Abstract
Long noncoding RNAs (lncRNAs) are recently defined as thousands of RNA molecules longer than 200 nucleotides and lacking an appreciable open reading frame in mammals. Although lncRNAs lack protein-coding function, they play critical roles in the regulation of almost all the protein-coding genes in a cell at various stages including chromatin modification, transcription and post-transcriptional processing. It is thus not surprising that lncRNAs may be the crucial regulators in the normal development, physiology and pathology. LncRNAs in neuroscience is a novel research field. Interestingly, recent studies have demonstrated that many lncRNAs are highly expressed in brain and their dysregulations occur in neurological disorders. In this review, we describe the current understanding of lncRNAs in neurobiology and neurological diseases including cerebral injury. LncRNAs could be novel biomarkers and could be potential new targets for new drugs for many neurological diseases in the future, although the related studies are still at in the early stages.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Next-generation sequencing technologies and genome-wide analysis of the eukaryotic transcriptome have revealed that up to 90% of the human genome is transcribed; however, GENCODE-annotated exons of protein-coding genes only cover 2.94% the genome (Wang et al. 2009a; Li et al. 2014; Meyer and Liu 2014; The ENCODE Project Consortium 2012). The results suggest the majority of the transcribed products of genome are noncoding RNAs (ncRNAs). According to their basic functions, non-coding RNAs (ncRNAs) are divided into the “housekeeping” RNAs and the regulatory RNAs (Fig. 1). “Housekeeping” ncRNAs usually considered constitutive include ribosomal, transfer, small nuclear and small nucleolar RNAs, whereas the regulatory RNAs are ncRNAs with strong regulatory effects on protein-coding gene expression. It is well-known that “housekeeping” ncRNAs have no regulatory effects on protein-coding gene expression, however, a recent study showed that tRNA-derived stress-induced RNAs (tiRNAs) cooperated with the translational repressor Y-box binding protein 1 (YB-1) to displace the cap-binding complex eIF4F from capped mRNA, inhibit translation initiation, and induced the assembly of stress granule (Lyons et al. 2016). Moreover, the regulatory ncRNAs are further divided into two categories according to their size. ncRNAs less than 200 nucleotides (nt) are defined as small ncRNAs (sncRNA), including microRNAs (miRNAs), small interfering RNAs(siRNAs) and Piwi-associated RNAs(piRNAs). In contrast, the regulatory ncRNAs longer than 200 nucleotides are named as long non-coding RNAs (lncRNAs), which lack an appreciable open reading frame (usually less than 100 amino acids) (Taft et al. 2010; Dey et al. 2014; Mattick 2001). However, it is far from being scientific and systematic in defining lncRNAs simply based on the aforementioned scales with the increasing acquaintance of lncRNAs. Based on their discovery, its size, sequencing and functions, each identified lncRNA has been tried to give a special name. The initial lncRNAs, X-inactive specific transcript (XIST) and H19 were firstly discovered and named in human and mouse in 1980s and 1990s (Brown et al. 1991; Bartolomei et al. 1991). However, unlike miRNAs, the nomenclature of lncRNAs is standard and uniform. This may give rise to the difficulty in retrieving and integrating the study results with the large amount of identified lncRNAs and increasing number of new discovered lncRNAs (Kapranov et al. 2007; Davey et al. 2011; Bras et al. 2012).
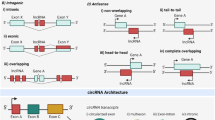
Paradigms for ncRNAs classification. Non-coding RNAs (ncRNAs) are functionally grouped into the “housekeeping” RNAs and the regulatory RNAs. The regulatory RNAs are further grouped into small ncRNAs and long ncRNAs (lncRNAs) based on their nucleotides (nt) size. Small ncRNAs are less than 200 nucleotides, whereas lncRNAs are longer than 200 nucleotides. Small ncRNAs and lncRNAs are further divided into some sub-groups
The central nervous system (CNS) is a complex biological system and is composed of many cell types working in concert. The precise spatial and temporal control of gene expression in the neural cells is responsible for the intricate development and functioning of this highly ordered structure. Cell fates and functions in CNS are tightly control by gene expression and the neurological disorders are depended upon failure to develop and/or maintain the intricate regulatory networks properly.
LncRNAs are highly expressed in CNS and play crucial roles in spatial-temporal control of gene expression in the developing and adult brain (Mercer et al. 2008; Mehler and Mattick 2007). A cohort of 13 lncRNA-null mutant mouse models and the knocked-in lacZ reporter gene were used to determine the spatiotemporal expression profiles of lncRNAs in the brain and the transcriptome alterations resulting from the loss of these lncRNA loci. It is surprising that several lncRNAs are differentially expressed both in time and space, with some presenting highly restricted expression in only selected brain regions. The expression of some neighboring proteincoding genes can be significantly affected by several lncRNAs in a cis-like manner. LncRNAs are emerging as important components of gene regulatory networks CNS, working in concert with transcription factors and epigenetic regulators of gene expression. Recent studies have demonstrated that lncRNAs are key regulators in neuron function, maintenance, differentiation, brain development and brain functions. Dysregulation and/or dysfunction of lncRNAs are involved in neurological disorders including cerebral ischemia reperfusion injury and hypoxic-ischemic brain damage (Dharap et al. 2012; Dharap et al. 2013; Kaur et al. 2014; Zhang et al. 2016a; Zhao et al. 2015). LncRNAs could be novel biomarkers and new therapeutic effects for many neurological diseases, although the related studies are still at early stages. In this review, we will describe the current understanding of lncRNAs and their roles in neurodevelopment and neurological diseases.
lncRNAs subgroup and classification
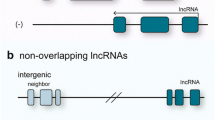
According to their transcription positions in the genome, lncRNAs are categorized into three major groups: (A) lncRNAs transcribed at location relative to the host protein coding gene (PCG), (B) lncRNAs transcribed from the gene regulator regions and (C) lncRNAs transcribed at some specific chromosomal regions. In addition, each group can be further divided into some subgroups (Wu et al. 2013). For lncRNAs transcribed relative to the host PCG, it can be placed into one of the following five subcategories: 1) sense RNAs (when overlapping one or more exons of another transcript on the same strand); 2) antisense RNAs (when overlapping one or more exons of another transcript on the opposite strand); 3) bidirectional RNAs (when the expression of it and a neighboring coding transcript on the opposite strand is initiated in close genomic proximity); 4) intronic RNAs (when it is derived wholly from within an intron of a second transcript); or 5) intergenic RNAs(when it lies within the genomic interval between two genes) (Ponting et al. 2009). For the lncRNAs transcribed from the gene regulator regions, the following subgroups are described: 1) 3-UTR associated RNAs (lncRNAs are derived from 3′-untranslated regions of protein-coding transcript, also referred to uaRNAs) (Mercer et al. 2011); 2) promoter associated RNAs (lncRNAs are transcribed from promoter domains of protein-coding genes) (Hung et al. 2011); 3) enhancers or enhancer-like lncRNAs (lncRNAs are transcribed from enhancer domains and expressed coordinately with, activity-dependent genes, or lncRNAs exhibiting enhancer activity) (Ørom et al. 2010). Specific chromosomal regions derived lncRNAs include telomeres, telomeric repeat-containing RNA (TERRA) (Azzalin et al. 2007) (Fig. 1).
Functions of lncRNAs
Unlike miRNAs, lncRNA functions can not only be inferred from their special sequences and/or structures. The broad spectrum of lncRNAs functions are involved in biological process, high-order chromosomal dynamics and subcellular structural organization from chromatin structure to the protein level (Wu et al. 2013; Amaral and Mattick 2008). Importantly, lncRNAs are able to regulate gene expression at the level of chromatin modification, transcription and post-transcriptional processing (Mercer et al. 2009). LncRNA functions also include genomic site-specific epigenetic reprogramming, long-range genomic interactions, scaffolds for nuclear macromolecular assemblies and maintenance of stoichiometry and molecular titration (Qureshi and Mehler 2012). Figure summarizes that current knowledge about the functions of lncRNAs, although the full functions of lncRNAs have not yet been totally identified. In respect to the gene regulation function, recent studies have clearly demonstrated that lncRNAs are able to regulate the expression of protein-coding genes by transcription interference, chromatin remodeling and histone modifications, alternative splicing patterns, and other post-transcriptional regulation (Fig. 2) (Wilusz et al. 2009). In addition, a recent study has shown that terminal differentiation-induced ncRNA (TINCR) as a key lncRNA, can control human epidermal differentiation by a post-transcriptional mechanism. Genome-scale RNA interactome analysis showed that TINCR-mRNA interaction occurs through a 25-nucleotide “TINCR box” motif that is strongly enriched in interacting mRNAs and required for TINCR binding (Kretz et al. 2013). It has been revealed that the most specific interaction was between lncRNA HOTAIR(HOX transcript antisense RNA) and the heterogeneous nuclear ribonucleoprotein (hnRNP) A2/B1, a member of a family of proteins involved in nascent mRNA processing and RNA matchmaking. Furthermore, this study also demonstrated that a direct RNA-RNA interaction between HOTAIR and a target transcript (Meredith et al. 2016).
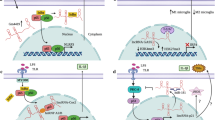
Paradigm for the mechanisms of lncRNAs functions. Transcription from an upstream noncoding promoter (tawny) can negatively or positively affect the downstream gene expression (purple) via transcriptional interference through inhibiting RNA polymerase II recruitment (1), or via inducing chromatin remodeling and histone modifications (2),respectively. Additionally, antisense transcripts (blue) are able to hybridize to their specific sense transcripts (purple), resulting in alternatively spliced transcript (3) or various endogenous siRNAs(4). With binding to specific protein partners(green), a noncoding transcript can modulate the activity of the protein(5), or allow the formation of a larger RNA–protein complex(6), or alter the cellular localization of the protein(7). LncRNAs (yellow) can be processed to yield small RNAs including miRNAs, piRNAs and others (8). Moreover, they can also act as miRNA sponges that affect the competitive endogenous RNAs(9) (Wilusz et al. 2009)
LncRNAs in the CNS
The CNS, a complex biological system, is composed of an enormous array of regionally distinct neuronal and glia subtypes, in which lncRNAs are highly expressed. A pioneering study utilizing data from the Allen Brain Atlas showed that 849 lncRNAs were expressed in the adult mouse brain (among the 1328 examined) and found that the majority were lncRNAs associated with specific regions, cell types, and subcellular compartments, indicating that they provide biological function (Mercer et al. 2008). Subsequently, recent studies analyzing with high-throughput transcriptomic methods have shed light on the expression patterns and roles of lncRNAs during neural cell fate determinations and neural differentiation (Guttman et al. 2009; Fantes et al. 2003; Amaral et al. 2009; Tochitani and Hayashizaki 2008; Sone et al. 2007; Rapicavoli et al. 2010; Ng et al. 2012), synaptic and neural plasticity (Iyengar et al. 2014; Francescatto et al. 2014; Aprea et al. 2013), brain development (Sauvageau et al. 2013), neurodegenerative disorders (Johnson 2012; Liu et al. 2014; Soreq et al. 2014) neuropsychiatric disease (Ziats and Rennert 2013), and cerebral injury (Dharap et al. 2012; Dharap et al. 2013; Kaur et al. 2014; Zhang et al. 2016a; Zhao et al. 2015). It is well-known that lncRNAs are critical regulators for those conditions.
LncRNAs in neural cell fate determinations and neural differentiation
LncRNAs exert the effects on mediating neural differentiation, including neural line restriction, neural cell fate determinations and progressive stage differentiation. LncRNA-AK053922 transcribed from the Gli3 locus has the ability of helping specify distinct neuronal cell types via acting as a bifunctional transcriptional switch (Meyer and Roelink 2003; Hashimoto-Torii et al. 2003). A study identified approximately 1600 evolutionarily conserved intergenic lncRNAs (lincRNAs) by analyzing chromatin signatures across four mouse cell types (Guttman et al. 2009). Intriguingly, further analyses of specific lncRNAs, which dynamically expressed in the CNS, demonstrated potential roles of these lncRNAs in mediating neural cell fate determinations.
It is well known that Sox2, a key transcription factor, is required for neural induction and maintenance of progenitor and neural stem cells. A recent study has shown that the lncRNA Sox2OT, which contains the Sox2 gene within one of its introns and is subsequently transcribed in the same direction (Fantes et al. 2003), was expressed in the regions of constitutive adult neurogenesis (Mercer et al. 2008; Amaral et al. 2009). In addition, Sox2OT was dynamically regulated in CNS structures during development, where it may be responsible for modulating Sox2 expression (Amaral et al. 2009).
Similarly, the lncRNA Nkx2.2AS could regulate Nkx2.2 (a transcription factor that is critical for oligodendrocyte (OL) lineage specification). Overexpression of Nkx2.2AS in neural precursor cells (NPCs) in vitro enhanced induction of their differentiation to the oligodendrocyte lineage by an increase in Nkx2.2 mRNA levels (Tochitani and Hayashizaki 2008). The result implies that Nkx2.2AS has a regulatory effect based not only on the transcription of Nkx2.2 in cis, but also on the other mechanisms responsible for OL lineage specification in trans. The lncRNA RNCR2, also known as Gomafu and Miat, is primarily expressed in neuronal cells and localized in nuclear subdomains. It is thought to play a critical role in regulating retinal cell specification (Sone et al. 2007; Rapicavoli et al. 2010). In addition, a study has also shown that the lncRNAs are important regulators of pluripotency and neurogenesis. Among then, lncRNA RMST, lncRNA N1 (AK124684), lncRNA N2 (AK091713) and lncRNA N3 (AK055040) are required for efficient neuronal differentiation (Ng et al. 2012).
LncRNAs play potential roles in adult neurogenesis. Knockdown of two lncRNAs such as Six3os and Dlx1as, indicates the roles of lncRNAs in the glial-neuronal lineage specification of multipotent adult stem cells (Ramos et al. 2013). Pinky (Pnky), as a neural-specific lncRNA, can interact with a key RNA processing factor and regulate neurogenesis from neural stem cells (NSCs) in the embryonic and postnatal brain (Ramos et al. 2015). lncRNA LOC646329 is enriched in single radial glia cells but is observed at low level in tissues. Knockdown of LOC646329 shows that lncRNA can regulate cell proliferation (Liu et al. 2016).
LncRNAs in synaptic plasticity, cognitive function and memory
LncRNAs are also implicated in the processes of gene regulations responsible for synaptic plasticity, cognitive function and memory. It is well known that the normal development of GABAergic inhibitory interneurons in the hippocampus is responsible for learning. LincRNA Evf-2, which transcribed from the Dlx-5/6 ultraconserved region, is essential for the development of GABAergic neuron. Evf-2 exerts its function via Dlx-2 transcriptional coactivator to increase the transcriptional activity of Dlx-5/6 and the glutamate decarboxylase 1(Gad1), and then regulates the gene expression of GABAergic interneurons in the developing mouse brain. Gad1 is necessary for the conversion of glutamate to GABA) (Feng et al. 2006). Evf-2 silence can result in anomalies synaptic activity by the aberrant formation of GABAergic circuitry in the hippocampus and dentate gyrus in mice (Bond et al. 2009).
The lncRNA, metastasis-associated lung adenocarcinoma transcript 1 (Malat1), is enriched in hippocampal neurons. Malat1 knockdown causes decrease in synaptic density, whereas its over-expression leads to a cell-autonomous increase in synaptogenesis (Bernard et al. 2010). The rodent-specific BC1 and primate-specific BC200 lncRNAs derived from transposable elements and transcribed by RNA polymerase III, are selectively targeted to postsynaptic dendritic compartments, where they regulate the synthesis of local protein via inhibiting the initiation of translation through an eIF4Adependent mechanism (Lin et al. 2008; Kondrashov et al. 2005; Zalfa et al. 2005). Although BC1 knockout in mice showed no overt phenotype in the cage, it produced behavioral changes including reduced exploration and increased anxiety, leading to reduction in survival rates (Lewejohann et al. 2004). Similarly, lncRNA Ntab is expressed in developing and adult rat brain and it may function within cellular processes (French et al. 2001).
Brain derived neurotrophic factor (BDNF) are essential for supporting neuronal growth, survival, synaptic plasticity, learning and memory process (Lynch et al. 2007; Xie et al. 2010; Kealy and Commins 2010). Several recent studies have found that the lncRNA antiBDNF (BDNF-AS, also named as BDNF-OS) which transcribed from the antisense of BDNF gene, could form dsRNA duplexes with BDNF mRNA in the brain. Losing function of BDNF-AS caused increase in BDNF mRNA and protein level, then promoted the neurite outgrowth and maturation (Modarresi et al. 2012).
LncRNAs in brain development
During brain development, an expanding inventory of different lncRNAs is expressed in species including fly, zebrafish, mouse and human (Lipovich et al. 2014; Ponjavic et al. 2009; Young et al. 2012; Ulitsky et al. 2011). Many lncRNAs expressed in the developing brain are associated with protein-coding genes involved in neural gene regulation and brain development due to their genomic context (Qureshi and Mehler 2013). A study on lncRNAs from zebrafish reported that knocking down of two zebrafish lincRNA resulted in encephalodysplasia including initially defects in ventricular morphology (i.e., forebrain ventricle contraction) and subsequently in enlarged ventricles (i.e., hydrocephalus), small heads and eyes and neurogenic differentiation-positive neurons loss in the retina (Ulitsky et al. 2011). Loss of the Dlx6 antisense RNA 1 (Dlx6as1) lncRNA could induce the deregulated expression of transcription factors involved in the development of gamma-aminobutyric acid (GABA)-ergic interneurons in mouse, as demonstrated by the decreased GABAergic interneurons within the early postnatal hippocampus and dentate gyrus (Bond et al. 2009). Interestingly,a recent study has highlighted that loss of linc–Brn1b results in specific reduction in the number of intermediate progenitor cells (IPCs) in the cerebral cortex. The study (18 lincRNA knockout mouse models) demonstrates that lncRNAs play critical roles in life and brain development in vivo (Sauvageau et al. 2013).
Importantly, compared with that in animals, some of these lncRNAs may exhibit similar expression patterns in human brain with similar functions. The highly accelerated region 1A/B (HAR1A/B) lncRNAs are transcribed from a Human Accelerated Region. For orchestrating forebrain development, HAR1A is expressed specifically in Cajal-Retzius cells of the developing human neocortex during gestational weeks 7–19, a crucial period for cortical neuron specification and migration. This is a pattern similar to that of reelin, a key protein orchestrating forebrain development (Pollard et al. 2006). The evidence has shown that some orthologous lncRNAs exhibit similar expression patterns, implying that they have played a role in the emergence of brain structure and function in the embryonic and postnatal brains of birds and eutherian mammals. Therefore, it is not surprised that lncRNAs have fundamental roles in normal brain development (Qureshi and Mehler 2013; Chodroff et al. 2010). Examples of specific lncRNAs that are dysregulated in neurological disorders are listed in Table 1.
LncRNAs in neurodegenerative diseases and neuroimmunological disorders
LncRNAs in Alzheimer’s disease
Alzheimer’s disease (AD), the most common neurodegenerative disease, is a slowly progressive disease of the brain that is characterized by the progressive loss of synapses and eventually by impairment of memory (Chartier-Harlin et al. 1991; Mattson 2004; Choi et al. 2014; Selkoe 2002). To date, it is a pity that the reasons of this disease have not yet been elucidated. Emerging studies suggest that a series of aberrant lncRNAs can influence the pathogenesis of AD (Liu et al. 2014; Kang et al. 2014; Yang et al. 2014; Faghihi et al. 2008). The β-site amyloid precursor protein-cleaving enzyme 1 (BACE1) is a crucial enzyme that cleaves amyloid precursor protein (APP) and generates amyloid β (Aβ) peptides, which form amyloid plaques on the neurons. Interestingly, although the mechanisms modulating the expression and function of BACE1 in AD are complex and have not completely understood, a conserved antisense transcript, BACE1-AS lncRNA is involved. BACE1-AS is highly expressed in AD patients and in amyloid precursor protein transgenic mice, and it plays the key role in increasing BACE1 mRNA stability, upregulating of BACE1 protein and then generating additional Aβ42 through a post-transcriptional feed-forward mechanism. Those findings highlight that a regulatory noncoding RNA can control the expression of BACE1 mRNA that may drive Alzheimer”s disease-associated pathophysiology (Faghihi et al. 2008).
Another lncRNA detected to be dysregulated in AD is the brain cytoplasmic (BC) RNAs. The BC1 RNA (in mice) and BC200 RNA (in humans) are lncRNA transcripts transported to dendritic processes, and modulate gene expression at translational level. One study has identified that that BC200 levels is increased in brain regions that are preferentially affected in AD (Mus et al. 2007). Several studies have reported that the primate BC200 is associated with synapse plasticity (Lin et al. 2008; Smalheiser 2014; Wang et al. 2002).
LncRNAs in Parkinson’s disease
Parkinson’s disease (PD), the second most common neurodegenerative disorder and the most common movement disorder, is characterized by progressive loss of muscle control. As symptoms worsen, it may become difficult to walk, talk, and complete simple tasks. PD is caused by a lack of dopamine due to the loss of dopamine-producing cells in the brain substantia nigra. Despite of focus research for many years, the pathologic process of PD is still not well understood. Recent studies highlight that some lncRNAs such as PINK-AS1 and UCHL1-AS1 are correlated with Parkinson’s disease (Gutschner and Diederichs 2012; Antoniou et al. 2014).
LncRNAs in Huntington’s disease
Huntington”s disease (HD) is a complex disorder that affect”s a person”s ability to feel, think, and move. HD results from genetically programmed degeneration of nerve cells, called neurons in certain areas of the brain. It is induced by an expansion of a CAG triplet repeat stretch in the Huntingtin(Htt) gene. The expansion can result in a mutant form of the Htt protein. It has been reported that Htt may regulate the nuclear translocation of the transcriptional repressor RE1-silencing transcription factor/neuron-restrictive silencer factor (REST/NRSF), then caused the disrupted expression of REST target genes (Johnson 2012; Shimojo 2008).
To discover the potential HD-associated ncRNAs, a study has determined the expression profile of ncRNAs in human HD brain tissues, and found HAR1 expression was significantly decreased in the striatum. The REST targeted to HAR1 locus by specific DNA regulatory motifs resulted in potent transcriptional repression (Johnson et al. 2010). NEAT1 is a lncRNA involved in cell death mechanisms. Importantly, NEAT1 controls target gene transcription by protein sequestration into paraspeckles (stress-responsive subnuclear structures). One recent study has found that the transcriptional up-regulation of NEAT1 causes the enlargement of these stress-responsive subnuclear structures after proteasome inhibition (Hirose et al. 2014).
LncRNAs in neuroimmunological disorders
Interestingly, accumulating researches proved that the dysregulation of lncRNAs implicated in neuroimmunological disorders (Qureshi et al. 2010; Pang et al. 2009). Multiple sclerosis (MS) is a complex autoimmune disease, and recent studies implicate that lncRNAs may also play an important role in the development and progression of MS. Actually, lncRNA transcripts derived from the mouse T early α (TEA) promoter are responsible, at least in part, for regulating downstream promoter usage (Abarrategui and Krangel 2007). In addition, a study identified a lot of lncRNAs that are dynamically expressed during the differentiation and the activation of T-cell (Pang et al. 2009). For example, a number of lncRNAs are nested within individual introns of the IL2RA gene, and the expression of one of these lncRNAs( M21981) is strongly upregulated with the activation of T-cell. These clues hint that lncRNAs are responsible, at least in part, for mediating neuroimmunological disorders. A study showed that there were 2353 upregulated lncRNAs, 389 downregulated lncRNAs in peripheral blood mononuclear cells of patients with MS (Zhang et al. 2016b). In another study, three lncRNAs were found to be up-regulated in relapsing-remitting MS patients respectively to controls: 7SK small nuclear (RN7SK RNA), taurine up-regulated 1 (TUG1), and nuclear paraspeckle assembly transcript 1 (NEAT1) (Santoro et al. 2016). Although the differentially expressed lncRNAs may be important in the process of MS, it need further study on the specific molecular mechanisms and biological functions of these lncRNAs. Interestingly, Zhang et al. (Zhang et al. 2016c) reported that linc-MAF-4 regulated Th1/Th2 differentiation and was associated with the pathogenesis of multiple sclerosis by targeting MAF. The functions of more lncRNAs should be explored in the future.
LncRNA in neuro-oncological disease
The glioma, a type of tumor, arises from glial cells mostly in the human brain. Several lncRNAs associated with glioma development have been indentified (Wang et al. 2012). Maternally Expressed Gene 3 (MEG3), an imprinted gene located at 14q32, is highly expressed in the human pituitary (Miyoshi et al. 2000) and encodes a lncRNA(approximately 1700 nucleotides) associated with tumorigenesis. There are twelve different MEG3 gene transcripts generated by alternative splicing. A study reported that MEG3 expression was lost in most human tumor cell lines and it was markedly downregulated in glioma tissues compared with adjacent normal tissues (Zhang et al. 2010). Moreover, it is interesting that the lncRNA MEG3 overexpression in human glioma cell lines can inhibit cell proliferation and promote cell apoptosis in U251 and U87 MG human glioma cell lines (Wang et al. 2012). MEG3 hypermethylation can lead to the loss of MEG3 expression, followed by the inhibition of the p53 pathways in gliomas (Li et al. 2016).
It has been reported that lncRNAs can control gene expression at the epigenetic level. Intriguingly, epigenetic regulation of gene expression also have a significant impact on glioma pathogenesis. HOTAIR, as an epigenetic gene regulator, is the most well-known lncRNA. It can indirectly silence HOXD genes by upregulating chromatin modifier complex PRC2 (Gao et al. 2016). A recent study has demonstrated that HOTAIR is overexpressed in glioblastoma multiforme (GBM), where it is key to sustain tumor cell proliferation, and is necessary to induce cell cycle arrest in GBM cells by inhibition of HOTAIR by I-BET151 (Pastori et al. 2015). HOTAIR also primarily serves as a prognostic factor for glioma patient survival, as well as a biomarker for identifying glioma molecular subtypes, a critical regulator of cell cycle progression (Zhang et al. 2013). MALAT1 can promotes tumor cell proliferation and migration (Han et al. 2016; Ma et al. 2016), however, knockdown of MALAT1 may reduce migration in glioblastoma cells, without effect on proliferation (Vassallo et al. 2016). Hypoxia inducible factor 1 alpha-antisense RNA 2 (HIF1A-AS2) up-regulated in mesenchymal GBM stem-like cells (GSCs),may contribute to speciation and adaptation to hypoxia within the tumor microenvironment (Mineo et al. 2016).
Additionally, a study also demonstrated that H19 regulates the development of glioma by deriving miR-675 and provided crucial clues for understanding the key roles of lncRNA-miRNA functional network in glioma (Shi et al. 2014). The lncRNA antisense transcript (anti-NOS2A RNA) is expressed in different types of brain tumors, including meningiomas and glioblastomas (Korneev et al. 2008). Importantly, a recent study has identified lncRNAs associated with cancer subtypes and clinical prognosis, and predicted those that are potential drivers of cancer progression (Du et al. 2013). LncRNAs in other neuro-oncological diseases need to be determined.
LncRNA in psychiatric disorders
Major depressive disorder (MDD), one of the most common psychiatric disorders, affects 10–15% of the general population with high levels of morbidity, disability, and mortality (Tsuang et al. 2004). A recent study showed that circulating lncRNAs were aberrantly changed in MDD. The result suggested that lncRNAs might contribute to the pathogenesis of MDD. Three lncRNAs located at chr10:874,695–874,794, chr10:75,873,456–75,873,642, and chr3:47,048,304–47,048,512 were identified to be associated with MDD through interactions with coding transcripts. Another study on schizophrenia highlights that lncRNAs are dynamically regulated by neuronal activation, including acute downregulation of the lncRNA Gomafu (Barry et al. 2014).
LncRNAs in cerebral injury
Stroke is induced by either cerebral ischemic or hemorrhagic damages in brain (Wang et al. 2009b; Donnan et al. 2008), whichaccounts for approximately 40% of severely disabled patients and is the second most common causes of death worldwide.
LncRNAs may be involved in stroke as the distinct temporal and spatial expression of specific lncRNAs has been detected during cerebral ischemia injury and hypoxic-ischemic brain damage (Dharap et al. 2012; Dharap et al. 2013; Kaur et al. 2014; Zhang et al. 2016a; Zhao et al. 2015). Dharap et al. (Dharap et al. 2012) showed, for the first time, that the cerebral lncRNAs were dysregulated in rats after focal ischemia via middle cerebral artery occlusion. The study also showed that the stroke-responsive lncRNAs were homologous to protein-coding genes including ribosomal complex formation, splicing, translation initiation, and nuclear import of mRNAs. However, the biological functions of these stroke-responsive lncRNAs are currently not determined.
In another study, Dharap et al. (Dharap et al. 2013) reported that 177 of the 2497 lncRNAs expressed in rat cerebral cortex showed significantly increased binding to either Paired amphipathic helix protein Sin3A (Sin3A) or corepressors of the RE-1 silencing transcription factor (coREST) following brain ischemia. For functional analysis, stroke-induced lncRNAs are associated with chromatin-modifying proteins (CMPs) to modulate the post-ischemic pathophysiology. Yet, a recent study reported that the expression of the cell adhesion molecules, Ncam1 and Negr1 mRNAs during neuronal development could be modulated by lncRNAs together with miR-377, which could maintain neuronal structure and function in a vitro model of ischemic-reperfusion injury (Kaur et al. 2014). In addition, it had been demonstrated that endothelial-selective lncRNAs might function as a class of novel master regulators in cerebrovascular endothelial pathologies after ischemic stroke (Zhang et al. 2016a). A recent study also showed that a total of 322 lncRNAs were found to be differentially expressed in hypoxic-ischemic brains. Importantly, lncRNA BC088414 correlated with apoptosis-related genes was one of the most significantly urpregulated lncRNAs (Zhao et al. 2015).
These 5 reports highlight that the lncRNA-based regulatory pathways are associated with cerebral ischemia injury. LncRNAs might provide a unique approach for therapeutic intervention in ischemic injury of brain.
Future prospects and challenge
Great progress has been made in the past 5 year to determine the potential involvement of lncRNAs in neurobiology and neurological diseases, although the study in this field is still at early stage. It is now clear that lncRNAs are highly expressed in brain and reside in distinct neuroanatomical regions, specific cell types, or subcellular components. During brain development, the spatial-temporal expression of lncRNAs is tightly controlled. Thus, different neurological disorders may have their unique expression profiles of lncRNAs. Thus, lncRNAs could be novel biomarkers and new diagnostic approaches for neurological diseases. This could be an important research direction in this research field.
Although thousands of lncRNAs have been identified in brain, the biological functions in CNS of only a few of them have been experimentally determined. Functional assay of these lncRNAs that are aberrantly expressed during neurological diseases should be a major focus in future studies. Based on the functional analysis of lncRNAs, novel therapeutic strategies can be envisioned, which focuses on targeting lncRNA genes, the epigenetic status of lncRNA gene loci, the expression of lncRNA transcript and lncRNA function modulation, etc. In addition, the circuitries involved in lncRNA–mRNA and lncRNA–miRNA, could be modulated by some approaches that change binding site stoichiometry (introducing sponges and masks) and affinities, effectively changing the topologies and functions of these complicated networks. To date, the most important discoveries in this research area are from animal studies. Human and clinical studies should be increased because many human lncRNAs are different from animal lncRNAs.
For the functional identification of lncRNAs, the most approaches current used are bioinformatics analysis and siRNA-based loss-of-function approach. However, the current bioinformatics data are far from satisfaction, and loss-of-function approach is not sufficient enough for any gene study. Virus vector-mediated gene transfer includes adenovirus and lentivirus should be used as a gain-of-function approach. Although adeno-associated virus(AAV) are successfully used to deliver miRNAs, it is difficult to be used for delivery of lncRNAs, because many lncRNAs are over the size limitation for the AAV. Moreover, gene knockout and/or transgenic animals should be developed and broadly used in future studies.
However, as a result, experiments involving lncRNAs require appropriate replicas and controls. Recently, the function of genes has been detected with the bacterial clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated 9 (Cas9) system, which has emerged as a promising method (Kashi et al. 2016; Hsu et al. 2014). CRISPR has been used to delete some MicroRNAs and lncRNAs in human cell lines or in mice in vivo (Ho et al. 2015). Notedly, the CRISPR system might become a promising tool in exploring lncRNA function in the future. CRISPR-Display (CRISP-Disp), a targeted localization method that uses Cas9 to deploy large RNA cargos to DNA loci, could also be applied to the relocalization of lncRNAs to probe their functions. There is no obvious limitation in the size of RNA that can be loaded for this technology (Shechner et al. 2015).
In summary, the importance of lncRNAs in neurobiology, brain development and neurological diseases are beginning to be unraveled. Although the study in this field is still at early stage, a growing body of evidence suggests that lncRNAs may play key roles in neurobiology.
References
Abarrategui I, Krangel MS (2007) Noncoding transcription controls downstream promoters to regulate T-cell receptor alpha recombination. EMBO J 26(20):4380–4390
Amaral PP, Mattick JS (2008) Noncoding RNA in development. Mamm Genome 19(7–8):454–492
Amaral PP, Neyt C, Wilkins SJ et al (2009) Complex architecture and regulated expression of the Sox2ot locus during vertebrate development. RNA 15(11):2013–2027
Antoniou D, Stergiopoulos A, Politis PK (2014) Recent advances in the involvement of long non-coding RNAs in neural stem cell biology and brain pathophysiology. Front Physiol 5:155
Aprea J, Prenninger S, Dori M et al (2013) Transcriptome sequencing during mouse brain development identifies long non-coding RNAs functionally involved in neurogenic commitment. EMBO J 32(24):3145–3160
Azzalin CM, Reichenbach P, Khoriauli L, Giulotto E, Lingner J (2007) Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318(5851):798–801
Barry G, Briggs JA, Vanichkina DP et al (2014) The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing. Mol Psychiatry 19(4):486–494
Bartolomei MS, Zemel S, Tilghman SM (1991) Parental imprinting of the mouse H19 gene. Nature 351(6322):153–155
Bernard D, Prasanth KV, Tripathi V et al (2010) A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J 29(18):3082–3093
Bond AM, Vangompel MJ, Sametsky EA et al (2009) Balanced gene regulation by an embryonic brain ncRNA is critical for adult hippocampal GABA circuitry. Nat Neurosci 12(8):1020–1027
Bras J, Guerreiro R, Hardy J (2012) Use of next-generation sequencing and other whole-genome strategies to dissect neurological disease. Nat Rev Neurosci 13(7):453–464
Brown CJ, Ballabio A, Rupert JL et al (1991) A gene from the region of the human X inactivation Centre is expressed exclusively from the inactive X chromosome. Nature 349(6304):38–44
Chartier-Harlin MC, Crawford F, Houlden H et al (1991) Early-onset Alzheimer's disease caused by mutations at codon 717 of the beta-amyloid precursor protein gene. Nature 353(6347):844–846
Chodroff RA, Goodstadt L, Sirey TM et al (2010) Long noncoding RNA genes: conservation of sequence and brain expression among diverse amniotes. Genome Biol 11(7):R72
Choi SH, Kim YH, Hebisch M et al (2014) A three-dimensional human neural cell culture model of Alzheimer's disease. Nature 515(7526):274–278
Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat Rev Genet 12(7):499–510
Dey BK, Mueller AC, Dutta A (2014) Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription 5(4):e944014
Dharap A, Nakka VP, Vemuganti R (2012) Effect of focal ischemia on long noncoding RNAs. Stroke 43(10):2800–2802
Dharap A, Pokrzywa C, Vemuganti R (2013) Increased binding of stroke-induced long non-coding RNAs to the transcriptional corepressors Sin3A and coREST. ASN Neuro 5(4):283–289
Donnan GA, Fisher M, Macleod M, Davis SM (2008) Stroke Lancet 371(9624):1612–1623
Du Z, Fei T, Verhaak RG et al (2013) Integrative genomic analyses reveal clinically relevant long noncoding RNAs in human cancer. Nat Struct Mol Biol 20(7):908–913
Faghihi MA, Modarresi F, Khalil AM et al (2008) Expression of a noncoding RNA is elevated in Alzheimer's disease and drives rapid feed-forward regulation of beta-secretase. Nat Med 14(7):723–730
Fantes J, Ragge NK, Lynch SA et al (2003) Mutations in SOX2 cause anophthalmia. Nat Genet 33(4):461–463
Feng J, Bi C, Clark BS, Mady R, Shah P, Kohtz JD (2006) The Evf-2 noncoding RNA is transcribed from the dlx-5/6 ultraconserved region and functions as a dlx-2 transcriptional coactivator. Genes Dev 20(11):1470–1484
Francescatto M, Vitezic M, Heutink P, Saxena A (2014) Brain-specific noncoding RNAs are likely to originate in repeats and may play a role in up-regulating genes in cis. Int J Biochem Cell Biol 54:331–337
French PJ, Bliss TV, O’Connor VN (2001) A novel non-coding RNA abundantly expressed in rat brain. Neuroscience 108(2):207–215
Gao YF, Wang ZB, Zhu T et al (2016) A critical overview of long non-coding RNA in glioma etiology 2016: an update. Tumour Biol 37(11):14403–14413
Gutschner T, Diederichs S (2012) The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol 9(6):703–719
Guttman M, Amit I, Garber M et al (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458(7235):223–227
Han Y, Zhou L, Wu T et al (2016) Downregulation of lncRNA-MALAT1 affects proliferation and the expression of Stemness markers in glioma stem cell line SHG139S. Cell Mol Neurobiol 36(7):1097–1107
Hashimoto-Torii K, Motoyama J, Hui CC, Kuroiwa A, Nakafuku M, Shimamura K (2003) Differential activities of sonic hedgehog mediated by Gli transcription factors define distinct neuronal subtypes in the dorsal thalamus. Mech Dev 120(10):1097–1111
Hirose T, Virnicchi G, Tanigawa A et al (2014) NEAT1 long noncoding RNA regulates transcription via protein sequestration within subnuclear bodies. Mol Biol Cell 25(1):169–183
Ho TT, Zhou N, Huang J et al (2015) Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Res 43(3):e17
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157(6):1262–1278
Hung T, Wang Y, Lin MF et al (2011) Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet 43(7):621–629
Iyengar BR, Choudhary A, Sarangdhar MA, Venkatesh KV, Gadgil CJ, Pillai B (2014) Non-coding RNA interact to regulate neuronal development and function. Front Cell Neurosci 8:47
Johnson R (2012) Long non-coding RNAs in Huntington's disease neurodegeneration. Neurobiol Dis 46(2):245–254
Johnson R, Richter N, Jauch R et al (2010) Human accelerated region 1 noncoding RNA is repressed by REST in Huntington's disease. Physiol Genomics 41(3):269–274
Kang MJ, Abdelmohsen K, Hutchison ER et al (2014) HuD regulates coding and noncoding RNA to induce APP → Aβ processing. Cell Rep 7(5):1401–1409
Kapranov P, Cheng J, Dike S et al (2007) RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316(5830):1484–1488
Kashi K, Henderson L, Bonetti A, Carninci P (2016) Discovery and functional analysis of lncRNAs: methodologies to investigate an uncharacterized transcriptome. Biochim Biophys Acta 1859(1):3–15
Kaur P, Karolina DS, Sepramaniam S, Armugam A, Jeyaseelan K (2014) Expression profiling of RNA transcripts during neuronal maturation and ischemic injury. PLoS One 9(7):e103525
Kealy J, Commins S (2010) Frequency-dependent changes in synaptic plasticity and brain-derived neurotrophic factor (BDNF) expression in the CA1 to perirhinal cortex projection. Brain Res 1326:51–61
Kondrashov AV, Kiefmann M, Ebnet K, Khanam T, Muddashetty RS, Brosius J (2005) Inhibitory effect of naked neural BC1 RNA or BC200 RNA on eukaryotic in vitro translation systems is reversed by poly(a)-binding protein (PABP). J Mol Biol 353(1):88–103
Korneev SA, Korneeva EI, Lagarkova MA, Kiselev SL, Critchley G, O'Shea M (2008) Novel noncoding antisense RNA transcribed from human anti-NOS2A locus is differentially regulated during neuronal differentiation of embryonic stem cells. RNA 14(10):2030–2037
Kretz M, Siprashvili Z, Chu C et al (2013) Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493(7431):231–235
Lewejohann L, Skryabin BV, Sachser N et al (2004) Role of a neuronal small non-messenger RNA: behavioural alterations in BC1 RNA-deleted mice. Behav Brain Res 154(1):273–289
Li S, Tighe SW, Nicolet CM et al (2014) Multi-platform assessment of transcriptome profiling using RNA-seq in the ABRF next-generation sequencing study. Nat Biotechnol 32(9):915–925
Li J, Bian EB, He XJ et al (2016) Epigenetic repression of long non-coding RNA MEG3 mediated by DNMT1 represses the p53 pathway in gliomas. Int J Oncol 48(2):723–733
Lin D, Pestova TV, Hellen CU, Tiedge H (2008) Translational control by a small RNA: dendritic BC1 RNA targets the eukaryotic initiation factor 4A helicase mechanism. Mol Cell Biol 28(9):3008–3019
Lipovich L, Tarca AL, Cai J et al (2014) Developmental changes in the transcriptome of human cerebral cortex tissue: long noncoding RNA transcripts. Cereb Cortex 24(6):1451–1459
Liu T, Huang Y, Chen J et al (2014) Attenuated ability of BACE1 to cleave the amyloid precursor protein via silencing long noncoding RNA BACE1-AS expression. Mol Med Rep 10(3):1275–1281
Liu SJ, Nowakowski TJ, Pollen AA et al (2016) Single-cell analysis of long non-coding RNAs in the developing human neocortex. Genome Biol 17:67
Lynch G, Kramar EA, Rex CS et al (2007) Brain-derived neurotrophic factor restores synaptic plasticity in a knock-in mouse model of Huntington's disease. J Neurosci 27(16):4424–4434
Lyons SM, Achorn C, Kedersha NL, Anderson PJ, Ivanov P (2016) YB-1 regulates tiRNA-induced stress granule formation but not translational repression. Nucleic Acids Res 44(14):6949–6960
Ma J, Wang P, Yao Y et al (2016) Knockdown of long non-coding RNA MALAT1 increases the blood-tumor barrier permeability by up-regulating miR-140. Biochim Biophys Acta 1859(2):324–338
Mattick JS (2001) Non-coding RNAs: the architects of eukaryotic complexity. EMBO Rep 2(11):986–991
Mattson MP (2004) Pathways towards and away from Alzheimer's disease. Nature 430(7000):631–639
Mehler MF, Mattick JS (2007) Noncoding RNAs and RNA editing in brain development, functional diversification, and neurological disease. Physiol Rev 87(3):799–823
Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS (2008) Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci U S A 105(2):716–721
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10(3):155–159
Mercer TR, Wilhelm D, Dinger ME et al (2011) Expression of distinct RNAs from 3′ untranslated regions. Nucleic Acids Res 39(6):2393–2403
Meredith EK, Balas MM, Sindy K, Haislop K, Johnson AM (2016) An RNA matchmaker protein regulates the activity of the long noncoding RNA HOTAIR. RNA 22(7):995–1010
Meyer CA, Liu XS (2014) Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. Nat Rev Genet 15(11):709–721
Meyer NP, Roelink H (2003) The amino-terminal region of Gli3 antagonizes the Shh response and acts in dorsoventral fate specification in the developing spinal cord. Dev Biol 257(2):343–355
Mineo M, Ricklefs F, Rooj AK et al (2016) The long non-coding RNA HIF1A-AS2 facilitates the maintenance of mesenchymal glioblastoma stem-like cells in hypoxic niches. Cell Rep 15(11):2500–2509
Miyoshi N, Wagatsuma H, Wakana S et al (2000) Identification of an imprinted gene, Meg3/Gtl2 and its human homologue MEG3, first mapped on mouse distal chromosome 12 and human chromosome 14q. Genes Cells 5(3):211–220
Modarresi F, Faghihi MA, Lopez-Toledano MA et al (2012) Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat Biotechnol 30(5):453–459
Mus E, Hof PR, Tiedge H (2007) Dendritic BC200 RNA in aging and in Alzheimer's disease. Proc Natl Acad Sci U S A 104(25):10679–10684
Ng SY, Johnson R, Stanton LW (2012) Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J 31(3):522–533
Ørom UA, Derrien T, Beringer M et al (2010) Long noncoding RNAs with enhancer-like function in human cells. Cell 143(1):46–58
Pang KC, Dinger ME, Mercer TR et al (2009) Genome-wide identification of long noncoding RNAs in CD8+ T cells. J Immunol 182(12):7738–7748
Pastori C, Kapranov P, Penas C et al (2015) The Bromodomain protein BRD4 controls HOTAIR, a long noncoding RNA essential for glioblastoma proliferation. Proc Natl Acad Sci U S A 112(27):8326–8331
Pollard KS, Salama SR, Lambert N et al (2006) An RNA gene expressed during cortical development evolved rapidly in humans. Nature 443(7108):167–172
Ponjavic J, Oliver PL, Lunter G, Ponting CP (2009) Genomic and transcriptional co-localization of protein-coding and long non-coding RNA pairs in the developing brain. PLoS Genet 5(8):e1000617
Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136(4):629–641
Qureshi IA, Mehler MF (2012) Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease. Nat Rev Neurosci 13(8):528–541
Qureshi IA, Mehler MF (2013) Long non-coding RNAs: novel targets for nervous system disease diagnosis and therapy. Neurotherapeutics 10(4):632–646
Qureshi IA, Mattick JS, Mehler MF (2010) Long non-coding RNAs in nervous system function and disease. Brain Res 1338:20–35
Ramos AD, Diaz A, Nellore A et al (2013) Integration of genome-wide approaches identifies lncRNAs of adult neural stem cells and their progeny in vivo. Cell Stem Cell 12(5):616–628
Ramos AD, Andersen RE, Liu SJ et al (2015) The long noncoding RNA Pnky regulates neuronal differentiation of embryonic and postnatal neural stem cells. Cell Stem Cell 16(4):439–447
Rapicavoli NA, Poth EM, Blackshaw S (2010) The long noncoding RNA RNCR2 directs mouse retinal cell specification. BMC Dev Biol 10:49
Santoro M, Nociti V, Lucchini M, De Fino C, Losavio FA, Mirabella M (2016) Expression profile of long non-coding RNAs in serum of patients with multiple sclerosis. J Mol Neurosci 59(1):18–23
Sauvageau M, Goff LA, Lodato S et al (2013) Multiple knockout mouse models reveal lincRNAs are required for life and brain development. Elife 2:e01749
Selkoe DJ (2002) Alzheimer's disease is a synaptic failure. Science 298(5594):789–791
Shechner DM, Hacisuleyman E, Younger ST, Rinn JL (2015) Multiplexable, locus-specific targeting of long RNAs with CRISPR-display. Nat Methods 12(7):664–670
Shi Y, Wang Y, Luan W et al (2014) Long non-coding RNA H19 promotes glioma cell invasion by deriving miR-675. PLoS One 9(1):e86295
Shimojo M (2008) Huntingtin regulates RE1-silencing transcription factor/neuron-restrictive silencer factor (REST/NRSF) nuclear trafficking indirectly through a complex with REST/NRSF-interacting LIM domain protein (RILP) and dynactin p150 glued. J Biol Chem 283(50):34880–34886
Smalheiser NR (2014) The RNA-centred view of the synapse: non-coding RNAs and synaptic plasticity. Philos Trans R Soc Lond B Biol Sci 369(1652). doi:10.1098/rstb.2013.0504
Sone M, Hayashi T, Tarui H, Agata K, Takeichi M, Nakagawa S (2007) The mRNA-like noncoding RNA Gomafu constitutes a novel nuclear domain in a subset of neurons. J Cell Sci 120(Pt 15):2498–2506
Soreq L, Guffanti A, Salomonis N et al (2014) Long non-coding RNA and alternative splicing modulations in Parkinson's leukocytes identified by RNA sequencing. PLoS Comput Biol 10(3):e1003517
Taft RJ, Pang KC, Mercer TR, Dinger M, Mattick JS (2010) Non-coding RNAs: regulators of disease. J Pathol 220(2):126–139
The ENCODE Project Consortium (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489(7414):57–74
Tochitani S, Hayashizaki Y (2008) Nkx2.2 antisense RNA overexpression enhanced oligodendrocytic differentiation. Biochem Biophys Res Commun 372(4):691–696
Tsuang MT, Taylor L, Faraone SV (2004) An overview of the genetics of psychotic mood disorders. J Psychiatr Res 38(1):3–15
Ulitsky I, Shkumatava A, Jan CH, Sive H, Bartel DP (2011) Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147(7):1537–1550
Vassallo I, Zinn P, Lai M, Rajakannu P, Hamou MF, Hegi ME (2016) WIF1 re-expression in glioblastoma inhibits migration through attenuation of non-canonical WNT signaling by downregulating the lncRNA MALAT1. Oncogene 35(1):12–21
Wang H, Iacoangeli A, Popp S et al (2002) Dendritic BC1 RNA: functional role in regulation of translation initiation. J Neurosci 22(23):10232–10241
Wang Z, Gerstein M, Snyder M (2009b) RNA-seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10(1):57–63
Wang Q, Peng Y, Chen S et al (2009a) Pretreatment with electroacupuncture induces rapid tolerance to focal cerebral ischemia through regulation of endocannabinoid system. Stroke 40(6):2157–2164
Wang P, Ren Z, Sun P (2012) Overexpression of the long non-coding RNA MEG3 impairs in vitro glioma cell proliferation. J Cell Biochem 113(6):1868–1874
Wilusz JE, Sunwoo H, Spector DL (2009) Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 23(13):1494–1504
Wu P, Zuo X, Deng H, Liu X, Liu L, Ji A (2013) Roles of long noncoding RNAs in brain development, functional diversification and neurodegenerative diseases. Brain Res Bull 97:69–80
Xie H, Leung KL, Chen L et al (2010) Brain-derived neurotrophic factor rescues and prevents chronic intermittent hypoxia-induced impairment of hippocampal long-term synaptic plasticity. Neurobiol Dis 40(1):155–162
Yang X, Gao L, Guo X et al (2014) A network based method for analysis of lncRNA-disease associations and prediction of lncRNAs implicated in diseases. PLoS One 9(1):e87797
Young RS, Marques AC, Tibbit C et al (2012) Identification and properties of 1,119 candidate lincRNA loci in the Drosophila melanogaster genome. Genome Biol Evol 4(4):427–442
Zalfa F, Adinolfi S, Napoli I et al (2005) Fragile X mental retardation protein (FMRP) binds specifically to the brain cytoplasmic RNAs BC1/BC200 via a novel RNA-binding motif. J Biol Chem 280(39):33403–33410
Zhang X, Rice K, Wang Y et al (2010) Maternally expressed gene 3 (MEG3) noncoding ribonucleic acid: isoform structure, expression, and functions. Endocrinology 151(3):939–947
Zhang JX, Han L, Bao ZS et al (2013) HOTAIR, a cell cycle-associated long noncoding RNA and a strong predictor of survival, is preferentially expressed in classical and mesenchymal glioma. Neuro-Oncology 15(12):1595–1603
Zhang J, Yuan L, Zhang X et al (2016c) Altered long non-coding RNA transcriptomic profiles in brain microvascular endothelium after cerebral ischemia. Exp Neurol 277:162–170
Zhang F, Gao C, Ma XF et al (2016a) Expression profile of long noncoding RNAs in peripheral blood mononuclear cells from multiple sclerosis patients. CNS Neurosci Ther 22(4):298–305
Zhang F, Liu G, Wei C, Gao C, Hao J (2016b) Linc-MAF-4 regulates Th1/Th2 differentiation and is associated with the pathogenesis of multiple sclerosis by targeting MAF. FASEB J. doi:10.1096/fj.201600838R
Zhao F, Qu Y, Liu J et al (2015) Microarray profiling and Co-expression network analysis of LncRNAs and mRNAs in neonatal rats following hypoxic-ischemic brain damage. Sci Rep 5:13850
Ziats MN, Rennert OM (2013) Aberrant expression of long noncoding RNAs in autistic brain. J Mol Neurosci 49(3):589–593
Acknowledgments
This work was supported by National Natural Science Foundation of China (No.81301055 and No.81271478), partly by Educational Scientific Research Projects of Sichuan Province (No.13ZB0273). We would like to thank Pro. Chunxiang Zhang for his excellent assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors confirm that this article content has no conflicts of interest.
Rights and permissions
About this article
Cite this article
Chen, Y., Zhou, J. LncRNAs: macromolecules with big roles in neurobiology and neurological diseases. Metab Brain Dis 32, 281–291 (2017). https://doi.org/10.1007/s11011-017-9965-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-017-9965-8