Abstract
With the development of high-throughput sequencing techniques, transcriptome sequencing projects which provide valuable resources for designing simple sequence repeat (SSR) primers have been carried out for many plants. However, the utility of SSRs for molecular breeding depends on genome-wide distribution and coverage, as well as moderately high allelic variability, in the available SSR library. In this study, we characterized the exonic SSRs developed from the publicly available Populus genome as a case study to determine their value for molecular breeding. As expected, our results confirmed that microsatellites occurred approximately three times less often in coding regions than in non-coding regions. Mutability test also showed that exonic SSRs contained less allelic variability than intronic SSRs. More importantly, exonic SSRs were unevenly distributed both among and within chromosomes. Large exonic SSRs deserts were observed on several chromosomes. Differential selection between paralogous chromosomes, at the gene level, appears to be responsible for these SSR deserts, though the mechanisms that cause chromosome-specific SSR deserts are not known. This work provides ample evidence that the candidate gene approach based on unigenes identified from transcribed sequences may not be the best strategy to identify highly polymorphic SSRs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Molecular breeding is the use of genetic markers to assist and accelerate conventional breeding and selection. Simple sequence repeats (SSRs) or microsatellites are tandem repeats of short DNA sequences which are abundant and widespread in all eukaryotic genomes (Toth et al. 2000). SSR markers are one of the most important and powerful genetic tools for molecular breeding. Microsatellites owe their variability to an increased rate of mutation compared to the other neutral regions of DNA (Li et al. 2002). Due to their high mutability, microsatellites are also thought to play a significant role in genome and gene evolution by creating and maintaining quantitative genetic variation (Kashi et al. 1997; Tautz et al. 1986). The priming sequences of microsatellites are normally conserved within species and may even be transferable among taxa within genera (Castillo et al. 2008; Yin et al. 2009). Their high transferability makes microsatellites one of the most powerful genetic markers for syntenic studies (Yin et al. 2004). Using SSRs, it is feasible to build a genotyping platform which facilitates the study of alternative individuals and/or species within the context of a macro-genetic system and to validate genetic discoveries among alternate pedigrees. Because of their high variability, ubiquity, and co-dominance, microsatellites have been used as versatile molecular markers that have wide-ranging applications in the field of genetics.
The sequencing capacity available to biological scientists has increased in an exponential manner, and with it the number of complete draft plant genomes continues to grow (Lyons and Freeling 2008). However, the genetic resources available to a plant researcher vary significantly depending on the focal species. Many plant researchers still struggle to study and manipulate valuable traits with limited genetic tools. For plants with limited genomic/genetic resources available, developing molecular markers is an important step for detecting genetic loci underlying traits of interest, and SSR markers are widely accepted as the most desirable molecular markers for molecular breeding. Compare to whole genome sequencing, transcriptome sequencing is an attractive and efficient alternative way for identification, discovery, and characterization of genes in different organisms (Lijoi et al. 2007). Transcribed sequences also provide valuable resources for developing SSR markers (Castillo et al. 2008; Khanal et al. 2008). However, the utility of SSR markers in molecular breeding is mainly determined by their distribution in the genome, coverage of the genome, and variability. Supported by the US Department of Energy (DOE), the genome of a black cottonwood (Populus trichocarpa) tree, Nisqually-1, has been completely sequenced and publicly released (Tuskan et al. 2006). The achievement of the Nisqually-1 genome sequence offers us an opportunity to characterize the exonic SSRs in the genome of a representative woody plant. In this paper, our objectives were to: (1) characterize the distribution and coverage of exonic SSRs in the Nisqually-1 genome, (2) reveal the exonic SSR deserts in the poplar genome, and (3) investigate the allelic variability of exonic SSRs compared to SSRs in the neutral regions of DNA.
Materials and methods
Genome sequences were obtained from the Populus genome browser (http://shake.jgi-psf.org/cgi-in/searchGM?db=Poptr1_1) and data for exonic SSRs were collected from the SSR primer resources developed by Yin et al. (2009). Among the 45,555 predicted protein-coding gene loci in the Populus genome (Tuskan et al. 2006), we examined approximately 30,000 gene models which were unambiguously mapped to sequence scaffolds along 19 chromosomes of the Populus genome (about 80% of the total genome length) and showed high conservation with the Arabidopsis gene set. The distribution of exonic microsatellites among chromosomes was evaluated by the observed number of exonic microsatellites compared to their expectations using a Poisson calculation. The expected number of exonic SSRs λ i in chromosome i would be a sample from a Poisson distribution:
where m i is the observed number of exonic SSRs on chromosome i and L i is the length of A, T, C, G designation on chromosome i. The probabilities of P(m i < λ i ) and P(m i > λ i ) were calculated at α = 0.05 and α = 0.01 significant levels. To explore the SSR distribution within each chromosome, we created 2-Mb sliding windows along each chromosome. The abundance or disparity of exonic SSRs within each window was determined based on the above statistical method, except that chromosome i was replaced with window i. Visual plotting of exonic SSRs on each chromosome was produced by the Mapchart software (Voorrips 2002).
The identification of paralogous chromosomal segments in the Populus genome resulting from “salicoid” duplication events was described by Tuskan et al. (2006). Blocks of the same color represented the paralogous chromosomal segments. Segmentally duplicated gene pairs were identified by BLASTp as described by Yang et al. (2008) with settings of e-values ≤1×10−25; minimum conserved region width ≥100 bp; and conservation identity ≥70%.
The efficacy of the SSR markers for genetic analyses was determined by genotyping of 24 genetically diverse Populus adenopoda genotypes sampled in the suburban areas of Nanjing, China. In total, 96 SSR primer pairs were synthesized and used in the test. Among these primer pairs, 48 amplified microsatellites in exons, and the rest amplified microsatellites in introns. To minimize the effect of confounding factors on allelic variation, priming sequences of all primer pairs were located in coding regions and the tested microsatellites had identical lengths in the SSR database. PCR reactions were performed as described by Yin et al. (2004). Polymorphic differences among the tested individuals were used to calculate the polymorphism information content (PIC) associated with each SSR marker using the following formula (Keim et al. 1992):
where n is the total number of alleles detected for a SSR marker, and P i is the frequency of the ith allele.
Results
Exonic SSRs distribution in Nisqually-1 genome
In an earlier study, a total of 148,428 SSR primer pairs were identified and created from the unambiguously mapped sequence scaffolds of the Nisqually-1 genome (Yin et al. 2009). In the present paper, we specifically collected the information and analyzed data for exonic SSRs from this comprehensive SSR database. In total, we identified 5,989 SSR primer pairs (about 4% of the total) that amplified microsatellites in transcribed sequences of the Populus genome. On average, exonic SSRs occurred approximately every 47 kb of DNA within the Populus genome. The Poisson distribution tests (Table 1) revealed that exonic SSRs were distributed unevenly among chromosomes. Ten chromosomes had more exonic SSRs than expected by chance alone; six chromosomes contained fewer than expected; and only three chromosomes had exonic SSRs that did not significantly depart from the expected numbers. By creating 2-Mb sliding windows along each chromosome, we further explored SSR distribution within chromosomes (Supplementary Table 1). The results showed that the exonic SSRs were also distributed unevenly within each chromosome. In total, we created 146 sliding windows on the 19 chromosomes of the Populus genome. Among these windows, 51 were overabundant with exonic SSRs, 46 were sparse with exonic SSRs, and 46 contained exonic SSRs that did not significantly depart from the expected numbers. Thus, the distribution of exonic SSRs was uneven both among and within chromosomes.
Large exonic SSR deserts observed in Nisqually-1 genome
Coverage of exonic SSRs was evaluated using heat plots for each chromosome (Fig. 1). In the plotting chart, large exonic SSR regions lacking SSRs (i.e., SSR “deserts”) were found on chromosomes III, V, X, XII, XVII, XVIII and XIX, encompassing DNA regions of several million base pairs. The extant Populus genome arose from an ancient whole-genome duplication event known as “salicoid” duplication and appears to have evolved from ten ancestral chromosomes (Tuskan et al. 2006). Based on the homology among chromosomes, we found that many of the exonic SSR deserts had duplicated segments on other chromosomes. In Fig. 1, it is noteworthy that exonic SSR densities diverged on some of the paralogous chromosomal segments. To explore whether the differential loss of exonic SSRs on the paralogous segments is due to loss of the corresponding genes or the corresponding motifs, we analyzed the genes containing SSRs on the paralogous segments between chromosome I and chromosome III (Supplementary Table 2), which shared two duplicated regions. In the upper duplicated region on chromosome I, 160 exonic repeat motifs were present. On its duplicated segment on chromosome III, 36 of these repeat motifs were maintained, 113 lost, and 11 of these repeat motifs were in paralogous genes on unmapped scaffolds. Among the 113 missing repeat motifs, 47 (41.6%) were due to loss of the corresponding genes, and 66 were due to loss of the corresponding motifs; whereas the lower paralogous region on chromosome I contained 65 exonic SSR repeat motifs. On its duplicate segment on chromosome III, three of these repeat motifs were maintained, 51 missed, and ten of these motifs were in paralogous genes on unmapped scaffolds. Among the 51 missing repeat motifs, 25 (49.0%) were due to loss of the corresponding genes, and the other ones were due to loss of the corresponding motifs. Thus, on average, 45.3% of exonic repeat motifs that were missing on the duplicated chromosomal segments were due to loss of the corresponding genes, and 54.7% of them were due to loss of the corresponding motifs.
Visual plotting of exonic SSRs along each Populus chromosome. The leftmost vertical bar is the physical length ruler scaling in Mb. In this figure, each chromosome corresponds to two vertical bars, with the left bar showing the chromosomal homology and the right bar displaying the positions of exonic SSRs. Fragments in the same color indicate the duplicated chromosomal segments resulted from the salicoid duplication event, as described by Tuskan et al. (2006)
Allelic variability of exonic SSRs
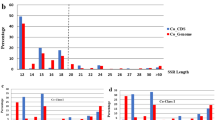
High mutability makes SSRs one of the most desirable molecular markers for genetic analyses. In this paper, we designed an experiment to test the allelic variability of exonic SSRs among 24 P. adenopoda genotypes and to compare their mutability with that of microsatellites located in introns. Out of the 96 selected exonic primer pairs from the Nisqually-1 resource, only seven failed to amplify product in the selected P. adenopoda genotypes (Supplemental Table 3). The successful amplification rate was 92.7%. In an earlier study, the transferable rate of SSR primer pairs developed from Nisqually-1 genome sequences to P. adenopoda was found to be relatively low (75%) (Yin et al. 2009). The high success rate in this study may be largely related to exonic-anchored primer sites among alternative Populus species. The allele numbers and PIC value revealed by each primer pair are demonstrated in Fig. 2. According to this Figure, over 70% of exonic SSRs generated one allele per locus; in contrast, more than 85% intronic SSRs generated two or more alleles per locus. The average allele number for genetic loci encompassing exonic microsatellites was 1.40 vs. 3.28 for genetic loci encompassing intronic microsatellites. PIC values were also significantly different between intronic and exonic microsatellites. According to the PIC values, intronic SSRs were ranked as moderately to highly informative with an average PIC of 0.403; whereas exonic SSRs were extremely uninformative with an average PIC of 0.072.
Comparison of allele numbers (a) and PIC values (b) between exonic SSRs and intronic SSRs, as determined by genotyping of 24 unrelated P. adenopoda genotypes. In each diagram, the vertical bars indicate the percentage of SSRs with the corresponding values on the x axis. The mean, the median and the maximum values of the allele numbers and PIC are also listed in the corresponding diagrams
Discussion
Microsatellites are generally found in genomic positions free from genetic selection; recent studies have attributed various functional roles to microsatellites (Jewell et al. 2006). There is direct evidence that microsatellites can function as transcriptional activating elements (Li et al. 2002), and there is some evidence suggests that even microsatellites in non-coding regions may have functional significance (Kashi et al. 1997). Microsatellites were found to play roles in many human neurodegenerative disorders (Reddy and Housman 1997) and in some human cancers (Lothe 1997). Thus, microsatellites may contain functional importance relative to the corresponding DNA sequence. Results from this study indicate that genes containing microsatellites are under selection. The evidence is as follows: first, the frequency of microsatellites is three times lower in the coding regions than in the neutral regions. Second, the allelic variability of exonic SSRs is much lower than that of intronic SSRs. Third, the exonic SSRs are found to be differentially lost in the segmentally duplicated gene pairs in the Populous genome.
In an earlier study on genomic SSRs, no large SSR deserts were observed across the Populus genome (Yin et al. 2009); in contrast, huge exonic SSR deserts were found on many poplar chromosomes in the present study. Meanwhile, we noted that the exonic SSRs were differentially lost on the paralogous segments among chromosomes. Most eukaryotic genomes have numerous duplicated genes, many of which appear to have arisen from one or more cycles of ancient polyploidy (paleopolyploidy). Following paleopolyploidy, there is extensive loss of duplicated genes (Adamsa and Wendel 2005) and/or gene subfunctionalization (Adamsa et al. 2004; Blanc and Wolfe 2004). Cytological studies reveal that all extant Populus species exist in the diploid form with a haploid numbers of chromosomes equal to 19 (Smith 1943). However, the Populus genome has experienced three separate ancient genome-wide duplication events, with the most recent event contained within the Salicaceae (Tuskan et al. 2006). Analyzing the exonic repeat motifs on the paralogous chromosomal segments in the Populus genome revealed that the majority of the exonic repeat motifs were missing on their duplicated segments. Nearly half of the missing motifs were due to loss of the corresponding genes and the rest were due to loss of the corresponding motifs. These results implied the extensive loss of duplicated genes and gene subfunctionalization in the Populus genome after salicoid duplication. The mechanism leading to and the functional relevance of this phenomenon are not known at this time.
The exonic SSRs are useful for syntenic studies and have an important use to anchor BACs because they are more conserved than the intronic and intergenic SSRs. The transcribed sequences provide valuable resources for developing SSR markers. However, the transcribed sequences are commonly redundant and only represent a portion of the putative genes (Lijoi et al. 2007; Susko and Roger 2004; Wang et al. 2005), which will lead to redundant amplification and limit the genome range covered by the exonic SSRs. Moreover, based on the allele numbers and PIC values derived from 24 unrelated P. adenopoda genotypes, exonic SSRs were found to be much less polymorphic than intronic SSRs. Normally, SSRs with PIC value >0.5 were considered as highly informative markers that would likely be useful in tracking the economically important traits in breeding populations (Ren et al. 2009). Results in this paper indicated that exonic SSRs with PIC value >0.5 were scarce (Fig. 2). To our knowledge, this is the first study to carry out extensive investigation specifically on exonic SSRs at the genome-wide level. While genome structure always varies among alternative genotypes and among related species, this study provides a valuable reference to evaluate exonic SSRs and their applications for molecular breeding.
References
Adamsa KL, Wendel JF (2005) Novel patterns of gene expression in polyploid plants. Trends Genet 21:539–543
Adamsa KL, Percifield R, Wendel JF (2004) Organ-specific silencing of duplicated genes in a newly synthesized cotton allotetraploid. Genetics 168:2217–2226
Blanc G, Wolfe KH (2004) Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. Plant Cell 16:1679–1691
Castillo A, Budak H, Varshney RK, Dorado G, Graner A, Hernandez P (2008) Transferability and polymorphism of barley EST-SSR markers used for phylogenetic analysis in Hordeum chilense. BMC Plant Biol 8:97–105
Jewell E, Robinson A, Savage D et al (2006) SSR primer and SSR taxonomy tree: biome SSR discovery. Nucl Acids Res 34:656–659
Kashi Y, King D, Soller M (1997) Simple sequence repeats as a source of quantitative genetic variation. Trends Genet 13:74–78
Keim P, Beavis W, Schupp J, Freestone R (1992) Evaluation of soybean RFLP marker diversity in adapted germplasm. Theor Appl Genet 85:205–212
Khanal S, Tang S, Beilinson V, Miguel PS et al (2008) ESTs are a rich source of polymorphic SSRs for genomics and molecular breeding applications in peanut [abstract]. In: Proceedings of the plant and animal genome XVI conference. San Diego, California, p 135
Li YC, Korol AB, Beiles A et al (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol Ecol 11:2453–2465
Lijoi A, Mena RH, Prünster I (2007) A Bayesian nonparametric method for prediction in EST analysis. BMC Bioinformatics 8:339–359
Lothe RA (1997) Microsatellite instability in human solid tumors. Mol Med Today 3:61–68
Lyons E, Freeling M (2008) How to usefully compare homologous plant genes and chromosomes as DNA sequences. Plant J 53:661–673
Reddy PS, Housman DE (1997) The complex pathology of trinucleotide repeats. Curr Opin Cell Bio 9:364–372
Ren Y, Zhang Z, Liu J, Staub JE, Han Y et al (2009) An integrated genetic and cytogenetic map of the cucumber genome. PLoS ONE 4(6):e5795
Smith EC (1943) A study of cytology and speciation in the genus Populus L. J Arnold Arbor 24:275–305
Susko E, Roger RJ (2004) Estimating and comparing the rates of gene discovery and expressed sequence tag (EST) frequencies in EST surveys. Bioinformatics 20:2279–2287
Tautz D, Trick M, Dover GA (1986) Cryptic simplicity in DNA is a major source of genetic variation. Nature 322:652–656
Toth G, Gaspari Z, Jurka J (2000) Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res 10:967–981
Tuskan GA, DiFazio SP, Hellsten U et al (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray ex Brayshaw). Science 313:1596–1604
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang JPZ, Lindsay BG, Cui LY et al (2005) Gene capture prediction and overlap estimation in EST sequencing from one or multiple libraries. BMC Bioinformatics 6:300–310
Yang XH, Kalluri UC, Jawdy S et al (2008) Comparative analysis of F-Box proteins in arabidopsis, poplar and rice suggests differential proteolytic pathway mediated through F-Box protein between woody perennial and herbaceous annual plants. Plant Physiol 148:1189–1200
Yin TM, DiFazio SP, Gunter LE, Riemenschneider D, Tuskan GA (2004) Large-scale heterospecific segregation distortion in Populus revealed by a dense genetic map. Theo Appl Genet 109:451–463
Yin TM, Zhang XY, Gunter L et al (2009) Microsatellite primers resource developed from the mapped sequence scaffolds of Nisqually-1 genome. New Phytol 181:498–503
Acknowledgments
We thank Han Jingling, Zhenghong Liu and Faying Yuan at Nanjing Forestry University for their help in sample collection. Special thanks go to the editor and anonymous reviewers for their help in formulating the revision. Funds for this research were provided by Natural Science Foundation of China (30971609) and Forestry Nonprofit Project (200904002). This work is also supported by the US DOE BioEnergy Science Center. The BioEnergy Science Center is a US Department of Energy Bioenergy Research Center supported by the Office of Biological and Environmental Research in the DOE Office of Science.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11032_2010_9413_MOESM1_ESM.xls
Table 1. Distribution test of exonic SSRs in the 2-Mb sliding windows along the Populus chromosomes. The valid reading length is the length of A, T, G, C designation in each window. Distribution test is performed by the Poisson calculation at P ≤ 0.05 and P ≤ 0.01 significance levels. “**+” Significantly overabundant at P ≤ 0.01; “*+” significantly overabundant at P ≤ 0.05; “**−” significantly sparse at P ≤ 0.01; “*−” significantly sparse at P ≤ 0.05 (XLS 78 kb)
11032_2010_9413_MOESM2_ESM.xls
Table 2. Exploring the loss of exonic SSR repeat motifs on the paralogous segments between chromosome I and chromosome III. Note: The two chromosomes share two duplicated segments; lines without shadow correspond to exonic SSRs in the upper duplicated region on chromosome I, whereas lines in gray shadow correspond to the exonic SSRs in the lower duplicated region on chromosome I (XLS 201 kb)
11032_2010_9413_MOESM3_ESM.xls
Table 3. Summary of primer information and amplification results in SSR variability test. In this table, the primer name is designated as “P” followed by “number of chromosome ID”, followed by “_”, and ends with “the start base of left primer”. Columns I to AF contain the amplification results of the tested primer pairs. In these columns, “M/M” stand for the missing data; “No amp” stand for no amplification products. The corresponding gene model and their definition are listed in the last two columns (XLS 165 kb)
Rights and permissions
About this article
Cite this article
Li, S., Yin, T., Wang, M. et al. Characterization of microsatellites in the coding regions of the Populus genome. Mol Breeding 27, 59–66 (2011). https://doi.org/10.1007/s11032-010-9413-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-010-9413-5