the gene expression related to inflorescence meristem identity
One of the striking phenotypes observed in the heterozygous taf5 mutants is the terminal flower-like phenotype with characteristic determinate inflorescence. A similar phenotype was observed in the plants that overexpress TAF10 (Furumoto et al. 2005; Tamada et al. 2007), a gene that could be functionally linked with TAF5. The terminal flower-like phenotype is mediated by the function of several genes including TFL1 and TFL2 (Shannon and Meeks-Wagner 1991; Takada and Goto 2003). To maintain the indeterminate inflorescence meristem, AP1/CAL and LFY must be activated only in floral primordia (Mandel and Yanofsky 1995; Weigel and Nilsson 1995). In the tfl mutant, ectopic expression of LFY and AP1 transforms the whole inflorescence meristem into a floral meristem (Bradley et al. 1997). In taf5 (+/−) mutants, AP1 is overexpressed, suggesting that TAF5 could function directly or indirectly in the regulatory mechanisms involved to prevent AP1 expression and the consequent termination of the inflorescence meristem.
In conclusion, our data indicates that TAF5 is an essential gene in the Arabidopsis life cycle. Moreover, TAF5 is required for male gamentogenesis and the regulatory mechanisms of pollen tube growth. Finally, TAF5 is also required in transcriptional mechanisms involved in maintenance of indeterminate inflorescence.
References
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Anzola JM, Sieberer T, Ortbauer M, Butt H, Korbei B, Weinhofer I, Müllner AE, Luschnig C (2010) Putative Arabidopsis transcriptional adaptor protein (PROPORZ1) is required to modulate histone acetylation in response to auxin. Proc Natl Acad Sci USA 107:10308–10313
Apone LM, Virbasius CA, Reese JC, Green MR (1996) Yeast TAFII90 is required for cell-cycle progression through G2/M but not for general transcription activation. Genes Dev 10:2368–2380
Baker SP, Grant PA (2007) The SAGA continues: expanding the cellular role of a transcriptional co-activator complex. Oncogene 37:5329–5340
Benhamed M, Bertrand C, Servet C, Zhou D-X (2006) Arabidopis GCN5, HD1, and TAF1/HAF2 interact to regulate histone acetylation required for light-responsive gene expression. Plant Cell 18:2893–2903
Bertrand C, Bergounioux C, Domenichini S, Delarue M, Zhou D-X (2003) Arabidopsis histone acetyltransferase AtGCN5 regulates the floral meristem activity through the WUSCHEL/AGAMOUS pathway. J Biol Chem 278:28246–28251
Bertrand C, Benhamed M, Li Y-F, Ayadi M, Lemonnier G, Renou J-P, Delarue M, Zhou D-X (2005) Arabidopsis HAF2 gene encoding TATA-binding protein (TBP)-associated factor TAF1 is required to integrate light signals to regulate gene expression and growth. J Biol Chem 280:1465–1473
Bhattacharya S, Takada S, Jacobson RH (2007) Structural analysis and dimerization potential of the human TAF5 subunit of TFIID. Proc Natl Acad Sci USA 104:1189–1194
Boavista LC, McCormick S (2007) Temperature as a determinant factor for increased and reproducible in vitro pollen germination in Arabidopsis thaliana. Plant J 52:570–582
Boavista LC, Shuai B, Yu H-J, Pagnussat GC, Sundaresan V, McCormick S (2009) A collection of Ds insertional mutants associated with defects in male gametophyte development and function in Arabidopsis thaliana. Genetics 181:1369–1385
Borges F, Gomes G, Gardner R, Moreno N, McCormick S, Feijo JA, Becker JD (2008) Comparative transcriptomics of Arabidopsis sperm cells. Plant Physiol 148:1168–1181
Bradley D, Ratcliffe O, Vincent C, Carpenter R, Coen E (1997) Inflorescence commitment and architecture in Arabidopsis. Science 275:80–83
Burley SK, Roeder RG (1996) Biochemistry and structural biology of transcription factor IID (TFIID). Annu Rev Biochem 65:769–799
Chalkley GE, Verrijzer CP (1999) DNA binding site selection by RNA polymerase II TAFs: a TAF(II)250-TAF(II)150 complex recognizes the initiator. EMBO J 18:4835–4845
Chen Z, Hafidh S, Poh SH, Twell D, Berger F (2009) Hyperproliferation caused by loss of Retinoblastoma prevents cell fate establishment during Arabidopsis male gametogenesis. Proc Natl Acad Sci USA 106:7257–7262
Cohen R, Schocken J, Kaldis A, Vlachonasios KE, Hark AT, McCain ER (2009) The histone acetyltransferase GCN5 affects the inflorescence meristem and stamen development in Arabidopsis. Planta 230:1207–1221
Daniel JA, Grant PA (2006) Multi-tasking on chromatin with the SAGA coactivator complexes. Mutat Res 618:135–148
Durso RJ, Fisher AK, Albright-Frey TJ, Reese JC (2001) Analysis of TAF90 mutants displaying allele-specific and broad defects in transcription. Mol Cell Biol 21:7331–7344
Freiman RN (2009) Specific variants of general transcription factors regulate germ cell development in diverse organisms. Biochim Biophys Acta Gene Regul Mech 1789:161–166
Furumoto T, Tamada Y, Izumida A, Nakatani H, Hata S, Izui K (2005) Abundant expression in vascular tissue of plant TAF10, an orthologous gene for TATA box-binding protein-associated factor 10, in Flaveria trinervia and abnormal morphology of Arabidopsis thaliana transformants on its overexpression. Plant Cell Physiol 46:108–117
Gao X, Ren F, Lu YT (2006) The Arabidopsis mutant stg1 identifies a function for TBP-associated factor 10 in plant osmotic stress adaptation. Plant Cell Physiol 47:1285–1294
Gasch A, Hoffmann A, Horikoshi M, Roeder RG, Chua NH (1990) Arabidopsis thaliana contains two genes for TFIID. Nature 346:390–394
Gouet P, Courcelle E, Stuart DI, Metoz F (1999) ESPript: multiple sequence alignments in PostScript. Bioinformatics 15:305–308
Grant PA, Schieltz D, Pray-Grant MG, Steger DJ, Reese JC, Yates JR III, Workman JL (1998) A subset of TAF(II)s are integral components of the SAGA complex required for nucleosome acetylation and transcriptional stimulation. Cell 94:45–53
Green MR (2000) TBP-associated factors (TAFIIs) multiple, selective transcriptional mediators in common complexes. Trends Biochem Sci 25:59–63
Gurley WB, O’Grady K, Czarnecka-Verner E, Lawit SJ (2006) General transcription factors and the core promoter: ancient roots. In: Grasser KD (ed) Regulation of transcription in plants. Annu Plant Rev 29:1–21, Blackwell, Oxford
Hahn S (1998) The role of TAFs in RNA polymerase II transcription. Cell 95:579–582
Hark AT, Vlachonasios KE, Pavangadkar KA, Rao S, Gordon H, Adamakis ID, Kaldis A, Thomashow MF, Triezenberg SJ (2009) Two Arabidopsis orthologs of the transcriptional coactivator ADA2 have distinct biological functions. Biochim Biophys Acta Gene Reg Mech 1789:117–124
Hiller Μ, Lin T-Y, Wood C, Fuller TM (2001) Developmental regulation of transcription by a tissue-specific TAF homolog. Genes Dev 15:1021–1030
Hiller Μ, Chen ΧM, Pringle J, Suchorolski M, Sancak Y, Viswanathan S, Bolival B, Lin T-Y, Marino S, Fuller TM (2004) Testis-specific TAF homologs collaborate to control a tissue specific transcription program. Development 131:5297–5308
Honys D, Twell D (2004) Transcriptome analysis of haploid male gametophyte development in Arabidopsis. Genome Biol 5:R85. doi:10.1186/gb-2004-5-11-r85
Johnsson-Brousseau SA, McCormick S (2004) A compendium of methods useful for characterizing Arabidopsis pollen mutants and gametophytically-expressed genes. Plant J 39:761–775
Kim HJ, Oh S-A, Brownfield L, Ryu H, Hwang I, Twell D, Nam H-G (2008) Control of plant male germline proliferation by SCFFBL17 degradation of cell cycle inhibitors. Nature 455:1134–1137
Kornet N, Scheres B (2009) Members of the GCN5 histone acetyltransferase complex regulate PLETHORA-mediated root stem cell niche maintenance and transit amplifying cell proliferation in Arabidopsis. Plant Cell 21:1070–1079
Lago C, Clerici E, Mizzi L, Colombo L, Kater MM (2004) TBP-associated factors in Arabidopsis. Gene 342:231–241
Lago C, Clerici E, Dreni L, Horlow C, Caporali E, Colombo L, Kater MM (2005) The Arabidopsis TFIID factor AtTAF6 controls pollen tube growth. Dev Biol 285:91–100
Lawit JS, O’Grady K, Gurley BW, Czarnecka-Verner E (2007) Yeast two-hybrid map of Arabidopsis TFIID. Plant Mol Biol 64:73–87
Lee TI, Young RA (1998) Regulation of gene expression by TBP-associated proteins. Genes Dev 12:1398–1408
Leurent C, Sanders SL, Demeny MA, Garbett KA, Ruhlmann C, Weil PA, Tora L, Schultz P (2004) Mapping key functional sites within yeast TFIID. EMBO J 23:719–727
Long JA, Ohno C, Smith ZR, Meyerowitz EM (2006) TOPLESS regulates apical embryonic fate in Arabidopsis. Science 312:1520–1523
Mandel AM, Yanofsky MF (1995) A gene triggering flower formation in Arabidopsis. Nature 377:522–524
McCormick S (1993) Male gametophyte development. Plant Cell 5:1265–1275
McCormick S (2004) Control of male gametophyte development. Plant Cell 16(Suppl):S142–S153
Pina C, Pinto F, Feijo JA, Becker JD (2005) Gene family analysis of the Arabidopsis pollen transcriptome reveals biological implications for cell growth, division control, and gene expression regulation. Plant Physiol 138:744–756
Poon D, Bai Y, Campbell AM, Bjorklund S, Kim Y-J, Zhou S, Kornberg RD, Weil PA (1995) Identification and characterization of a TFIID-like multiprotein complex from Saccharomyces cerevisiae. Proc Natl Acad Sci USA 92:8224–8228
Pray-Grant MG, Schieltz D, McMahon SJ, Wood JM, Kennedy EL, Cook RG, Workman JL, Yates JR III, Grant PA (2002) The novel SLIK histone acetyltransferase complex functions in the yeast retrograde response pathway. Mol Cell Biol 22:8774–8786
Qin Y, Leydon AR, Manziello A, Pandey R, Mount D, Denic S, Vasic B, Johnson MA, Palanivelu R (2009) Penetration of the stigma and style elicits a novel transcriptome in pollen tubes, pointing to genes critical for growth in a pistil. PloS Genet 5:8. doi:10.1371/journal.pgen.1000621
Robles LM, Wampole JS, Christians MJ, Larsen PB (2007) Arabidopsis enhanced ethylene response 4 encodes an EIN3-interacting TFIID transcription factor required for proper ethylene response, including ERF1 induction. J Exp Bot 58:2627–2639
Roeder RG (1996) The role of general initiation factors in transcription by RNA polymerase II. Trends Biochem Sci 21:327–335
Servet C, Conde E, Silva N, Zhou D-X (2010) Histone acetyltransferase AtGCN5/HAG1 is a versatile regulator of developmental and inducible gene expression in Arabidopsis. Mol Plant 34:670–677
Sessions A, Burke E, Presting G et al (2002) A high-throughput Arabidopsis reverse genetics system. Plant Cell 14:2985–2994
Shannon S, Meeks-Wagner DR (1991) A mutation in the Arabidopsis TFL1 gene affects inflorescence meristem development. Plant Cell 3:877–892
Sieberer T, Hauser MT, Seifert GJ, Luschnig C (2003) PROPORZ1, a putative Arabidopsis transcriptional adaptor protein, mediates auxin and cytokinin signals in the control of cell proliferation. Curr Biol 13:837–842
Smith TF, Gaitatzes C, Saxena K, Neer EJ (1999) The WD40 repeat: a common architecture for diverse functions. Trends Biochem Sci 24:181–185
Takada S, Goto K (2003) TERMINAL FLOWER2, an Arabidopsis homolog of HETEROCHROMATIN PROTEIN1, counteracts the activation of FLOWERING LOCUS T by CONSTANS in the vascular tissues of leaves to regulate flowering time. Plant Cell 15:2856–2865
Tamada Y, Nakamori K, Nakatani H, Matsuda K, Hata S, Furumoto T, Izui K (2007) Temporary expression of the TAF10 gene and its requirement for normal development of Arabidopsis thaliana. Plant Cell Physiol 48:134–146
Tao Y, Guermah M, Martinez E, Oelgeschlager T, Hasegawa S, Takada R, Yamamoto T, Horikoshi M, Roeder RG (1997) Specific interactions and potential functions of human TAFII100. J Biol Chem 272:6714–6721
Tora L (2002) A unified nomenclature for TATA box binding protein (TBP)-associated factors (TAFs) involved in RNA polymerase II transcription. Genes Dev 16:673–675
Verrijzer CP, Yokomori K, Chen JL, Tjian R (1994) Drosophila TAFII150: similarity to yeast gene TSM-1 and specific binding to core promoter DNA. Science 264:933–941
Vlachonasios KE, Thomashow MF, Trizenberg SJ (2003) Disruption mutations of ADA2b and GCN5 transcriptional adaptor genes dramatically affect Arabidopsis growth, development, and gene expression. Plant Cell 15:626–638
Walker KA, Blackwell TK (2003) A broad but restricted requirement for TAF-5 (Human TAFII100) for embryonic transcription in Caenorhabditis elegans. J Biol Chem 278:6181–6186
Wang Y, Zhang WZ, Song LF, Zou JJ, Su Z, Wu WH (2008) Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis. Plant Physiol 148:1201–1211
Weigel D, Nilsson O (1995) A developmental switch sufficient for flower initiation in diverse plants. Nature 377:495–500
Wright KJ, Marr MT II, Tjian R (2006) TAF4 nucleates a core subcomplex of TFIID and mediates activated transcription from a TATA-less promoter. Proc Natl Acad Sci USA 103:12347–12352
Yadegari R, Drews GN (2004) Female gametophyte development. Plant Cell 16(Suppl):S133–S141
Acknowledgments
We thank Drs Amy Hark (Muhlenberg College) and Vasilis Gegas (John Innes Centre) for thoughtful discussions during the course of this work. We also thank Dr Maria Moustaka and Savvas Genitsaris (Aristotle University of Thessaloniki) for the assistance with fluorescence microscopy. This research was supported by Greek General Secretary of Research and Technology grant number 82337 and by funds from Aristotle University of Thessaloniki, School of Biology to KV.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mougiou, N., Poulios, S., Kaldis, A. et al. Arabidopsis thaliana TBP-associated factor 5 is essential for plant growth and development. Mol Breeding 30, 355–366 (2012). https://doi.org/10.1007/s11032-011-9626-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-011-9626-2