Abstract
Primary or acquired drug resistance accounts for the failure of chemotherapy and cancer recurrence in esophageal squamous cell carcinoma (ESCC). However, the aberrant mechanisms driving drug resistance are not fully understood in ESCC. In our previous study, FAT Atypical Cadherin 1 (FAT1) was found to inhibit the epithelial–mesenchymal transition (EMT) process in ESCC. EMT plays a critical role in the development of drug resistance in multiple cancer types. Besides, it equips cancer cells with cancer stem cell (CSC)-like characters that also are associated with chemotherapy resistance. Whether FAT1 regulates the stemness or drug resistance of ESCC cells is worth being explored. Here we found that FAT1 was downregulated in ESCC spheres and negatively correlated with stemness-associated markers including ALDH1A1 and KLF4. Knocking down FAT1 enhanced the sphere-forming ability, resistance to cisplatin and drug efflux of ESCC cells. Additionally, FAT1 knockdown upregulated the expression of drug resistance-related gene ABCC3. Furtherly, we found FAT1 knockdown induced the translocation of β-catenin into nucleus and enhanced its transcriptional activity. The result of ChIP showed that β-catenin was enriched in ABCC3 promoter. Furthermore, β-catenin promoted expression of ABCC3. In conclusion, FAT1 knockdown might enhance the stemness and ABCC3-related cisplatin resistance of ESCC cells via Wnt/β-catenin signaling pathway. FAT1 and its downstream gene ABCC3 might be potential targets for overcoming chemoresistance in ESCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Esophageal cancer ranks the seventh in cancer incidence and sixth in cancer deaths worldwide [1]. The two main types of esophageal cancer are esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC), while ESCC accounts for the majority of esophageal cancer in China [2]. Chemotherapy is one of the main methods to treat ESCC. Chemotherapy drugs applied in ESCC treatment included cisplatin (DDP), 5-fluorouracil (5-FU), adriamycin (ADR) and paclitaxel (PTX) [3,4,5]. However, the response rate remains as low as 40%. The recurrence rate is around 80% and the 5-year survival rate is less than 30% [6, 7]. Primary or acquired drug resistance is the main reason that accounts for the failure of chemotherapy and cancer recurrence. Therefore, understanding the molecular basis and reversing chemoresistance are urgent in terms of ESCC treatments.
Chemoresistance has been attributed to various factors, including increased drug efflux, decreased drug uptake, drug inactivation, target gene mutations, apoptosis defects, and phenotypic transformation [8]. The role of epithelial–mesenchymal transition (EMT) in drug resistance has been well elucidated. For example, analysis of responses to chemotherapy in breast cancer patients revealed a close association between treatment resistance and increased gene expression primarily expressed by stromal cells, and this transcriptional upregulation appears to be caused partly by activation of the EMT program within cancer cells [9]. Besides, EMT equips cancer cells with cancer stem cell (CSC) phenotypes by influencing extracellular and intracellular molecular mechanisms [10, 11]. CSCs are generally in G0 phase. CSCs are self-renewing cells with high tumorigenic ability and associated with chemotherapy resistance.
FAT Atypical Cadherin 1 (FAT1) is a member of the cadherin superfamily, which contains 34 tandem cadherin-type repeats, 5 epidermal growth factor (EGF)-like repeats and laminin A-G domain. It functions as an adhesion molecule and signaling receptor. FAT1 has been identified as one of the significantly mutated genes in multiple ESCC cohort studies by whole genome sequencing [12,13,14,15,16], indicating that FAT1 mutation might be one of the important driver events in tumorigenesis and progression of ESCC. FAT1 has been reported to inhibit the progression of ESCC through regulating the expression of CDK4/CDK6/CCND1 complex or cellular mechanical properties [17,18,19]. Besides that, we have validated that decreased FAT1 induced EMT in ESCC [20, 21], and EMT is closely associated with the development of drug resistance and stemness in various cancers [22]. Consequently, whether FAT1 regulates the stemness or sensitivity of ESCC cells to chemotherapy is worth being further explored.
Materials and methods
Cell culture and sphere formation
Human ESCC cell lines KYSE450 and KYSE180 were collected by our lab. TE-1 and TE-9 were purchased from RIKEN Cell Bank. These cell lines were cultured in RPMI-1640 containing 10% FBS at 37℃, 5% CO2. The lentiviruses for stable overexpression and knockdown of FAT1 were constructed and packaged by Shanghai Genechem Co. (Shanghai, China). Sequence of FAT1 shRNAs were cloned into the pGLV-H1-GFP-Puro and specific sequence of FAT1 wild type was cloned into the Ubi-MCS-flag-CMV-GFP-IRES-Puro. The lentiviruses for stable knockdown of FAT1 were used to infect KYSE450 and TE-9 (named as FAT1-shRNA). The lentiviruses for stable overexpression of FAT1 were used to infect KYSE180 and TE-1 (named as FAT1-WT). The matching empty vector viruses were used as negative control (named as NC). Stable cell lines strains were then treated with 4 μg/mL puromycin (Invitrogen) for positive clone screening. The efficiency of knockdown and overexpression was detected by real-time quantitative PCR (RT-qPCR) and western blot, respectively.
For drug sensitivity assay, 3 × 103 cells were seeded into 96-well plate (Corning) and cultured overnight. Cisplatin was added into cells in concentration 0 μg/mL, 1 μg/mL, 2 μg/mL, 4 μg/mL, 6 μg/mL. After 48 h of cisplatin treatment, the cell viability for each group was measured by MTT.
For sphere-forming assay, cancer cells were plated into 6-well ultra-low attachment plate (Corning) and cultured with DMEM/F12 containing 20 ng/mL human epidermal growth factor (PEPROTECH), 20 ng/mL basic fibroblast growth factor (PEPROTECH), 5 μg/mL insulin (Gibco) and 1 × B27 (Gibco). 1 mL fresh culture medium was added to the culture plate every 3 days. Cells were cultured in suspension for 7 to 14 days. The diameter of spheres was measured under the inverted microscope. The sphere whose diameter was larger than 50 μm was counted.
RNA isolation and RT-qPCR
Total RNA was extracted from cells using the RNAiso Plus (Takara). PrimeScript™ RT Master Mix (Takara) was used to reverse transcript RNA into complementary DNA (cDNA). TB Green® Premix Ex Taq™ II kit (Takara) was used to conduct RT-qPCR. The primers of genes were synthesized in Sangon (Shanghai, China). The expression of genes was normalized by GAPDH and calculated based on \(2^{{ - \Delta \Delta C_{{\text{t}}} }}\) formula.
The primers for FAT1 were as follows: 5′-GAGGCGTTTGATGTGGTTGT-3′ (Forward) and 5′-AACAACGGCGTAGTAGGGAA-3′ (Reverse). The primers for GAPDH were as follows: 5′-TCGTGGAAGGACTCATGACC-3′ (Forward) and 5′-ATGATGTTCTGGAGAGCCCC-3′ (Reverse).
Western blot
ESCC cells were lysed by RIPA lysis buffer. The concentration of total protein was tested by BCA kit (BOSTER). 50 μg protein was used to western blot. Nonspecific protein binding sites of PVDF membrane were blocked by 5% skim milk for 2 h at room temperature. Primary antibodies were incubated with PVDF membrane overnight at 4℃, including ALDH1A1 (Proteintech, 1:1000), KLF4 (Proteintech, 1:1000), GAPDH (Proteintech, 1:5000), FLAG (CST, 1:1000), ABCC3 (Abcam, 1:50), β-catenin (Proteintech, 1:1000). Secondary antibodies were similarly incubated with PVDF membrane for 2 h at room temperature. ECL kit (Beyotime) was used to visualize the protein bands on the PVDF membrane.
Quantification of apoptosis
ESCC cells in each group were treated with 6 μg/mL cisplatin for 48 h and digested with trypsin without EDTA. ESCC cells were washed twice with PBS. ESCC cells were then suspended in binding buffer at a density of 5 × 105 cells/500 μL. 5 μL Annexin V-APC and 5 μL 7-AAD staining solution were added to the cell suspension. The mixture was then incubated for 30 min away from light at room temperature. Finally, apoptosis was evaluated via flow cytometry.
Rhodamine 123 (Rho 123) accumulation assay
Stable FAT1 knockdown or overexpression cell lines emit green fluorescence, so we transfected siRNA of FAT1 into ESCC cells. 36 h after transfection, the cells were digested and counted. 2 mL medium containing 2 × 106 cells was added to 6 cm culture dish. Rho 123 were added with a final concentration of 1 μg/mL and incubated with cells for 60 min at 37℃, 5% CO2. Cells were then washed twice by cold PBS to terminate efflux of Rho 123 and collected under dark conditions. The excitation wavelength of Rho 123 is 480 nm, emission wavelength is 530 nm. The intensity of green fluorescence of Rho 123 was detected by flow cytometry FL1 channel.
Co-immunoprecipitation (Co-IP)
Co-IP was used to investigate whether FAT1 binds to β-catenin. Because the protein encoded by FAT1 gene contains 4588 amino acids with a molecular weight of 506 kDa, FAT1 is not maneuverable. FAT1 plasmid was generous gift from Dr. Luc GT Morris (Human Oncology and Pathogenesis Program, Memorial Sloan-Kettering Cancer Center). It was amenable to experimental manipulation but retained all key functional domains including the N terminus, first two cadherin domains, all five EGF repeats, the trans-membrane region and the cytoplasmic domain. To overexpress stably the wild type FAT1, specific sequence of FAT1 was cloned into the Ubi-MCS-flag-CMV-GFP-IRES-Puro lentivirus expression vector. The infection of lentivirus produced the predicted protein product (named as FAT1-WT). Co-IP was carried out in KYSE180 in which the expression of FAT1 was over-expressed stably. Flag was fused with FAT1 and Flag antibody was used to detect FAT1 protein in KYSE180. 50 μL slurry of anti-DYKDDDDK agarose (Abmart) was added into cell lysate and then they were mixed overnight at 4℃. Furthermore, the precipitation was boiled with SDS-loading buffer and analyzed by western blot using β-catenin antibody.
Immunofluorescence (IF)
Coverslip and 3 × 105 ESCC cells were plated into 6-well plate successively. Then we used 4% paraformaldehyde and 0.25% Triton-100/PBS to fix and permeabilize simultaneously ESCC cells attaching to coverslip for 1 h at room temperature. Nonspecific protein binding sites were blocked by 5% BSA overnight at 4℃. Primary antibodies against β-catenin were added and incubated with cells for 2 h at room temperature. Secondary antibodies were similarly incubated with cells for 2 h at room temperature away from light. Moreover, nucleus was stained by DAPI. In the end, the laser scanning confocal microscopy was used to capture pictures of ESCC cells.
Dual-luciferase reporter assay
TOP flash/FOP flash dual-luciferase reporter system was used to analyze the transcriptional activity of β-catenin. TOP flash/FOP flash plasmids and PRL-Tk plasmid (Generous gift from Dr. Canhua Huang, Sichuan University) were transfected simultaneously into ESCC cells. 48 h after transfection, luciferase reaction reagent (TRANS) was used to measure the activity of the firefly luciferase and renilla luciferase. Data were normalized by renilla luciferase activity, and then we get TOP flash/FOP flash ratios.
For the classical Wnt/β-catenin signaling pathway, β-catenin enters the nucleus and combines with transcription factor TCF/LEF to form a complex to start the transcription of downstream regulatory genes. It is reported that there are two TCF binding sites in ABCC3 promoter region [23]. TCF binding sites were amplified by DNA polymerase (Takara) from genomic DNA. And they were cloned into pGL3-Basic vector via Homologous recombination Kit (YEASEN) to test the effect of FAT1 on ABCC3 transcriptional activity. Then ESCC cells were seeded in 24-well plate and transfected with the pGL3-Basic and PRL-Tk plasmid. The measurement method is same as TOP flash/FOP flash reporter assay.
Chromatin immunoprecipitation (ChIP)
ChIP was performed by the ChIP assay kit (CST) according to the manufacturer’s instructions. DNA fragments bound to β-catenin were precipitated by β-catenin antibody (Proteintech). ChIP was followed by PCR, and the sequences for primers were as follows: 5′-TAGGGCCCTTCAAAGCTTAG-3′ and 5′-CTGTCTGGGGACTACAGGCG-3′ for TCF binding site 1, 5′-TAGGTGAAGAAACAGGGGAA-3′ and 5′-CTCCCAAA TTGCTCCTGAAT-3′ for TCF binding site 2.
Immunohistochemistry (IHC)
We previously constructed subcutaneous tumor-bearing model to detect the effect of FAT1 on subcutaneous tumorigenesis of ESCC cells in nude mice [20]. In this study, ESCC cells KYSE450, TE9, KYSE180 and TE1 which do not have the ability of subcutaneous tumorigenesis in nude mice, were used to detect the effect of FAT1 on stemness and cisplatin resistance of ESCC cells. Therefore, IHC was performed to detect the effect of FAT1 on the subcellular localization of β-catenin in tumor tissue sections of previous project. Antigen retrieval was performed in sodium citrate buffer (pH 6.0). Tissue sections were incubated with FAT1 antibody (Abcam, 1:200), β-catenin antibody (Proteintech, 1:200) at 4℃ overnight. Tissue sections were then incubated with the second antibody (MXB) at 37℃ for 20 min. Tissue sections were then stained with DAB and counterstained with hematoxylin. All images were captured and analyzed with Aperio Cytoplasma 2.0 software.
Statistical analysis
Statistical analysis was performed using SPSS 22.0 software. The differences between two groups were compared by t-test. And one-way ANOVA was used to analyze three or more three data sets. p-value < 0.05 was considered statistically significant.
Results
FAT1 is associated with ESCC cell stemness
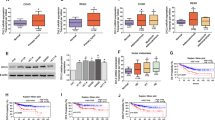
ESCC spheres were isolated by suspension culture in ultra-low attachment plates for KYSE450 and TE1 (Fig. 1A). The expression of stemness-associated biomarkers in ESCC spheres and the corresponding adherent cells was detected by RT-qPCR and western blot. In ESCC spheres, stemness-associated biomarkers including KLF4 and ALDH1A1 were significantly higher than that in the corresponding adherent ESCC cells (Fig. 1B–D). At the same time, the expression of FAT1 in ESCC spheres was remarkably lower than that in corresponding adherent ESCC cells (Fig. 1E). We have also found that the mRNA of FAT1 was negatively correlated with stemness-associated biomarkers including ALDH1A1 and KLF4 by analyzing GSE53625 data set (Fig. 1F, G).
Differentially expressed stemness-associated biomarkers between ESCC spheres and adherent cells. A Representative images of adherent cells and spheres for KYSE450 and TE1. B and C The mRNA expression of stemness-associated biomarkers between ESCC spheres and adherent cells. D The protein expression of stemness-associated biomarkers between ESCC spheres and adherent cells. E The mRNA expression of FAT1 between ESCC spheres and adherent cells. F The relationship between FAT1 and ALDH1A1. G The relationship between FAT1 and KLF4 (*p < 0.05, **p < 0.01, ***p < 0.001)
FAT1 knockdown enhances stemness of ESCC cells
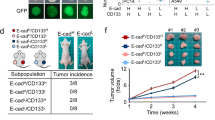
To assess the effect of FAT1 on the stemness of ESCC cell, we knocked down FAT1 in TE-9 and KYSE450 ESCC cells, and overexpressed it in TE-1 and KYSE180 ESCC cells (Fig. 2A–D). Sphere formation assay and colony formation assay were used to assess the influence of FAT1 on the self-renewal ability of ESCC cells. As expected, FAT1 knockdown significantly increased the number and the average diameter of spheres and colonies of ESCC cells. In contrast, FAT1 overexpression decreased the number and the average diameter of the spheres and colonies (Fig. 2E, F). These results indicated that FAT1 knockdown might enhance the stemness of ESCC cells.
FAT1 inhibited sphere formation and colony formation of ESCC cells. A The basal expression of FAT1 in 4 ESCC cell lines. B–D FAT1 was knocked down and overexpressed respectively via lentivirus infection. E FAT1 inhibited the sphere formation ability of ESCC cells. F FAT1 inhibited the colony formation ability of ESCC cells (*p < 0.05, **p < 0.01, ***p < 0.001)
FAT1 knockdown enhances cisplatin resistance of ESCC cells
EMT and stemness play crucial roles in drug resistance in various cancers [22]. Therefore, we tested the effect of FAT1 on drug resistance in ESCC cells. The results of drug sensitivity assay showed that FAT1 knockdown reduced the sensitivity of ESCC cells to cisplatin and increased IC50 value of cisplatin in ESCC cells (Fig. 3A, B). FAT1-overexpressed ESCC cells were more sensitive to cisplatin and the IC50 of cisplatin was lower than that in the negative control (Fig. 3C). Consistent with the result of MTT, the proportion of apoptotic ESCC cells was significantly decreased when FAT1 was knocked down in ESCC cells treated with cisplatin (Fig. 3D–F). These results suggested that FAT1 knockdown could enhance the resistance of ESCC cells to cisplatin.
FAT1 knockdown enhances the resistance of ESCC cells to cisplatin. A–C FAT1 inhibited cell viability in the different concentrations of cisplatin. D–F FAT1 knockdown reduced the fold changes of apoptosis rate in cisplatin treatment. G The intracellular fluorescence intensity of Rho 123 decreased with the decrease of FAT1 (*p < 0.05, **p < 0.01, ***p < 0.001)
The increase of drug efflux is an important factor leading to chemotherapy resistance. Rho 123 accumulation assay is a classic method to reflect the drug efflux of ATP binding cassette (ABC) family members [24]. The results of Rho 123 accumulation assay showed that the intracellular fluorescence intensity was remarkably decreased with the downregulation of FAT1 (Fig. 3G). Therefore, we speculated that FAT1 knockdown might increase the drug efflux, thus enhancing the drug resistance of ESCC cells to cisplatin.
FAT1 knockdown increases ABCC3 expression
ABC transporters are the main drug pumps. They locate on cell membranes and remove chemotherapeutic drugs and cytotoxic agents effectively, which account mainly for the resistance to chemotherapy. Consequently, the expression of ABC transporter family genes including ABCB1, ABCC1, ABCC2, ABCC3, ABCC4, ABCC5, ABCC6, and ABCG2 were detected. The result indicated that FAT1 knockdown significantly increased the expression of ABCC3 in both KYSE450 and TE9, but it did not affect the ABC members as listed above (Fig. 4A, B). Besides, ABCC3 knockdown weaken the influence of FAT1 on the resistance of ESCC cells to cisplatin (Fig. 4C). The results indicated that FAT1 might be involved in the chemoresistance by regulating the expression of ABCC3.
Decreased FAT1 promotes the release of β-catenin by unbinding to it
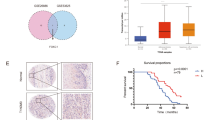
To determine how FAT1 knockdown enhances the cisplatin resistance of ESCC cells via ABCC3, we re-analyzed RNA sequencing data of FAT1 knockdown [20]. 80 differentially expressed genes (DEGs) were identified that were changed simultaneously in both FAT1-knockdown KYSE150 and KYSE180 cells. KEGG was used to analyze the enriched pathway of DEGs. And the pathway included Wnt/β-catenin signaling pathway, NF-κB signaling pathway, and NOD-like receptor signaling pathway (Fig. 5A, B). In the above pathways, Wnt/β-catenin signaling pathway have been reported to play important roles in regulating drug resistance of cancer cells [25].
FAT1 regulated Wnt/β-catenin signaling pathway. A and B Altered signaling pathways were shown by KEGG analysis in FAT1 knockdown cells including the Wnt/β-catenin signaling pathway. C FAT1 knockdown increased the transcriptional activity of β-catenin. D The interaction between FAT1 and β-catenin was revealed by Co-IP. E The result of IF showed that FAT1 knockdown increased the expression level of β-catenin in nucleus of ESCC cells. F The result of IHC showed that FAT1 knockdown increased the expression level of β-catenin both in nucleus and cytoplasm in tumor tissue sections (*p < 0.05, **p < 0.01, ***p < 0.001)
Firstly, the activity of Wnt/β-catenin signaling pathway was detected by TOP flash/FOP flash dual-luciferase reporter system. The results showed that the value of TOP flash/FOP flash was increased significantly after FAT1 knockdown (Fig. 5C), indicating that knocking down FAT1 can increase the transcriptional activity of β-catenin. β-Catenin is an essential component of cell adhesion through its interaction with E-cadherin on the cytoplasmic side of cell membrane [26]. FAT1, a member of cell membrane-cadherin complexes, maintains the polarity and morphology of cells by combining with stress fibers, Ena/Vasp, and β-catenin that binding to the cytoplasmic domain of FAT1 [27,28,29]. Similarly, the result of Co-IP indicated that FAT1 bound to β-catenin in ESCC cells (Fig. 5D). And the result of IF showed that FAT1 knockdown increased the expression level of β-catenin in nucleus of ESCC cells (Fig. 5E). The result of IHC also showed that FAT1 knockdown increased the expression level of β-catenin in both nucleus and cytoplasm in tumor tissue sections (Fig. 5F). Therefore, FAT1 knockdown might release more β-catenin by unbinding to it, which was subsequently translocated from cell membrane into cell nucleus.
β-Catenin mediates the function of FAT1 and regulates ABCC3
We further detected the effect of β-catenin on FAT1 function. β-Catenin is encoded by CTNNB1 gene. The results showed that β-catenin knockdown rescued the influence of FAT1 knockdown on the resistance of ESCC cells to cisplatin (Fig. 6A). And β-catenin knockdown increased the proportion of apoptotic ESCC cells in cisplatin treatment (Fig. 6B). Furthermore, β-catenin knockdown significantly decreased the expression of ABCC3 (Fig. 6C, D). To clarify whether FAT1 regulated the expression of ABCC3 via β-catenin in ESCC, we cloned ABCC3 promoter with potential TCF binding sites (sites 1 and 2) into pGL3-Basic vector (Fig. 6E). ChIP-PCR results revealed that β-catenin was enriched in TCF binding site 1, but not in TCF binding site 2 (Fig. 6F). And the results of dual-luciferase reporter assay showed that FAT1 knockdown significantly increased the transcriptional activity of ABCC3 promoter region with TCF binding site 1 (Fig. 6G). Consequently, FAT1 might regulate the expression of ABCC3 via β-catenin in ESCC.
β-Catenin mediates the function of FAT1 and regulates ABCC3. A β-Catenin knockdown decreased the cell viability induced by FAT1 knockdown. B β-Catenin knockdown increased the fold changes of apoptosis rate reduced by FAT1 knockdown with cisplatin treatment. C and D β-Catenin knockdown decreased the expression of ABCC3. E Documented TCF binding sites of ABCC3 were cloned into the pGL3-Basic vector. F The result of ChIP revealed that β-catenin was enriched in the TCF binding site 1 of ABCC3 promoter region. G FAT1 knockdown increased the transcriptional activity of ABCC3 by TCF binding site 1 (*p < 0.05, **p < 0.01, ***p < 0.001)
Correlation between FAT1 and cisplatin resistance in cancer patients
To further confirm that FAT1 is involved in cisplatin resistance of cancer patients, we collected the clinical data of cisplatin treatments from TCGA, including 277 patients across 14 cancer types. Cisplatin treatment response can be divided into complete response, partial response, stable disease and clinical progressive disease. FAT1 expression was different in the four groups of pan-cancer patients (Fig. Supplement A), especially higher in patients of complete response group than that in patients of partial response group (Fig. 7A). We further analyzed FAT1 expression in patients with pan-squamous cell carcinoma (pan-SCC), head and neck squamous cell carcinoma (HNSC) and cervical squamous cell carcinoma (CSCC). Similarly, FAT1 expression was different in the four groups (Fig. Supplement B–D), and higher in complete response group than that in partial response group, especially for HNSC (Fig. 7B–D). Therefore, the higher the expression of FAT1, the better the response of patients to cisplatin treatment.
FAT1 expression is different in complete response group and partial response group. A FAT1 expression was higher in complete response group than that in partial response group for 225 patients pan-cancer. B FAT1 expression was significantly higher in complete response group than that in partial response group for 100 patients with pan-SCC. C FAT1 expression was significantly higher in complete response group than that in partial response group for 39 patients with HNSC. D FAT1 expression was higher in complete response group than that in partial response group for 48 patients with CSCC. E A proposed model for FAT1 regulatory mechanism in ESCC progression
Discussion
In this study, we found the regulation of FAT1 on the stemness and cisplatin resistance of ESCC cells and explored its underlying mechanism. We found that FAT1 was negatively correlated with stemness-associated markers including ALDH1A1 and KLF4. Besides that, FAT1 was downregulated in ESCC spheres compared with ESCC cells. FAT1 knockdown enhanced the sphere-forming ability, resistance to cisplatin and drug efflux of ESCC cells. Additionally, FAT1 knockdown upregulated the expression of drug resistance-related gene ABCC3. Furtherly, we found FAT1 knockdown induced the translocation of β-catenin into nucleus and enhanced its transcriptional activity. The result of ChIP showed that β-catenin was enriched in ABCC3 promoter. Furthermore, β-catenin increased the expression of ABCC3. The TCGA data further confirmed the correlation between FAT1 and cisplatin resistance in cancer patients.
Whole genome sequencing of ESCC cohorts have shown FAT1 is a significantly mutated gene in ESCC. And multiple studies have shown that FAT1 could act as an oncogene or suppressor gene in tumor progression [12,13,14,15,16]. Most of all, we have demonstrated that FAT1 prevented EMT via MAPK/ERK signaling pathway in ESCC [20]. EMT is one of the crucial reasons for stemness and drug resistance in various cancer. Thus, we further explored the effect of FAT1 on stemness and drug resistance in ESCC cells. Our experiments showed that FAT1 was downregulated in ESCC spheres. And its knockdown enhanced the stemness of ESCC cells and resistance of ESCC cells to cisplatin. Analysis of RNA sequencing data showed Wnt/β-catenin signaling pathway was activated by decreased FAT1.
For the β-catenin-dependent Wnt signaling pathway, unphosphorylated β-catenin migrates into the nucleus and binds with TCF/LEF to activate transcription of downstream target genes involving in the progression of cancer [30,31,32]. TOP flash/FOP flash dual-luciferase reporter system has depicted that FAT1 knockdown increased the transcriptional activity of the β-catenin in ESCC cells. And we have verified that FAT1 bound to β-catenin by Co-IP and FAT1 knockdown facilitated β-catenin translocated into the nucleus by IF and IHC. β-Catenin knockdown rescued the resistance ability of ESCC cells to cisplatin induced by FAT1 knockdown in vitro.
TCF7L2 binds to the ABCC3 locus and cooperates with β-catenin regulates ABCC3 expression in colorectal cancer [23]. And we have found that β-catenin knockdown decreased the expression ABCC3 in ESCC cells. Consequently, we performed ChIP and cloned ABCC3 promoter with potential TCF binding sites (sites 1 and 2) into pGL3-Basic vector. We ultimately found that β-catenin enriched in promoter of ABCC3. And FAT1 knockdown increased the transcriptional activity of ABCC3.
In conclusion, FAT1 downregulation enhances stemness and cisplatin resistance of ESCC cells. FAT1 knockdown might increase the transcription and expression of ABCC3 by releasing more β-catenin into cell nucleus. FAT1 and its downstream gene might be potential targets for the new molecular targeting treatment to improve the diagnosis and prognosis of ESCC.
Data availability
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Arnold M, Soerjomataram I, Ferlay J, Forman D (2015) Global incidence of oesophageal cancer by histological subtype in 2012. Gut 64:381–387. https://doi.org/10.1136/gutjnl-2014-308124
Wang T, Yu J, Liu M, Chen Y, Zhu C, Lu L, Wang M, Min L, Liu X, Zhang X, Gubat JA, Chen Y (2019) The benefit of taxane-based therapies over fluoropyrimidine plus platinum (FP) in the treatment of esophageal cancer: a meta-analysis of clinical studies. Drug Des Dev Ther 13:539–553. https://doi.org/10.2147/DDDT.S189514
Ando N, Kato H, Igaki H, Shinoda M, Ozawa S, Shimizu H, Nakamura T, Yabusaki H, Aoyama N, Kurita A, Ikeda K, Kanda T, Tsujinaka T, Nakamura K, Fukuda H (2012) A randomized trial comparing postoperative adjuvant chemotherapy with cisplatin and 5-fluorouracil versus preoperative chemotherapy for localized advanced squamous cell carcinoma of the thoracic esophagus (JCOG9907). Ann Surg Oncol 19:68–74. https://doi.org/10.1245/s10434-011-2049-9
Ter Veer E, Haj Mohammad N, van Valkenhoef G, Ngai LL, Mali RMA, Anderegg MC, van Oijen MGH, van Laarhoven HWM (2016) The efficacy and safety of first-line chemotherapy in advanced esophagogastric cancer: a network meta-analysis. J Natl Cancer Inst. https://doi.org/10.1093/jnci/djw166
Ilson DH (2008) Esophageal cancer chemotherapy: recent advances. Gastrointest Cancer Res 2:85–92
Altorki N, Harrison S (2017) What is the role of neoadjuvant chemotherapy, radiation, and adjuvant treatment in resectable esophageal cancer? Ann Cardiothorac Surg 6:167–174. https://doi.org/10.21037/acs.2017.03.16
Shibue T, Weinberg RA (2017) EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat Rev Clin Oncol 14:611–629. https://doi.org/10.1038/nrclinonc.2017.44
Farmer P, Bonnefoi H, Anderle P, Cameron D, Wirapati P, Becette V, Andre S, Piccart M, Campone M, Brain E, Macgrogan G, Petit T, Jassem J, Bibeau F, Blot E, Bogaerts J, Aguet M, Bergh J, Iggo R, Delorenzi M (2009) A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nat Med 15:68–74. https://doi.org/10.1038/nm.1908
Scheel C, Eaton EN, Li SH, Chaffer CL, Reinhardt F, Kah KJ, Bell G, Guo W, Rubin J, Richardson AL, Weinberg RA (2011) Paracrine and autocrine signals induce and maintain mesenchymal and stem cell states in the breast. Cell 145:926–940. https://doi.org/10.1016/j.cell.2011.04.029
Ni T, Li XY, Lu N, An T, Liu ZP, Fu R, Lv WC, Zhang YW, Xu XJ, Grant Rowe R, Lin YS, Scherer A, Feinberg T, Zheng XQ, Chen BA, Liu XS, Guo QL, Wu ZQ, Weiss SJ (2016) Snail1-dependent p53 repression regulates expansion and activity of tumour-initiating cells in breast cancer. Nat Cell Biol 18:1221–1232. https://doi.org/10.1038/ncb3425
Zhang L, Zhou Y, Cheng C, Cui H, Cheng L, Kong P, Wang J, Li Y, Chen W, Song B, Wang F, Jia Z, Li L, Li Y, Yang B, Liu J, Shi R, Bi Y, Zhang Y, Wang J, Zhao Z, Hu X, Yang J, Li H, Gao Z, Chen G, Huang X, Yang X, Wan S, Chen C, Li B, Tan Y, Chen L, He M, Xie S, Li X, Zhuang X, Wang M, Xia Z, Luo L, Ma J, Dong B, Zhao J, Song Y, Ou Y, Li E, Xu L, Wang J, Xi Y, Li G, Xu E, Liang J, Yang X, Guo J, Chen X, Zhang Y, Li Q, Liu L, Li Y, Zhang X, Yang H, Lin D, Cheng X, Guo Y, Wang J, Zhan Q, Cui Y (2015) Genomic analyses reveal mutational signatures and frequently altered genes in esophageal squamous cell carcinoma. Am J Hum Genet 96:597–611. https://doi.org/10.1016/j.ajhg.2015.02.017
Lin DC, Hao JJ, Nagata Y, Xu L, Shang L, Meng X, Sato Y, Okuno Y, Varela AM, Ding LW, Garg M, Liu LZ, Yang H, Yin D, Shi ZZ, Jiang YY, Gu WY, Gong T, Zhang Y, Xu X, Kalid O, Shacham S, Ogawa S, Wang MR, Koeffler HP (2014) Genomic and molecular characterization of esophageal squamous cell carcinoma. Nat Genet 46:467–473. https://doi.org/10.1038/ng.2935
Gao YB, Chen ZL, Li JG, Hu XD, Shi XJ, Sun ZM, Zhang F, Zhao ZR, Li ZT, Liu ZY, Zhao YD, Sun J, Zhou CC, Yao R, Wang SY, Wang P, Sun N, Zhang BH, Dong JS, Yu Y, Luo M, Feng XL, Shi SS, Zhou F, Tan FW, Qiu B, Li N, Shao K, Zhang LJ, Zhang LJ, Xue Q, Gao SG, He J (2014) Genetic landscape of esophageal squamous cell carcinoma. Nat Genet 46:1097–1102. https://doi.org/10.1038/ng.3076
Song Y, Li L, Ou Y, Gao Z, Li E, Li X, Zhang W, Wang J, Xu L, Zhou Y, Ma X, Liu L, Zhao Z, Huang X, Fan J, Dong L, Chen G, Ma L, Yang J, Chen L, He M, Li M, Zhuang X, Huang K, Qiu K, Yin G, Guo G, Feng Q, Chen P, Wu Z, Wu J, Ma L, Zhao J, Luo L, Fu M, Xu B, Chen B, Li Y, Tong T, Wang M, Liu Z, Lin D, Zhang X, Yang H, Wang J, Zhan Q (2014) Identification of genomic alterations in oesophageal squamous cell cancer. Nature 509:91–95. https://doi.org/10.1038/nature13176
Cui Y, Chen H, Xi R, Cui H, Zhao Y, Xu E, Yan T, Lu X, Huang F, Kong P, Li Y, Zhu X, Wang J, Zhu W, Wang J, Ma Y, Zhou Y, Guo S, Zhang L, Liu Y, Wang B, Xi Y, Sun R, Yu X, Zhai Y, Wang F, Yang J, Yang B, Cheng C, Liu J, Song B, Li H, Wang Y, Zhang Y, Cheng X, Zhan Q, Li Y, Liu Z (2020) Whole-genome sequencing of 508 patients identifies key molecular features associated with poor prognosis in esophageal squamous cell carcinoma. Cell Res 30:902–913. https://doi.org/10.1038/s41422-020-0333-6
Wang Y, Wang G, Ma Y, Teng J, Wang Y, Cui Y, Dong Y, Shao S, Zhan Q, Liu X (2019) FAT1, a direct transcriptional target of E2F1, suppresses cell proliferation, migration and invasion in esophageal squamous cell carcinoma. Chin J Cancer Res 31:609–619. https://doi.org/10.21147/j.issn.1000-9604.2019.04.05
Hu XL, Zhai YF, Li GD, Xing JF, Yang J, Bi YH, Wang J, Shi RY (2018) FAT1 inhibits cell proliferation of esophageal squamous cell carcinoma through regulating the expression of CDK4/CDK6/CCND1 complex. Zhonghua Zhong Liu Za Zhi 40:14–20. https://doi.org/10.3760/cma.j.issn.0253-3766.2018.01.003
Hu X, Zhai Y, Shi R, Qian Y, Cui H, Yang J, Bi Y, Yan T, Yang J, Ma Y, Zhang L, Liu Y, Li G, Zhang M, Cui Y, Kong P, Cheng X (2018) FAT1 inhibits cell migration and invasion by affecting cellular mechanical properties in esophageal squamous cell carcinoma. Oncol Rep 39:2136–2146. https://doi.org/10.3892/or.2018.6328
Hu X, Zhai Y, Kong P, Cui H, Yan T, Yang J, Qian Y, Ma Y, Wang F, Li H, Cheng C, Zhang L, Jia Z, Li Y, Yang B, Xu E, Wang J, Yang J, Bi Y, Chang L, Wang Y, Zhang Y, Song B, Li G, Shi R, Liu J, Zhang M, Cheng X, Cui Y (2017) FAT1 prevents epithelial mesenchymal transition (EMT) via MAPK/ERK signaling pathway in esophageal squamous cell cancer. Cancer Lett 397:83-93. https://doi.org/10.1016/j.canlet.2017.03.033
Pastushenko I, Mauri F, Song Y, de Cock F, Meeusen B, Swedlund B, Impens F, Van Haver D, Opitz M, Thery M, Bareche Y, Lapouge G, Vermeersch M, Van Eycke YR, Balsat C, Decaestecker C, Sokolow Y, Hassid S, Perez-Bustillo A, Agreda-Moreno B, Rios-Buceta L, Jaen P, Redondo P, Sieira-Gil R, Millan-Cayetano JF, Sanmatrtin O, D’Haene N, Moers V, Rozzi M, Blondeau J, Lemaire S, Scozzaro S, Janssens V, De Troya M, Dubois C, Perez-Morga D, Salmon I, Sotiriou C, Helmbacher F, Blanpain C (2021) Fat1 deletion promotes hybrid EMT state, tumour stemness and metastasis. Nature 589:448–455. https://doi.org/10.1038/s41586-020-03046-1
Ashrafizadeh M, Zarrabi A, Hushmandi K, Kalantari M, Mohammadinejad R, Javaheri T, Sethi G (2020) Association of the epithelial–mesenchymal transition (EMT) with cisplatin resistance. Int J Mol Sci. https://doi.org/10.3390/ijms21114002
Kobayashi M, Funayama R, Ohnuma S, Unno M, Nakayama K (2016) Wnt-beta-catenin signaling regulates ABCC3 (MRP3) transporter expression in colorectal cancer. Cancer Sci 107:1776–1784. https://doi.org/10.1111/cas.13097
Altenberg GA, Vanoye CG, Horton JK, Reuss L (1994) Unidirectional fluxes of rhodamine 123 in multidrug-resistant cells: evidence against direct drug extrusion from the plasma membrane. Proc Natl Acad Sci USA 91:4654–4657. https://doi.org/10.1073/pnas.91.11.4654
Kumar V, Vashishta M, Kong L, Wu X, Lu JJ, Guha C, Dwarakanath BS (2021) The role of Notch, Hedgehog, and Wnt signaling pathways in the resistance of tumors to anticancer therapies. Front Cell Dev Biol 9:650772. https://doi.org/10.3389/fcell.2021.650772
Nelson WJ, Nusse R (2004) Convergence of Wnt, beta-catenin, and cadherin pathways. Science 303:1483–1487. https://doi.org/10.1126/science.1094291
Tanoue T, Takeichi M (2004) Mammalian Fat1 cadherin regulates actin dynamics and cell–cell contact. J Cell Biol 165:517–528. https://doi.org/10.1083/jcb.200403006
Moeller MJ, Soofi A, Braun GS, Li X, Watzl C, Kriz W, Holzman LB (2004) Protocadherin FAT1 binds Ena/VASP proteins and is necessary for actin dynamics and cell polarization. EMBO J 23:3769–3779. https://doi.org/10.1038/sj.emboj.7600380
Morris LG, Kaufman AM, Gong Y, Ramaswami D, Walsh LA, Turcan S, Eng S, Kannan K, Zou Y, Peng L, Banuchi VE, Paty P, Zeng Z, Vakiani E, Solit D, Singh B, Ganly I, Liau L, Cloughesy TC, Mischel PS, Mellinghoff IK, Chan TA (2013) Recurrent somatic mutation of FAT1 in multiple human cancers leads to aberrant Wnt activation. Nat Genet 45:253–261. https://doi.org/10.1038/ng.2538
Gordon MD, Nusse R (2006) Wnt signaling: multiple pathways, multiple receptors, and multiple transcription factors. J Biol Chem 281:22429–22433. https://doi.org/10.1074/jbc.R600015200
Kohn AD, Moon RT (2005) Wnt and calcium signaling: beta-catenin-independent pathways. Cell Calcium 38:439–446. https://doi.org/10.1016/j.ceca.2005.06.022
Bilic J, Huang YL, Davidson G, Zimmermann T, Cruciat CM, Bienz M, Niehrs C (2007) Wnt induces LRP6 signalosomes and promotes dishevelled-dependent LRP6 phosphorylation. Science 316:1619–1622. https://doi.org/10.1126/science.1137065
Funding
This work was supported by funding from the National Natural Science Foundation of China (81802825 to X.H.).
Author information
Authors and Affiliations
Contributions
Conceptualization: XC, XH, YZ; Methodology: CS; Validation: CS, YZ; Formal analysis: PK, HZ; Investigation: HZ; Resources: XH; Data curation: LZ, CS; Writing original draft: YZ; Writing review and editing: YW; Supervision: PK, XC; Project administration: XH, XC; Funding acquisition: XH. All authors have contributed significantly and approved the submitted version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
All experimental samples were authorized by the Ethics Committee of Shanxi Medical University. All institutional and national guidelines for the care and use of laboratory animals were followed. No human subjects were involved in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11010_2022_4475_MOESM1_ESM.tif
Supplementary file1 (TIF 1675 kb) Fig. Supplement FAT1 expression is different in the four groups. A–D FAT1 expression was different in the four groups for 277 patients across cancer types, 115 patients with pan-SCC, 42 patients with HNSC and 57 patients with CSCC. E The relationship between FAT1 and ABCC3 by analyzing GSE53625 data set. F The relationship between CTNNB1 and ABCC3 by analyzing GSE53625 data set
Rights and permissions
About this article
Cite this article
Zhai, Y., Shan, C., Zhang, H. et al. FAT1 downregulation enhances stemness and cisplatin resistance in esophageal squamous cell carcinoma. Mol Cell Biochem 477, 2689–2702 (2022). https://doi.org/10.1007/s11010-022-04475-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-022-04475-4